Abstract
This study elucidated the potential molecular mechanism of chrysin in hepatocellular carcinoma (HCC) treatment using network pharmacology and in vitro experiments. Chrysin and candidate targets of HCC were obtained from the TCMSP and DrugBank databases, followed by mapping and screening of chrysin and HCC targets to identify the core targets of chrysin in HCC treatment. The interaction of chrysin and its targets, including CDK1, CDK5, as well as MMP9, were evaluated by molecular docking. The STRING database and Cytoscape (version 3.8.2) software were used to construct protein interactions and component-target networks of the core targets. Gene Ontology (GO) annotation and Kyoto Encyclopedia of Genes and Genomes (KEGG) signaling pathway enrichment analysis of the core target genes were performed using the DAVID database. Network pharmacology results showed that chrysin treatment of HCC was mainly related to cell proliferation and cell cycle. Accordingly, the cell counting kit-8 method and flow cytometry were used to detect the cell viability and cell cycle of hepatocarcinoma cells HCCLM3 and BEL-7402 in vitro. A total of 142 compound targets of chrysin, 12,179 HCC-related targets, and 116 intersecting targets were screened. The first 20 GO biological annotations of 17 core targets and the first 20 KEGG pathways mainly involved cell proliferation and cell cycle. In vitro experiments showed that chrysin inhibits the proliferation of human hepatocarcinoma cells (HCCLM3 and BEL-7402) in a dose-dependent manner. Moreover, chrysin induced cell cycle arrest in HCCLM3 and BEL-7402 cells in the G2 phase, and the expression was downregulated of cyclin-dependent kinases (CDKs), CDK2 and CDK4. Chrysin can offset HCC mainly by regulating the cell cycle and inhibiting cell proliferation. The network pharmacology results were verified, providing the basis for further study on the mechanism of chrysin intervention in HCC.
Introduction
Hepatocellular carcinoma (HCC) is one of the most aggressive and lethal cancers worldwide.1,2 HCC exhibits no obvious symptoms in the early stage as the symptoms often manifest in the late stage. HCC exhibits characteristics of high malignancy and poor prognosis, which causes severe harm to the health of the population. Some treatments for HCC include surgery, liver transplantation, chemotherapy, and radioembolization3–5. However, due to the high recurrence rate and lack of appropriate treatment, approximately two-thirds of patients with HCC have poor physical conditions and prognoses. Therefore, new antitumor drugs with low toxicity and high efficacy are required for the treatment of HCC6–8. Natural sources are important resources for developing antitumor drugs. Several natural pharmaceutical ingredients have been used in treating hepatocellular carcinoma 9 . Flavone is a natural polyphenol that is widely distributed in fruits, vegetables, and other plants. Flavone is also the active ingredient of many traditional Chinese medicines. Early studies have found that flavonoids exhibit various pharmacological activities such as anti-inflammatory, antioxidant, and antitumor effects. Additionally, flavonoids can induce apoptosis and inhibit the proliferation of HCC, breast cancer, lung cancer, and other cancer cells in a dose- and time-dependent manner10–12.
Chrysin (5,7-dihydroxyflavone) is a bioactive flavonoid obtained from plants and their extracts, such as passionflower, propolis, and honey, which are widely used as herbal medicines in China. Besides its diverse antioxidant 13 , anti-inflammatory 14 and antibacterial 15 activities, chrysin also exhibits strong antitumor effects16–19. Studies have shown that chrysin exerts antitumor effects by inducing cell-cycle arrest and apoptosis via different mechanisms, such as the activation of exogenous apoptotic pathways 20 and modification of cyclins and cyclin-dependent kinases (CDKs) 21 . Moreover, chrysin has been shown to regulate the Ras-Raf-MAPKs, PI3K-Akt, STAT, NF-κB, Wnt-β-catenin, and Notch signaling pathways in tumor cells to inhibit cell proliferation, angiogenesis, invasion, and metastasis22–24. However, the specific molecular mechanism and material basis of chrysin in HCC treatment require further elucidation. Therefore, in this study, we used a network pharmacology approach to construct a biological network and employed network visualization analysis for the complex interactions between drug components and diseases. We explored the potential molecular mechanism of HCC treatment with chrysin and conducted in vitro experiments to verify the network pharmacological prediction results. Furthermore, we clarified the molecular mechanism of chrysin in HCC treatment. To the best of our knowledge, this is the first study to use network pharmacology and experimental verification methods to study the potential mechanism of chrysin intervention in HCC.
Results
Construction and Analysis of the Relationship Between Chrysin and HCC Target Network
The secondary structure of chrysin was determined by searching the TCMSP and PubChem databases to obtain the corresponding SMILE number. This was then imported into the Swiss Target and Prediction databases to obtain the candidate targets of chrysin components. After deleting the duplicates, a total of 119 targets were obtained. Then, a query of DisGenet, OMIM, TTD, and other databases was conducted to obtain HCC targets, which were then entered into the Uniprot database for correction. After sorting, 17,355 disease targets were included. Venny2.1 (Venny 2.1.0 – BioinfoGP; bioinfogp.cnb.csic.es › tools › venny) was used to map the targets of chrysin and HCC, and the results showed that 116 intersection targets of the two sets of genes occurred, as shown in Figure 1 (A). The components and targets were visualized using the Cytoscape version 3.8.2 (Cytoscapecytoscape.org) software, and a component-target network diagram was illustrated, as shown in Figure 1 (B). The red diamond represents the composition of salicylic acid, and the yellow circle represents the intersection target of salicylic acid and HCC, which is the potential target of salicylic acid in HCC treatment.

Construction and analysis of the relationship between chrysin and HCC target network. (A) Wayne, diagram of chrysin and hepatocellular carcinoma candidate targets. (B) Chrysin targeting-hepatocellular carcinoma network. The red diamond represents the Chrysin component, and the light-yellow circle represents the potential target of Chrysin in the treatment of hepatocellular carcinoma. (C) Protein-protein interaction (PPI) network consisting of 116 potential targets. The “node” in the network represents the target protein, and the “edge” represents the interaction between the target proteins. The greater the number of edges, the more important is the role of the target protein corresponding to the node in the network.
Protein Interaction Network (PPI) of Protein Interactions
To clarify the interaction between the intersection target proteins, we logged into the STRING database, uploaded the target of chrysin intervention in HCC, and set the protein type as “Homo Sapiens” to exclude proteins that are independent of the network and provide associations between two different nodes (gene/protein). The constructed PPI network is shown in Figure 1 (C), and a confidence score of > 0.95 was considered to be highly confident. Table 1 lists the acting nodes with high confidence scores, such as TOP2A, PTPN1, CDK5, CDK1, CalM1, F2, and MMP9. The highly connected nodes in the network usually involved the same biological function or pathway, so the key target proteins were determined according to the topological characteristics of network nodes in the PPI. Notably, the top target proteins CDK1, CDK5, CDK6, and TOP2A are all related to the proliferation and cycle of tumor cells, which may essentially affect the pharmacological effects of chrysin in HCC treatment.
Nodes From HCC Targets’ PPI Network (Score ≥0.95).

Gene Ontology (GO) Annotation and KEGG Analysis of Potential Targets for HCC Treatment With Chrysin
To clarify further the biological processes and pathways involved in HCC treatment with chrysin, we used the DAVID database to perform GO annotation analysis on the core potential targets of chrysin intervention in HCC; the three options of biological process, cell composition, and molecular function were selected; and the species was set as “Homo sapiens", with a threshold limited to
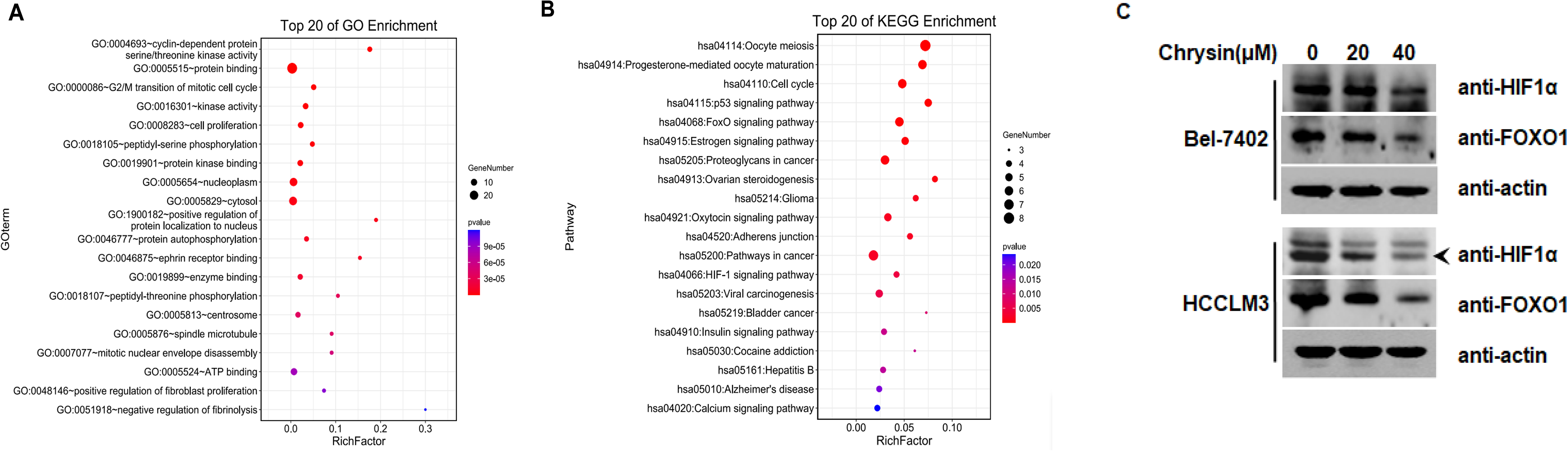
GO annotation and KEGG analysis of potential targets for HCC treatment with chrysin. (A) GO enrichment analysis of 20 hypothetical targets. (B) KEGG pathway enrichment analysis of 20 putative targets. (C) Chrysin inhibits the expression of HIF-1α and FOXO1 in Bel-7402 and HCCLM3 cell lines.
Analysis of Docking Results of Chrysin and HCC Cell Cycle Related Hub Genes
Based on the biological annotation, chrysin functions in regulating the cell cycle; thus chrysin and hub genes (CDK1, CDK5, MMP9) related to the HCC cell cycle were analyzed by molecular docking. It is generally believed that a lower energy will ensue and the binding of two molecules will occur if a ligand binds to a receptor in a stable conformation. The molecular docking results show that the molecular docking affinity of the 3 core genes related to the cell cycle of chrysin is less than −5 kJ/mol (Figure 3); the conformation of interaction of chrysin and the indicated genes is shown in Figure 4, which indicates that chrysin has good binding activity and could interact with CDK1, CDK5, and MMP9 directly, thereby contributing to regulating the HCC cell cycle and proliferation.

Molecular docking conformation of chrysin and hub genes. (A) CDK1. (B) CDK5. (C) MMP9.

Chrysin induced cell cycle arrest in G2/M phase and decreased the levels of CDK2 and CDK4. (A) HCCLM3. (D) Bel-7402. (B, E) Cell cycle phase distribution maps. (C, F) CDK2 and CDK4 expression.
Effect of Chrysin on the Proliferation of HCCLM3 and BEL-7402 Cells
To detect the effect of chrysin on the proliferation of human hepatocellular carcinoma cells (HCCLM3, BEL-7402 cells) and reveal the toxic effect of chrysin on hepatocellular carcinoma cells, the cell counting kit-8 (CCK-8) method was used to determine the different concentrations of chrysin (0, 10, 20, 40, 80, and 160 μmol/L) for the growth of HCCLM3 and BEL-7402 cells at 48 h. As shown in Figure 5 (A, B), chrysin inhibited the growth of HCCLM3 and BEL-7402 cells in a dose-dependent manner. In the HCCLM3 group, the inhibitory percentage of chrysin at 60 μmol/L was 75%, and approximately 50% of that was at 50 μmol/L, indicating that the lethal 50% concentration of chrysin against HCCLM3 cells was 50 μmol/L. In the BEL-7402 experimental group, 40 μmol/L of chrysin inhibited the proliferation of colorectal cancer cell lines by 75%, implying that approximately 30 μmol/L chrysin inhibited cell proliferation by 50%. The LC50 concentration of chrysin on BEL-7402 cells is 30 μmol/L. Therefore, for the subsequent experiments, 10, 20, and 40 μmol/L were selected as the concentrations for chrysin treatment.

The effect of chrysin on the proliferation of HCCLM3 and BEL-7402 cells. The cell viability was detected by the CCK-8 method. Different concentrations of chrysin-treated HCCLM3 (A) and BEL-7402 (B), respectively.
Chrysin Induced Cell Cycle Arrest in the G2/M Phase and Decreased the Levels of CDK2 and CDK4
According to the previous network pharmacology prediction, we found that the biological processes and pathways involved in HCC treatment by chrysin include the cell cycle, which suggests that chrysin may interfere with HCC via the cell cycle process. Therefore, flow cytometry was used to detect the regulation of chrysin on the cell cycle of HCCLM3 and BEL-7402, and similar results were obtained. As shown in Figure 4 (A-D), compared with the control group (DMSO), chrysin increases the proportion of G2/M phase cells in HCCLM3 and BEL-7402 cells in a dose-dependent manner, compared with G1 and S phase cells. CDK2 and CDK4 are involved in the G2/M phase of tumor cell cycle arrest25,26; this was verified in Western blot results, as shown by the downregulated levels of CDK2 and CDK4 after the treatment of BEL-7402 and HCCLM3 cells with chrysin, (Figure 4 E and F), suggesting the mechanism of chrysin-induced cell cycle arrest.
Discussion
Primary liver cancer (eg, HCC) is a malignant tumor with one of the highest incidences and second highest mortality rate worldwide 27 . Treatment of patients at early-stage HCC is mainly via surgical resection and liver transplantation supplemented by chemotherapy. However, for patients who relapse after surgery, develop late metastases, and cannot tolerate surgery, chemotherapy has become the main treatment method 28 , and most of the current improvements in the efficacy of liver cancer are reliant on drug treatment. Existing conventional chemotherapeutic drugs have low selectivity and serious toxic effects, and a single drug efficiency rarely exceeds 25% 29 . Therefore, the search for highly selective anti-liver cancer drugs remains an important research focus in liver cancer prevention and treatment. Chrysin is a natural flavonoid with widely ranging pharmacological activities. Chrysin is found abundantly in propolis and honey, and it inhibits cell proliferation, regulates the cell cycle, and induces apoptosis in various malignant tumors, such as breast, lung, and liver cancer30,31, which indicates the important role of chrysin in HCC treatment. However, the exact molecular mechanism of chrysin in HCC remains unclear. Therefore, the use of network pharmacology methods and experimental verification to clarify fully the possible mechanism of chrysin intervention in HCC provides a basis for further research.
In this study, the candidate targets of chrysin were obtained through the TCMSP database, after which the disease database was used to extract the relevant HCC targets. After the two were mapped, 134 common targets were obtained. On this basis, we constructed a PPI of chrysin targeting the HCC network. The PPI successfully identified the core targets of chrysin intervention in HCC, including SRC, PTPN1, CDK5, CDK1, CALM1, F2, and MMP9. To explore further the specific mechanism of chrysin on HCC, biological annotation and pathway enrichment were performed. These analyses shows that chrysin has a certain effect on protein, ATP synthesis, cell proliferation, cell cycle, and other biological processes; signal pathways such as cell cycle, oocyte meiosis, cancer pathways, and FOXO signaling pathway are relatively important, and they may be important targets and pathways for chrysin intervention in HCC.
In the PPI, CDKs exhibited a strong degree of connectivity. Several studies have shown that CDK family members fundamentally affect tumor cell cycle progression. When either CDKs or CDK inhibitors are absent, tumor cell proliferation stagnates and even death may occur32–34. As one of the important flavonoids, chrysin exhibits extensive cytotoxicity to breast, colon, and prostate cancers35,36, and it can also regulate various types of CDKs by inhibiting their activity. Cyclins cause tumor cell cycle arrest, thereby inhibiting cancer cell proliferation. The downregulation of CDK affects the expression of enzymes, genes, and proteins involved in DNA repair and cell cycle progression, effectively preventing HCC development 34 . Besides CDKs, cell cycle proteins B1 and B2 (CCNB1, CCNB2), and DNA topoisomerase 2α (TOP2A) are also related to chrysin intervention in HCC. Cyclin B1 (CCNB1) significantly contributes to regulating cell cycle progression. Many studies have confirmed that CCNB1 and CCNB2 are expressed in various malignant tumors, and are involved in the growth, differentiation, apoptosis, and metastasis of tumor cells. Studies have found that after inhibiting CCNB1 expression in hepatocellular carcinoma, the proliferation ability of liver cancer cell lines is decreased; cell migration ability, invasion ability, and tumor formation ability is significantly inhibited; and the apoptosis rate is significantly increased 37 . Moreover, the critical role of CCNB1 was also revealed in HCC proliferation. It is reported that CCNB1 was overexpressed in HCC tissues, and inhibition of CCNB1 suppressed the proliferation of SMMC7721 as well as HepG2 cells 38 . This result shows that the knockdown of CCNB1 can significantly inhibit the proliferation of the two HCC cell lines and the cloning ability of tumor cells. TOP2A is a sensitive and specific marker in the late S phase, the transition from G2 to M phase, and the development and regulation of normal cells, and is overexpressed in many human tumors. Wong et al found that TOP2A overexpression in liver cancer was associated with an early onset, short patient survival time, and chemoresistance39,40. Notably, chrysin can increase apoptosis directly and indirectly. Apoptosis proteins BAD and BAX reduce the expression level of antiapoptotic proteins. The expression levels of MCL-1 and BCL2 genes promote cancer cell apoptosis and change the trend of TOP2A and CDC45 gene expression in cancer tissues and affect the therapeutic effect of tumor diseases and prognosis 41 . Thus, chrysin may directly regulate related targets to achieve the effect of HCC treatment.
GO terms and KEGG pathway enrichment analysis is a key step after obtaining the components and disease mapping the targets. The results show that in GO function annotation, chrysin mainly regulates mitosis, cell cycle G2/M phase, cell proliferation, and the proteins orthophosphate and pyruvate orthophosphate dikinase. This shows that chrysin may act on ATP, protein synthesis, and various enzyme activities, affecting cell cycle, proliferation, and apoptosis, and consequently improving HCC. The KEGG pathways were mainly enriched in the cell cycle, oocyte meiosis, cancer pathways, FOXO signaling pathway, p53 signaling pathway, and HIF-1 signaling pathway. Chrysin may interfere with HCC via multiple pathways. Simultaneously, we found that the pathway analysis for chrysin also includes cell cycle and mitosis. It has been reported that mitosis can be inhibited by chrysin, and chrysin regulates the expression levels of cell cycle genes CCNB1, CCNB2, CCND1, and CCND2 to block the tumor cell cycle and inhibit tumor cell growth42,43. GO and KEGG analysis both revealed that the cell cycle is involved in chrysin-mediated HCC treatment, and both biological function and pathway analysis play an important role. Therefore, to verify the results of network pharmacology, we performed flow cytometry and found that the growth cycle of HCCLM3 and BEL-7402 cells was blocked at G2 after chrysin administration. Given that the cyclin-dependent kinases CDK2 and CDK4 play a key role in the G2/M phase of the tumor cell cycle, we also detected the expression of the two kinases through Western blotting; the results showed that chrysin can downregulate the expression of CDK2 and CDK4 in HCCLM3 and BEL-7402 cells. Simultaneously, CCK-8 test results show that chrysin can inhibit the proliferation of HCCLM3 and BEL-7402 cells. In summary, our experimental results verify the network pharmacology prediction, suggesting that chrysin can directly regulate the growth cycle of HCC and inhibit proliferation, which provides a basis for in-depth study of the mechanism of action of chrysin in HCC treatment.
Materials and Methods
Extraction of Potential Chrysin Therapeutic Targets
Extraction of Potential HCC Disease Targets
By mining the target genes of HCC in the TTD, OMIM, and Disgenet databases, and searching the keyword “Hepatocellular Carcinoma", we downloaded the HCC database and extracted the standardized gene list from it. A Venn diagram was used to map the active ingredient targets and HCC targets, and the intersection genes of the two were obtained.
Construction of the Component-target Network
Targets obtained after the mapping of chrysin and HCC are candidate genes of chrysin in HCC treatment. We inputted the “target of drug action-HCC target gene” obtained after the above mapping into the Cytoscape software version 3.8.2 to build a “component-target” relationship network. The entire network was constructed, which demonstrated the connection between the component and the target. Through this network, the mechanism of action of chrysin in HCC treatment was deeply analyzed.
Construction of the PPI
Homo Sapiens was selected as the “Multiple Proteins” in the STRING database, and the genes at the intersection of the active ingredient and HCC targets were introduced into the system. The confidence score > 0.90 was used for retrieval, and the independent nodes in the network were hidden to construct the chrysin-HCC target interaction network.
Biological Function and Pathway Enrichment Analysis
The key targets of chrysin in HCC treatment were inputted into the DAVID online analysis tool for GO function enrichment analysis and KEGG pathway enrichment analysis. The information was sorted and stored in the omicshare (https://omicshare.com/) database. With
Composition-target Molecular Docking
By searching the Pubchem website, the SDF structure file of chrysin was obtained, and the Open Babel version 2.3.2 software was used to convert the SDF files into PDB files retrieved from the Protein Data Bank (http://www.rcsb.org/pdb) database to obtain receptor proteins CDK1 (PDBID: 4YC6), CDK5 (PDBID: 4AU8), and MMP9 (PDBID: 6ESM). The PYMOL version 2.3.4 software was used to perform operations such as removing water and ligands on the receptor protein., AutoDock Vina version 1.1.2 was used for molecular docking of receptor protein and ligand small molecule. A binding energy of less than zero indicates that the ligand and the receptor can bind spontaneously. A binding energy of − 5.0 kJ/mol was selected as the basis for screening of therapeutic targets in chrysin for HCC.
Reagents and Equipment
Chrysin (purity > 98%) was purchased from Yuanye Biotechnology Co., Ltd (Shanghai, China), high-glucose Dulbecco's modified Eagle's medium (DMEM), RPMI Medium 1640, and fetal bovine serum from Gibco Biosciences, phosphate-buffered saline (PBS), porcine trypsin, penicillin, and streptomycin from Hyclone Life Sciences (Bangalore, India), and CCK-8 from Boshi Biotechnology Co., Ltd (Dongguan, China). Propidium iodide (PI) was purchased from Dingguo Changsheng Biotechnology Co., Ltd (Beijing, China), and anti-cyclin-dependent kinase (CDK), anti-CDK2, anti-CDK4 primary antibodies, and horseradish peroxidase-conjugated secondary antibodies from Protein Tech Group (Wuhan, China).
Cell Culture
Human hepatocellular carcinoma cell lines, HCCLM3 and BEL-7402 were purchased from the American Type Culture Collection. Cells were cultured in DMEM and 1640 containing 10% serum and 1% each of penicillin and streptomycin, at 37 °C and 5% CO2 in a saturated humidity incubator.
CCK-8 Detection
Cell proliferation and viability were assessed using a CCK-8 assay. Briefly, cell lines were inoculated into a 96-well plate (5000 cells/well for HCCLM3 and BEL-7402) and incubated at 37 °C with 5% CO2 until the cells adhered (24 h). Cells were then treated with chrysin at varying concentrations (0, 10, 20, 40, 80, and 160 μmol/L) for 48 h, followed by the addition of 10 μL CCK-8 solution per well and incubating for 30 min in the dark. Finally, per-well absorbance (A) was assessed at a wavelength of 450 nm using a microplate reader (Meigu Molecular Instruments Co., Ltd, Shanghai, China).
Flow Cytometry
Cells harvested during the logarithmic growth phase were inoculated into six-well plates (2 × 105 cells/well for HCCLM3 and 2 × 105 cells/well for BEL-7402), which were incubated for 24 h before treatment with either vehicle (control group) or vehicle containing chrysin (20 μmol/L or 40 μmol/L; experimental group) for 48 h. After the designated treatment, the cells were collected by trypsinization (2 × 105), washed twice in cold PBS, and then fixed overnight at −20 °C in 70% ethanol. After adding 1 mL of PI solution to each sample, gently mixing, and incubating for 30 min in the dark, 10,000 stained nuclei were analyzed with a flow cytometer (Beckman Coulter Biotechnology Co. Ltd, Suzhou, China). DNA histograms were prepared using the ModFit analysis program (BD Biosciences).
Western Blot Analysis
Following chrysin treatment, HCCLM3 and BEL-7402 cells were harvested and washed twice with cold PBS. Then, a quantitative RIPA buffer was added to the cell pellet, and the lysate was sonicated and centrifuged at 12,000 rpm for 30 min at 4 °C to obtain the total protein supernatant. The BCA kit was used to estimate the total protein content of the lysate according to the manufacturer's instructions. The lysate was subjected to sodium dodecyl sulfate polyacrylamide gel electrophoresis, and the protein was transferred to a polyvinylidene fluoride membrane. The membrane was blocked with 5% skim milk for 1.5 h, then incubated overnight at 4 °C with CDK2 and β-actin primary antibodies. After incubation with either peroxidase-conjugated goat anti-rabbit IgG or anti-mouse IgG at room temperature for 90 min, the color was developed using the ECL kit (Abcam).
Footnotes
Author Contributions
H.W. and D.L. designed the experiments; J.W., Z.S., L.S. and S.H. performed the experiments; D.L., J.W. contributed to manuscript preparation.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the National Natural Science Foundation of China (Grant No. 82003985, 81973712). Science and technology research project of Jilin Provincial Department of Education (Grant No. JJKH20210995KJ, JJKH20210995KJ), Jilin Province Science and Technology Development Project in China (Grant No. 20200404060YY, 20210204013YY).
