Abstract
To investigate the mechanism of action of components of Yinma Jiedu granules in the treatment of coronavirus disease 2019 (COVID-19) using network pharmacology and molecular docking. The main chemical components of Yinma Jiedu granules were collected in the literature and Traditional Chinese Medicine Systems Pharmacology Database and Analysis Platform database. Using the SwissTargetPrediction database, the targets of the active component were identified and further correlated to the targets of COVID-19 through the GeneCards database. The overlapping targets of Yinma Jiedu granules components and COVID-19 were identified as the research target. Using the Database for Annotation, Visualization and Integrated Discovery database to carry out the target gene function Gene Ontology enrichment and Kyoto Encyclopedia of Genes and Genomes pathway annotation and Cytoscape 3.6.1 software was used to construct a “component-target-pathway” network. The protein-protein interaction network was built using Search Tool for the Retrieval of Interacting Genes/Proteins database. Using Discovery Studio 2016 Client software to study the virtual docking of key protein and active components. One hundred active components were screened from the Yinma Jiedu Granules that involved 67 targets, including mitogen-activated protein kinase 3 (MAPK3), epidermal growth factor receptor, tumor necrosis factor, tumor protein 53, and MAPK1. These targets affected 109 signaling pathways including hypoxia-inducible factor-1, apoptosis, and Toll-like receptor signaling pathways. Molecular docking results showed that the screened active components have a strong binding ability to the key targets. In this study, through network pharmacology and molecular docking, we justified the multicomponent, multitarget, and multipathways of Yinma Jiedu Granules in the treatment of COVID-19.
Keywords
Since December 2019, coronavirus disease 2019 (COVID-19) caused by the SARS-CoV-2 virus broke out in Wuhan, China, and then quickly spread all over the world. SARS-CoV-2 is highly contagious and infectious and belongs to the β genus of coronaviruses, 1 infecting all age groups, but particularly the elderly. The clinical symptoms are mainly fever, dry cough, fatigue, and dyspnea. Severe cases exhibit complications manifested as acute respiratory distress syndrome due to cytokine storm. 2 At present, there is no specific medicine for this disease. In China, we use traditional Chinese medicine to treat COVID-19. Chinese medicine accounted for 91.5% of the total treatment, and the total effective rate of Chinese medicine reached over 90% and Traditional Chinese Medicine (TCM) has shown its unique advantages in being able to participate in the prevention, treatment, and rehabilitation of COVID-19 patients.
Yinma Jiedu Granules are composed of Licorice (
Network pharmacology was systematically explored using a variety of systematic biological databases and software, for the multicomponent, multipathway, multitarget mechanisms of TCM to analyze and explain the mechanism of drug action. High-throughput molecular docking technology simulates the interaction between receptors and drug molecules and is usually used to study the active sites of drugs, thus playing an important role in the study of natural products. This study uses network pharmacology and molecular docking technology to explore the possible mechanism of Yinma Jiedu Granules at the molecular level and to provide new ideas for the treatment strategies of COVID-19.
Materials and Methods
Collection and Screening of Active Components
We searched for all the chemical components of portulaca, licorice, rhubarb, honeysuckle, and plantain in the database of the Traditional Chinese Medicine Systems Pharmacology Database and Analysis Platform (TCMSP) (http://lsp.nwu.edu.cn/tcmsp.php). The relevant literature was downloaded and combined with the chemical structures from PubChem and saved in Spatial Data File (SDF) format. With oral bioavailability (OB) ≥30% and drug likeness (DL) ≥0.18 as the screening thresholds, the active components in Yinma Jiedu Granules were screened. 6
Target Prediction of Active Components
SDF format of the active components was imported into the SwissTargetPrediction database (http://www.swisstargetprediction.ch/), and the targets were collected with a probability value >0.
Screening of COVID-19 Related Targets
Disease-related target genes were collected in the GeneCards database with the keywords of “coronavirus disease 2019,” “new coronavirus pneumonia,” and “new coronavirus 2019.” These were compared with the potential gene targets of the active components using a Venn diagram. Finally, the potential target genes of Yinma Jiedu Granules for the treatment of new coronary pneumonia were identified.
Gene Ontology and Pathway Enrichment
Both the Database for Annotation, Visualization and Integrated Discovery (DAVID, https://david.ncifcrf.gov/) and KEGG Orthology Based Annotation System (KOBAS 3.0) (http://kobas.cbi.pku.edu.cn/anno_iden.php)provide comprehensive biofunctional annotations for a large number of genes. The potential targets were introduced into the DAVID database and KOBAS 3.0 system. Then we selected the identifier as “GENE OFFICIAL SYMBOL” and species as “Homo sapiens” for Gene Ontology (GO) enrichment and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway annotation. All results were screened with
Network Construction
Cytoscape 3.6.1 software (https://cytoscape.org/download.html) was used to construct a component-target-pathway network. Different nodes were used to represent the active components, targets, and pathways. The association between the two nodes is represented by an edge. The 2 groups of networks were merged to obtain the component-target-pathway network. In this, the high-degree components were regarded as important active components. The node degree value was set greater than the median of twice as the standard.
Constructing a Protein-Protein Interaction Network
The interaction relationship between proteins was constructed through the Search Tool for the Retrieval of Interacting Genes/Proteins database (STRING) (https://string-db.org/), imported into Cytoscape 3.6.1 software, and processed using the Network Analysis function.
Molecular Docking
The virtual docking of key proteins and active components was performed using Discovery Studio 2016 software to study their interaction. The structures of related proteins were obtained from the RSCB PDB database (https://www.rcsb.org/) and component structures were obtained from the PubChem database (https://pubchem.ncbi.nlm.nih.gov/). These files were imported into Discovery Studio 2016 for docking. It is generally believed that LiDockscore ≥90 has a strong binding ability when docking with the processed protein. 8
Results
Screening of Active Components
Figure 1 shows a research process of network pharmacology and molecular docking employed for mechanistic investigation of anti-COVID 19 mechanisms of Yinma Jiedu Granules. The five medicinal substances used in the formulation of Yinma Jiedu Granules, namely, purslane, licorice, rhubarb, honeysuckle, and portulaca, were searched through the TCMSP database. With OB ≥30% and DL ≥0.18, 132 active components were selected, of which 9, 13, 16, 84, and 10 were from purslane, rhubarb, honeysuckle, licorice, and portulaca, respectively. After removing duplicates and components lacking target prediction data, 100 active components were further analyzed. The basic information of some of the active components is shown in Table 1. These active components were processed into the SwissTargetPrediction database, and 804 potential targets were identified for Yinma Jiedu Granules.

Research process of network pharmacology and molecular docking for Yinma Jiedu granules. COVID-19, coronavirus disease 2019; C-T-P, component-target-pathway; GO, geneontology; KEGG, Kyoto Encyclopedia of Genes and Genomes; PPI, protein-protein interaction; TCMSP, Traditional Chinese Medicine Systems Pharmacology Database and Analysis Platform.
Potential Active Components of Prescription.

Abbreviations: DL, drug likeness; OB, oral bioavailability; TCM, traditional Chinese medicine.
Screening of Potential Targets
A Venn diagram was used to analyze and compare the targets of 100 potentially effective components with the 262 COVID-19 related genes (Figure 2). The analysis of 67 potential target genes of each active component for the treatment of COVID 19 is shown in Table 2.

Venn diagram of coincidence targets.
Targets That Associate Components With Disease.

Target Biological Function Analysis
The KEGG pathway annotation results showed that 67 potential target genes involve 115 pathways, of which 109 are significantly related to target genes (

Bubble chart of the results of KEGG pathway enrichment analysis of the identified target proteins by Database for Annotation, Visualization and Integrated Discovery. HIF-1, hypoxia-inducible factor-1; KEGG, Kyoto Encyclopedia of Genes and Genomes; TNF, tumor necrosis factor.
GO enrichment analysis includes Cell Component (CC), Biological Process (BP), and Molecular Function (MF). The top ones in BP are as follows: lipopolysaccharide-mediated signaling and extrinsic apoptotic signaling pathway in the absence of ligand (Figure 4(A)). The top most in MF are MAP kinase activity and ATP binding (Figure 4(B)), whereas the top ranking in CC involves cytosol and the external side of the plasma membrane (Figure 4(C)). This shows that Yinma Jiedu Granules play a significant role in the treatment of new coronary pneumonia by participating in the regulation of various biological processes.

The enrichment analysis in biological processes, cellular components, and molecular functions of the identified target proteins by Database for Annotation, Visualization and Integrated Discovery.
Construction of a Component-Target-Pathway Network
The 100 active components, 67 target genes, and the top 10 pathways were analyzed by KEGG pathways to construct a component-target-pathway network diagram (Figure 5). There are 177 nodes and 824 edges. In Figure 5, amaranth represents the pathway, green represents the active components, and purple represents the target. The network diagram shows that Yinma Jiedu Granules treat COVID-19 through coordinated regulation of multiple components, multiple targets, and multiple pathways. By analyzing the degree value of the active components, the component MLS000849829 and baicalein with higher degree values were selected for molecular docking experiments.
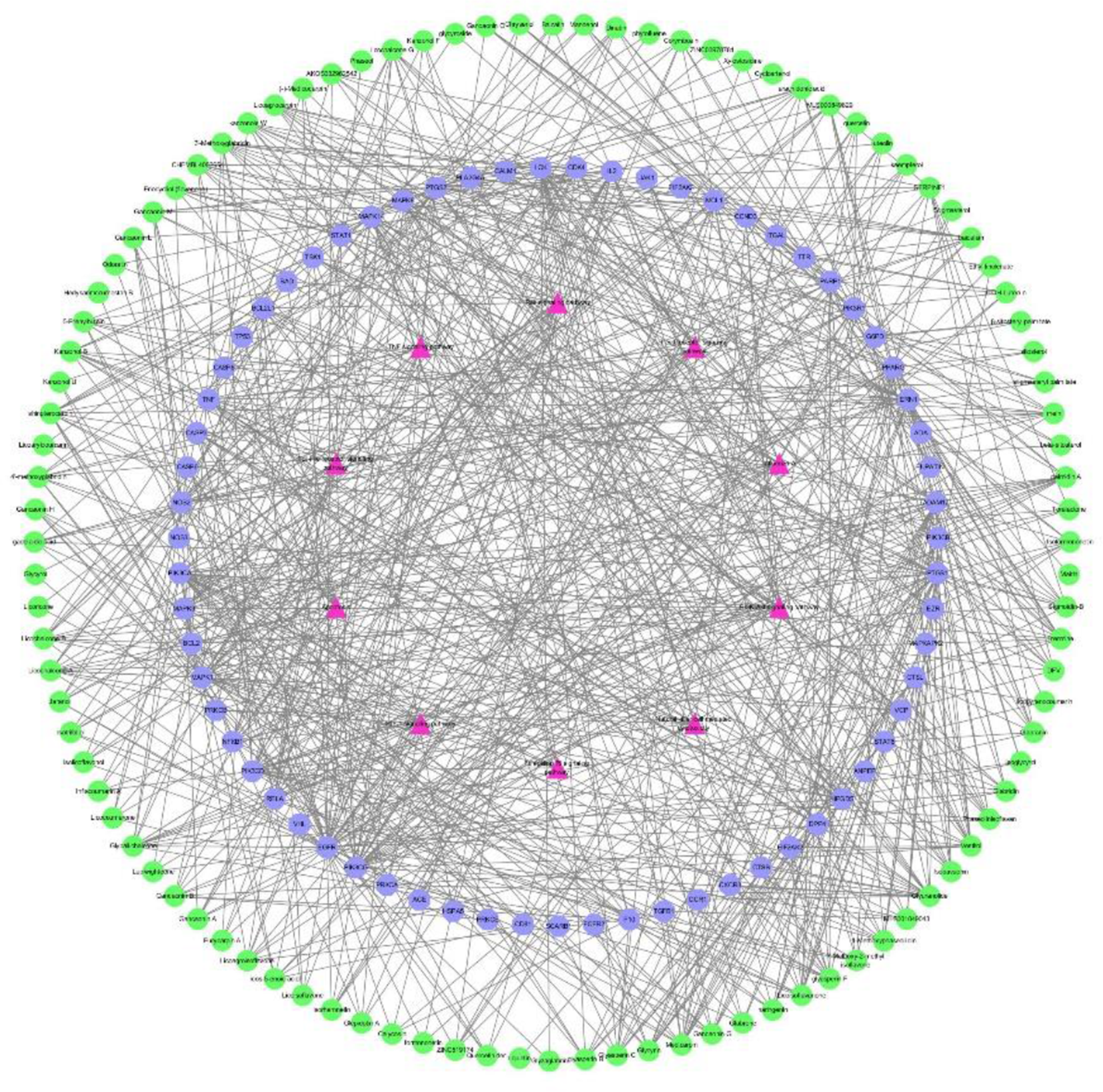
Components-Targets-Pathways network.
Protein-Protein Interarction Network Construction
The target genes of the active component were imported into the STRING database to obtain the interaction network of the corresponding protein (Figure 6(A)). The network was analyzed using the Network Analyse function, and mitogen-activated protein kinase 3 (MAPK3) and epidermal growth factor receptor (EGFR) were selected as the targets for molecular docking verification. It is reported in the literature that the two target proteins of angiotensin-converting enzyme-2 (ACE2) and SARS-COV-2 3 Cl are closely involved in COVID-19 infection. The virus S protein binds to the host cell ACE2 receptor, allowing the virus particles to enter the cells and thus blocking the ACE2 receptor reveals a potential therapeutic target for drug discovery to prevent SARS-CoV-2 transmissibility. Additionally, a coronavirus large-polyproteins-encoded cysteine protease, designated 3-chymotrypsin-like protease 3 Cl, was previously considered as an important target to combat the SARS and MERS coronavirus epidemics. Therefore, these two proteins, along with the active components, were further subjected to molecular docking studies. The protein-protein interaction (PPI) network between ACE2 and the top 20 target proteins is shown in Figure 6(B).

Protein-protein interaction (PPI) interaction network. (A) Components and disease-related targets PPI network. (B) The top 20 targets that interact with the angiotensin-converting enzyme.
Molecular Docking
The protein structure for molecular docking was downloaded from the Protein Data Bank (PDB) database (https://www.rcsb.org/), and Discovery Studio Client 2016 was used for molecular docking. The docking scores of proteins and selected components are shown in Table 3 and the corresponding 3D and 2D views are shown in Figures 7 and 8, respectively. The LiDockScores of baicalein and MLS000849829 had a higher docking score with SARS-COV-2 3 Cl and ACE2 was over 90. In the molecular docking with the protein, a LiDockScore > 90 is considered as effective binding. Thus, these components could be the important potential effective components of this prescription responsible for the therapeutic effects. The results of molecular docking experiments showed that the docking scores of the selected active components and key targets reached effective binding scores. This proved that the predicted active components are likely to bind to the predicted key target proteins, and the network pharmacology prediction results are reliable and accurate.

Results of MLS000849829 molecular docking. (A) MLS000849829-4QTB, (B) MLS000849829-6DUK, (C) MLS000849829-6M2N and (D) MLS000849829-3D0G.
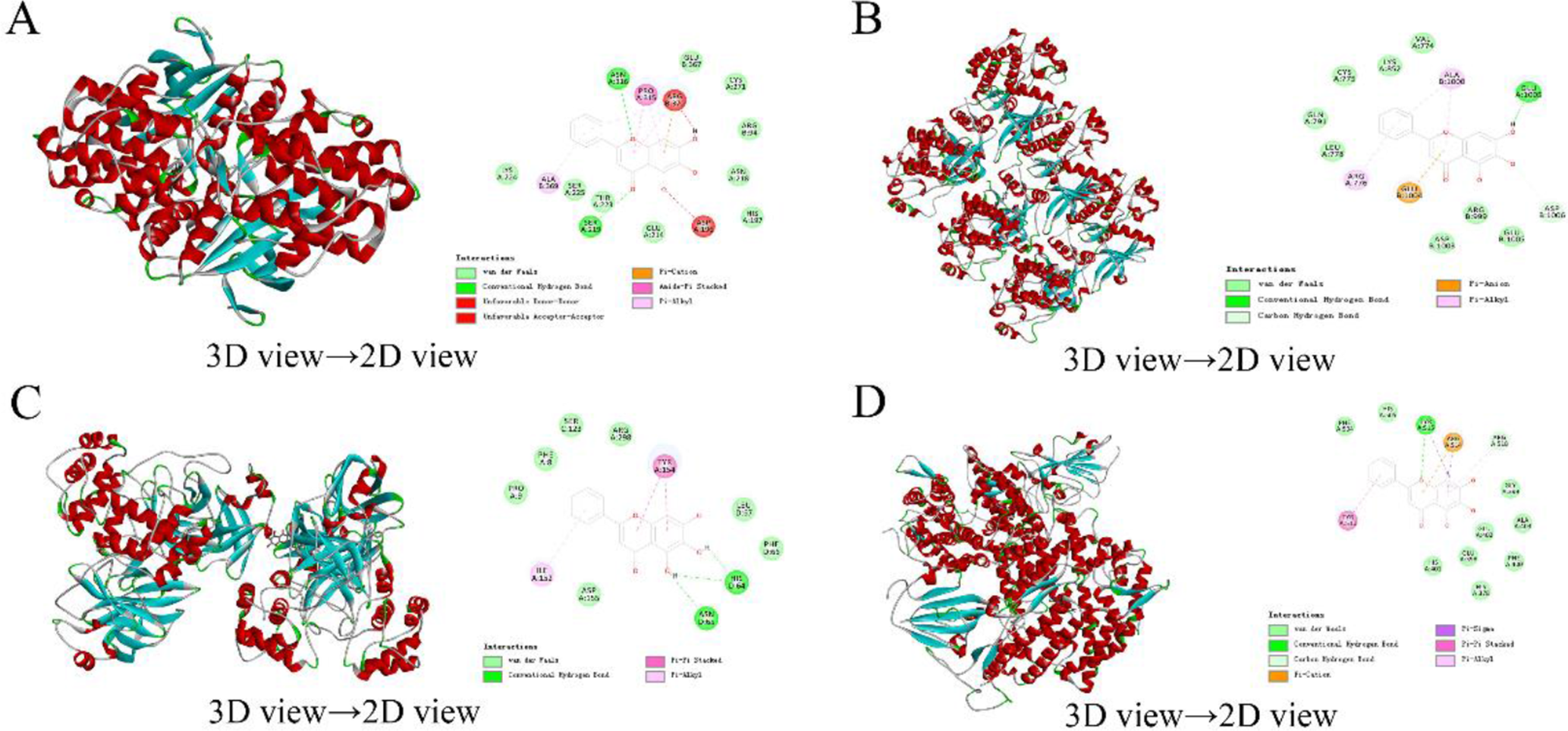
Results of baicalein molecular docking (A) baicalein-4QTB, (B) baicalein-6DUK, (C) baicalein-6M2N, and (D) baicalein-3D0G.
LiDockScore of the Active Components and Target Proteins.

Discussion
Yinma Jiedu Granules are pure Chinese medicine preparations made from 5 different medicinal herbs, licorice, honeysuckle, rhubarb, portulaca, and plantain. In this combination formulation, licorice is the main medicine for the treatment of heat, sores, ulcer, fever, and other syndromes. 9 Several triterpenoids in licorice, including glycyrrhizic acid, glycyrrhizin, glycyrrhetinic acid, and its derivatives, also have potential applicability in the treatment of viral infections. It can be considered as the main medicine in the prescription. Bioactive constituents of honeysuckle, such as rutin, quercetin, and oleanolic acid, have shown significant therapeutic benefits in the treatment of upper respiratory tract infection. 10 Portulaca has shown strong antioxidant and anti-inflammatory effects in a lipopolysaccharide induced lung injury model. 11 Honeysuckle and portulaca can work together to clear heat and detoxify, clear the lungs, and strengthen the efficacy of licorice. Rhubarb is used together with plantain to cause heat toxins to flow out of the body in the form of stools and at the same time to reduce the side effects of licorice, so both are adjuvants. The combination of all the medicines in the whole prescription has the therapeutic effects of clearing away heat and detoxification, reducing swelling and inflammation, relieving cough, and expectorating. 3,4 COVID-19 belongs to the category of “warm epidemic” in TCM. The disease is located in the lungs and the basic pathogenesis is characterized by “dampness, heat, poison, and stasis.” TCM has unique advantages in dialectical treatment. In this study, the network pharmacology method was used to analyze systematically the potential treatment mechanism of Yinma Jiedu Granules on COVID-19.
Through the PPI network analysis, we found that the main targets of Yinma Jiedu Granules in the treatment of COVID-19 were tumor necrosis factor (TNF), tumor protein 53 (TP53), MAPK1, MAPK3, EGFR, cysteine-aspartic acid protease 3 (CASP3), MAPK8, and interleukin 2 (IL2). TNF is the main cytokine that mediates inflammation and innate immunity. Overexpression causes cytokine storm, dysfunction of cytokines, and disrupts the immune balance. 12 TP53 is a cell cycle-related gene, which maintains cell stability by regulating the synthesis of cell cycle-related proteins and induces cell apoptosis, thereby inhibiting tumor growth. 13 MAPK1, MAPK3, and MAPK8 are activated during oxidative stress, DNA damage, cancer development, and viral infections. The MAPK family is composed of extracellular signal-regulated kinase (ERK), p38, and c-JUN N-terminal kinase, which mediate intracellular signals related to cell proliferation, differentiation, survival, death, and transformation. 14,15 Studies have shown that human cytomegalovirus, human herpesvirus type 8, Epstein-Barr virus, HIV, influenza virus, and other viral infections are all related to p38 MAPK. 16,17 EGFR is a member of the type I tyrosine kinase receptor gene family, mainly involved in cell signal transduction. After being activated, it causes cell differentiation, proliferation, infiltration, and angiogenesis. 18 During viral infection, by inhibiting the high expression of CASP3 protein in virus-infected cells, it antagonizes virus-induced cell apoptosis to achieve an antiviral effect. 19,20 IL-2 is secreted by Th1 cells, which can promote the proliferation and differentiation of T lymphocytes, and killing activity. The levels of IL-2 can reflect, to a certain extent, the function of the body’s cellular immunity. 21
In this study, 100 active components of Yinma Jiedu Granules were screened through network pharmacology, and their target action genes were predicted. By intersecting with the targets related to COVID-19, 67 targets were identified for the treatment of new coronary pneumonia. GO enrichment analysis showed that the biological processes were mainly involved lipopolysaccharide-mediated signaling pathway, extrinsic apoptotic signaling pathway lacking in absence of ligand, response to cytokine, and angiogenesis, indicating that these biological processes played a key role in the treatment of COVID-19 by Yinma Jiedu Granules. KEGG enrichment shows that the signal pathways used by Yinma Jiedu Granules to treat COVID-19 include the HIF-1, TLR, and TNF signaling pathways. HIF-1α is a key factor in response to hypoxic stress. It can form different signaling pathways with various upstream and downstream proteins, mediate hypoxia signals, and regulate cells to produce a series of compensatory responses to hypoxia. The role of HIF-1α has already been identified in regulating influenza A virus replication and is considered as a novel therapeutic target for combating viral infection. 22 The HIF-1 signaling pathway can participate in the body’s immune response and regulate neovascularization. 23,24 TLRs are involved in the early interplay of host cells with invading viruses, which regulate viral replication and/or host responses, ultimately impacting viral pathogenesis. At present, TLR1-TLR10 is found in the human body. TLR4, TLR7, and TLR8 receptors can induce the activation of related inflammatory factors, leading to excessive inflammation and severe pneumonia, and TLR7/8 activated downstream transcription factor nuclear factor kappa-light-chain-enhancer of activated B cells has a significant effect on antivirus and induction of lung immune injury. 25 TNF is a key mediator and regulator of the immune response of mammals under healthy biological and disease conditions. It can induce a variety of intracellular signaling pathways, including apoptosis and cell survival, inflammation, and immunity. 26,27 The above results indicate that Yinma Jiedu Granules may be effective in treating COVID-19 by inhibiting inflammation and regulating immunity.
Conclusion
In summary, we screened the active components of TCM Yinma Jiedu Granules and explored and predicted their pharmacological mechanisms for the therapy of COVID-19 through network pharmacology. Network pharmacology and the molecular docking method confirm the “multicomponent, multipathway, multitarget mechanisms” for the therapeutic actions of Yinma Jiedu Granules in the treatment of COVID-19. The present work may provide valuable evidence for further clinical application of Yinma Jiedu Granules for treating COVID-19 and may lay a good theoretical foundation for further experimental verification and facilitate the widespread application of the TCM in treating the serious and life-threatening pulmonary infection caused by SARS-CoV-2.
Footnotes
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was supported by the National Natural Science Foundation of China (No. 31570343).
