Abstract
To elucidate the molecular mechanisms underlying the therapeutic activity of ginsenoside Rg3 (Gs-Rg3) in the context of hepatocellular carcinoma (HCC).
Hepatocellular carcinoma (HCC) is a common malignancy associated with a high metastatic rate, 1,2 as well as increasing incidence. 3 Due to the long initial incubation period, it often becomes clinically apparent only in the middle to late stages of development, 4 resulting in delayed diagnosis. Thus, hepatic cancers typically exhibit a high mortality rate. 5 At present, clinical treatment mainstays include surgery, radiotherapy, and chemotherapy, 6 all of which are associated with significant potential adverse effects, 7,8 and prognosis typically remains poor. Therefore, the identification of chemotherapeutic or radio/chemo-adjuvant drugs with fewer adverse effects and superior prognostic profiles is highly desirable. 9
Ginseng (
This study used network pharmacology to predict protein targets and pathways relevant to the therapeutic impact of Gs-Rg3 in HCC as well as investigate mechanisms underlying the regulatory effect of Gs-Rg3 on HCC cell cycling. Specifically, cell cycle regulation resulting in inhibited proliferation was confirmed using the methyl thiazolyl tetrazolium (MTT) assay and flow cytometry; fluorescence microscopy in conjunction with Western blotting also demonstrated that altered sirtuin 2 (SIRT2)-mediated target acetylation may be responsible for this effect.
Materials and Methods
Identification of Shared Disease and Drug Targets
The chemical structure, molecular weight, and two dimensional structure of Gs-Rg3 were obtained from PubChem (https://pubchem.ncbi.nlm.nih.gov/; Pubchem CID: 9918693). A list of known/predicted Gs-Rg3 protein targets was obtained from the Swiss Target Prediction database (http://www.swisstargetprediction.ch/). Lists of known/predicted proteins either dysregulated during HCC or implicated in its pathogenesis were obtained from GeneCards (https://www.genecards.org/), Online Mendelian Inheritance in Man (OMIM) (https://omim.org/), Drugbank (https://www.drugbank.ca/), and Disgenet (https://www.disgenet.org/) databases using “liver cancer” as the search term. Duplicates were deleted to produce the final disease target list. The drug and disease target lists were compared to identify shared target proteins (duplicates were deleted).
Network Model Construction and Analysis
The list of shared targets was inputted into the STRING database (http://string-db.org/cgi/input.pl), specifying
Functional Annotation and Biological Pathway Enrichment Analysis
The resulting list of the top 30 most highly connected nodes was inputted into the Database for Annotation, Visualization, and Integrated Discovery (https://david.ncifcrf.gov/) for annotation with their Gene Ontology (GO) terms (cellular components, molecular functions, and biological processes) and for batch-conversion of protein to entry IDs. The R Project for Statistical Computing (The R Foundation, Vienna, Austria) in conjunction with the Bioconductor package org.Hs.eg.db was used for genome-wide annotation. The resulting gene lists were used for Kyoto Encyclopedia of Genes and Genomes (KEGG) biological pathway enrichment analysis. Relevant bar charts of the most enriched GO terms and biological pathways were constructed.
Cells and Reagents
The human liver cell line LO2 and human hepatoma cell lines Bel-7402 and HCCLM3 were obtained from Hufeng Biology (China). Dulbecco’s Modified Eagle’s Medium (DMEM), Roswell Park Memorial Institute (RPMI)-1640 medium, combined with penicillin/streptomycin, trypsin, and phosphate-buffered saline (PBS) was purchased from Hyclone (Logan, UT, USA), fetal bovine serum (FBS) from Gibco (Thermo Fisher Scientific, UK), standard Gs-Rg3 from Chengdu Must Bio-Technology Co., Ltd. (Chengdu, China), dimethyl sulfoxide (DMSO) and ethanol from Beijing Chemical Reagent Research Institute Co., Ltd. (Beijing, China), MTT salt, Triton X-100, and bovine serum albumin (BSA) from Solarbio (Beijing, China), propidium iodide (PI) from Beijing Dingguo Changsheng Biotechnology Co., Ltd. (Beijing, China), bicinchoninic acid (BCA) and electrochemiluminescence (ECL) kits from Biyuntian Biotechnology Co., Ltd. (Shanghai, China), paraformaldehyde from Coolaber Science & Technology (Beijing, China), anti-SIRT2, anti-cyclin-dependent kinase (CDK)2, anti-CDK4, anti-cyclin D1 primary antibodies, and horseradish peroxidase (HRP)-conjugated secondary antibodies from Proteintech (Wuhan, China), antibodies for the identification of acetylated H3, H3K18, and H4K16 from PTM BIO (Hangzhou, China), and slide sealant containing 4’,6-diamidino-2-phenylindole (DAPI) from Haoran Bio Technologies Co., Ltd. (Nanchang, China).
Cell Proliferation Assay
The LO2, Bel-7402, and HCCLM3 cell lines were seeded into 96-well plates (5000 cells/well) in RPMI-1640 and DMEM containing 10% FBS. The cells were incubated for 24 hours at 37 °C in an atmosphere containing 5% CO2 prior to addition of Gs-Rg3. The cells were incubated with a range of Gs-Rg3 concentrations (10, 20, 40, 80, 160, and 240 µM dissolved in DMSO) for 48 hours, followed by determination of cell viability using the MTT colorimetric assay. Briefly, MTT-containing culture medium was added to each well, the cells were incubated at room temperature for 4 hours, the supernatants were discarded, 150 µL/well DMSO was added, and absorbance at 490 nm was measured with Multi-Mode Detection Platform (Molecular Devices Company, San Jose, CA). The percentage cell viability was determined and compared between treated and untreated cells.
Cell Cycle Analysis
The cell lines Bel-7402 and HCCLM3 were evenly dispersed and seeded into 6-well plates (1 × 106 cells/well) in RPMI-1640 medium and DMEM containing 10% FBS. The cells were incubated for 24 hours at 37 °C in an atmosphere containing 5% CO2 prior to addition of Gs-Rg3. The cells were incubated with a range of Gs-Rg3 concentrations (0, 40, 80, and 160 µM) for 48 hours, followed by PI colorimetry to determine the proportion of cells in each cell cycle phase. Briefly, the harvested cells were fixed in 70% ice-cold ethanol solution overnight at −20 °C; thereafter, the cells were incubated with PI, and cell cycle changes were detected using a Beckman Coulter flow cytometer (Beckman CytoFLEX; Brea, CA, USA). A cell cycle phase distribution map was generated using GraphPad Prism 7 software (GraphPad Software, Inc., San Diego, CA, USA).
Western Blotting
The cell lines Bel-7402 and HCCLM3 were cultured, exposed to Gs-Rg3, and lysed. The lysate total protein content was estimated using a BCA kit according to the manufacturer’s instructions. Lysates were subjected to sodium dodecyl sulfate polyacrylamide gel electrophoresis, and proteins were transferred to a polyvinylidene difluoride membrane. The membrane was incubated with 1% BSA solution for 1.5 hours at room temperature to block non-specific binding sites, followed by overnight incubation with the primary antibody at 4 °C. The membrane was then rinsed thrice with PBS/deionized water, incubated for 1.5 hours with the secondary antibody at room temperature, and color was developed using an ECL kit.
Fluorescence Microscopy
The cell lines Bel-7402 and HCCLM3 were seeded into 6-well plates (1 × 105 cells/well) in RPMI-1640 medium and DMEM containing 10% FBS. The cells were incubated for 48 hours at 37 °C in an atmosphere containing 5% CO2. The cells were then exposed to Gs-Rg3, harvested, and mounted onto glass slides. The cells were fixed using 4% paraformaldehyde (15 minutes) and permeabilized using 0.3% Triton X-100 (5 minutes), and non-specific binding sites were blocked using 1% BSA (1 hour). After addition of the anti-SIRT2 primary antibody, the cells were incubated at 37 °C for 1 hour, rinsed thrice with PBS, and incubated with goat anti-rabbit IgG/fluorescein isothiocyanate (1:500 dilution) at 37 °C for 1 hour. Subsequently, the cells were sealed onto glass slides using a sealant containing DAPI and subjected to fluorescence microscopy using an inverted fluorescence microscope (Olympus, Tokyo, Japan).
Statistical Analysis
Unless otherwise specified, each experiment was performed at least 3 times. Differences between the Gs-Rg3-treated and untreated groups were analyzed via one-way analysis of variance using the GraphPad Prism 7 software (GraphPad Software, Inc.). All data are expressed as means ± standard deviations.
Results
Shared Disease- and Drug-Target Network Model Construction and Analysis
Upon searching the Swiss Target Prediction database with “ginsenoside Rg3” as a keyword, 105 target genes of Gs-Rg3 were obtained. Meanwhile, using “HCC” as a keyword in the GeneCards, OMIM, Drugbank, and Disgenet databases, 1644 HCC target genes were obtained after removing duplicates. A total of 104 shared drug and disease targets were identified.
The resulting STRING-generated PPI (Figure 1(B)) and Cytoscape-generated network models (Figure 1(A)) are shown. The top 30 most highly represented protein targets (Figure 1(C)) represent the core list of proteins modulated by Gs-Rg3 during HCC.

Network pharmacology of Gs-Rg3 in hepatocellular carcinoma (HCC) treatment. (
Functional Annotation and Biological Pathway Enrichment Analysis
The top 10 enriched cellular components and molecular functions, as well as the top 50 enriched GO terms are presented in Tables 1 -3. Regarding cellular component and molecular function terms, both plasma membrane and cell-cell signaling were closely associated with the HCC cell cycle. Regarding biological processes, multiple terms (eg, negative regulation of multicellular organism growth, positive regulation of extracellular signal-regulated kinase (ERK)1 and ERK2 cascade, positive regulation of endothelial cell proliferation, positive regulation of cell proliferation, negative regulation of apoptotic process, transcription, negative regulation of cell proliferation, positive regulation of angiogenesis, and positive regulation of cell division) suggest that Gs-Rg3 may regulate the HCC cell cycle, proliferation, and apoptosis. The top 20 enriched KEGG biological pathways (Table 4) demonstrate the likely involvement of Gs-Rg3 in regulating HCC-relevant pathways, such as mitogen-activated protein kinase signaling, focal adhesion, measles, and other pathways relevant to cell cycling, proliferation, and apoptosis. Thus, Gs-Rg3 potentially targets multiple signaling pathways relevant to HCC pathogenesis.
Analysis of the Composition of Potential Target Cells in the Treatment of HCC With Gs-Rg3.

Biological Process Analysis of Potential Targets of Gs-Rg3 Therapy for HCC.

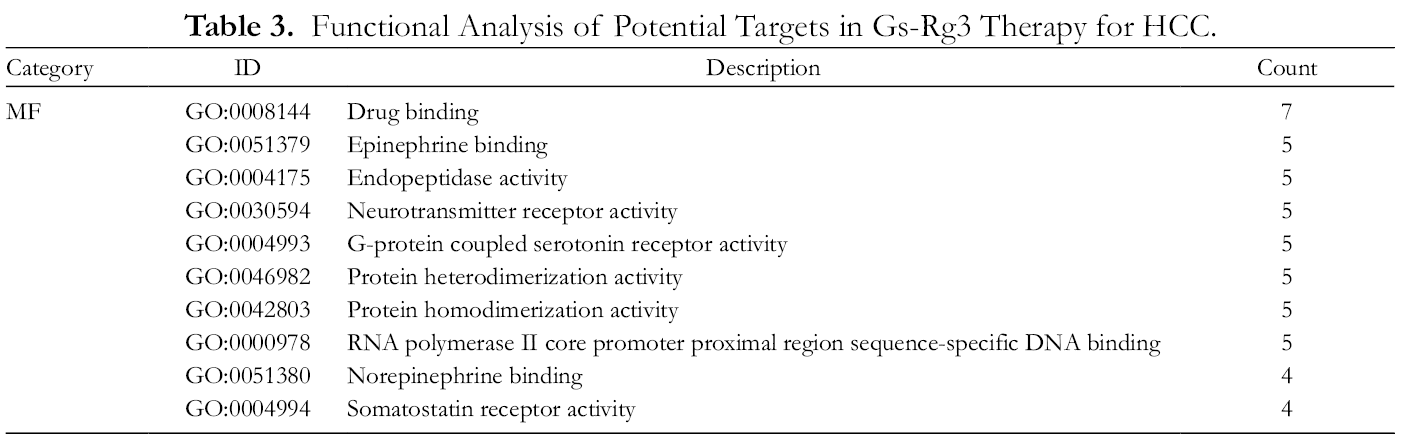
Functional Analysis of Potential Targets in Gs-Rg3 Therapy for HCC.

Analysis of KEGG Pathway of Potential Targets in the Treatment of HCC With Gs-Rg3.

Exposure to Gs-Rg3 Inhibits HCC Cell Line Viability
To verify the inhibitory effect of Gs-Rg3 on the proliferation and viability of human liver cells and HCC, the MTT assay was performed. The results demonstrated that, relative to the blank control, Gs-Rg3 significantly and dose-dependently inhibited the cell proliferation and viability of both Bel-7402 (Figure 2(B)) and HCCLM3 (Figure 2(C)) cells, and the half-maximal inhibitory concentration values were 287.6 µM and 462.1 µM, respectively. Moreover, there was no significant inhibitory effect on the proliferation and viability of the normal liver cell line LO2 (Figure 2(A)), indicating that Gs-Rg3 specifically suppresses liver cancer cell viability.

Gs-Rg3 inhibits hepatocellular carcinoma (HCC) cell viability and proliferation. (
Exposure to Gs-Rg3 Induces G1 Phase Cell Cycle Arrest, Concomitant With Decreased CDK2 and Cyclin D1 Levels
In this study, HCC cells were treated with different concentrations of Gs-Rg3. Flow cytometry analysis showed that Gs-Rg3 increased the ratio of cells in the G1 phase relative to both the G2 and S phases in a dose-dependent manner, indicating that Gs-Rg3 induces G1 phase cell cycle arrest. It has been reported that cyclin D1 is involved in the G1–S phase transition, 16 which is consistent with the result that Gs-Rg3 treatment concomitantly decreased cyclin D1 and CDK2 levels, whereas CDK4 was downregulated in Bel-7402 cells, but only slightly decreased in HCCLM3 cells, by Gs-Rg3 treatment, according to the results of Western blot analysis (Figure 3), suggesting the mechanism by which Gs-Rg3 induces G1 phase cell cycle arrest.
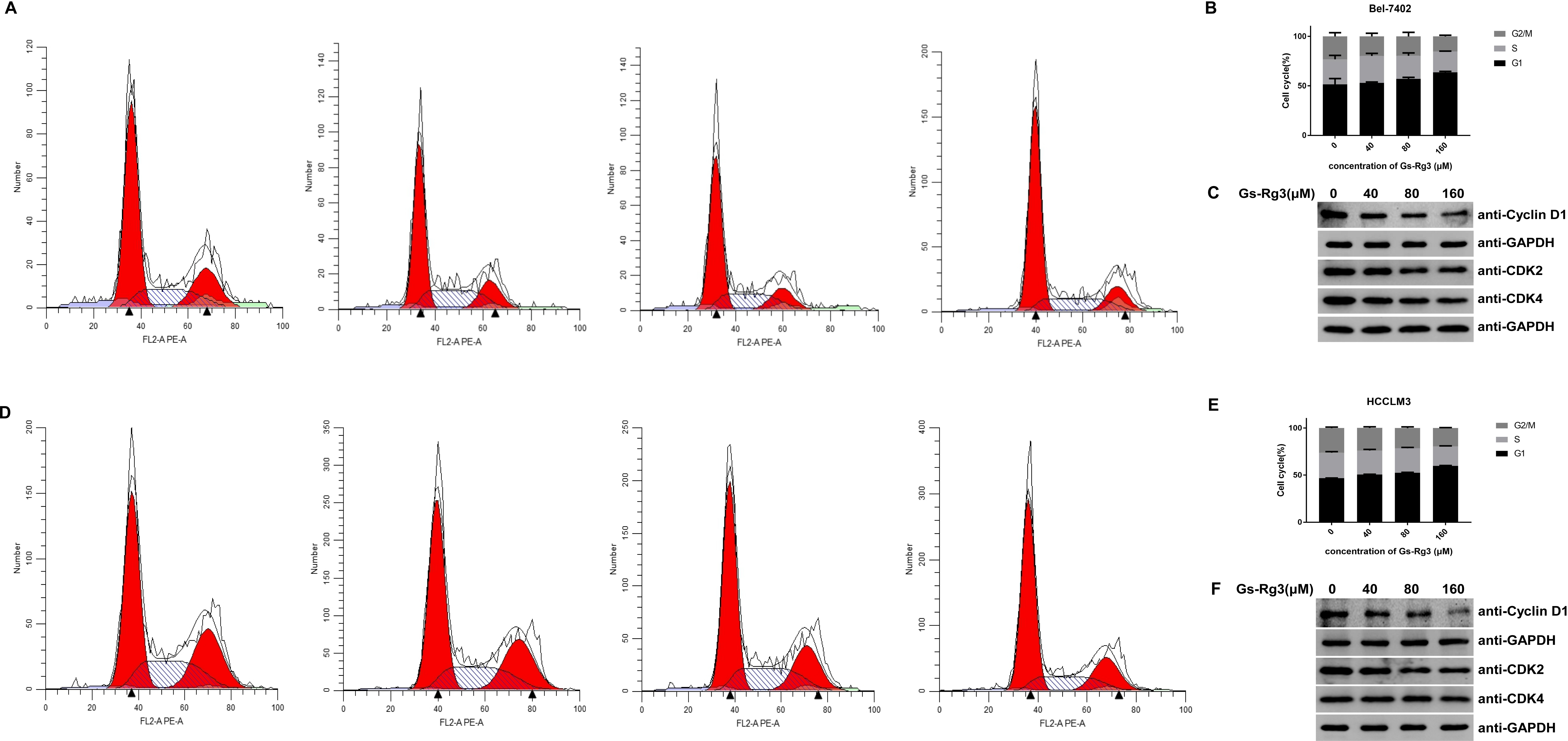
Gs-Rg3 induces G1 phase cell cycle arrest and decreases cyclin D1 levels. (
Exposure to Gs-Rg3 Decreases SIRT2 Levels, Concomitant With Increased Histone Acetylation
It has been reported that G1 phase cell cycle arrest is associated with the inhibition of SIRT2 expression. 17 Fluorescence microscopy demonstrated that Gs-Rg3 decreased SIRT2 levels in a dose-dependent manner (Figure 4(A)), which was confirmed through Western blot analysis. Furthermore, Gs-Rg3-mediated SIRT2 level reductions increased the global cellular acetylation of H3K18 and H4K16 (Figure 4(B) and (C)), indicating that Gs-Rg3 suppresses SIRT2 levels and promotes the acetylation of SIRT2 target sites, thereby regulating the HCC (Bel-7402 and HCCLM3) cell cycle and proliferation.

Gs-Rg3 reduces SIRT2 levels and modulates acetylation of its histone targets. (
Discussion
This study used network pharmacology to identify potential proteins, GO terms, and signal transduction pathways associated with the therapeutic effects of Gs-Rg3 in the context of HCC. Our results indicate that the impact of Gs-Rg3 is likely attributable to multi-target and pathway synergy, including cell cycle modulation. In vitro experiments validated several network pharmacology-derived hypotheses: the MTT assay demonstrated that Gs-Rg3 inhibits the viability and proliferation of 2 types of HCC cell lines, flow cytometry and Western blot analysis demonstrated that Gs-Rg3 induces G1 phase cell cycle arrest and decreases cyclin D1 protein levels, and fluorescence microscopy and Western blot analysis demonstrated that Gs-Rg3 decreases SIRT2 levels and upregulates global cellular H3K18ac and H4K16ac.
It was previously shown that Gs-Rg3 inhibits tumor cell proliferation, induces apoptosis, inhibits cell migration and invasion, and inhibits tumor angiogenesis. 18 Additional studies have demonstrated that a combination of Gs-Rg3 and doxorubicin inhibits late autophagy, concomitant with decreased growth and increased apoptosis of HCC cells, and without enhanced toxicity to normal hepatocytes. 19 Finally, Gs-Rg3 is known to inhibit cancer cell growth via the in vitro and in vivo induction of apoptosis, including ovarian, breast, colon, prostate, and other cancers (including melanoma). 20 -24 The present study found that Gs-Rg3 induces G1 phase cell cycle arrest in HCC cell lines. Other studies have indicated that G1 phase cell cycle arrest is typically associated with a downward trend in cyclin D1 expression. 25 Cyclin D1 can promote the transition from G1 to S by binding and activating CDK2 and CDK4. 26 The present study confirmed this occurrence in response to Gs-Rg3 in 2 types of HCC cell lines. The results suggest that Gs-Rg3 induces G1 phase cell cycle arrest via a decrease in cyclin D1 and CDK2 levels, thereby inhibiting HCC cell proliferation.
SIRT2 belongs to the family of class III histone deacetylases, 27 which play important roles in various biological processes. Alterations to these pathways can increase the risk of HCC and impact HCC patient survival intervals. 28,29 The present study found that nodes exhibiting a high degree of freedom within the shared drug- and disease-target network model during topology analysis were associated with SIRT2, consistent with prior findings. For example, SIRT2 directly inhibits the expression of phosphorylated signal transducer and activator of transcription 3 (STAT3), which inhibits vascular endothelial growth factor A (VEGFA) expression, indicating that the function of SIRT2 in tumor angiogenesis depends on the STAT3-VEGFA signaling pathway. 30 Furthermore, inhibiting SIRT2 expression via the RAS/ERK/c-Jun N-terminal kinase/matrix metalloproteinase-9 pathway promotes migration and invasion in the context of gastric cancer. 31 Given the network pharmacology predictions and existing literature, the present study examined the impact of Gs-Rg3 on SIRT2 expression in HCC cell lines and confirmed that it inhibits SIRT2 expression. In addition, since SIRT2 catalyzes the acetylation of histones H3 (at K18/K56) and H4 (at K16), 32 this study investigated the impact of Gs-Rg3 on global cellular histone modifications H3K18ac and H4K16ac, demonstrating an increase in the levels of these modifications, concomitant with decreased SIRT2 expression. This observation suggests that decreased SIRT2 expression may be responsible for altered histone acetylation profiles and thus altered gene expression. Furthermore, it has been reported that in colon cancer cells, SIRT2 functions in the G1-S phase transition, 33 which implies that suppressed SIRT2 levels induced by Gs-Rg3 might contribute to G1 phase arrest. Thus, detection of the Gs-Rg3 pharmacological activity of suppress the proliferation and migration of HCC in vivo could be further performed. Moreover, the substrate of SIRT2, the acetylation level of which is regulated by Gs-Rg3, could be identified in the future.
Conclusions
Network pharmacology indicated that Gs-Rg3 impacts HCC via its action on multiple targets (such as SIRT2) and pathways (such as cell cycle). These hypotheses were empirically validated by confirming that Gs-Rg3 inhibits HCC cell line proliferation (likely via decreased cyclin D1 and CDK2 levels, thereby inducing G1 phase cell cycle arrest) and decreases SIRT2 levels, concomitant with altered global cellular histone acetylation levels (which likely alter gene expression). This study provides a theoretical basis for further elucidation of the potential therapeutic mechanisms of Gs-Rg3 in the context of HCC.
Footnotes
Acknowledgments
We would like to thank Editage (www.editage.cn) for English language editing.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was funded in part by the National Natural Science Foundation of China (grant no. 81903876) and the Jilin Province Science and Technology Development Project in China (grant nos. 20170309005YY and 20200504005YY).
