Abstract
Osteoclasts are large, multinucleated, bone-absorbing cells and play a crucial role in osteolytic bone diseases such as osteopetrosis and rheumatoid arthritis. Therefore, controlling osteoclast differentiation and activation has been considered a promising strategy to prevent and treat osteolytic diseases. In this study, we demonstrate, using the mouse monocyte-derived macrophage-like cell line RAW 264, that extract from Nasturtium officinale or watercress, an herb of European origin, suppresses receptor activator of nuclear factor-κB ligand-induced osteoclast differentiation in vitro. N. officinale extract decreased the emergence of tartrate-resistant acid phosphatase-positive differentiated multinuclear cells and inhibited their bone-absorbing activity. The extract decreased expression of genes associated with osteoclast differentiation and function. Induction of nuclear factor of activated T cells c1 (NFATc1), the master transcriptional regulator of osteoclastogenesis, was blunted by N. officinale extract. Activation of nuclear factor-κB and mitogen-activated protein kinases pathways, both of which are necessary for NFATc1 induction and osteoclast differentiation, was also suppressed by the extract. Among upstream kinases, activity of IκB-kinase β (IKKβ), but not that of TGFβ-activated kinase 1, was inhibited by N. officinale extract in vitro. Pharmacological inhibition of IKKβ by a specific inhibitor PS1145 in RAW 264 cells mostly recaptured the inhibitory action of N. officinale extract. These findings provide a novel pharmacological action of N. officinale and its potential usefulness for the prevention of osteoporosis.
Bone remodeling is an important biological process to maintain normal bone mass and calcium metabolism throughout life. 1 Bone remodeling results from osteoblastic bone formation and osteoclastic bone resorption; normal bone mass requires a balance of these processes. 2 In bone disorders such as osteoporosis and rheumatoid arthritis, bone resorption exceeds bone formation. 3,4 A major cause of osteoporosis is excess activation of osteoclasts due to the postmenopausal decrease in estrogen production. 5 Patients affected by excessive bone resorption suffer from bone fractures, pain, and impaired mobility. Therefore, controlling osteoclast differentiation and activation is considered a promising target for preventive and therapeutic strategies in osteolytic diseases.
Osteoclasts are bone-resorbing, multinucleated giant cells formed by the fusion of hematopoietic monocyte/macrophage-derived cells. Osteoclast differentiation is regulated by 2 essential cytokines, macrophage colony-stimulating factor (M-CSF) and receptor activator of nuclear factor-κB ligand (RANKL). 1 M-CSF supports the survival and proliferation of osteoclast precursor cells, whereas RANKL triggers differentiation and maturation of osteoclasts. 6,7 RANKL is a member of the tumor necrosis factor superfamily and is expressed on osteoblasts and bone marrow stromal cells. Upon RANKL binding to its receptor, RANK, which is expressed on osteoclast precursor cells, intracellular signaling is transduced by recruitment of adaptor molecules such as tumor necrosis factor receptor-associated factor 6 (TRAF6). 8 This then activates multiple signal transducing kinases including TGFβ-activated kinase 1 (TAK1) and IκB kinase (IKK). 9 Subsequent activation of downstream nuclear factor-κB (NF-κB) and mitogen-activated protein kinases (MAPKs) (extracellular signal-regulated kinase, ERK; c-Jun N-terminal kinase, JNK; and p38) 9 culminates in the induction of nuclear factor of activated T cells c1 (NFATc1), 10 the master transcriptional regulator of osteoclast differentiation. 11 Despite these findings, the mechanism of how RANKL activates NFATc1 is not fully understood.
Plant-derived bioactive metabolites are a treasure-trove of organic compounds, which have various chemical structures and biological activities. Elucidation of their biological activities and identification of their cellular targets will lead not only to discovery of novel components, but also understanding of previously unknown biochemical mechanisms. An herb of European origin, Nasturtium officinale or watercress, belongs to the Brassicaceae family. This plant has long been used in traditional medicine in, for example, Iran and Azerbaijan. 12 It has also been experimentally proven that N. officinale has various pharmacological activities including anti-carcinogenic, anti-oxidative, anti-bacterial, anti-inflammatory, anti-psoriatic, and cardioprotective actions. 12
To investigate potential novel pharmacological actions of N. officinale, we examined whether the extract of this herb could affect osteoclastogenesis. Using the RANKL-stimulated mouse monocyte-derived macrophage-like cell line RAW 264, a widely used in vitro model of osteoclastogenesis, we demonstrate that N. officinale extract suppresses NFATc1 induction and osteoclast differentiation by inhibiting kinase activity of IKKβ.
Results
N. Officinale Extract Inhibits Osteoclastogenesis in Vitro
To examine the effect of N. officinale extract on osteoclastogenesis, cultured RAW 264 cells were stimulated to differentiate into osteoclasts with RANKL in the presence of an increasing concentration of N. officinale extract or vehicle (DMSO). Five days after RANKL stimulation, osteoclast differentiation was evaluated by tartrate-resistant acid phosphatase (TRAP) activity staining and cell morphology. As previously reported, RANKL stimulation induced emergence of TRAP-positive multinucleated cells. We found that treatment of the cells with N. officinale extract reduced the number of TRAP-positive multinucleated cells in a dose-dependent manner (Figure 1(A) and (B)).
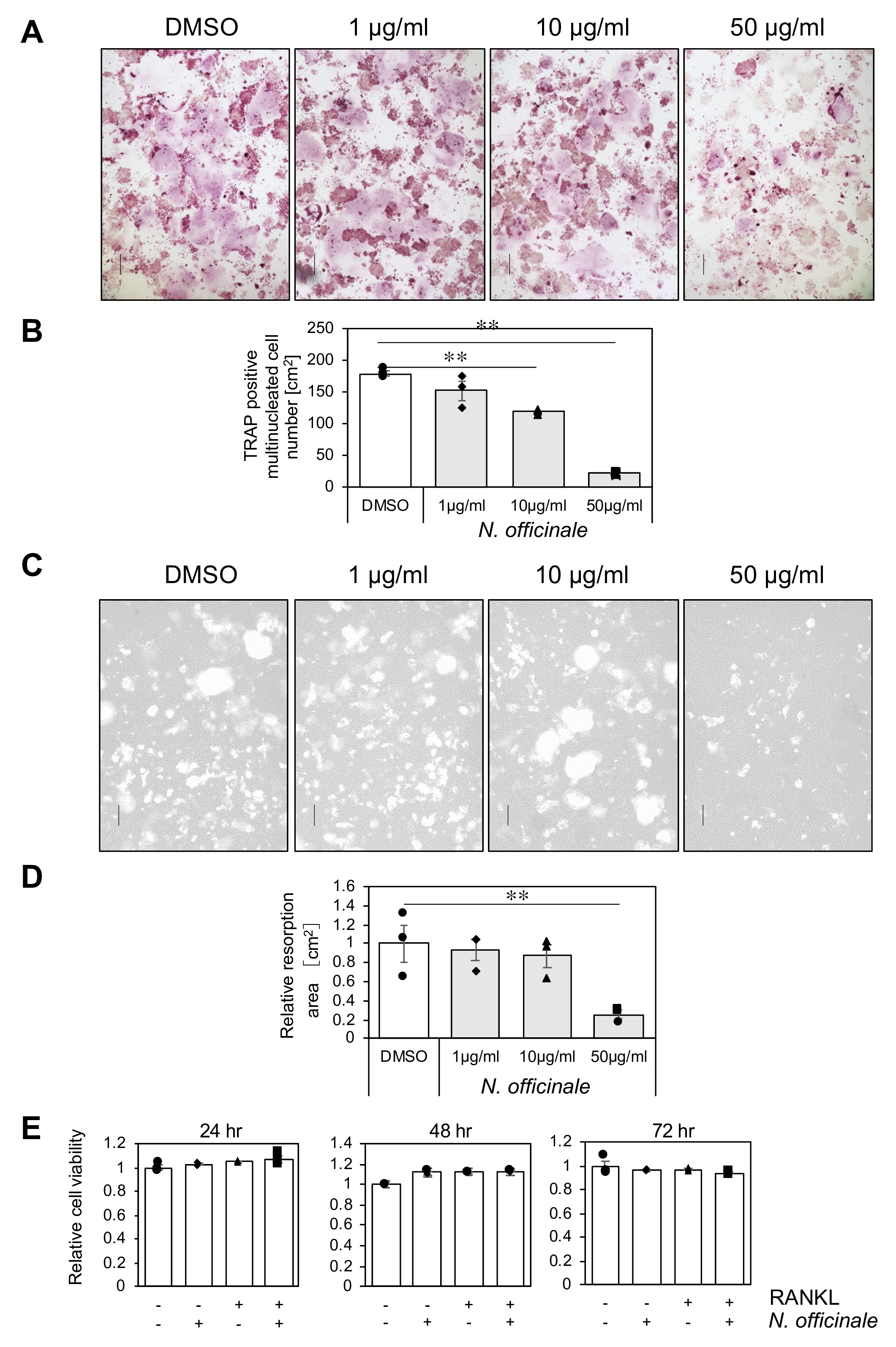
N. officinale extract inhibits osteoclastogenesis in vitro. (
We also assessed whether N. officinale extract affects osteoclast function with a bone resorption assay. RAW 264 cells seeded on the bone biomimetic synthetic surface were treated with RANKL in the presence of N. officinale extract for 5 days, and bone resorption activity of the cells was measured. The absorbed pit area was reduced by treatment with N. officinale extract in a dose-dependent manner (Figure 1(C) and (D)). We also demonstrated that N. officinale extract treatment did not affect osteoclast precursor cell viability (Figure 1(E)). These results indicate that N. officinale extract inhibited osteoclast differentiation without exhibiting cytotoxicity to the precursor cells.
N. Officinale Extract Suppresses RANKL-Induced NFATc1 and Osteoclast-Specific Gene Expression
The observation that N. officinale extract suppressed RANKL-induced osteoclastogenesis prompted us to examine whether the extract affects osteoclast-specific gene expression. RAW 264 cells were stimulated with RANKL in the absence or presence of N. officinale extract for 3 days, and mRNA expression of osteoclast-specific genes was examined by quantitative RT-PCR analysis. In accordance with our finding that the extract inhibited RANKL-induced emergence of TRAP-positive cells (Figure 1(A)), induction of trap mRNA was suppressed by N. officinale extract treatment (Figure 2(A)). We also examined genes encoding proteins associated with osteoclast function (dc-stamp, atp6v0d2, and cathepsin K), which are regulators of cell fusion, proton pump regulating acidification of extracellular space, and secreted cysteine protease, respectively. RANKL-induced expression of dc-stamp and atp6v0d2 was significantly decreased by treatment with N. officinale extract (Figure 2(B–D)).

N. officinale extract inhibits RANKL-induced expression of osteoclastic genes. (
NFATc1 is a transcriptional regulator of these osteoclastic genes 8 and is the master regulator of osteoclastogenesis. We thus examined expression levels of nfatc1 and found that N. officinale extract significantly inhibited RANKL-induced nfatc1 mRNA expression (Figure 3(A)). Expression of the NFATc1 protein was also compromised by N. officinale extract treatment in a dose-dependent manner (Figure 3(B)), whereas expression of RANK was not affected. These findings indicate that N. officinale extract inhibited osteoclastic differentiation and gene expression by suppressing NFATc1 induction.

N. officinale extract inhibits RANKL-induction of NFATc1. (
N. Officinale Extract Suppresses NF-κB and MAPK Activation by RANKL Stimulation
Previous reports have shown that RANKL-triggered induction of NFATc1 requires preceding activation of NF-κB and MAPK signaling pathways. 13 -15 We thus evaluated whether N. officinale extract affects these pathways. RAW 264 cells were preincubated with N. officinale extract and then stimulated by RANKL for 15, 30, and 60 minutes. Nuclear extracts were prepared and analyzed by electrophoretic mobility shift assay (EMSA) using oligonucleotide probe containing κB-binding site (Figure 4(A)) or Oct1-binding site as a control (Figure 4(B)). RANKL quickly and transiently stimulated NF-κB DNA-binding activity, which peaked at 15 minutes. On the other hand, N. officinale extract suppressed the magnitude of NF-κB induction and delayed its peak to 30 minutes after RANKL stimulation.

Inhibition of the canonical NF-κB pathway and MAPK pathways by N. officinale extract. (
NF-κB is a heterodimeric protein complex that is formed by RelA and p50 (canonical complex) or RelB and p52 (non-canonical complex), both of which are potentially stimulated by RANKL through distinct signaling pathways. 1 To determine the component of RANKL-stimulated NF-κB in our assay system, nuclear extract of RAW 264 cells that had been stimulated by RANKL for 15 minutes were analyzed by EMSA in the presence of anti-RelA or anti-RelB antibodies (Figure 4(C)). The κB-binding complex was supershifted by addition of anti-RelA antibody but not by anti-RelB antibody. This indicates that the canonical RelA-p50 complex of NF-κB was activated in RANKL-treated RAW 264 cells.
The canonical NF-κB complex is tethered in the cytoplasm in complex with IκBα. Upon RANKL stimulation, IκBα is phosphorylated by the trimeric IκB-kinase (IKK) complex (IKKα, IKKβ, and NEMO) and degraded by Ubiquitin-proteasome system to release RelA-p50 into the nucleus. We therefore examined the amount and phosphorylation status of IκBα (Figure 4(D)). Upon RANKL stimulation, the IκBα protein is rapidly and transiently phosphorylated (5 minutes) and degraded (15 minutes), and then re-induced at later time points (60 minutes). By contrast, in the presence of N. officinale extract, induction of phosphorylation levels of IκBα were lower, and its protein levels were not significantly reduced upon RANKL stimulation. These findings indicate that N. officinale extract perturbed the canonical NF-κB activation pathway.
We also analyzed MAPK signaling in the induction of NFATc1. RAW 264 cells preincubated with N. officinale extract were stimulated with RANKL for 5, 15, and 30 minutes, and total cell extracts were analyzed by immunoblotting (Figure 4(E)). All 3 MAPK components, ERK, JNK, and p38, were phosphorylated by RANKL stimulation as reported, and N. officinale extract treatment reduced the activating phosphorylation of these kinases. These results indicated that N. officinale extract inhibited NFATc1 expression by attenuating activation of canonical NF-κB and ERK, JNK, and p38 MAP kinases.
Kinase Activity of IKKβ Is Inhibited by N. Officinale Extract
Previous reports have demonstrated that, upon RANKL binding to its receptor RANK, the adaptor molecule TRAF6 is recruited to RANK, which triggers activation of TAK1 kinase. Subsequently, activated TAK1 phosphorylates and activates IKKβ, the predominant catalytic component of trimeric IKK complex, 16 and MAPKs. We therefore analyzed whether N. officinale extract inhibits these signaling events. First, recruitment of TRAF6 to RANK was evaluated by GST-pull down assay using GST-tagged intracellular domain of RANK and FLAG-tagged TRAF6 expressed in HEK293T cells. We found that N. officinale extract had no effect on interaction of RANK and TRAF6 (Figure 5(A)).

Inhibition of IKKβ by N. officinale extract. (
We next examined the effect of N. officinale extract on kinase activities of TAK1 and IKKβ using in vitro kinase assay. N. officinale extract did not affect kinase activity of TAK1 (Figure 5(B)), whereas it inhibited kinase activity of IKKβ (Figure 5(C)).
To determine whether IKKβ inhibition by N. officinale extract is related to its ability to attenuate osteoclastogenesis, the extract was divided into 5 tractions (fraction 1 to 5) using HPLC (Figure 6(A)). Each fraction was concentrated, resuspended into DMSO, and assayed for IKKβ inhibition in vitro at 10 µg/mL concentration (Figure 6(B)). IKKβ inhibitory activity was mainly detected in fractions 4 and 5, and marginally in fraction 3. By contrast, fractions 1 and 2 had no inhibitory activity. The fractions were then assayed for their ability to attenuate osteoclastogenesis of RANKL-stimulated RAW 264 cells (Figure 6(C) and (D)). Emergence of TRAP-positive multinucleated cells were reduced by addition of fractions 4 and 5 and to a lesser extent by fraction 3, whereas fractions 1 and 2 had no effect. Therefore, the ability of the fractions to inhibit IKKβ is well correlated with their ability to inhibit osteoclast differentiation. This suggests that chemical component(s) in N. officinale extract that inhibits IKKβ is responsible for the ability of the extract to inhibit osteoclast differentiation.
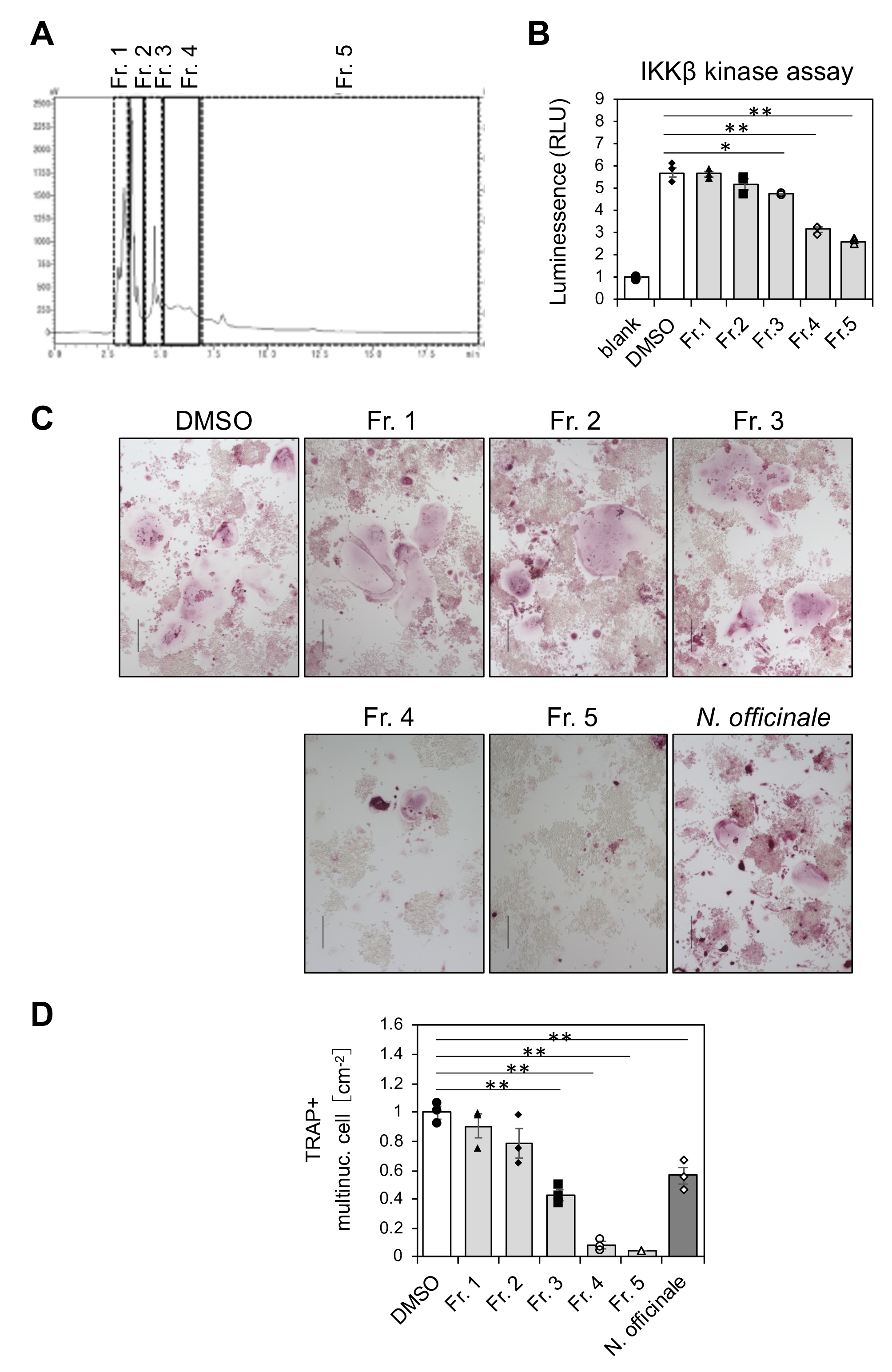
Analysis of N. officinale extract by fractionation. (
Selective Inhibitor of IKKβ Recaptures the Action of N. Officinale Extract
The finding that N. officinale extract has ability to inhibit IKKβ prompted us to examine whether pharmacological inhibition of IKKβ could recapture effects of N. officinale extract on osteoclastogenesis and RANKL-RANK signaling events. PS1145, a potent and selective inhibitor of IKKβ, effectively inhibited its kinase activity in vitro (Figure 7(A)). Treatment of RANKL-stimulated RAW 264 cells with PS1145 resulted in inhibition of osteoclast differentiation in a dose-dependent manner (Figure 7(B) and (C)). Expectedly, PS1145 inhibited RANKL-induced phosphorylation of IκBα in RAW 264 cells (Figure 7(D)). PS1145 treatment also inhibited phosphorylation of ERK and JNK, but not p38 MAP kinase (Figure 7(D)). These results indicated that IKKβ activity is required for not only IκBα phosphorylation but also ERK and JNK activation by RANKL. Altogether, IKKβ inhibitor PS1145 recaptures action of N. officinale extract, namely, inhibition of RANKL stimulated phosphorylation of IκBα, ERK and JNK (but not that of p38), and osteoclast differentiation of RAW 264 cells (Figure 7(E)).

Effect of IKKβ inhibitor PS1145 on RANKL-stimulated osteoclastogenesis and signaling events. (
Discussion
In this study, we demonstrate a novel pharmacological action of N. officinale extract, that is, attenuation of osteoclastogenesis in vitro. Gene expression profiles suggested that the extract suppressed RANKL-stimulated osteoclastic gene induction by inhibiting NFATc1 upregulation. NF-κB and MAPK pathways, the 2 essential components of NFATc1 induction by RANKL–RANK signaling, were attenuated by N. officinale extract. In a search to identify molecular target of N. officinale extract that is responsible for the attenuation of osteoclast differentiation, we found that N. officinale extract directly inhibits kinase activity of IKKβ in vitro. We also showed that PS1145, a selective inhibitor of IKKβ, mostly mimicked the action of N. officinale extract during RANKL-induced osteoclastogenesis, namely, inhibition of phosphorylation of IκBα, ERK, and JNK. By contrast, PS1145 did not inhibit p38 MAPK as N. officinale extract did. A previous study has also shown that PS1145 treatment inhibited osteoclastogenesis of human osteoclast progenitor cells in vitro. 17 These findings together suggest that N. officinale extract inhibits osteoclast differentiation through attenuating IKKβ activity (Figure 7(E)).
The wide variety of pharmacological actions of N. officinale are due to its rich chemical composition, which includes glucosinolates, isothiocyanates, polyphenols, vitamins, and carotenoids. For example, polyphenols and vitamins have an anti-oxidative capacity, 12 and isothiocyanates exhibit cancer-preventing and anti-allergic activities. 18,19 It has been reported that phenethyl isothiocyanate suppresses osteoclast differentiation of RAW 264 cells. 20 However, our analysis of the chemical composition of N. officinale extract revealed that the extract used in this study contained phenethyl glucosinolate but not phenethyl isothiocyanate (Y. T., unpublished observations). We also observed that the phenethyl glucosinolate isolated from the extract did not suppress osteoclast differentiation of RAW 264 cells (Y. T., unpublished observations). Hence, the active ingredient(s) in the extract contained in our HPLC fractions 3, 4, and 5 responsible for inhibiting IKKβ and suppressing osteoclastic differentiation remains to be identified.
Since osteoporosis is a disease with few subjective symptoms, there are many cases in which the disease condition worsens without symptoms being noticed until a bone fracture occurs. 21 Therefore, preventive medicine is important, and recent clinical studies have shown that functional foods and supplements have complementary medical effects. 22 The findings in this study suggest that N. officinale is potentially useful in preventing osteoporosis. Further exploration of the active ingredient(s) in the extract is necessary to understand the precise mechanism of anti-osteoclastogenic activity of N. officinale.
Experimental
Preparation of N. Officinale Extract
The above-ground parts of N. officinale dry powder (Bulgaria) were extracted with 50% ethanol. The crude extract was filtered (5 µm), lyophilized, and resuspended in dimethyl sulfoxide (DMSO). The extraction ratio from dry powder of N. officinale were about 10 to 15%.
Cell Culture
A monocyte-derived macrophage-like cell line, RAW 264, was purchased from RIKEN cell bank (RCB0535). The cells were maintained in Eagle’s minimum essential medium (MEM) (Sigma-Aldrich, MO, USA) supplemented with 10% (v/v) heat-inactivated fetal bovine serum (FBS) (Thermo Fisher Scientific, MA, USA), 2 mM L-glutamine, non-essential amino acids, and penicillin/streptomycin (FUJIFILM Wako Pure Chemical Corporation, Osaka, JAPAN) in a humidified atmosphere containing 5% CO2 at 37 °C.
In Vitro Osteoclastogenesis and TRAP Staining
RAW 264 cells (1 × 10⁴) were seeded into 24-well plates in MEM supplemented with 10% FBS containing 5 nM RANKL (Orientalbio, Tokyo, JAPAN). N. officinale extract or vehicle (DMSO) was also added when indicated. After 4‐5 days, cells were fixed with 5% paraformaldehyde in PBS at room temperature for 15 minutes and permeabilized by incubation with acetone-ethanol (1:1 v/v) at room temperature for 1 minute. The cells were then stained for acid phosphatase by incubating with tartrate-resistant acid phosphatase (TRAP) staining solution (0.01% Naphthol AS-MX phosphate [Sigma-Aldrich], 0.06% Fast red violet LB salt [Sigma-Aldrich], 50 mM sodium tartrate, 90 mM sodium acetate [pH 5.0]) and were rinsed twice with PBS. The number of TRAP-positive multinucleated (>3 nuclei) cells were counted as osteoclasts. Cell viability was assessed using the Cell Counting Kit-8 (Dojindo Molecular Technologies, Kumamoto, JAPAN), according to the manufacturer’s instruction.
Bone Resorption Assay
RAW 264 cells (1 × 10⁴) were seeded in a 24-well Osteo Assay Plate (Corning, NY, USA). The cells were cultured with or without N. officinale extract in the presence of 5 nM RANKL for 5‐6 days. Then, the cells were removed by adding 10% sodium hypochlorite treatment for 5 minutes, and the plate was dried. The pit area was observed under a light microscope (magnification, 20×; KYENCE, Osaka, JAPAN) and measured using ImageJ software.
Quantitative RT-PCR Analysis
Total RNA was extracted using the ReliaPrep RNA Cell Miniprep System (Promega, WI, USA), and first-strand cDNA was produced by the High-Capacity cDNA Reverse Transcription Kit (Thermo Fisher Scientific). mRNA encoding trap, dendritic cell-specific transmembrane protein (dc-stamp), ATPase, H + transporting, lysosomal 38 kDa, V0 subunit d2 (atp6v0d2), cathepsin K, and nfatc1 and gapdh was quantified using SYBR green and the CFX96 Real-Time System (Bio-Rad, CA, USA). Primer sets used were as follows: trap 5’-ACTTCCCCAGCCCTTACTACCG-3’ and 5’-TCAGCACATAGCCCACACCG-3’, dc-stamp 5’-TGTATCGGCTCATTCTCTCCAT-3’ and 5’-GACTCCTTGGGTTCCTTGCTT-3’, atp6v0d2 5’-GAAGCTGTCAACATTGCAGA-3’ and 5’-TCACCGTGATCCTTGCAGAAT-3’; cathepsin K 5’-GCCAGGATGAAAGTTGTATG-3’ and 5’-CAGGCGTTGTTCTTATTCC-3’, nfatc1 5’-GGTAACTCTGTCTTTCTAACCTTAAGCTC-3’ and 5’-GTGATGACCCCAGCATGCACCAGTCACAG-3’, and gapdh mRNA, 5’-ACCACAGTCCATGCCATCAC-3’ and 5’-TCCACCACCCTGTTGCTGTA-3’. All data were normalized to gapdh mRNA expression.
Immunoblotting
Whole-cell extracts were prepared by direct addition of 4 × sodium dodecyl sulfate (SDS) sample buffer (200 mM Tris-HCl [pH 6.8], 8% SDS, 400 mM dithiothreitol, 0.2% bromophenol blue, 40% glycerol) to the cells, or by cell lysis in RIPA buffer (50 mM Tris-HCl [pH 7.5], 1% Triton-X, 1% sodium deoxycholate, 0.1% SDS, 150 mM NaCl, 0.2 mg/mL PMSF). The extracts were separated by SDS-PAGE and analyzed by immunoblotting using the following antibodies: anti-NFATc1 (7A6) (Santa Cruz Biotechnology, CA, USA [sc-7294]), anti-RANK (H-7) (Santa Cruz Biotechnology [sc-374360]), anti-β-Actin (Cell Signaling Technology, MA, USA [4970]), anti-Phospho-IκBα (nuclear factor of kappa light polypeptide gene enhancer in B-cells inhibitor α; Cell Signaling Technology [2859]), anti-IκBα (Cell Signaling Technology [4814]), anti-Phospho-ERK (Cell Signaling Technology [4370]), anti-ERK (Cell Signaling Technology [4695]), anti-Phospho-JNK (Cell Signaling Technology [4668]), anti-JNK (Cell Signaling Technology [9252]), anti-Phospho-p38 (Cell Signaling Technology [4511]), and anti-p38 (Cell Signaling Technology [8690]).
Electrophoretic Mobility Shift Assay (EMSA)
RAW 264 cells were pretreated with DMSO or N. officinale extract for 3 hours and then incubated with 5 nM RANKL for 0, 15, 30, or 60 minutes. The cells were washed twice with PBS and suspended in 1 ml buffer CE (10 mM Hepes-KOH [pH 7.9], 60 mM KCl, 1 mM EDTA, 0.5% NP-40, 1 mM DTT, 0.6 mg/mL PMSF), and lysed by vortexing. Nuclei were pelleted by centrifugation for 5 minutes at 4,000 × g and rinsed with 500 µL buffer CE. The nuclei were resuspended in 30 µL buffer NE (250 mM Tris-HCL [pH 7.8], 60 mM KCl, 1 mM EDTA, 1 mM dithiothreitol, 1 mg/mL PMSF) and lysed by 3 freeze–thaw cycles. The lysates were cleared by centrifugation for 5 minutes at 14,000 rpm at 4 °C. The supernatant was used as the nuclear extract. The binding reaction mixture, containing 3 µg nuclear protein and IRDye700-modified double-stranded oligonucleotide probe, was incubated for 30 minutes at room temperature in binding buffer (10 mM Tris-HCl [pH 7.5], 50 mM NaCl, 10% glycerol, 1% NP-40, 1 mM EDTA, and 1 µg poly(dI-dC)). The oligonucleotide probes used were as follows: organic cation transporter 1 (Oct1) 5´-TGTCGAATGCAAATCACTAGAA-3´ and 3´-ACAGCTTACGTTTAGTGATCTT-5´, and NF-κB 5′-AGTTGAGGGGACTTTCCCAGGC-3′ and 3′-TCAACTCCCCTGAAAGGGTCCG-5′. The reaction mixture was resolved on a non-denaturing 4% polyacrylamide gel containing 24.8 mM Tris, 190 mM glycine, and 1 mM EDTA in 0.5 mM TBE buffer. The IRDye700-modified probes were detected by infrared intensity using ChemiDoc MP Imaging System (Bio-Rad). For supershift analysis, the following antibodies were used: anti-RelA (F-6) (Santa Cruz Biotechnology [sc-8008]) and anti-RelB (D-6) (Santa Cruz Biotechnology [sc-48366]).
GST-Pull Down Assay
HEK293T cells were transfected with an expression vector for FLAG-tagged TRAF6 (pCIneo-FLAG-TRAF6) and with an expression vector for GST (pCIneo-GST) or GST-tagged RANK cytoplasmic domain (pCIneo-GST-RANK-IC). Twenty-four hours later, cells were pretreated with DMSO or N. Officinale extract for 3 hours and then incubated with 5 nM RANKL for 5, 15 and 30 minutes. The cells were lysed in L-buffer (50 mM Tris-HCl [pH7.4],150 mM NaCl, 5 mM EDTA [pH8.0], 1% Triton-X, 0.6 mg/mL PMSF, Protease inhibitor cocktail [Roche]) and cleared by centrifugation at 14,000 rpm. The supernatant was incubated with glutathione-Sepharose 4B (GE Healthcare, IL, USA) for 2 hours at 4 °C. The precipitates were then washed 5 times with the L-buffer and eluted the GST fusion protein complexes by boiling in SDS sample buffer. The eluate was subjected to SDS-PAGE and analyzed by immunoblot analysis.
Fractionation of N. Officinale Extract
N. officinale extract (0.5 g) was diluted with 4.5 ml of 50% (v/v) methanol/water. Then it was loaded to preparative HPLC (Shimadzu, Kyoto, Japan) using L-column ODS (10 mm i.d. ×250 mm, particle size 5 µm, Chemicals Evaluation and Research Institute, JAPAN). Elution was performed using mobile phase composed of water/acetonitrile = 95/5 at flow rate 4.5 mL/min. The UV spectrum at 244 nm was monitored. Eluates with retention times of 2.9 to 3.65, 3.65 to 4.4, 4.4 to 5.15, 5.15 to 6.75, and 6.75 to 20 minutes, respectively, were collected as fractions 1 to 5. Each fraction was concentrated under reduced pressure and freeze dried. Yields for fractions 1‐5 were 108.7, 15.7, 2.3, 3.3 and 8.4 mg, respectively.
Footnotes
Acknowledgments
We are grateful to Mr. Shohei Makino and Ms. Tomoko Sumi (Inabata Koryo Co., Ltd.) for chemical component analysis of N. officinale extract.
Declaration of Conflicting Interests
The author(s) declared the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article: K.K. receives research funding from IVY Cosmetics Corporation. Other authors declare no conflict of interest.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
