Abstract
Objective
Jinhua Qinggan Granules (JQGs) have achieved certain results in the prevention and treatment of COVID-19 in China during this coronavirus storm. In this study, we aimed to analyze the common mechanisms of JQG in the treatment of coronavirus-induced diseases, such as severe acute respiratory syndrome (SARS), Middle East respiratory syndrome (MERS), and COVID-19 via network pharmacology and molecular docking.
Methods
The active compounds of JQG were collected through Traditional Chinese Medicine Systems Pharmacology Database and Analysis Platform. The common targets associated with these 3 diseases were screened from GeneCards database. Gene ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) analysis of JQG’s core targets were analyzed using The Database for Annotation, Visualization, and Integrated Discovery and KOBAS 3.0 system. Further, the protein-protein interaction network was built using STRING database. The compound-target- signaling pathway network was constructed using Cytoscape 3.7.2. The core components of JQG were docked with core targets, COVID-19 coronavirus 3 Cl hydrolase, and angiotensin-converting enzyme 2 (ACE2) via Discovery Studio 2016 software.
Results
A total of 139 active compounds, 50 core targets, and 122 signaling pathways were screened out. The results of molecular docking showed that arctiin and linarin had a higher docking score with 3 Cl, ACE2, and core targets of JQH for antiviral effect.
Conclusion
The potential mechanism of action of JHQ in the treatment of MERS, SARS, and COVID-19 may be associated with the regulation of genes co-expressed with ACE2 and immune- related signaling pathways.
CoVs are single-stranded RNA viruses which belong to the order Nidovirales, family Coronaviridae, and subfamily Coronavirinae. 1 A CoV was first isolated from chickens in 1937, and then the first human CoV was isolated in 1965. CoVs are a large group of viruses that now exist widely in nature. Seven types of CoVs have been found that could infect humans, of which 4 mainly cause upper respiratory tract infection. 2 From Severe Acute Respiratory Syndrome (SARS) in 2002, Middle East Respiratory Syndrome (MERS) in 2012, and then to the current novel CoV disease (COVID-19), CoVs have caused 3 outbreaks of viral infectious diseases around the world and have become a major threat to global public health. On comparing the 3, COVID-19 has to date caused the worst impact. According to statistics, as of December 13th, 2020, 71,683,406 individuals had been infected with SARS- CoV-2 worldwide, of which 1,628,823 have died; the data were from WHO, official reports and authoritative media reports of various countries. SARS, MERS, and COVID-19 are similar in epidemiology, biological characteristics and clinical symptoms, that is, the patients will show symptoms of fever, cough, and upper respiratory tract infection. At the time of the 3 outbreaks, China has vigorously used traditional Chinese medicine for disease prevention and treatment, and achieved certain results. 3,4 Chinese medicine has thus shown its unique advantages. 5
In Chinese medicine, Jinhua Qinggan Granules (JQG) have been used for fever, sore throat, nasal congestion, runny nose, thirst, and cough. JQG were included in the treatment of CoV-induced pneumonia on January 27, 2020 by China’s National Health Commission and the State Administration of Traditional Chinese Medicine, and has achieved results. 6 -8 In fact, JQG had been previously included in the diagnosis and treatment plan for viral pneumonia and influenza several times in China. They were widely used for treatment of H1N1 influenza in 2009 in China. JQG consists of Menthae Herba (Bohe), Anemarrhenae Rhizoma (Zhimu), Fritillariae Thunbrgii Bulbus (Zhebeimu), Artemisia annua (Qinghao), Arctium lappa, (Niubangzi), Ephedra Herba (Mahuang), Forsythia suspensa (Lianqiao), Amygdalus communis (Kuxingren), Lonicerae Japomicae Flos (Jinyinhua), Scutellariae Radix (Huangqin), Licorice (Gancao), and Plaster (Shigao). According to TCM theory, the pathogenesis of CoV-induced pneumonia mainly indicates damp pathogens caused by cold and dampness outside the lungs and spleen, leading to Qi disorder and heat stagnation. Jinyinhua, Lianqiao, Qinghao, Bohe, and Huangqin in JQG have the functions of clearing heat, detoxifying dampness, and relieving asthma, respectively. 9 -12 In the clinical treatment of COVID-19 in China, JQG have also been used for treating critically ill patients, and the results show that it could have an anti-CoV effect and enhance immunity. However, the mechanism of JQG treatment for COVID-19 is unclear. Further, the efficacy of the main compound of each herb is not known.
Network pharmacology is the use of multiple technologies such as systems biology, high-throughput screening, network analysis, and network visualization to reveal the complex system of "disease-gene-target-drug" interactions, and analyze drug effects from a holistic and systematic perspective. Based on the similarities in the structure and efficacy of drugs, combined with the complex interaction relationships of the body’s target molecules and biological effects, by constructing drug-drug, drug-target, and other networks, it can effectively predict the drug function or the drug corresponding to a specific function. Under the background of molecular network regulation, it has become a new trend to apply the ideas and methods of network biology to the study of the pharmacological action and mechanism of traditional Chinese medicine, and a series of new data, methods and results have been produced. Molecular docking technology achieves the purpose of assisting drug screening based on the combination of the three dimensional structure of the receptor macromolecule with the ligand compound and the calculation of physical and chemical parameters. 13 It has been widely used in the study of active ingredients of traditional Chinese medicine and compound prescriptions based on specific targets.
In this study, we predicted the core compounds and core targets of JQG and the signaling pathways for treatment of SARS, MERS, and COVID-19 through network pharmacology and molecular docking, and the common effects and mechanisms of JQG in the treatment of these 3 kinds of CoV infection are discussed. The flow chart is shown in Figure 1.

The flow chart of this chart. TCMSP, PubChem, and PhamMapper were used to collect the active compounds of JQG and the compounds-related targets. GeneCards database was used to collect the targets associated with SARS, MERS, and COVID-19. JQG’s core targets were collected for GO and KEGG analysis, and PPI networks, compounds-pathways-targets network were constructed. Finally, molecular docking between the compound and the key target was carried out for verification.
Materials and Methods
Identification and Screening of Active Compounds
Traditional Chinese Medicine Systems Pharmacology Database and Analysis Platform (TCMSP, http://tcmspw.com/) was used to authenticate all compounds of the 12 Chinese medicinal herbs in JQG. TCSMP provides information on 6511 drug molecules and 3987 targets, as well as the interactions between them. Oral bioavailability (OB) and drug- likeness (DL) are the most significant pharmacokinetic parameters. OB indicates the speed and degree at which the active components or active groups of an oral drug are absorbed into systemic circulation, and DL refers to the similarity of a compound with a known drug, and the potential of this class of compounds to become drugs. 14,15 In this study, the names of herbs were used as the key words to retrieve all compounds, and the active compounds in JQG were selected according to the criterion of OB ≥30% and DL ≥0.18. 14,15
Identification of Protein Targets
The protein targets associated with active compounds were retrieved from the TCMSP database. The targets, including the gene names and gene ID, were further extracted using UniProtKB (http://www.uniprot.org), which comprised 54,247,468 sequence items.
Predicting the Common Targets of SARS, MERS, and COVID-19
GeneCards (https://www.genecards.org/) was used to gather target information concerning SARS, MERS, and COVID-19, which is a comprehensive database of functions involving proteomics, genomics, and transcriptomics. The keywords “novel coronavirus pneumonia,” “cough,” “MERS” “SARS,” and “fever” were utilized to screen these 3 disease-associated targets. Then, we obtained the intersection of the 3 disease targets through Venny2.1.0 database.
Gene Ontology and KEGG Pathway Enrichment Analysis
The results of Gene Ontology (GO) enrichment analysis were processed and visualized through the Database for Annotation, Visualization, and Integrated Discovery (DAVID, https://david.ncifcrf.gov/). GO enrichment analysis included biological processes (BPs), molecular functions (MFs), and cellular components (CCs). 16 Further, KOBAS3.0 system (http://kobas.cbi.pku.edu.cn/anno_ iden.php) was used to perform Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analysis. All results were screened with P < .05. The top 20 relevant results from the KEGG pathway and GO enrichment analysis were plotted using the Omicshare tool (http://www.omicshare.com) as bubble and enriched circle plots. 17
Construction of Protein-Protein Interactions (PPI) Network
Online SRING (https://string-db.org/cgi/input.pl) was used to build the PPI network interaction of the core targets of JQG for treating the 3CoVs. This database is mainly for predicting functional correlations between proteins. It is a pre-calculated global resource that can be used to browse and analyze these correlations, and it provides more than 2000 substances and their protein interaction relationships. 18
Construction of Compound-Target-Pathway Network
A visual compound-target-pathway network was built by the aforementioned data sets through Cytoscape3.7.2 (http://www.cyto-scape.org/) to reflect the complex relationships between compounds, targets and pathways. 19 At the same time, we have also established a network diagram of the top 10 compounds with degree value and their related targets in the same way; the 3D diagram of the compound was obtained from Pubchem (https://pubchem.ncbi.nlm.nih.gov/).
Molecular Docking
The core compounds selected from active compounds of JQG were docked with some core targets and Angiotensin-converting enzyme 2 (ACE2), which was confirmed to be closely related to SARS-CoV-2 in human cells. 20 At the same time, these core compounds were also docked with Remdesivir as a positive control. The structures of the related proteins were downloaded from the RSCB PDB database (https://www.rcsb.org/). A LiDockscore ≥90 is generally believed to represent a strong binding ability when docking with the processed protein. 21
Results
Active Compounds of JQG
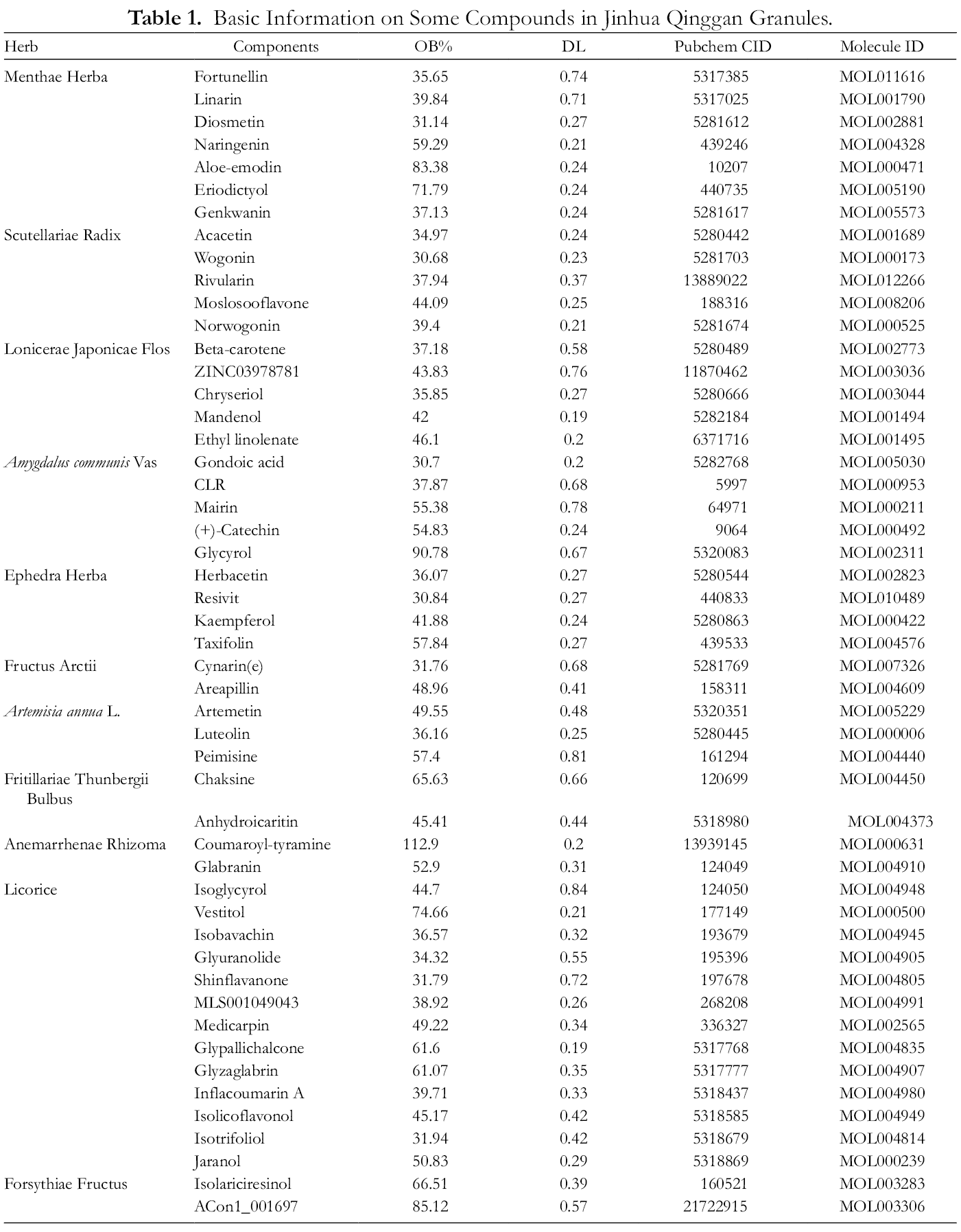
A total of 139 active compounds were collected with OB ≥30 and DL ≥0.18 from the TCMSP database of which 9 were from Menthae Herba, 5 from Anemarrhenae Rhizoma, 4 from Fritillariae Thunbrgii Bulbus, 11 from Artemisia annua, 5 from Fructus Arctii, 7 from Ephedra Herba, 5 from Forsythiae Fructus, 14 from Amygdalus communis, 6 from Lonicerae Radix, 12 from Scutellariae Radix, and 61from Licorice. Specific information of some active compounds is shown in Table 1.
Basic Information on Some Compounds in Jinhua Qinggan Granules.
The Core Targets of JQG for Treatment of the 3 Coronaviruses
A total of 1282 possible common targets of SARS, 1398 of MERS, and 259 of COVID-19 were screened through GeneCards. All targets of the 3 diseases were imported into Venny2.1.0 and 133 common targets were obtained, as shown in Figure 2(A). Among the 133 shared disease targets, there were 50 intersections with compound targets, and these were considered to be core targets of JQG for the 3 types of CoVs, as shown in Figure 2(B). Specific information of these 50 core targets is shown in Supplemental Table S1.

Wayne diagram for coincidence targets. (A) Common targets of 3 diseases: SARS, MERS and COVID-19; (B) Wayne for compound predicted target and common targets of 3 diseases illustration.
GO and KEGG Enrichment Analyze
GO functional enrichment analysis yielded 404 GO entries, including 298 (73.8%) BPs, 44 (10.9%) CCs, and 62 (15.3%) MFs. The enriched circle plot was used to visualize the enrichment results of the top 20 GOs, as shown in Figure 3(A). The outermost circle shows the classification of GO enrichment, and the yellow represents BP, the blue MF, and the green CC. The second circle shows the P value, with the smaller the value, the darker the color. The third circle shows the total number of foreground genes. The fourth circle shows the RichFactor value of each classification. KEGG pathway enrichment was conducted to cluster the major effects that are associated with the JHQG. A total of 122 pathways were screened out; the top 20-ranking ones are shown in Figure 3(B). The main pathways included hepatitis B, HIF-1 signaling pathway, pathways in cancer, TNF signaling pathway, tuberculosis, and apoptosis, as well as some related to cancer. Among them, hepatitis B involved 17 genes, including transforming growth factor β1 (TGFβ1), BAD, tumor necrosis factor (TNF), PI3KCG, caspase (CASP)8, CASP3, mitogen-activated protein kinase (MAPK)1, and nuclear factor kappa B subunit (NFκB)1. The TNF signaling pathway involved 15 genes, including epidermal growth factor receptor (EGFR), NFκB1, MAPK1, interleukin (IL)6, and B cell lymphoma (BCL)2. Pathways in cancer involved 22 genes, including TGFβ1, EGFR, PIK3CG, NFκB1, IL6, MAPK8, CASP8, and BCL2. We predicted that some compounds of JHQG could act on TNF, NFκB1, IL6, MAPK, and other genes to regulate the inflammation pathway, apoptosis, and other signaling pathway, to achieve an anti-CoV effect. Specific information of gene degree is shown in Supplemental Table S1.

Bubble chart and circle plot of the results of the identified target proteins by Database for Annotation, Visualization, and Integrated Discovery. (A) The top 20 significantly enriched terms in GO enrichment; (B) The top 20 KEGG pathways enrichment analysis.
PPI Network
The PPI network of 50 core targets is shown in Figure 4. We found that IL6, CASP 6, BCL2, EGFR, and NFκB1 have the most associations with other genes. This suggested that the mechanism of action of JQG in the treatment of the 3 CoV diseases may be related to the regulation of those genes.

Protein-protein interaction of core potential targets of JQG for the treatment of 3 coronaviruses.
Topology Analysis Network
The compound-target-pathway network is shown in Figure 5(A), including 209 nodes (139 compound nodes, 50 target nodes, 20 pathway nodes), and 1061 edges. The orange color represents the pathways, green the targets, and blue the active compounds. According to the degree analysis, the top 10 compounds in PubChem CID were 392442 (glyasperin F), 5317385 (fortunellin), 100528 (arctiin), 195396 (glyuranolide), 5317025 (linarin), 21722915 (pinoresinol monomethyl ether), 392443 (licoisoflavanone), 10026486 (asperglaucide), 161294 (peimisine), and 441298 (hyperforin). Figure 5(B) showed the 3D structure of the compound and the relationship of compounds and targets. We have discovered a phenomenon in which different compounds interact with the same target, reflecting the mechanism of interaction between multiple components in traditional Chinese medicine and multiple targets. This is mutually corroborated with the multi-target therapeutic mechanism of Chinese medicine.

Network constructed by cytoscape. (A) Compound-target-pathway network. The orange square was the active compounds of JQG, and the green circle was the core target proteins of JQG for treatment of the 3 coronaviruses, the blue hexagon was the pathway signaling. (B) Top 10 compounds-targets network. The blue hexagon was the compounds, the green square was the targets.
Molecular Docking Display
Through topological analysis results and literature review, we selected arctiin and linarin to be docked with the top 3 targets (EGFR, NFκB1, and IL6) mentioned in the PPI network. Further, the 2 compounds were docked with SARS- CoV-2 3 Cl protein and ACE2 protein. Compared with other targets, the docking score of NFκB1 with the core compounds was not too high. In addition, via molecular docking, linarin had a higher docking score than arctiin, with 5 targets. This suggested that linarin was more likely to be an important compound in JQG. The docking scores of Remdesivir with the 5 targets were greater than 130, which implied that our molecular docking data were accurate and true. From the 2D diagram, it could be found that van der Waals, conventional hydrogen bonds, and carbon hydrogen bonds were the most important chemical bonds in the docking of the compounds with the targets. The details of docking are shown in Figures 6 -8, and Table 2.

Results of molecular docking of each targets with linarin. (A) EGFR; (B) ACE2; (C) 3 Cl; (D) NFκB1; (E) IL6.

Results of molecular docking of each targets with arctiin. (A) EGFR; (B) ACE2; (C) 3 Cl; (D) NFκB1; (E) IL6.
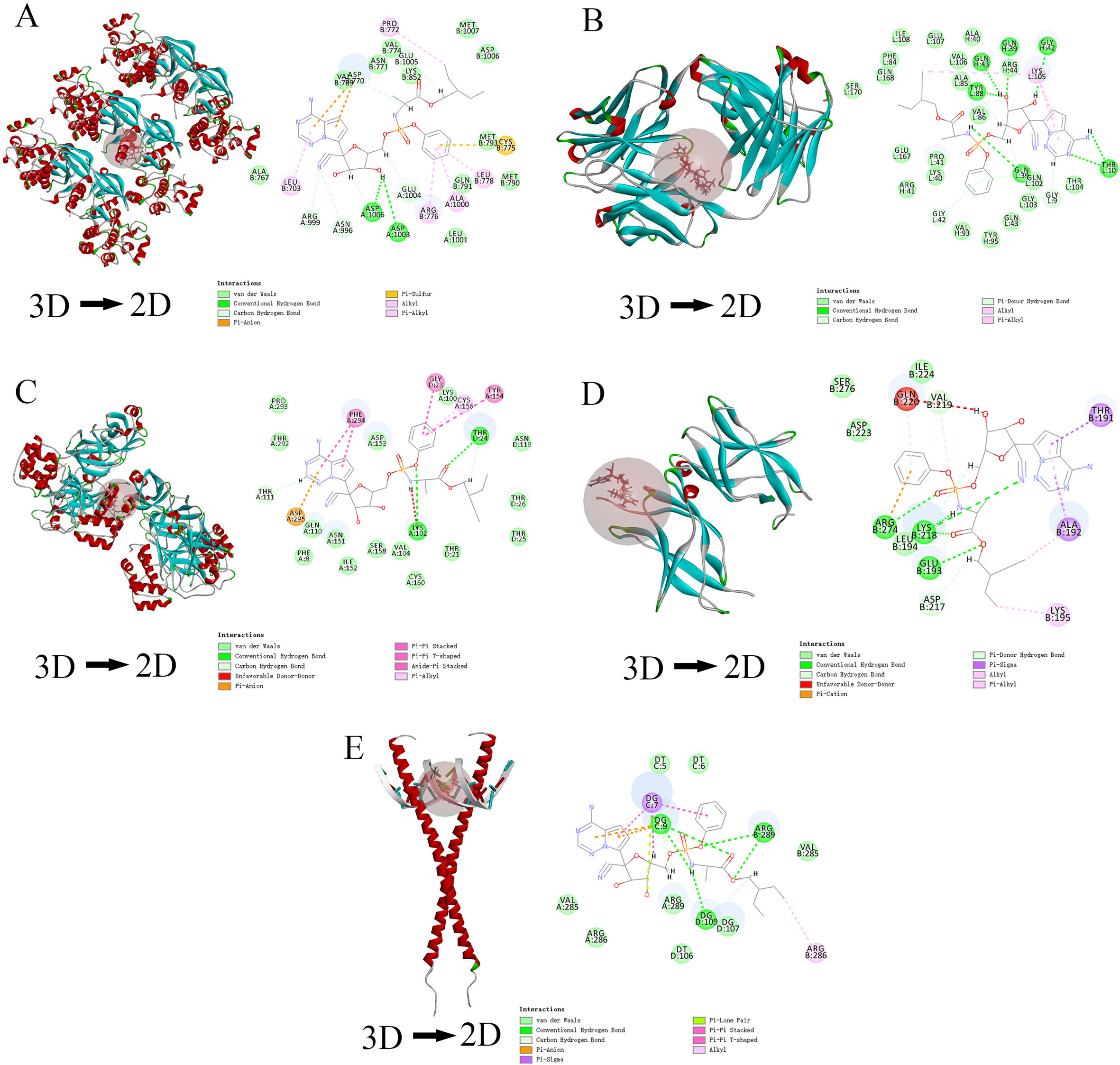
Results of molecular docking of each targets with remdesivir. (A) EGFR; (B) ACE2; (C) 3 Cl; (D) NFκB1; (E) IL6.
Results of Molecular Docking.

Discussion
In this study, the potential mechanism of JQG for treatment of CoV infection was studied by network pharmacology and molecular docking, and 139 active compounds, 50 core targets, and 122 signaling pathways were screened out. Glyasperin F, fortunellin, arctiin, glyuranolide, and linarin were found to be the main active compounds in JQG for treatment of CoV infection. Based on known literature, we chose arctiin and linarin for molecular docking with 3 Cl and ACE2; both showed strong affinity. Arctiin is the main active lignan in Arctium lappa L, which has anti-inflammatory, antiviral, antidiabetic and antitumor activities, in addition to other pharmacological properties. 22 At present, controlling the expression of inflammatory factors to improve the body’s immunity is considered to be an important method for treatment of viral pneumonia. As previously reported, arctiin exhibited potent anti-inflammatory activities by inhibiting inducible nitric oxide synthase (iNOS) via modulation of several cytokines. 23 Li et al. have reported that arctiin exhibits potent anti-inflammatory and anti-allergic activities via down-regulating the activation of MAPKs and AKT pathways. 24 In addition, Kyoko Hayashi et al. found that arctiin had a direct antiviral property, with a therapeutic effect in immunocompetent and immunocompromised mice infected with influenza A virus. 25 Linarin, a flavonoid, is a constituent of various plants and has antibacterial, anti-aging, anti-anoxia, anti-adverse irritation, anti-fatigue and other pharmacological effects. Xiang et al. found that linarin could prevent LPS-induced acute lung injury by suppressing oxidative stress and inflammation by inhibition of TXNIP/NLRP3 and NF-κB pathways. 26 Bomi Kim found that linarin had an anti-inflammatory effect, in part through the suppression of phagocytosis, cytokine production, and antigen presentation in macrophages. 27 Therefore, arctiin and linarin may be effective against CoV infection-induced pneumonia through their anti-inflammatory or antiviral pharmacological effects.
The HIF-1 signaling pathway, pathways in cancer, TNF signaling pathway and apoptosis were screened to be the main pathways of JQG for treatment of the 3 CoV diseases. KEGG enrichment results showed that the core targets included TGFβ1, EGFR, PIK3CG, NFκB1, IL6, MAPK8, CASP8, and BCL2, among others. Based on the network pharmacology and molecular docking results, we predicted that JQG could treat CoV-induced diseases by regulating NFκB1, IL6, and EGFR in the following ways. NFκB1 is a nuclear transcription factor that exists in a variety of cells and has a multi-directional regulatory function. Once the human body is infected with the coronavirus, the inhibitory factor κB of NFκB1 is phosphorylated, which leads to the ubiquitination of κB and proteasome-mediated degradation, which releases NFκB1 from the cytoplasm, causing inflammation storms in the body and accelerating viral cells transfer in the patient, and finally aggravates the patient’s infection. Severe cases of COVID-19 are characterized by a strong inflammatory process that may ultimately lead to organ failure and patient death. IL-6 in the NLRP3 inflammasome was considered an important marker in the sera correlated with COVID-19 severity. 28 The core compounds in JQG may play an antiviral role by regulating factors in inflammatory pathways such as NFκB1 and IL6. EGFR is a member of the type I tyrosine kinase receptor gene family, which is mainly involved in cell signal transduction. 29 Once activated, it leads to cell differentiation, proliferation, infiltration, and angiogenesis; inhibits the high-expression CASP3 protein in virus- infected cells; and achieves an antiviral effect.
However, there are several limitations in our study. First, a more comprehensive TCM target genes database is needed, which would make the results of network pharmacology analysis more reliable. Second, even if the results of network pharmacology and molecular docking were combined, we still could not completely understand the accurate therapeutic mechanism of JQG. At present, some docking software still has false positive results, and, therefore, there may be inaccuracies in our docking results. In further research, pharmacodynamics and pharmacokinetics are necessary to understand the mechanism of JQG for treating CoV-induced diseases.
Conclusion
In this study, network pharmacology showed that the main active compounds of JQG, particularly arctiin and linarin, could act on multiple targets, such as EGFR, IL6, and NFκB1. JQG had an effect on the treatment of CoV mainly through the TNF signaling and HIF-1signaling pathways, and pathways in cancer. Molecular docking showed that arctiin and linarin were the top 2 compounds, which indicated that they might play an important role in the treatment of CoV-induced diseases.
Supplemental Material
Table S1 - Supplemental material for Investigation of Anti-SARS, MERS, and COVID-19 Effect of Jinhua Qinggan Granules Based on a Network Pharmacology and Molecular Docking Approach
Supplemental material, Table S1, for Investigation of Anti-SARS, MERS, and COVID-19 Effect of Jinhua Qinggan Granules Based on a Network Pharmacology and Molecular Docking Approach by Ying Zhang, Yunfeng Yao, Yanfang Yang and Hezhen Wu in Natural Product Communications
Footnotes
Acknowledgments
The authors would like to thank all the colleagues and students who contributed to this study.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was supported by the National Natural Science Foundation of China [No.31570343], and the Hubei University of Chinese Medicine Funding Project on COVID-19 Emergency Science and Technology Research.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
