Abstract
The process of investigating a possible cure for coronavirus disease 2019 (COVID-19) in vitro and in vivo may take a long time. For this reason, several in silico studies were performed in order to produce preliminary results that could lead to treatment. Extract of Juniperus procera Hochst is used as a traditional medicine for recovery from flu in Saudi Arabia. In the present study, more than 51 phytochemicals of J. procera were docked against the main protease of COVID-19. Rutin gave the highest interaction score among all the phytochemicals and the commercially available antiviral drugs. Lopinavir showed the second highest binding score. Rutin and lopinavir were further investigated using homology models of COVID-19. Rutin showed a better inhibition score in 9 of the 11 of homology models compared with lopinavir. Analysis of ligand-protein interaction contacts revealed that 3 residues (Glu166, Gly143, and Thr45) of the main protease formed hydrogen bonds with rutin. This simulation study suggests that rutin could be a possible effective inhibitor of several COVID-19 protein targets, including the main protease. Rutin, already available for commercial use, was evaluated for its ability as a possible drug. To our knowledge, this is the first study that suggests rutin having a possible strong inhibitory role against several protein targets of COVID-19.
Coronaviruses are a family of viruses that are linked to several diseases in animals and humans. These diseases range from the common cold to more severe ones. At the end of December 2019, coronavirus disease 2019 (COVID-19), also known as severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), appeared in Wuhan City in China. No specific therapies are available for COVID-19 until now. COVID-19 symptoms are similar to those of common flu and in some cases can develop into pneumonia. At the time of writing this report, 5 704 736 cases and 357 736 deaths were confirmed by the World Health Organization (WHO) in 216 countries.
Juniperus procera Hochst (Arar) is one of more than 1200 medicinal plants found in Saudi Arabia. 1 Arar, found in the south of Saudi Arabia, is known for its possible ability for treating many diseases, including infections caused by viruses like the cold. 2 -4 The aim of this study was to investigate the possible inhibitory action of many phytochemicals of Arar against the solved crystal structure of the main protease of COVID-19 and 11 homology models of other proteins.
Results and Discussion
The results of docking indicate a strong possible interaction between rutin and the main protease enzyme, as shown in Table 1. Among 52 phytochemicals of Arar and 4 controls, rutin recorded the minimum binding energy of −136.47 indicating a strong interaction with the main protease. Supplemental Table S1 provides more details about the docked compounds. Rutin interacts with the main protease by forming several atomic contacts, as shown in Figure 1 and Supplemental Table S2. Ten hydrogen bonds were formed in the rutin-main protease complex compared with one found in 6-(ethylamino)pyridine-3-carbonitrile (RZS) (the native inhibitor)-main protease complex. This may explain the strong interaction between rutin and the main protease. For further confirmation, rutin was docked to the main protease using the COVID-19 Docking server; rutin gave a better inhibition effect than the other compounds. Further investigation was performed with rutin and lopinavir in order to know if these 2 compounds have an inhibition effect against potential targets of COVID-19 other than the main protease, as shown in Table 2. Rutin gave a better interaction score with 9 homology models of 11.
Docking Score of Juniperus procera Phytochemicals, the Native Inhibitor, and Controls Against Main Protease of COVID-19.
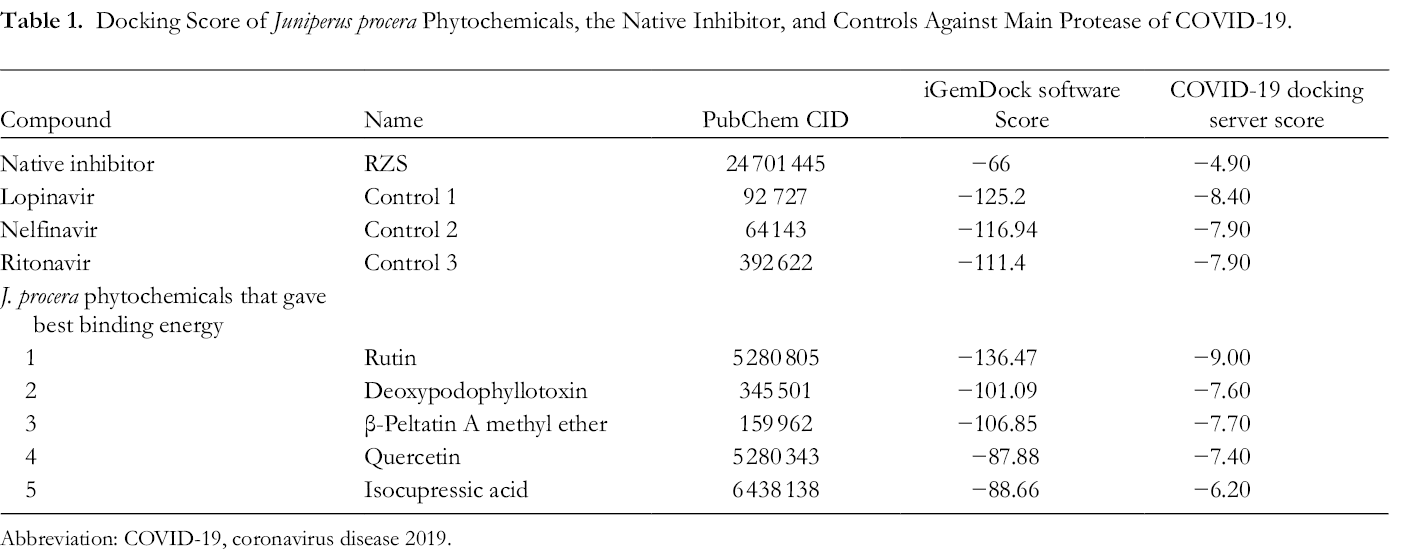
Abbreviation: COVID-19, coronavirus disease 2019.

Docked compounds located in the binding pocket. Pink is the main protease, green is the native inhibitor, red is the docked inhibitor (it was almost superimposed with the relevant co-crystallized inhibitor indicating the accuracy of docking), orange is the lopinavir, and blue is the rutin. The box on the left presents the hydrogen bond contacts formed by rutin-main protease interaction.
Docking Score of Rutin and Lopinavir Against Homology Models of COVID-19.
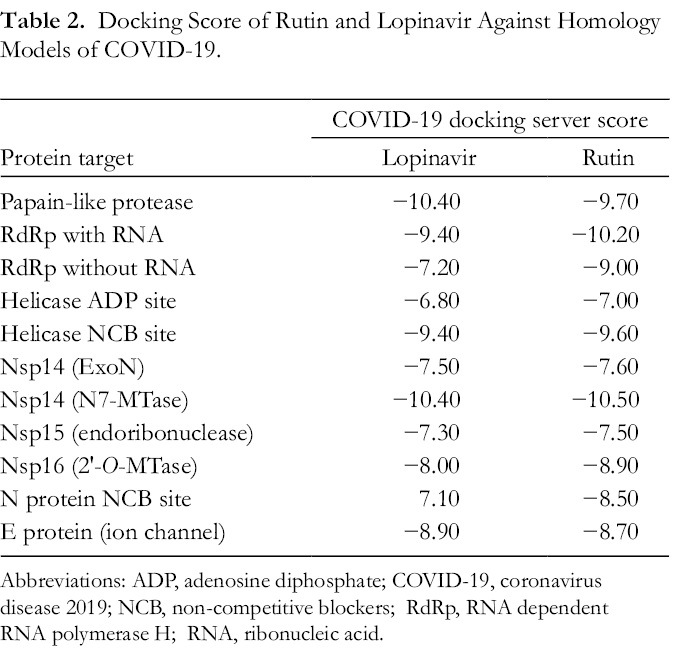
Abbreviations: ADP, adenosine diphosphate; COVID-19, coronavirus disease 2019; NCB, non-competitive blockers; RdRp, RNA dependent RNA polymerase H; RNA, ribonucleic acid.
The molecular dynamics results showed that the root-mean-square deviation (RMSD) per residue of the X-ray structure of the main protease with no ligands was in a good agreement with the docked rutin-main protease complex (Figure 2). MD results revealed a nonsignificant difference in RMSD of all atoms of each residue pairs of the 2 comparing proteins. Based on MD results, the rutin-main protease complex is stable, and the ligand has no significant effect on the stability of the main protease.
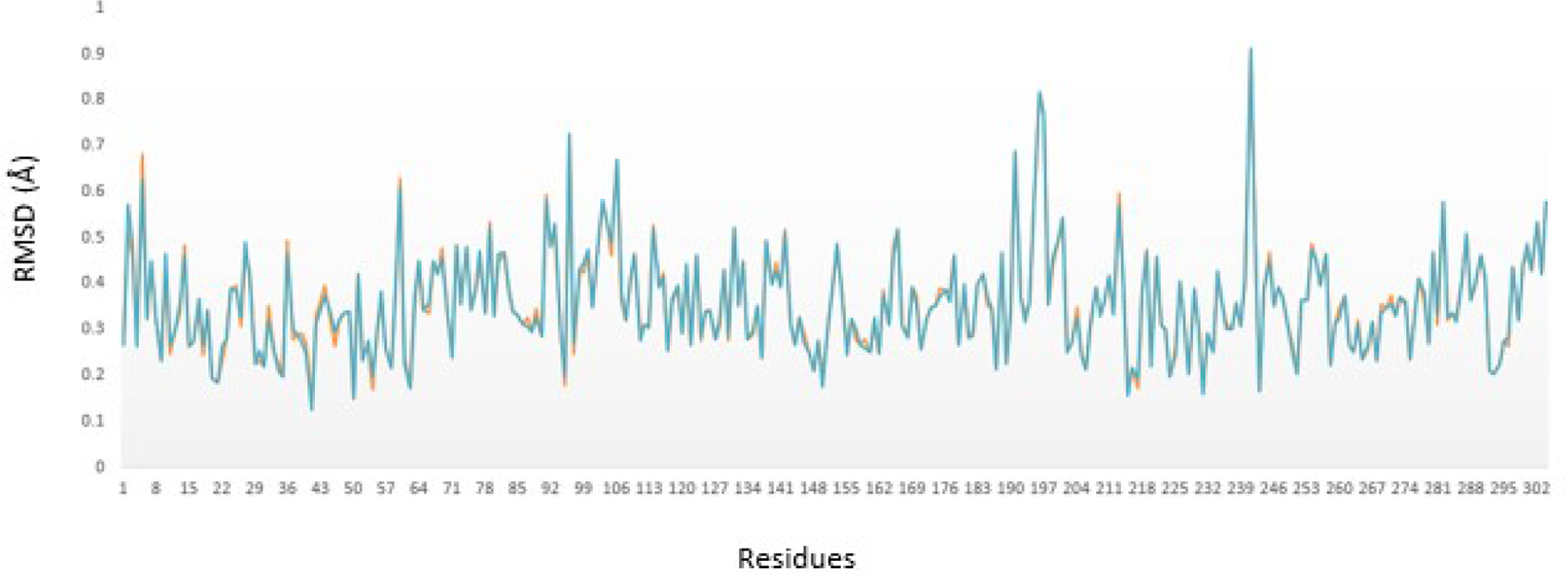
RMSD per residue of the X-ray structure of DP4 (orange) and docked rutin-main protease (blue). RMSD indicates root-mean-square deviation.
Several in silico studies have examined the interaction between natural products and antiviral drugs with the main protease. 5 -8 However, no previous study has investigated the interaction between the constituents of J. procera with the main protease of COVID-19. More details about the possible importance of the main protease as an inhibition target of COVID-19 were presented by Khaerunnisa et al. 6 Some studies have investigated the role of rutin as an inhibitor of other viruses. Dengue virus type-2 can be inhibited by rutin. 9 A study by Lin et al 10 reported the antiviral effects of rutin against the 3C protease of enterovirus A71 (EV-A71). Another study 11 observed the antiviral activity of rutin against herpes simplex virus type 1 in vitro. An extract of Euphorbia milii inhibited the growth of a ruminant virus. The study highlighted the possible link between the presence of rutin in the extract and the virus inhibition. 12
Experimental
Protein Docking
The COVID-19 main protease crystal structure (protein data bank [PDB] ID: 5r82) was obtained from PDB, in pdb format. Chain A of the 5r82 structure was used for receptor preparation and docking. Homology models of COVID-19 based on the protein structures of SARS-CoV were used for rutin-protein docking, as shown in Table 2. The phytochemicals and other ligands were obtained from the PubChem database. Docking was carried out with iGEMDOCK 2.1 software 13 and COVID-19 Docking server. 14 Preparation of input files for docking was reported by 15
Molecular Dynamics
Chain A of the main protease with no ligands and the docked rutin-main protease complex was used for the molecular dynamics (MD) study in order to investigate the effect of rutin on enzyme stability. MD simulation was performed using MDWeb. 16 The MD simulation parameters and methods were previously reported. 15
Conclusions
Rutin, a natural compound, showed a possible inhibition effect on the main protease and other key proteins in the life cycle of COVID-19. The use of a natural product as a treatment has its advantages over synthetic drugs in terms of reducing the side effects. Lopinavir showed the second highest interaction score after rutin and can be considered as a possible available drug for COVID-19. More in vitro and in vivo studies are needed for investigating this in silico study.
Supplemental Material
Supplementary Material 1. - Supplemental material for Rutin as a Promising Inhibitor of Main Protease and Other Protein Targets of COVID-19: In Silico Study
Supplemental material, Supplementary Material 1., for Rutin as a Promising Inhibitor of Main Protease and Other Protein Targets of COVID-19: In Silico Study by and Ateeq Ahmed Al-Zahrani in Natural Product Communications
Footnotes
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: this work was supported financially by the Deanship of Scientific Research at Umm Al- Qura University (Grant Code. 18-SCI-1-01- 0028).
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
