Abstract
Various computational studies, including in silico ones, have identified several existing compounds that could serve as effective inhibitors of the SARS-CoV-2 main protease (Mpro), and thus preventing replication of the virus. Among these, rutin has been identified as a potential hit, having prominent binding affinity to the virus. Moreover, its presence in several traditional antiviral medicines prescribed in China to infected patients with mild to moderate symptoms of COVID-19 justify its promise as a repurposed bioactive secondary metabolite against SARS-CoV-2.
Rutin, also called sophorin, rutoside, and quercetin-3-rutinoside, is a polyphenolic flavonoid, abundantly present in various dietary sources such as buckwheat, onions, oranges, lemons, grapes, limes, berries, peaches, plums, apples, and tomatoes, as well as in drinks such as wine and black tea. 1 -8 Chemically, it is a glycoside having rutinose (α-L-rhamnopyranosyl-(1→6)-β-D-glucopyranose) linked to OH-3 of quercetin (3,3',4',5,7-pentahydroxyflavone), as shown in Figure 1.
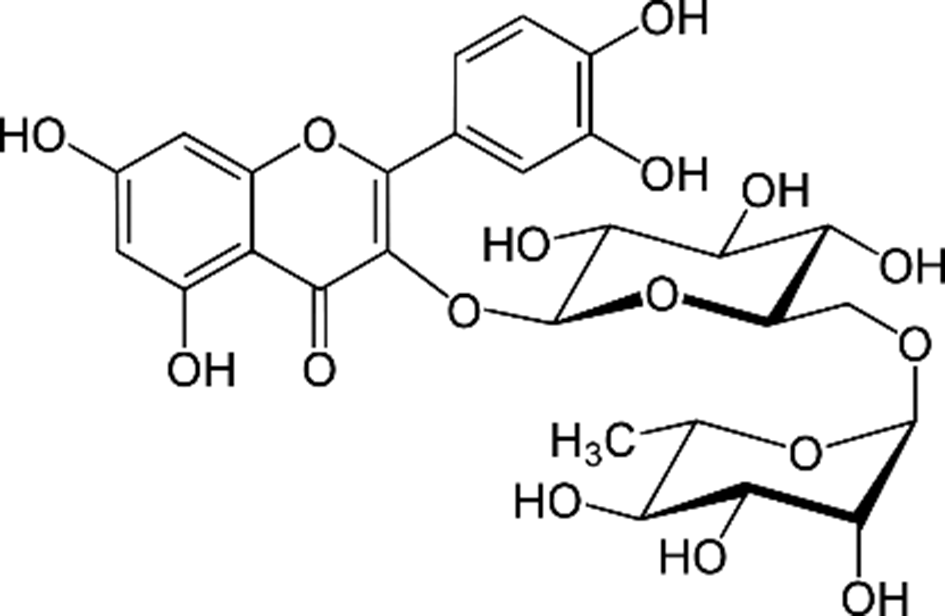
Structure of rutin.
Pharmacological Significance of Rutin
Rutin, a multifunctional phenolic natural product, is vital in diets and is of great interest due to its numerous reported beneficial activities. These include antibacterial, 9 -11 antiprotozoal, 12 antimalarial, 13 antimicrobial, 14 -16 anthelmintic, 17 antitumor, 18 -20 antiallergic, 21 analgesic, 22,23 antiinflammatory, 24 -26 antiulcer, 27 antiaging, 28,29 antioxidant, 30 -34 antidiabetic, 35 -39 myocardial protecting, 40 Alzheimer’s disease, 41,42 inflammatory brown disease, 43 prevention of hypertension, 44 vasodilator, 45 radioprotector, 46 nephroprotective, 47 hepatoprotective, 48,49 anticoagulant, 50 -52 combating blood toxicity, 53 cardioprotective, 54 immunomodulator, 55 ancillary therapeutic agent for snakebites, 56 beneficial effects on skin aging, 57 supporting a healthy immune system, 58,59 thyroid function, 60 neuroactive, 61 -63 for treating leishmaniasis, 63 insecticide, 64 amelioration of gut dysbiosis, 65 and cognitive impairment prevention. 66 Several review articles on the pharmacology of rutin have been published in recent years. 5,67 -86
Antiviral Significance of Rutin
In addition to the above-mentioned pharmacological properties, rutin has also been reported to have antiviral activity. For example, it has been tested against vesicular stomatitis virus on mouse fibroblasts and protected cells for about 24 hours. 87 In the case of canine distemper virus infection, rutin afforded immense viral embarrassment when added at the times of adsorption and penetration in the viral replicative cycle. 88 Rutin, a chief constituent of Capparis sinaica Veill, was tested for its in vitro antiviral activity against avian influenza strain H5N1 using a plaque inhibition assay in Madin-Darby canine kidney. The test showed a significant inhibition of the virus by about 73%. 89 Zuo et al. have reported the activity of rutin against the NS3 serine protease, in vitro. 90 Rutin from plum (Prunus domestica L.) was suggested as a strong inhibitor of hepatitis C virus (HCV) entry by inhibiting the early entry stage of the HCV lifecycle. 91 Rutin was able to inhibit the replication stage of infection with an IC50 of 110 µM against Enterovirus-A71 strain CMUH01 (B5). 92 In another study, rutin was found to reduce the infectivity of the C4 subgenotype EV-A71 with an IC50 of 200 µM. 93 Sulfated rutin was revealed in vitro to possess an antiviral effect against HIV-1 × 4 virus IIIB, and HIV-1 R5 isolates Ada-M and Ba-L strains by blocking viral entry and virus-cell fusion through interacting with the HIV-1 envelope glycoprotein. 94 Rutin, along with other flavonoid glycosides present in the ethanolic extract of Ficus benjamina L., exhibited activity against HSV-1 and HSV2. 95 The probable mechanism proposed by investigators was the inhibition of the HIV glycoprotein-mediated cell-cell fusion step. 95 Rutin has also been tested against Dengue virus type 2 96 -99 and influenza virus. 100 Antiviral testing for selected compounds against murine norovirus infection revealed that rutin has anti-murine norovirus activity in cell-based assays. 101 Rutin also showed effects on hepatic monooxygenase activities in experimental influenza virus infection (EIVI). 102 Molecular docking studies suggested that rutin can act as a novel hepatitis B virus inhibitor. 103 Thus, rutin has a broad antiviral activity against a diverse group of viruses. The antiviral activity of rutin, along with other flavonoids, has been reviewed in several publications. 104 -110
SARS-CoV-2
SARS-CoV-2 is an enveloped, non-segmented, positive-sense, single stranded RNA virus containing ~30,000 nucleotides encoding for 9860 amino acids. 111 The virus membrane has a crown-like appearance, due to its decoration with glycoprotein ‘spikes’. Its genome encodes several structural proteins such as spike (S) protein, membrane (M) protein, envelope (E) protein, and nucleocapsid (N) protein. The spike (S) glycoprotein plays a significant role in viral infectivity as it functions as a major inducer of host immune responses and identifies the target receptor. It has been noted that the S-glycoprotein spike on the outer surface of corona viruses and the host transmembrane serine protease 2 (TMPRSS2) are responsible for the attachment to the angiotensin-converting enzyme 2 (ACE2) receptor and subsequent entry into the cell targets of the human host. 112 -114 Thus, ACE2 plays a significant role during endocytosis and is expressed in the lungs, arteries, heart, kidneys, and other tissues. 115 -121 The viral particle is internalized in a vesicle, whose envelope is then removed, allowing the genomic RNA to be released into the cytoplasm. 122,123
The genome of SARS-CoV-2 contains two open reading frames ORF1a and ORF1ab translated to two respective viral replicase polyproteins pp1a and pp1ab through a 1-ribosomal frameshift mechanism. 124 -126 This is followed by proteolytic processing of polyproteins by two virally encoded cysteine proteases, SARS-CoV-2 papain-like protease (PLpro) and SARS-CoV-2 main protease, Mpro, also referred to as the 3C-like protease (3CLpro). Both of these proteases are involved in a proteolytic process of polyproteins. 127 -130 While PLpro digests the first three cleavage sites of its polyprotein, Mpro is responsible for cleavage of the remaining 11 locations resulting in the release of the functional non-structural proteins nsp4 through to nsp16 that are part of the replicase-transcriptase complex. The proteolytic processing of the 16 nsps by Mpro and PLpro is essential for virus replication and maturation because the nsps are involved in downstream binding and replication events, including the formation of the replicase complex, which is essential for viral replication and transcription of the genome. 131,132 This is followed by virus assembly, and subsequently virions are released from the infected cell through exocytosis.
The new copies of the virus spread into the environment and infect other cells and organs in the body, in a chain expansion. When the viral load is high and the cell is invaded by many viral particles, all its protein synthesis apparatus is dedicated to viral replication, up to cell death. The last phase can take place with either the “apoptosis” mechanism (if death is slow and controlled) or following “energetic-metabolic chaos,” such as to cause the breakdown of the various cell membranes, including lysosomes, and a total loss of structural integrity. Possibly, autoimmune phenomena are also involved in the attack on the infected cell by T-lymphocytes and antibodies. Eventually, both in the tissue where many cells have died (first of all in the lung), and systemically (lymph, blood, immune system, coagulation, kidney, liver), an inflammatory reaction develops, which can be clinically very serious, especially in patients with comorbidities. Excessive and “vicious” inflammation can be mediated by a distorted activation of the cytokine network, by clotting disorders, even by a paradoxical excess of the immune reaction (autoimmunity, cytotoxic lymphocytes). A brief description of the virus’s infectivity and its pathology is highlighted in Figure 2.
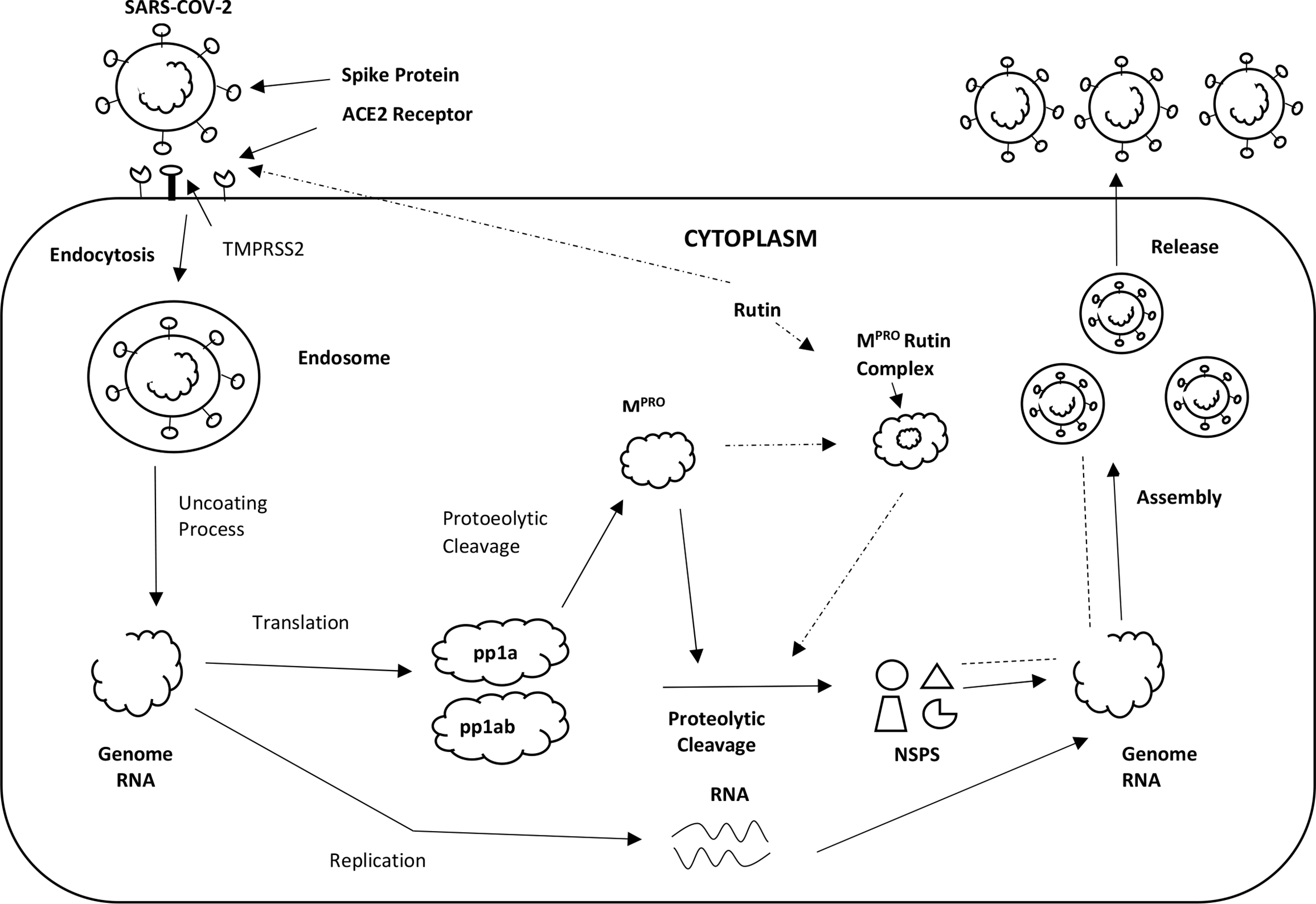
Replication cycle of the Severe acute respiratory syndrome coronavirus-2 (SARS-COV-2). The virus invades a human cell by attaching its spike protein to a cell surface receptor ACE 2. Upon entering the cell, the virus breaks up to release its genomic material. Mpro produces non-structural proteins (NSPS), that are essential for assembly of the viral transcription complex needed for RNA synthesis. Rutin binds to Mpro, resulting in failure of virion assembly and inhibition of release of functional NSPS. Rutin also binds to ACE2 receptors thus also inhibiting virus entry.
Based on this concise description, substances with a possible beneficial effect in coronavirus infection may act in various stages: (a) preventing either the binding of the virus to the receptors or inhibiting the function of the receptor itself, when it sets in motion the internalization process, (b) inhibiting viral replication by blocking, for example, RNA polymerase, proteases or new particle assembly, (c) helping the cell to resist viral attack, that is, stopping the cytotoxicity process, (d) blocking the spread of the virus in the body, (e) modulating the inflammation when, starting as an innate defensive mechanism, it becomes offensive and cytotoxic.
SARS-CoV-2 Main Protease (Mpro)
Mpro is found to be integral to the expression of viral gene and replication and is encoded by nsp5 in the viral genome. 127 The Mpro protease has a mass of approximately 33.8 kDa, 128 and is characterized by a self-cleavage protein. 129,130 It consists of a homodimer subdivided into two protomers (A and B) that have 3 distinct domains. 131 Domains I and II comprise β-barrels forming a chymotrypsin structure and harboring catalytic dyad histidine 41 (His41), and cysteine 145 (Cys145). 126,132 Domain III consists of α-helices. 133 For its catalytic activity, Mpro needs to dimerize, forming contacts with both the N- and C-terminal domains of the other protomer. 134
It is well known that the main protease (Mpro) of SARS-CoV-2 plays an important role in maturation of many viral proteins. The 12 viral non-structural proteins (Nsp4-Nsp16), including critical proteins like the RNA-dependent RNA polymerase (RdRp, Nsp12) and helicase (Nsp13), require cleavage through Mpro. It has also been demonstrated experimentally that inhibition of Mpro prevented viral replication in multiple cell models with COVID-19 pseudovirus as well as SARS2-CoV-2 virus. 128 The virus enters the cell via different proteases and receptors in different organs, however, Mpro activities are required for virus proliferation in all cell types.
The dependence of the virus on Mpro, and given that there are no similar human proteases, make this protein a promising drug target, 135 -137 and it is highly conserved among coronaviruses. 138 Therefore, Mpro has emerged as one of the key targets for the development of antiviral therapies aimed at blocking the life cycle of the coronavirus. 139 Besides the fact that this enzyme only exists in the virus and not in humans, 139 the high conservation of Mpro (96.1% similarity) among the related SARS-CoV viruses and its importance in the replication of the virus makes this enzyme an attractive target for potential antiviral drug design. Hence, there have been efforts to identify therapeutic candidates targeting Mpro using various virtual screening methods based on pharmacophore, molecule docking, and molecular simulations. 139 -146
Drug Repurposing
Drug repurposing, also known as drug repositioning or drug re-profiling, works as an alternative, systematic method in drug discovery that can aid in determining new indications for existing drugs. While the discovery and development of new antivirals that can efficiently treat COVID-19 should be actively pursued, drug repositioning stands out as an attractive strategy to be explored for SARS-CoV-2. Toward this end, many preclinical studies and investigational clinical trials have been conducted worldwide to determine which therapeutic options readily available can serve not only as an antiviral therapy to cure COVID-19, but also that can efficiently reverse the associated inflammatory conditions. Since de novo drug design can be time-consuming (usually 10 to 15 years to get new drugs to market), therefore either repurposing of FDA approved drugs or the search for small molecules from natural consumable resources which are generally considered to be either safe or having negligible toxicity, are considered to be appropriate approaches. 147 -151 The computational or in silico techniques have made them an exciting approach to the drug repurposing world. 152
There are two main approaches to the computational drug repurposing process: target-based and disease based. 147 The former allows the drug and the target to interact with each other leading to the establishment of drug-target interactions. The latter utilizes datasets to determine new indications for already approved drugs from comparisons of characteristics of diseases.
Computer-aided drug design techniques represent a very efficient approach to provide promising results in drug discovery. A structure-based pharmacophore approach has proven to be useful for supporting in silico hit discovery, hit-to-lead expansion, and lead optimization. 153,154 The protein-ligand complex-based approach is specialized to gain insights into ligand-protein interactions and to identify the essential pharmacophore features necessary for optimal interaction and biological activity. 155 Furthermore, structure-based pharmacophore models obtained using single ligand-target structures have proven their worth as an alternative to the traditional approaches. This is particularly evident by their role in the virtual screening process, where they have taken center stage in many studies. 153,156 An in silico approach provides researchers with a method to design promising drug or lead candidates or repurpose existing drugs that show inhibitory activity based on the rationale of structure-based drug designing which targets the essential structural features of a protein and identifies the potent inhibitors based on the concept of estimated free energy of binding and the formation of various intermolecular interactions such as hydrogen bonds, hydrophobic interactions and van der Waals interactions. 146
Rutin as a SARS-CoV-2 Mpro Inhibitor
The SARS-CoV- main protease (Mpro), a chymotrypsine like hydrolase, is an attractive target for anti-CoV drug design, due to its responsibility for the maturation of key functional enzymes such as replicase and helicase. 157 It is crucial for the processing of the coronavirus replicase polyprotein (P0C6U8), cleaving it at 11 conserved sites. It employs a cys-his catalytic dyad in its active site where the cysteine sulfur is the nucleophile and the histidine imidazole ring is a general base. This nonstructural protein of coronavirus is responsible for processing the polyprotein translated from viral RNA. 158 It has been confirmed that viral replication can be inhibited by Mpro inhibitor in SARS-CoV. 159,160
It has also been shown that strong Mpro inhibitors can substantially reduce SARS-Cov-2 virus titers, reduce weight loss, and improve survival in mice, 161 making Mpro an emerging and promising drug target for structure-based drug discovery. 161
A screening of 38 natural flavonoids against the active site of SARS–CoV-2 Mpro protein by molecular docking and molecular dynamics suggested that strong interaction occurs between flavonoids with sugar moieties and Mpro. 162 In this study, it has also been concluded that compounds that form bonds with the highest strength usually bind to Thr26, Ty54, His41, Asn142, Gly143, Cys145, and Glu166 in the substrate binding subsite for the proteolytic activity, 163,164 leading to strong binding due to hydrogen bonds. Mpro consists of 3 domains and substrate binding site residues in a cleft between domain I and II, while domain III is involved in catalytic function. Dimerization of the protein is required for its catalytic regulation. 165 Glu166 residue is a key amino acid involved in the dimerization of Mpro and the creation of the substrate binding pocket. 166,167 Cys141 and His41 residues form a catalytic dyad on the active site of the protein essential for its catalytic function. Similar results were recently obtained by molecular docking with different molecules against the Mpro protein. 165,168 Rutin has been found to interact with Thr26, Tyr54, Leu141, and Glu166 via conventional H-bonds, π-cation interactions with His41, π-alkyl interactions with Met49, π-sulfur interactions with Cys145, and C-H bond interactions with Met165.These results indicate that rutin possesses high docking scores with – 9.2 kcal/mol. 162 The binding of the flavonoids at the protease active site is clearly structure dependent, and the nature of the sugar is also primordial to the activities, as flavonoids with a rhamnose possess the strongest binding affinity. As flavonoids with such a moiety undergo either no or little modification in the intestine, they might be a good candidate for COVID-19 treatment. 162
A blind molecular docking approach to identify possible inhibitors of the SARS-CoV-2 main protease (PDB: 6Y84) was undertaken by screening 33 molecules, which included natural products, antivirals, antifungals, nematocidals, and antiprotozoals; rutin was identified as one of the important compounds. This undergoes several non-covalent interactions with the residues within the active site and is stabilized through hydrogen bonding with HIS41, LEU141, ASN142, GLU166, THR190, and GLN192; furthermore, it undergoes π-sulfur and π-alkyl interactions with CYS145 and PRO168, respectively. 169
According to virtual binding strategy with Mpro, rutin was found to have a docking score of −9.16 kcal/mol and an AUC (Area Under ROC Curve) of 0.990, and, therefore, was considered to be a potential Mpro inhibitor. Rutin was predicted to form hydrogen bonds involving Cys145 (2.63 Å), Asn142 (2.1 Å), Gly143 (2.3 Å), and Thr190 (2.35 Å), with additionally the possible formation of σ-π stacking interaction with Gln189. Notably, the major binding affinity was based on the presence of a hydroxyl group, which presented the key to anchoring and blocking the substrate into the active pocket of the catalytic center. 170
LeDock results demonstrated that rutin depicts the lowest predicted binding energy in the active pocket of Mpro (−8.67 kcal/mol), even lower than that of the reported positive inhibitor remdesivir. This may be because the phenolic hydroxyl group in flavonoids, especially that in the flavonoid glycosides, binds more easily with the heteroatoms of the amino acids of Mpro. Rutin forms multiple hydrogen bonds with the main chain of residues like Phe-140, Glu-166, Thr-26, Leu-141, Ser-144, Cys-145, and His-163. In particular, Asn-142 and Gln-189 were thought to contribute to the hydrophobic interactions with rutin. 171
Docking studies with more than 51 constituents of Juniperus procera Hochst, used as a traditional medicine for recovery from flu in Saudi Arabia, showed that, of all the tested compounds, rutin gave the highest interaction score with Mpro. It showed a better inhibition score in 9 of the 11 homology models. Analysis of ligand-protein interaction contacts revealed that 3 residues (Glu166, Gly143, and Thr45) of Mpro formed hydrogen bonds with rutin, thus suggesting it to be a possible effective inhibitor of several COVID-19 protein targets. 172
Five bioactive molecules from FuFang Zhenzhu Tiaozhi (FTZ) (lithospermic acid B, specnuezhenide, neonuezhenide, rutin, and neodiosmin) were predicted to target Mpro. Docking results suggested that the flavonol group of rutin forms hydrophobic bonds with MET49 and CYS145 as a key force to bind with the 3 Mpro protein with a CDOCKER interaction energy of −69.38 kcal/mol. 160 In addition, the hydroxyl groups of rutin interact with THR26, LEU141, GLU166, and ASP187 through hydrogen bonds, whereas the carbonyl group of rutin can form hydrogen bonds with MET165 and GLU166 residues. Moreover, rutin can also form van der Waals interactions with 17 amino acid residues, including THR25, LEU27, TYR54, and THR190. Based on the above observations, involvement of rutin in inhibiting SARS-CoV-2 Mpro was hypothesized. 173
Virtual screening (obtained by AutoDock VINA), molecular docking (obtained by AutoDock 4.2.6) results, and ROC probability of compounds binding to spike protein reflect that rutin exhibits hydrogen bonding with amino acid residues Asn343, Ser373, and Asn437, and interactions with Phe374, Ala372, Ser371, and Val367. 163 In the same study, for binding to 2’-o-ribosemethyltransferase, rutin showed hydrogen bonding with Leu100, Tyr132, and Lys135, and interactions with Gly148, Ph149, Cys115, Asp114, Asp99, and Gly71. 174 In another study, after evaluating 80 flavonoids for MD docking with the “SARS-CoV-2 protease 3CLpro (PDB: GLU7),” hesperidin, rutin, and diosmin were the top 3 candidates for docking at the active site. 175 In a detailed study of 72 flavonoids, rutin has been identified as a top hit from multiple in vitro inhibition and in silico docking investigations of the main protease 3CLpro of SARS-CoV and SARS-CoV-2. 176 Using docking and molecular dynamics simulation methods, the dynamics and energetics of rutin when interacting with the Mpro of SARS-CoV-2 has also been investigated suggesting that the highly hydrophilic rutin molecule can be bound inside the Mpro pocket (active site) and possibly inhibit its biological functions. 177
Rutin as a Helicase Inhibitor
Based on structure modeling of helicase protein, rutin was predicted to be a helicase inhibitor with high mfScores through virtual ligand screening, and thus its antiviral significance against SARS-CoV-2 infections has been suggested. 178
The presence of rutin in several traditional antiviral medicines prescribed in China for treating early stage COVID-19 infected patients justifies its antiviral significance. For example, Lianhuaqingwen (LHQW) has been used to treat COVID-19 patients and demonstrated outstanding control of the disease, which is related to inhibition of the replication of SARS-CoV-2 in vitro. 179 Based on a data mining approach, strong connections between rutin and quercetin with several elements within the network associated with the drug/compound-host molecules interaction network, the use of Hedera helix L. and other species containing flavonoids has been suggested for use against influenza, as folk medicines. 180
Rutin as an ACE2 Inhibitor
Several studies have reported that SARS-CoV–2 enters its target cells through angiotensin- converting enzyme 2 (ACE2). 115,165 This is highly expressed in the mouth and tongue, which facilitates the entry of the virus into the body. 115 Blocking of the ACE2 receptor could reduce its binding ability to viral spike protein attachment. Rutin has been identified as one of the three compounds (glycyrrhizin, rutin, and pheophorbide A) with a high binding affinity –6.9 kcal/mol to the ACE2 receptor. 166 It created an interaction with the following residues: PHE4, LEU29, ASP30, ASN33, VAL93, ALA99, LEU100, ALA387, ASP350, ALA387, GLN388, PRO389, LEU391, and ARG393. Some of them (such as ASP30) have an essential role in RBD binding. 167
Rutin as a Host Toll-Like Receptor Inhibitor
The host toll-like receptors (TLRs) play an important role in mediating the inflammatory response and host-based anti-CoV activity. The pocket site of TLR2, TLR6, and TLR7 presented potential combinations between rutin with binding energies of <-8 kcal/mol. 171 TLRs generally stimulate pro-inflammatory and antiviral host pathways. These potential bindings indicate two possible activities: antagonistic and stimulatory. For patients with COVID-19, this may provide a dual benefit, both preventing over-inflammation and restoring innate antiviral immunity. 123
Therefore, herbal medicines containing rutin and other flavonoids, such as quercetin, 107 diosmin, 181 and hesperidin 177 as major components might be meaningful for the treatment of SARS-CoV-2 infections. Since rutin is already present in FDA approved vitamin C supplements, it can be taken as a preventive medicine.
Conclusion
With regard to SARS-CoV-2 (COVID-19), it is well known that its main protease (Mpro) plays an important role in maturation of many viral proteins. The molecular dynamics (MD) simulation and docking studies of rutin binding with Mpro are encouraging and offer potential for its drug repurposing. Overall, rutin matched very well with the 6GLU7 binding pocket, indicating that it may be a potential inhibitor as it is able to form several hydrogen bonds and σ-π stacking interactions with various amino acids of Mpro in anchoring and blocking the substrate into the active pocket of the catalytic center. The Mpro inhibiting activity of rutin should therefore be considered potential for antiviral clinical trials and/or for combined therapy involving antiviral +rutin (or other flavonoids), assuming synergistic effects of combined therapy.
Footnotes
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
