Abstract
Cortex Lycii, the root barks of Lycium barbarum and L. chinense, known as “di gu pi” in traditional Chinese herbal drugs, is an important ingredient of formulations used for treating a variety of diseases. During the last 3 decades, more than 70 chemical entities have been separated and purified from either the aqueous or aqueous ethyl alcohol extracts of Cortex Lycii. In this study, high-performance liquid chromatography together with quadrupole-time-of-flight mass spectrometry (MS) was employed to explore new analog structures from aqueous ethyl alcohol extracts (50%, v/v), which led us to discover 4 new phenolic amides and a new cyclic peptide. The structure-based manual screening method, on the basis of the analysis of the fragmentation pathway of the previously known compounds, was used to make a preliminary analysis of the negative total ion chromatography and negative extract ion spectra. Three ions at m/z 472.1, 314.1, and 445.2 were assigned to phenolic amides, and by further analysis of their MS/MS data, the structure of 1, corresponding to one of them (m/z 314.1), was illustrated as an analog of the known compound KN1. A parent ion at m/z 856.1 was assigned to a cyclic peptide analog (
The dried root barks of Lycium barbarum L. and L. chinense Mill. (Solanaceae), called Cortex Lycii, and known in China as “di gu pi”, are important herbal drugs in traditional Chinese medicine (TCM). They are used as ingredients of formulations to treat various diseases, such as diabetes mellitus, night sweats, coughs, hematemesis, hypertension, and ulcers in China, Japan, and Korea. 1,2 Previous phytochemical studies on this drug have reported more than 70 compounds assigned to various classes including alkaloids, 3 -5 phenolic amides, 6,7 cyclic peptides, 8,9 dipeptide, 10 cyclic diterpenes, 11 and pentacyclic triterpenoids 12 from either the water or (and) ethanol extracts (Figure 1). The crude extracts and chemical monomeric entities from Cortex Lycii showed various pharmacological properties including antioxidant, free radical scavenging, 6,13 antitumor, 14 antimicrobial, antifungal, 7,9,15 antiviral, 14 antidiabetic, 3 and antihypotensive 4,8 activities.

The chemical structures of known compounds.
For the acceleration of modernization of the TCM process and better understanding the mechanism of pharmacological action at the molecular level, studies on the complex chemical constituents of TCMs have become one of the hottest subjects. Previous works indicated that the cyclic peptides, lyciumins A and B from Cortex Lycii, showed activities against angiotensin converting enzyme and angiotensin 4,8 and were expected to be applied in treating clinical hypertension. Lyciumins, bearing a unique cyclic chemical skeleton with a rare linkage between the indole N1 of tryptophan (Trp) and the Cα of glycine, has continuously attracted the attention of chemists and pharmacologists for a long time.
Natural products have continually been the most important source for the development of new medicines for their attractive complex structures and noticeable pharmacological activities. 16 -19 Discovering novel chemical structures from natural resources could play critical roles in enriching the chemical molecular library of natural products and also could provide fundamental skeletons for the designing of new structural drug molecules by chemical synthesis. In the past decades, with the development of mass spectrometry (MS), especially tandem mass spectrometry (MS/MS), numerous natural product structures have been discovered. In our previous work, 20 new structures were discovered on the basis of a manual screening and automatic networking workflow through a reanalysis of the liquid chromatography (LC)-MS/MS data for the fermentation broth of Streptomyces sp. SS. 20,21
Early studies on Cortex Lycii provided fundamental chemical structures for discovering novel components in trace amounts by high-performance liquid chromatography (HPLC)-(MS). Especially based on the MS/MS fragment patterns of known compounds, the analogs’ structures could be established by analysis of their similar secondary fragment ions or neutral losses, and by comparisons of their MS/MS fragmentation behaviors with those of known compounds (Figure 1). In this study, an HPLC-quadrupole-time-of-flight (Q-TOF)-MS method was employed to explore novel structures from the 50% ethanol (EtOH) aqueous extract of Cortex Lycii; 4 new phenolic amides (
Experimental Materials and Methods
Chemicals and Materials
Methanol of HPLC grade was supplied by Titan Scientific Co., Ltd. Shanghai, China. Watsons water repurified with a Milli-Q academic water purification system (Millipore, USA) was used for HPLC analysis. Acetic acid, ammonia, and ammonium acetate were of analytical reagent grade from Beijing Chemical Reagent Plant (Beijing, China).
A Sep-Pack C18 cartridge, the solid phase extraction column, was purchased from Waters Co., Ltd. (Milford, MA, USA). A CAPCELL PAK C18 AQ (4.6 × 250 mm, 5.0 µm, Shiseido, Kyoto, Japan) chromatography column was used for HPLC–MS/MS analysis. HPLC–MS/MS data were acquired using an Agilent 1100 (LC) in tandem with an Agilent 6510 Q-TOF MS (Agilent Technologies, Santa Clara, CA, USA).
Plant Material
Cortex Lycii, the root barks of Lycium barbarum and L. chinense (50 Kg), was purchased from Tong-Ren-Tang Pharmacy, Xihongmen shop, Xihongmen Town, Daxing District, Beijing City of People’s Republic of China, in December 2018. The material was identified by Professor Yufeng Ren of the school of biological sciences and engineering of North Minzu University, and the voucher specimen was numbered as NMUDGP-20181218 in plant specimen room.
Sample Preparation
Air dried, powdered Cortex Lycii (100 g) was extracted on an ultrasonic water bath with 100 mL of 50% aqueous EtOH at 20°C for 3 × 2 hours. The extract was evaporated to dryness under reduced pressure, and the residue, as a 2.0 mL water solution, was passed through a Sep-Pack C18 cartridge; the adsorbed components were eluted with purified water, 50% methanol aqueous solution, and HPLC grade methanol to yield samples 1-3. All samples were centrifuged for 10 minutes at 12 000 rpm at 4°C, and the supernatant used as the test samples.
HPLC-MS/MS Analysis
A 2-phase gradient elution system including phase 1 and phase 2 was used for the HPLC-Q-TOF-MS procedure. HPLC grade methanol was used in phase 1, while in phase 2, a solution of 0.1% ammonium acetate solution (pH 8.0) was employed. Linear gradient started with 10% mobile 1 and then reached 90% over 70 minutes. The flow rate was set to 0.8 mL/min, and the effluent was equally separated by a 3-way joint into the diode array detector (DAD) detector and the Q-TOF detector, respectively. The DAD was monitored at 254 nm, and the column temperature was maintained at 25°C. Twenty microliters of the test sample was injected.
The MS data were acquired with an electrospray ionization source; the capillary voltage was 4 kV, the capillary outlet voltage 135 V, the taper hole voltage 65 V, the desolvent temperature 350°C, the dry airflow speed 10 L/min, the atomizer pressure 40 psi, the collision energy 8-45 V, the scanning range m/z 100-2000, and the detection mode was negative ion (ESI−).
The molecular network was created following standard GNPS’s workflows provided in the website of https://gnps.ucsd.edu/ProteoSAFe/static/gnps-splash.jsp. In brief, raw data were transformed into mzXML format using the MS Convert program of ProteoWizard 3.0 and then analyzed using GNPS. The parameters are presented in supplemental Table S1.
Results and Discussion
Extraction of the Molecular Ions of Known Compounds
In order to investigate the new structural analogs, the accurate molecular ions of known phenolic amides (KN1–KN9, series 1, Figure 1) and cyclic peptides (KN10–KN13, series 2, Figure 1) were extracted from the negative total ion chromatograms (−TIC) of samples 1–3. The negative extracted ion overlaid spectra (Figure 2) showed that almost all the known compounds could be extracted (abundances > 105), except compounds KN3 and KN4. The abundances of compounds KN1 and KN9 were higher than other known compounds, which verified that phenolic amides are the main chemical components of Cortex Lycii (Figure 2(a)).

In samples 1-3, extract ion spectra (EIC) of known compounds of (a) m/z 300.1(KN1), 312.1(KN2), 632.2(KN3), 639.2(KN4), 634.2(KN5), 618.2(KN6), 641.2(KN7), 609.2(KN8), 476.1(KN9) and (b) 874.3(KN10), 895.3(KN11), 898.3(KN12), 962.3(KN13), 443.2(KN14) (b) in negative mode [M − H]−.
Figure 2(b) shows the extracted ions belonging to 4 cyclic peptides (KN10–KN13). The different abundance of peaks corresponding to KN10–KN13 indicated that lyciumin A (KN10) is the main cyclic peptide in Cortex Lycii. The ordinate difference between Figure 2(a) (<3.0 × 106) and 2(b) (<1.0 × 106) indicated that the contents of cyclic peptides were lower than those of phenolic amides. The peak for the dipeptide of lyciumamide (KN14) was lower than that of the cyclic peptides, indicating that the dipeptide is in trace amount in the extract of Cortex Lycii.
Fragmentation Patterns of Phenolic Amides and Cyclic Peptides in Negative Mode
Fragmentation behaviors in positive MS mode of phenolic amides and cyclic peptides have been discussed in the literature. 22 The fragment sites of phenolic amides mainly occurred at or around the amido bond in positive mode. However, for cyclic peptides, the losses of side chains were the main MS/MS features. Herein, we established the fragmentation behaviors of those compounds in negative mode by detailed analysis of the MS/MS of known compounds, especially on the basis of the MS/MS analysis of KN1, KN2, KN7, KN9, and KN10.
The fragmentation behaviors of KN1and KN2
The chemical structure of KN1 is very similar to that of KN2 except for the difference of a methoxyl group and a double bond, which is conjugated to the carbonyl group (Figure 1). Herein, the MS/MS of the ions at m/z 300.1 and 312.1 for KN1 and KN2 were extracted, respectively (Figure 3(a) and (b)), and the fragmentation patterns for KN1 and KN2 were established based on their reasonable neutral losses (Figure 3(a-1) and (b-1)). For KN1 (Figure 3(b)), the higher abundant fragment ions at m/z 178.1 and 121.1 indicated that the losses in the negative mode were mainly induced by the negative center of the 3,4-dihydroxyphenyl group, while for KN2, a 3-methoxyl-4-hydroxyl-substituted mode reduced the acidity that was caused by the intramolecular hydrogen bond of 4-OH with 3-OMe. A special fragment ion at m/z 197.1, which was produced by losing a CH3 +, gave support to the negative charge center around this phenyl also being an important active site. Also, the ions at m/z 190.1 and 135.1 could be determined by the neutral losses of a 3-methoxyl-4-hydroxylphenyl and 3-methoxyl-4-hydroxylphenyl acryloyl group, respectively (Figure 3(b-1)). The fragment ions at m/z 178.1 and 148.1 should be induced by the negative center around the 4-hydroxylphenyl group.

The tandem mass spectra of m/z (a) 300.1 (KN1) and (b) 312.1 (KN2) and the fragment patterns for (a-1) KN1 and (b-1) KN2.
The fragmentation behaviors of KN5 and KN7
The MS/MS extracted chromatograms of the molecular ions of KN5 and KN7 are presented in Figure 4(a) and (b), respectively, and the fragment patterns for KN5 and KN7 in Figure 4(a-1) and (b-1). Comparison of the chemical structures of KN5 and KN7 indicated that the acetylbutylammonia group in KN5 could provide another negative charge center that reduces the cleavages concentrated on the terminal sites and to produce ions at m/z 427, 442, 455, and 471 in a relatively lower abundance. Fragment ions at m/z 478, 463, and 299 were the common ions of KN5 and KN7, which indicated that these could be used as the characteristic ions for determining the chemical analogs of phenolic amides bearing the identical substituted stilbene group.

The tandem mass spectra of m/z 634.2 (KN5, a) and 641.3 (KN7, b) and the fragment patterns for (a-1) KN5 and (b-1) KN7.
The fragmentation behaviors of cyclic peptides
The negative mode fragmentation behaviors of cyclic peptides in Cortex Lycii have not been studied before. Herein, the MS/MS of the molecular ions at m/z 872.4, 895.4, and 962.4 for KN10, KN11, and KN13 were extracted as shown in Figure 5(a), (b), and (c), respectively. Due to the lower abundance of the ion for KN12 (<10−4), the MS/MS data for KN12 were not collected. Based on the reasonable losses for peptide compounds, the fragmentation patterns for KN10, KN11, and KN13 were provided as Figure 5 (a-1) (for KN10), (b-1) (for KN10), and (c-1) (for KN13). The loss of HCHO from the threonine (Thr) residue in the cyclotetrapeptide skeleton was the dominant neutral loss in the fragmentation patterns of cyclic peptides. The neutral loss of 208 Da in KN10 (M-664), KN11 (M-687), and KN13 (M-754) could be used as the evidence for determining the presence of a proline dipeptide moiety in the branched peptide chain. The 90 Da difference between the ions at m/z 591 and 574 in KN13 with those at m/z 501 and 484 in KN10 and KN11 could be caused by the different amino acids in the cyclotetrapeptide skeleton, a phenylalanine (Phe) in KN13 replacing the glycine (Gly) in KN10 and KN11. For the same reason, the difference of 23 Da corresponding to the chain peptide residue of 371 Da (M-501) in KN10 with that of 394 Da (M-501) in KN11 was identical to the difference in the molecular weight (MW) of tryptophan (Try, 204 Da) and tyrosine (Tyr, 181 Da).
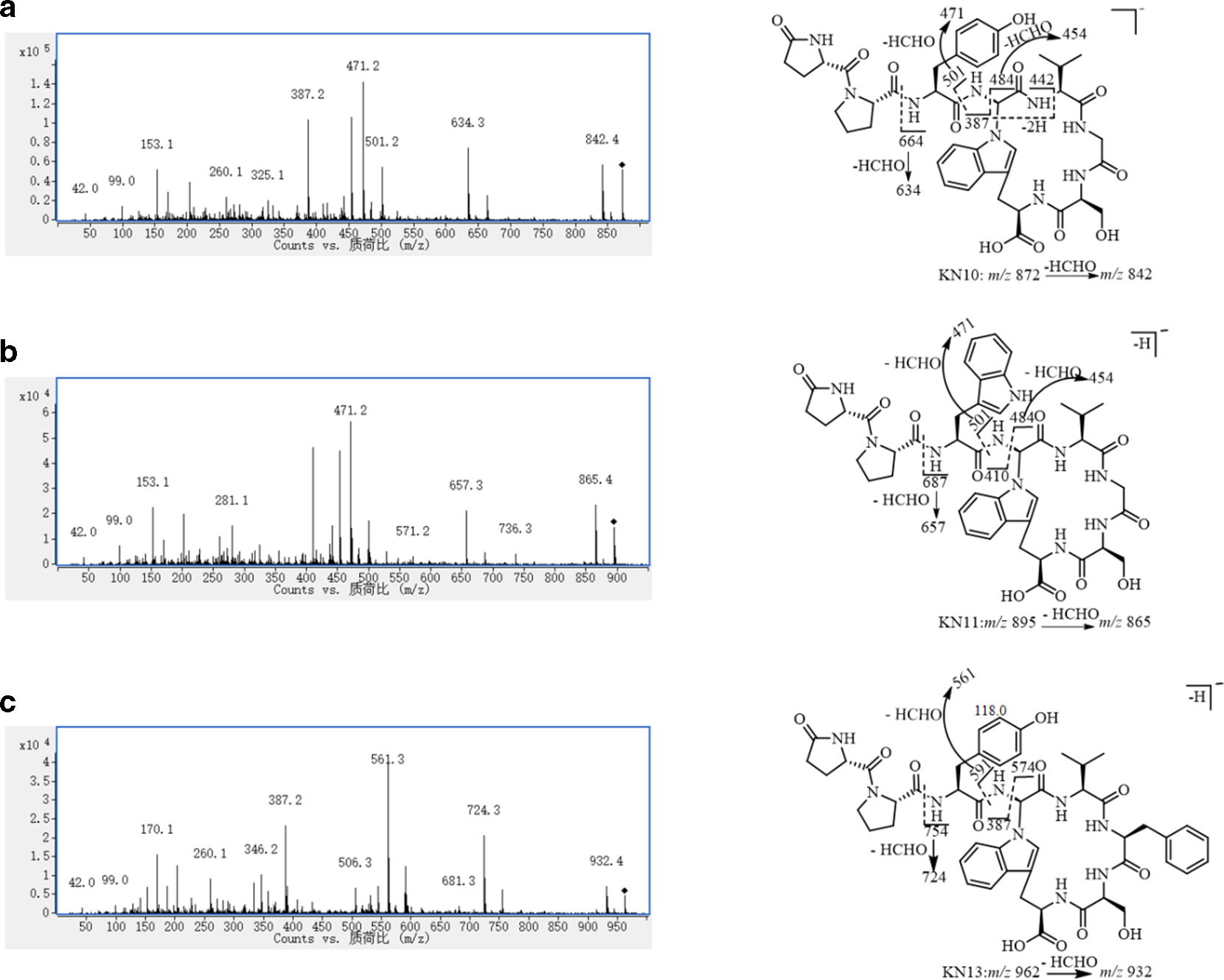
The tandem mass spectra of m/z (a) 872.4 (KN10), (b) 895.4 (KN11), and (c) 962.4 (KN13) and the fragment patterns for (a-1) KN10, (b-1) KN11, and (c-1) KN13.
Manual Exploration of the New Structural Chemical Constituents
The characteristic ions at m/z 478.1, 463.2, and 299.1 were extracted in the total ion chromatography (TIC) of samples 1–3 for exploring new structural analogs of phenolic amides (Figure 6(a)). The peaks with retention times of 23.5 and 29.9 minutes corresponded to compounds KN1 and KN7, respectively. In addition, 3 new peaks at 22.5, 26.0, and 30.0 minutes with molecular ions of m/z 472.1, 314.1, and 445.2 displayed similar MS/MS fragment ions to those of known phenolic amides. Detailed analysis of their MS/MS spectra, especially of comparisons of their neutral losses and characteristic ions, a structure corresponding to one of them (m/z 314.1) was determined, as shown in Figure 7(a). However, the chemical structures corresponding to the ions at m/z 472.1 and 445.2 could not be illustrated on the basis of their MS data. The MS/MS for the ions at m/z 472.1 and 445.2 are provided in the supplemental Figures S1 and S2, respectively.

Extract fragmentation ions for exploring analogs of phenolic amides (a) and cyclic peptides (b).

The tandem mass spectra for compounds 1 (a) and 2 (b) and the fragmentation for (a-1) 1 and (b-1) 2.
The characteristic ions, including those at m/z 501.2, 471.2, 387.1, and 410.1, were used as common ions for exploring cyclic peptides. Figure 6(b) shows the extracted peaks for the known KN10, KN11, and KN13 at retention times of 22.5, 24.1, and 27.3 minutes, respectively. In addition, a new ion at m/z 856.2, corresponding to the peak at 25.2 minutes in sample 2 (P4 marked *), displayed similar fragment ions to those of the known compounds in Figure 5, especially the presence of the ions at m/z 501.2, 484.2, 471.2, and 454.2 in relative clear abundance, which indicated that this compound also has a cyclotetrapeptide skeleton identical to that of KN10 and KN11 (Figure 7(b)). However, a fragmentation ion at m/z 371.1 corresponding to the side chain was 16 Da lower than that in KN10 (m/z 387.1) suggesting the lack of 1 oxygen atom in this new compound. A similar neutral loss of 208 Da (M-648) corresponding to the diproline residue indicated that the new compound also bore this structure at the terminal of the side chain. Therefore, the difference between compound P4 with KN10 should occur at the moiety of the Tyr, most likely replaced by a phenylalanine (Phe). This speculation was supported by the fragmentation ion of a Phe-cyclotetrapeptide at m/z 648.2 in this compound rather than a Tyr-cyclotetrapeptide (m/z 664.2) in KN10.
Automatic Exploration of New Structural Chemical Constituents Through Networking Workflow
The HPLC-MS/MS data were reanalyzed on the GNPS platform, and the results disposed in visualization networking (Figure 8). A total of 527 nodes were profiled in the molecular net and 33 sets were composed of at least 3 molecular nodes. In this networking map, the molecular ions corresponding to KN1–KN5, KN7, KN10, KN11, KN13, and KN14 appeared in different sets. Moreover, the complex ions of KN10-Cl− (m/z 908.4), KN11-Cl− (m/z 931.4), and KN13-Cl− (m/z 998.4) were also detected. By detailed analysis of this network, new compounds

The network (a) and the selectively amplifying sets signed in red cycle in (a), for discovering new structures (b).
The molecular ion for

The tandem mass spectrum of compound 3 (a) and the fragmentation pathway for 3 (b).
The molecular ion of
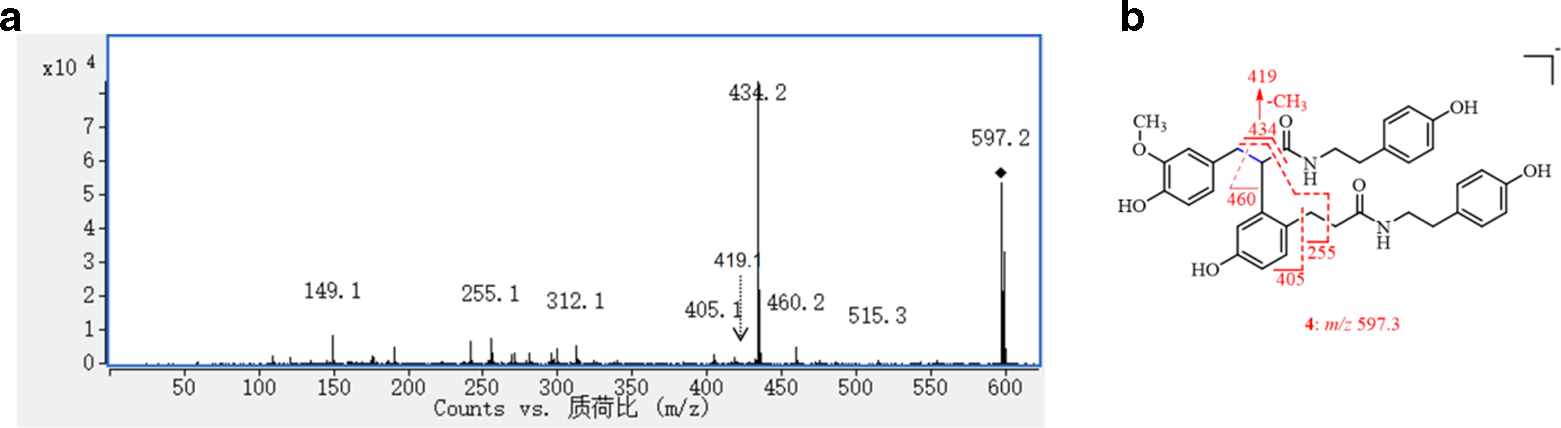
The MS/MS spectrum of compound 4 (a) and the fragmentation pathway for 4 (b).
The 30 Da lower molecular weight of

The tandem mass spectrum of compound 5 (a) and the fragmentation pathway for 5 (b).
The above work described two methods of profiling MS/MS data for exploring new structures. To date, numerous new technologies and methods have been developed for a detailed investigation of natural products with new structures. Systematic isolation and separation of natural products from natural sources (especial of plant materials) have been intensively investigated during the last century, and with the help of modern structural analysis technologies, the absolute chemical structures of numerous natural products have been determined, and quite a few of them are in clinical use, either as they are or after structural modification. However, it is becoming more and more difficult to discover new series of chemical structures from known resources, most likely due to repeated discoveries and due to their very trace contents. Therefore, using more sensitive and efficient equipment and technology has become the main choice to explore new natural products. HPLC–NMR, 23 HPLC–MS–NMR, 24 and HPLC–MS/MS 25 were designed to analyze systematically TCM, and microbial secondary metabolites, as well as drug metabolic pathways of important steps in drug discovery.
Automatic profiling of huge MS/MS data in the network is an auxiliary tool for the rapid screening of new compounds by clustering compounds with specific MS/MS ions and neutral losses. 26 The molecular networking method is a visualization tool and was developed as a public networking platform (https://gnps.ucsd.edu/ProteoSAFe/static/gnps-splash.jsp). This aims to be an open-access knowledge base for community-wide organizations and the sharing of raw, processed, or identified MS/MS spectrometric data. In this work, by manual analysis, together with the aid of the GNPS platform, the HPLC-Q-TOF-MS data of Cortex Lycii was investigated that has led us to discover 4 new phenolic amides and 1 new cyclic peptide analog. Although in the past work, more than 70 compounds have been found from extracts of Cortex Lycii, there are still a large number of mass spectra in the visualization map (Figure 8) that have not been assigned and may correspond to new types of structures.
Conclusions
A structure-based manual screening and automatic networking of HPLC-Q-TOF-MS data developed in the analysis of a TCM for systematically exploring new structures led to the discovery of 4 new phenolic amides (
Supplemental Material
Supplementary material - Supplemental material for Exploring Chemical Structures From Cortex Lycii, Based on Manual and Automatic Analysis of the HPLC-Q-TOF-MS Data
Supplemental material, Supplementary material, for Exploring Chemical Structures From Cortex Lycii, Based on Manual and Automatic Analysis of the HPLC-Q-TOF-MS Data by Zhi-Bo Jiang, Yong-Xin Chen, Jing-Zhi Chen, Xing Lu, Xin Guo, Bing-Zhen Ma, Chong-Long Li, Xuan Fang, Yong-Hong Tang and Xiao-Li Ma in Natural Product Communications
Footnotes
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was co-funded by: the Key Scientific Research Projects in 2017 at North Minzu University [grant number 2017KJ16]; the National Natural Science Foundation of China [grant number 81603006]; the Natural Science Foundation of Ningxia Province [grant numbers 2019AAC03113 and 2018AAC03116] and the Scientific Research Start-up project for Recruitment Talents of North Minzu University in 2019 [Financial Number 113159150].
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
