Abstract
A phytochemical study on the bark of Endiandra kingiana Gamble (Lauraceae) led to the isolation of a new benzofuranone, 4-hydroxy-6-(9,13,17-trimethyldodeca-8,12,16-trienyl)-2(3H)-benzofuranone (
Endiandra, a genus belonging to the family Lauraceae, comprises approximately 125 species found throughout the tropics. 1 Only 8 species, Endiandra kingiana, Endiandra anthropophagorum, Endiandra oligandra, Endiandra introrsa, Endiandra jonesii, Endiandra monothyra, Endiandra baiilonii, and Endiandra xanthocarpa, had undergone scientific studies that resulted in the isolation of endiandric acids, lignans, neolignans, and sesquiterpenes. 1 -4 The previous studies on E. kingiana by Azmi et. al 5 led to a series of endiandric acid analogs named kingianic acids A to I. These compounds were screened for Bcl-xL and Mcl-1 binding affinities, as well as cytotoxic activity on various cancer cell lines. 6
To the best of the authors’ knowledge, there is no scientific report as yet on this species regarding dengue virus related studies. Dengue virus is among the most widespread mosquito-borne diseases and is endemic in most tropical and subtropical parts of the world. 7,8 Malaysia with a population of 31.1 million and a population density of 95 per km2 has seen increasing cases of dengue over the past two decades with 49 726 cases. 9 There are 4 serotypes of dengue viruses (DENV1, DENV2, DENV3, and DENV4) that cause dengue hemorrhagic fever. Dengue virus type 2 is the most prevalent one of all serotypes in Malaysia. 9
In our search for bioactive compounds from Endiandra species against dengue virus, we carried out a phytochemical study of the bark of E. kingiana. A new benzofuranone (
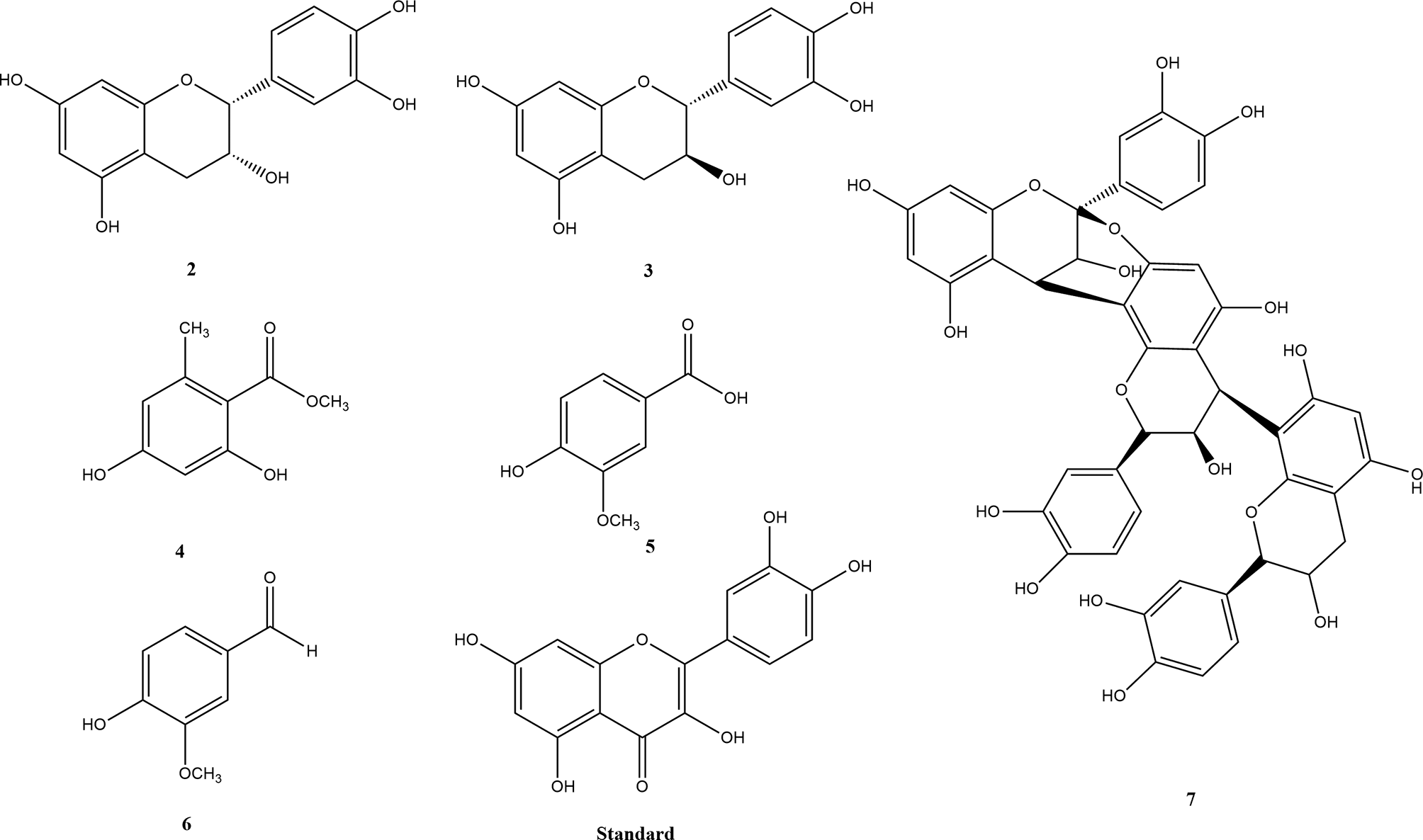
These compounds were evaluated against dengue virus type 2 NS2B/NS3 serine protease. In addition, molecular docking simulations were implemented to probe the binding modes of the active ligands to the amino acids in the active site of serine protease. Herein, we report the isolation and structural elucidation of these compounds, their dengue inhibitions, and their molecular docking.
Compound
The 1H NMR spectrum of
In the correlated spectroscopy (COSY) experiment (Figure 2), the methine proton of H-8 was cross-correlated to the methylene protons of H-7, suggesting that they were adjacent to each other. The in-depth analysis of HMBC experiments supported the location of 4 isoprene units which are connected to the aromatic ring through the correlation of H-7 with C-9, C-5, and C-1. The HMBC correlations of H-3 with C-2′ confirmed the presence of the furanone group which is fused to the aromatic ring through C-1 and C-2. Compound

(a) Structure of 1 from the bark of Endiandra kingiana, (b) selected HMBC (arrows) and COSY (bold lines) correlations of 1.
Compounds
Percentage Inhibition of the Crude Extract and Compounds Isolated From Endiandra kingiana Against DENV2 NS2B/NS3 Serine Protease.

Subsequently, all compounds that exhibited more than 60% inhibition toward the DENV2 NS2B/NS3 protease were further evaluated to determine their respective IC50 values. The IC50 values showed that
Molecular docking studies were also attempted in order to better understand how the chemical groups in these molecules (

Molecular docking of the active compounds with the DENV2 NS2B/NS3 serine protease: (a) 4-hydroxy-6-(9,13,17-trimethyldodeca-8,12,16-trienyl)-2(3H)-benzofuranone 1, (b) (−)-epicatechin 2, (c) (+)-catechin 3, and (d) quercetin (standard).
In conclusion, the phytochemical investigation of E. kingiana bark extract led to the isolation of 1 new benzofuranone compound, 4-hydroxy-6-(9,13,17-trimethyldodeca-8,12,16-trienyl)-2(3H)-benzofuranone (
Experimental
General
Preparative thin layer chromatography (PTLC) and analytical thin layer chromatography (TLC) were performed on commercially precoated aluminum supported silica gel 60 F254 TLC sheets (Merck 5554). The NMR spectra were recorded on BRUKER Advance III NMR spectrometers (400 MHz). The mass spectra were performed on Agilent 6530 LCMS-IT-TOF. The UV spectra were obtained on Jasco V530 UV-Vis spectrophotometer. The IR spectra were recorded on a Perkin-Elmer System 400 Fourier Transform Infrared Spectroscopy with methanol.
Plant
The bark of E. kingiana was collected from Kuala Lipis, Pahang. The identity of the plant was determined by Mr Teo Leong Eng and Mr Din Mat Nor, University of Malaya. Voucher specimens were numbered as KL4828 and deposited at the Herbarium of the Department of Chemistry, University of Malaya, Kuala Lumpur, Malaysia.
Extraction and Isolation
The extraction process was carried out using simple maceration method. The dried, ground bark of E. kingiana was first defatted with hexane for 3 days. The hexane extract was filtered and dried on the rotary evaporator. The plant material was dried followed by extraction with dichloromethane (CH2Cl2) for 48 hours and repeated twice. The liquid extracts were dried under reduced pressure using a rotary evaporator to get brown crude extract (15.2 g). The crude extract (8.0 g) was purified by an open column chromatography (CC) eluting with gradient solvent system 90:10 to 30:70 CH2Cl2:CH3OH to give 5 major fractions (A-E). Fraction A (52 mg) was separated by PTLC eluted with a CH2Cl2: CH3OH (98:2) solvent system to furnish vanillic acid (1.1 mg) and vanillin (1.0 mg). Fraction B (162.0 mg) was also purified by PTLC eluted with a CH2Cl2:CH3OH (97:3) solvent system to furnish methyl orsellinate (
In-Vitro Analysis Toward DENV2 NS2B/NS3 Serine Protease
The protease activity assay was conducted at constant concentrations of the protease, substrate, and the compounds with the values of 0.5 µM, 10 mM in dimethyl sulfoxide (DMSO) solution, and 200 ppm, respectively. The reaction mixtures were prepared in black 96-well plates. Each reaction mixture consisted of 200 mM Tris buffer with the total volume of 100 µL. Tris buffer was pipetted to the wells, followed by compounds and by the enzyme. The mixtures were pre-incubated at 37°C, shaken at 200 rpm for 10 minutes, the substrate added to the wells and incubated at 37°C and shaken at 200 rpm for 60 minutes. After the incubation time was completed, the well plate was analyzed, and the fluorescence was detected using the Promega Glomax Multi Detection System microplate reader with excitation and emission wavelengths at 365 and 410 to 460 nm, respectively.
Molecular Docking
A homology model of the DENV2 NS2B/NS3 protease was obtained from the Research Collaboratory for Structural Bioinformatics-Protein Data Bank (RCSB-PDB) database for structure-based design purposes. The selected 3D structure was downloaded in a PDB format file. The enzyme was then prepared under the protein preparation protocol implemented in Discovery Studio 2.5 (Accelry Inc., CA, United States) suite of program. After that, the geometry optimization and energy minimization of DENV2 NS2B/NS3 protease was conducted by removing the water molecules. Then, the protonation of the DENV2 NS2B/NS3 protease was done through “Protonate 3D” feature, followed by optimization of the partial charge and energy minimization. The optimized, minimized 3D structure of DENV2 NS2B/NS3 protease was saved in the .pdbqt format. For the ligands, the two-dimensional structure of
4-Hydroxy-6-(9,13,17-Trimethyldodeca-8,12,16-Trienyl)-2(3H)-Benzofuranone (1)
Pale-yellow amorphous powder.
IR (KBr): 3402, 1725, 1470, 1457 cm−1.
UV/Vis λmax (MeOH) nm (log ε): 210, 275.
1H and 13C NMR: Table 2.
1H (400 MHz) and 13C (100 MHz) NMR Data of Compound 1 in CDCl3.

LCMS-IT-TOF: m/z [M+H]+ calcd. for C23H31O3: 355.2228; found 355.2243
Supplemental Material
Supplementary material - Supplemental material for Chemical Constituents From Endiandra kingiana (Lauraceae) as Potential Inhibitors for Dengue Type 2 NS2B/NS3 Serine Protease and its Molecular Docking
Supplemental material, Supplementary material, for Chemical Constituents From Endiandra kingiana (Lauraceae) as Potential Inhibitors for Dengue Type 2 NS2B/NS3 Serine Protease and its Molecular Docking by Syazreen N. Sulaiman, Maywan Hariono, Haslinda Mohd Salleh, Soon-Lim Chong, Liew Sook Yee, Azeana Zahari, Habibah A. Wahab, Severine Derbré and Khalijah Awang in Natural Product Communications
Footnotes
Acknowledgment
This work was carried out within the framework of the LIA (Laboratoire International Associé) programme established between CNRS, ICSN and the University of Malaya.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was financial supported by University of Malaya research grant, PPP PG137-2013A, High Impact Research - Ministry of Education (HIR-MOE), UM.C/625/1/HIR/MOE/SC/37 and International Funding, the French Embassy in Malaysia on Collaborative Projects On Phytochemical Analysis On Malayan Flora, project number 57-02-03-1007.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
