Abstract
Baicalein is one of the main bioactive compounds in
Diabetes mellitus is a common chronic metabolic disease that causes many complications such as coronary artery, stroke, amputation, renal failure, and blindness, which aggravates our economic and social burden.
1
-3
Although the traditional synthetic α-glucosidase inhibitors (acarbose, voglibose, and miglitol) can effectively control the postprandial blood glucose level of diabetics, their side effects sometimes limit their acceptance.
4
-6
Some naturally occurring flavonoids (eg, rutin, quercetin, and taxifolin) were reported to be strong α-glucosidase inhibitors with potential application.
7
-9
Baicalein (Figure 1) is one of the main flavonoids in

Chemical structure of baicalein.
α-Glucosidase can hydrolyze

HPLC profile for the α-glucosidase assay.

α-Glucosidase inhibitory activities of baicalein (a) and acarbose (b).
To study the interaction behavior between baicalein and α-glucosidase, the effect of baicalein on the fluorescence spectrum (FS) of α-glucosidase at different temperatures (25°C, 30°C, and 37°C) was investigated (Figure 4). Compared with its initial fluorescence intensity (
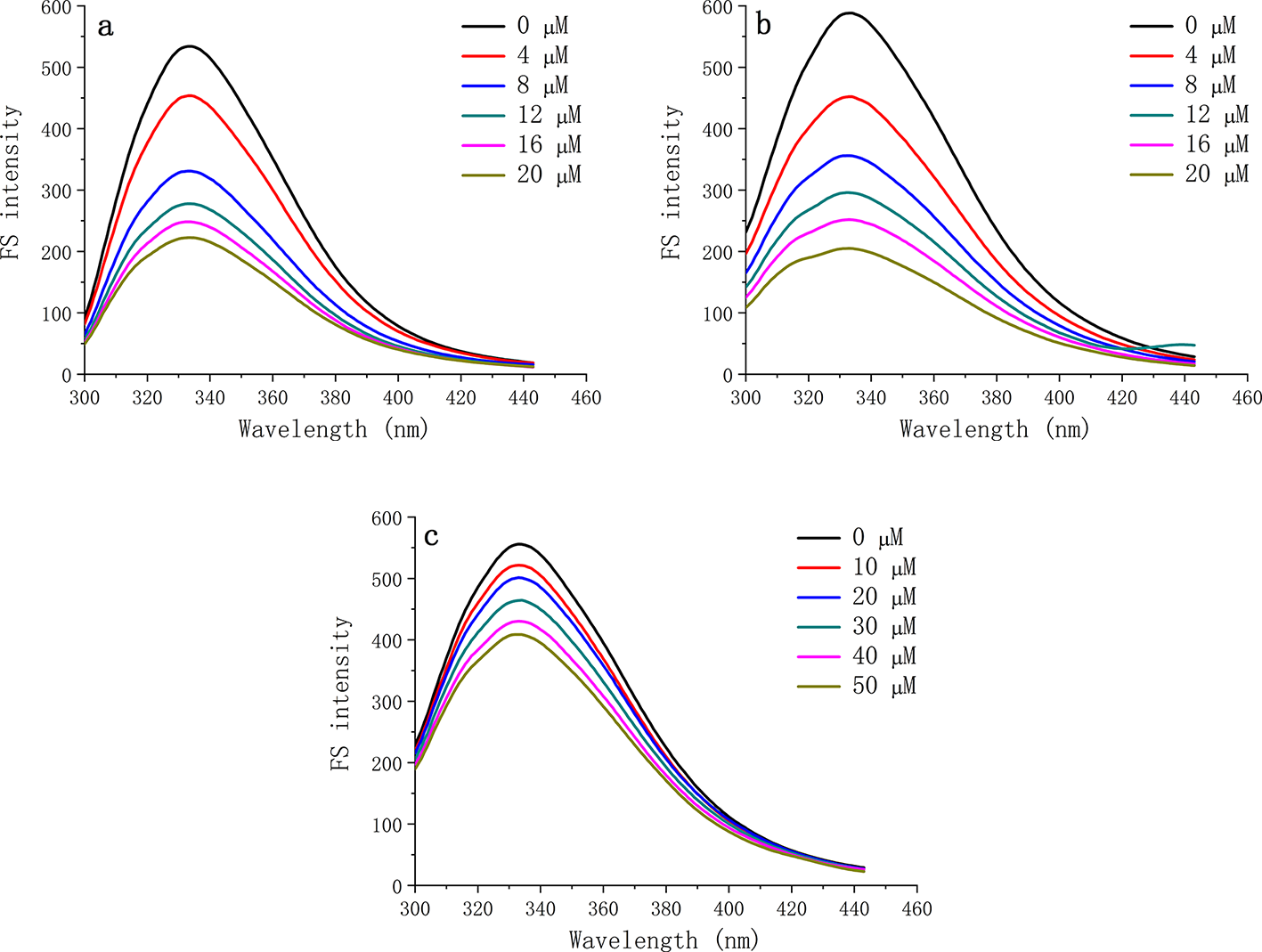
Effects of baicalein on the fluorescence spectrum of α-glucosidase at 25°C (a), 30°C (b), and 37°C (c).

At the designed temperatures, all the
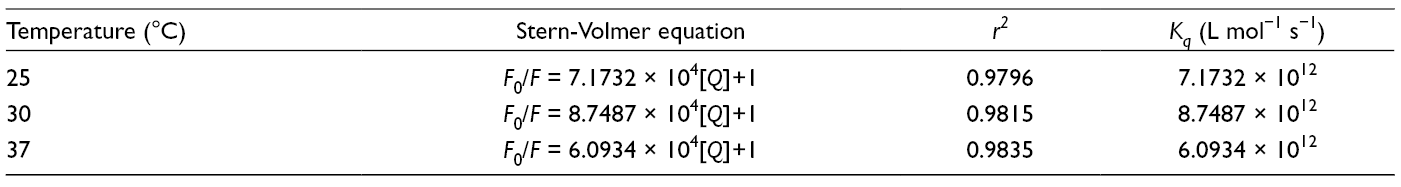
Stern-Volmer Quenching Constants Between Baicalein and α-Glucosidase.

Binding Parameters Between Baicalein and α-Glucosidase.
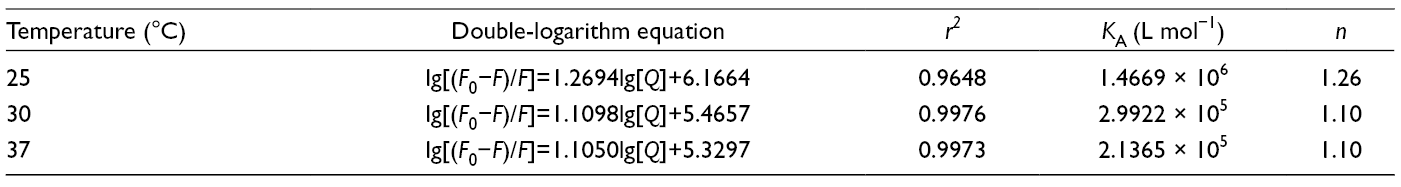
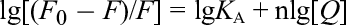
It was found that all the binding site numbers were near to 1. However, the
The thermodynamic parameters describing the interaction were further calculated based on the Vant Hoff equation and Gibbs free energy equation 14 :

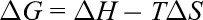
In Table 3, all the free energy changes (Δ
Thermodynamic Parameters Between Baicalein and α-Glucosidase.

Based on the above experimental results, molecular docking was carried out to establish the binding mode. Figure 5(a) provided the binding mode with the lowest binding energy (−7.17 kcal mol−1). Baicalein formed 6 hydrogen bonds with Trp391 (2), Arg428, Gly566, and Glu771 (2) of α-glucosidase. The lengths of these hydrogen bonds were in the range from 1.8 to 3.3 Å. Figure 5(b) shows the hydrophobic interaction around baicalein. The hydrophobic interaction between baicalein and some amino acid residues (Phe385, Phe389, Arg387, Glu429, Phe444, Trp789, and Trp710) could be found, which coincided well with the fluorescence results. Our results show that baicalein is a potential α-glucosidase inhibitor for controlling the postprandial blood glucose level of diabetics, which may be used in the fields of functional foods and medicines.

Binding mode (a) and noncovalent interaction (b) between baicalein and α-glucosidase provided by molecular docking analysis.
Experimental
Chemicals
Baicalein, acarbose, and
α-Glucosidase Inhibition Assay
In total, 0.25 mL of 0.3 U mL−1 α-glucosidase solution (0.1 M PBS, pH 6.8), 0.25 mL of baicalein at different concentrations (0.05-0.3 mg mL−1), and 0.25 mL PBS (0.1 M, pH = 6.8) were mixed and incubated at 37°C for 10 minutes. Then, 0.25 mL of 1 mM pNP-G was added. The obtained mixture was oscillated at 37°C for 20 minutes before 1.0 mL ethanol was added to terminate the reaction. The decrease of pNP-G and the formation of PNP were detected at 405 nm at 25°C by using an Agilent 1260 HPLC system (Santa Clara, CA, USA) with a DIKMA C18 Diamonsil Plus column (250 × 4.6 mm, 5 µm particle size, Beijing, China). The mobile phase consisted of methanol and water (60:40) with a flow rate of 1.0 mL min−1. The injection volume was 10 µL. Both the PNP areas of sample (

Fluorescence Spectra
These were obtained according to a previous report. 15 One milliliter of baicalein at different concentrations (0-50 μM) and 4 mL 0.4 U mL−1 α-glucosidase solution were mixed and incubated at different temperatures (25°C, 30°C, and 37°C) for 10 minutes. Then, the FS in the range of 290 to 450 nm was recorded at the excitation wavelength of 280 nm. Both the slit widths of excitation and emission were 5 nm.
Molecular Docking
The chemical structure of baicalein was optimized by the PM6-D3H4 method of MOPAC 2016, and the crystal structure of α-glucosidase (PDB ID: 4J5T) was used. The molecular docking was performed by using Auto-dock 4.2 software (The Scripps Research Institute, La Jolla, USA).
16
The grid was run with Auto-grid module. The grid spacing was set at 0.375 Å, and all the numbers of grid points in
Statistical Analysis
The results are expressed as mean ± standard deviation of triplicate measurements. Student’s
Footnotes
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the National Natural Science Foundation of China (No. 31771941).
