Abstract
Background:
Poor data availability and accessibility characterizing some research areas in biomedicine are still limiting potentialities for increasing knowledge and boosting technological advancement. This phenomenon also characterizes the field of diabetes research, in which glycemic data may serve as a basis for different applications. To overcome this limitation, this review aims to provide a comprehensive analysis of the publicly available data sets related to dynamic glycemic data.
Methods:
Search was performed in four different sources, namely scientific journals, Google, a comprehensive registry of clinical trials and two electronic databases. Retrieved data sets were analyzed in terms of their main characteristics and on the typology of data provided.
Results:
Twenty-five data sets were identified including data from challenge tests (5 of 25) or data from Continuous Glucose Monitoring (CGM, 20 of 25). As for the data sets including challenge tests, all of them were freely downloadable; most of them (80%) related only to oral glucose tolerance test (OGTT) with standard duration (2 h), but varying for timing and number of collected blood samples, and variables collected in addition to glucose levels (with insulin levels being the most common); the remaining 20% of them also included intravenous glucose tolerance test (IVGTT) data. As for the data sets related to CGM, 7 of 20 were freely downloadable, whereas the remaining 13 were downloadable upon completion of a request form.
Conclusions:
This review provided an overview of the readily usable data sets, thus representing a step forward in fostering data access in diabetes field.
Introduction
The so-called “data tsunami” phenomenon—being defined as the possibility to generate data from multiple sources—is increasingly impacting several fields and in particular the health care field. 1 However, this potential data availability does not always translate into an authentic opportunity for data access; thus, fostering the open data has become a priority in policy initiatives, especially to potentiate the impact of the current artificial intelligence (AI) solutions. 2
Diabetes care is one of the health care areas for which the data tsunami may have a considerable impact. As an example, it is worth mentioning the great amount of data generated from wearable devices, such as continuous glucose monitoring (CGM) devices, insulin pumps, or other devices, which can be incorporated into electronic health records and exploited in digital health technologies like AI or telehealth solutions. 3 However, at the same time, this area suffers remarkably from the lack of available data, 4 thus limiting the potentialities for technological advancements in personalized patient care. 5 This may be due to multiple reasons connected to the nature of the relevant data and the issues in producing and sharing them. First, data acquisition involves sampling of blood or interstitial fluid to quantify metabolites and/or hormone concentrations, thus implying invasive or minimally invasive procedures performed only in clinical settings or under the supervision of medical personnel. Second, data sets including several variables are usually collected through procedures that are demanding in terms of time and/or money, and for this reason, they are performed only during clinical trials rather than in routine clinical practice. This often translates into data colletion on a limited number of subjects. Finally, data sets from routine clinical practice can be large in terms of number of subjects, but they typically include a limited number of variables and they are difficult to be accessed for administrative issues. In order to support research on diabetes and help data retrieval, this review aimed at conducting a comprehensive search to locate and categorize data sets containing glycemic data that are accessible to the public. In particular, this review focused on dynamic glycemic data, which are of interest for the development of many technological applications. 6
Methods
Eligibility Criteria
This review targeted scientific articles, websites, and clinical trials that provide free access to data sets including dynamic glycemic data (both human and animal). Both immediately downloadable data sets and data sets downloadable after completion of a request form were included.
Exclusion Criteria
This analysis excluded scientific articles, websites, and clinical trials: (1) that do not contain any type of data set; (2) that contain an unavailable data set or a data set available upon payment of subscription fees; (3) in which the data of interest are mainly expressed as a fasting glucose or glycated hemoglobin A1c (HbA1c) as a single value, being very often collected within a basic routine examination and also as secondary variables (ie, belonging to a data set whose primary aim is not collecting glycemic data useful for diabetes-related research). This latter criterion was necessary to exclude data sets with some diabetes-related information but with limited usefulness for novel diabetes-related research, being them likely already exploited in the original studies for which they were collected.
Literature Search Strategy
This scoping review was conducted according to the Arksey and O’Malley methodological framework 7 and the guidelines provided by Daut et al 8 and Peters et al. 9 The PRISMA extension for Scoping Reviews (PRISMA-ScR) completed checklist 10 was provided as Supplementary Material. To discover all the available data sets, a systematic search of scientific articles, websites, and clinical trials was carried out utilizing four different sources, namely: (1) high-ranking scientific journals in the topic domain of interest, (2) Google, (3) a comprehensive clinical trial registry, and (4) two electronic databases. To perform the search, two groups of terms were considered and combined; in details, the first group included the terms referring to the resource of interest (eg, “Repository,” “Biobank,” “Platform,” and “Dataset”) and the second group included the terms concerning the topics domain of interest (eg, “Diabetes,” “Glucose,” “Glycemia,” “Glycaemia,” “Physiology”). Details on the Boolean logic operators used to combine the various keywords and on the strategy applied for each of the four sources are provided in the following subsections. The search was initially conducted in the period from February 2023 to May 2023, but the last update to search was performed in December 2024. The English language was set as a filter.
Scientific journal search
High-ranking scientific journals were selected according to the publicly available portal Scimago Journal & Country Rank (SJR). 11 Scientific journals appropriate for this research were selected by considering “Medicine” as subject area and “Endocrinology, Diabetes and Metabolism” as subject category or considering “Engineering” as subject area and “Biomedical engineering” as subject category. Only journals ranked in the first two quartiles (Q1, Q2) were considered. For each selected journal, its homepage was accessed and a second search for the relevant journal articles was performed. For journals providing the opportunity to select the type of results (ie, insert the filter “Dataset” or “Data paper”) directly on their homepage, a second search was performed using the terms of the second group connected with the comma or the “OR” (the choice depending on the portal instructions). If this option was not available, a search was performed by linking through the “AND” operator all the terms of the second group (connected with “OR”) and the search string (“Dataset” OR “Data Paper”). Steps for the scientific journal search strategy are summarized in Figure 1.
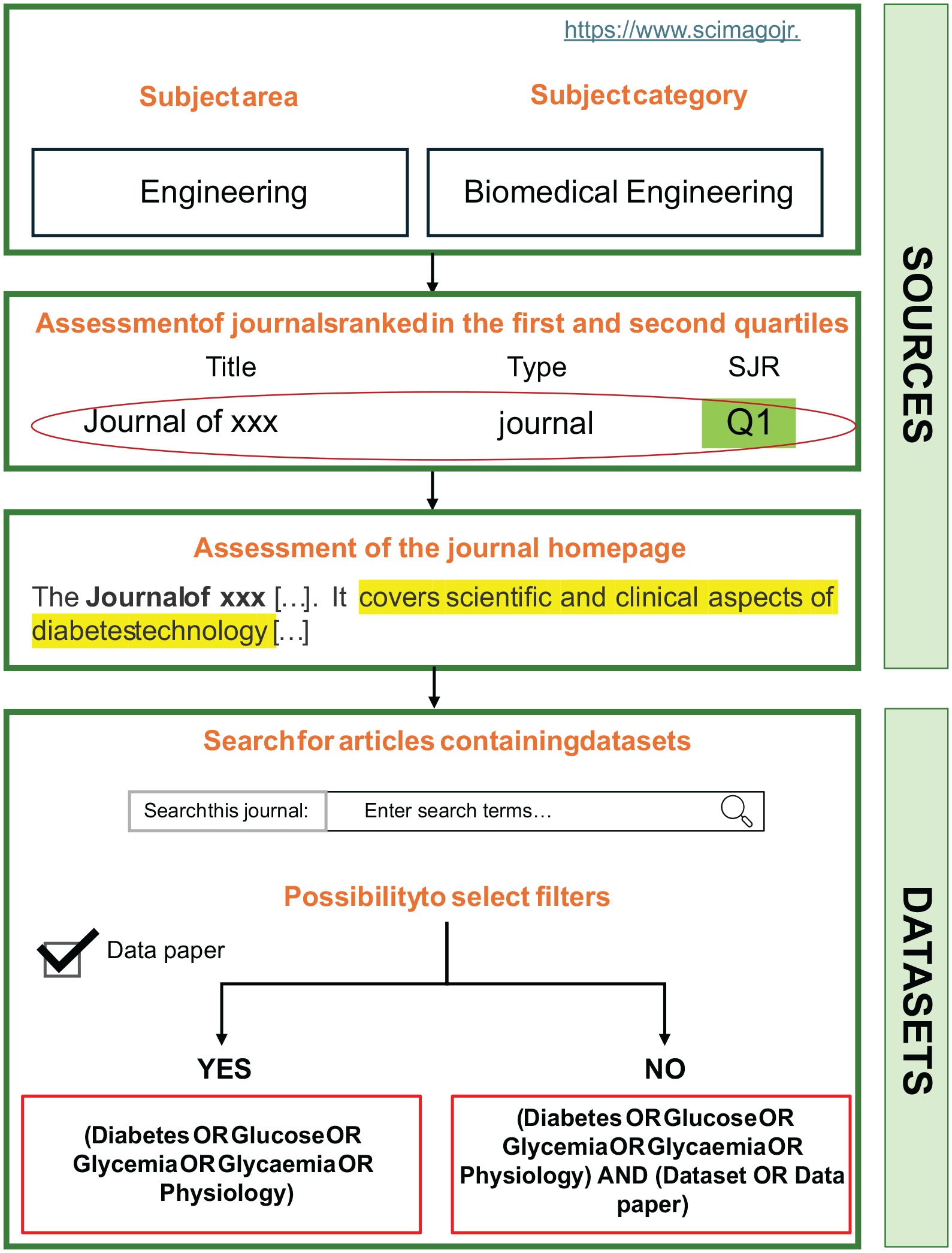
Overview of the steps for the search and screening strategy in scientific journals.
Google search
Five different searches of websites were performed according to the following strategy: for each search, all the terms belonging to the first group were considered and only one of the five terms of the second group was chosen; the terms within the first group were linked by the Boolean operator “OR” and then combined with the Boolean operator “AND” with the term chosen in the second group (eg, “Repository OR Biobank OR Platform OR Dataset” AND “Diabetes”). For each search, the first 50 websites ranked in order of relevance were evaluated in relation to the typology of their content (ie, data set, paper with associated data set, repository/portal, etc) similarly to what indicated as the last step of the scientific journal search. Steps for the Google search strategy are summarized in Figure 2.
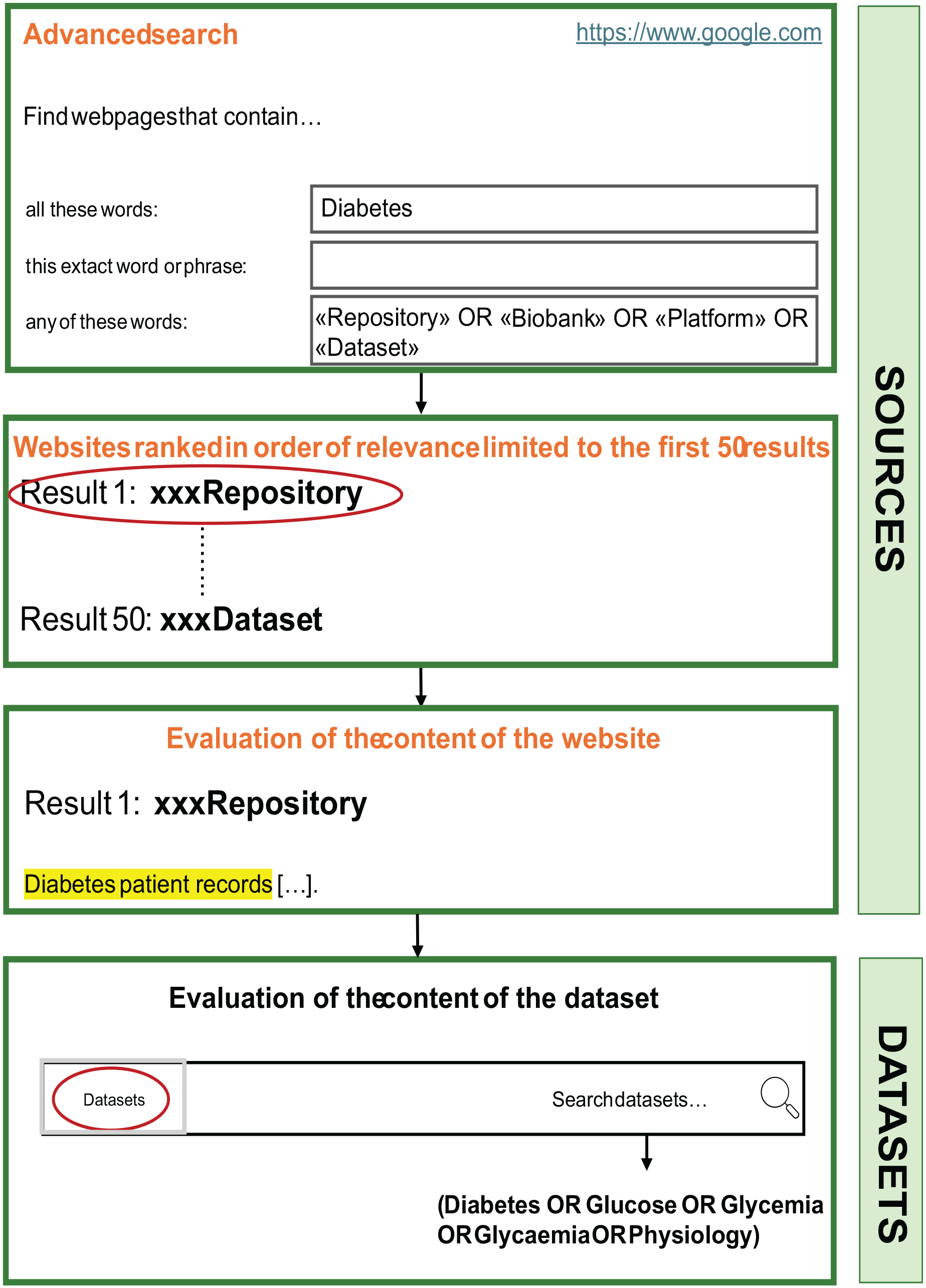

Overview of the steps for the search and screening strategy in Google.
Clinical trials search
The considered clinical trial registry was ClinicalTrials.gov (https://www.clinicaltrials.gov/). The domain “Condition of Disease” was set to “Diabetes” and in the field “Other Terms” the words “glucose, glycaemia, insulin” were inserted to indicate glycemic data and those data mostly related to it, thus usually collected in diabetes-related studies. The filter “Study with results” was applied. Steps for the search strategy in ClinicalTrials.gov are summarized in Figure 3.
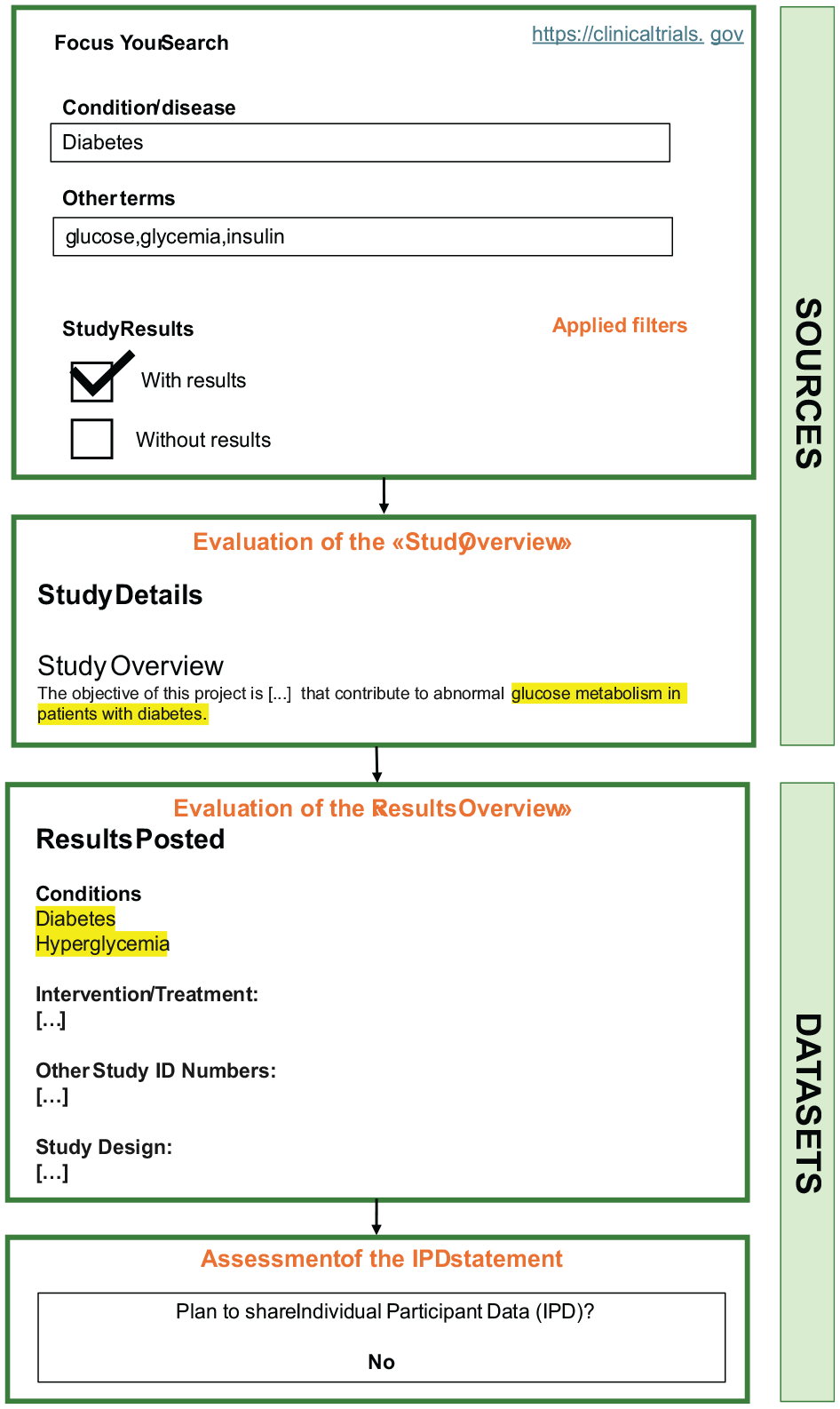
Overview of the steps for the search and screening strategy in the clinical trial registry ClinicalTrials.gov.
Electronic databases search
The SCOPUS and PubMed were selected as electronic databases, and the advanced search strategy was implemented. The following search query was applied: (“Diabetes” OR “Glucose” OR “Glycemia” OR “Physiology”) AND (“Repository” OR “Biobank” OR “Platform” OR “Dataset”). The filters “data paper” or “associated dataset” were applied depending on the database’s instructions.
Screening Strategy
For each typology of search, a two-level screening strategy was performed as detailed in the following.
Screening strategy for journal search
For each selected journal, the journal’s scope was first screened; if the journal was retained based on its scope, a second screening was performed at the article level considering also supplementary materials when applicable, according to eligibility/exclusion criteria detailed above (Figure 1).
Screening strategy for Google search
The 50 websites resulting for each of the five searches were first screened without fully accessing them. If the website was deemed appropriate, a more advanced screening was performed with detailed access of the website (Figure 2).
Screening strategy for clinical trials
Screening for the clinical trials was performed first by details reported in “Study overview” and then by those reported in “Results overview” (Figure 3).
Screening strategy for electronic databases
After removing duplicates between the two sources, all the record titles were assessed as a first screening step. Then, a second screening was performed at the full-text and data set level.
Data Analysis
Each data set was analyzed in terms of the publication year, the study design, the population involved (humans or animals), the number of subjects participating in the study, information on gender and age, the type of diabetes, the type of dynamic glycemic data (ie, oral or intravenous glucose tolerance test [IVGTT], CGM), the test duration, the number of acquired samples and variables, or, if applicable, the devices employed for the measurement and the condition of the study (laboratory/hospital or free-living conditions).
Results
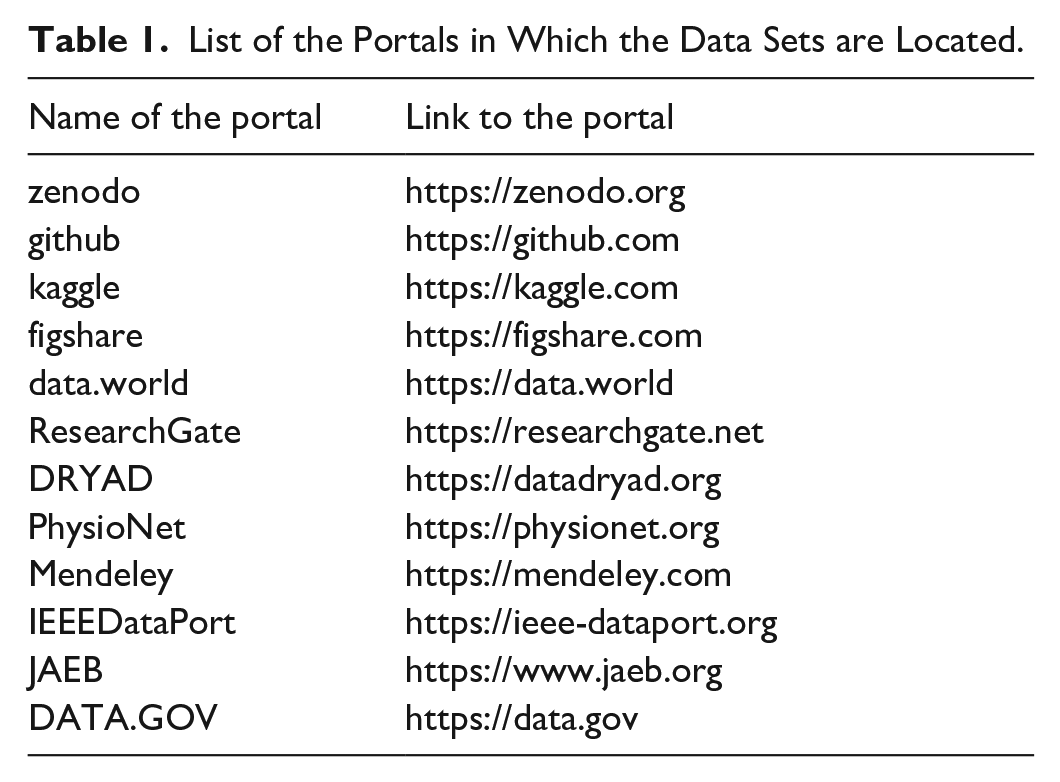
A total of 25 data sets were included in the review, following screening and selection as detailed in the flowchart in Figure 4. The portals in which the data sets are located are listed in Table 1. Retrieved data sets included two main typologies of dynamic glycemic data, namely data from challenge tests (5 data sets)12-16 and data from CGM (20 data sets).17-36 As for the first typology, data were related to oral (oral glucose tolerance test [OGTT]) and, in a small number of cases (1 of 5) also to IVGTT, as detailed in Table 2. The duration of the OGTT in the considered data sets is equal to 2 hours, whereas the number of blood samples acquired showed variability among the data sets. Also, the number of variables acquired during the test can vary depending on the objectives of the study, as it is shown in Table 3; a common characteristic in the majority of the challenge test data sets (3 of 5) is the assessment of both glycemia and insulinemia. All the challenge test data sets are freely available and downloadable.
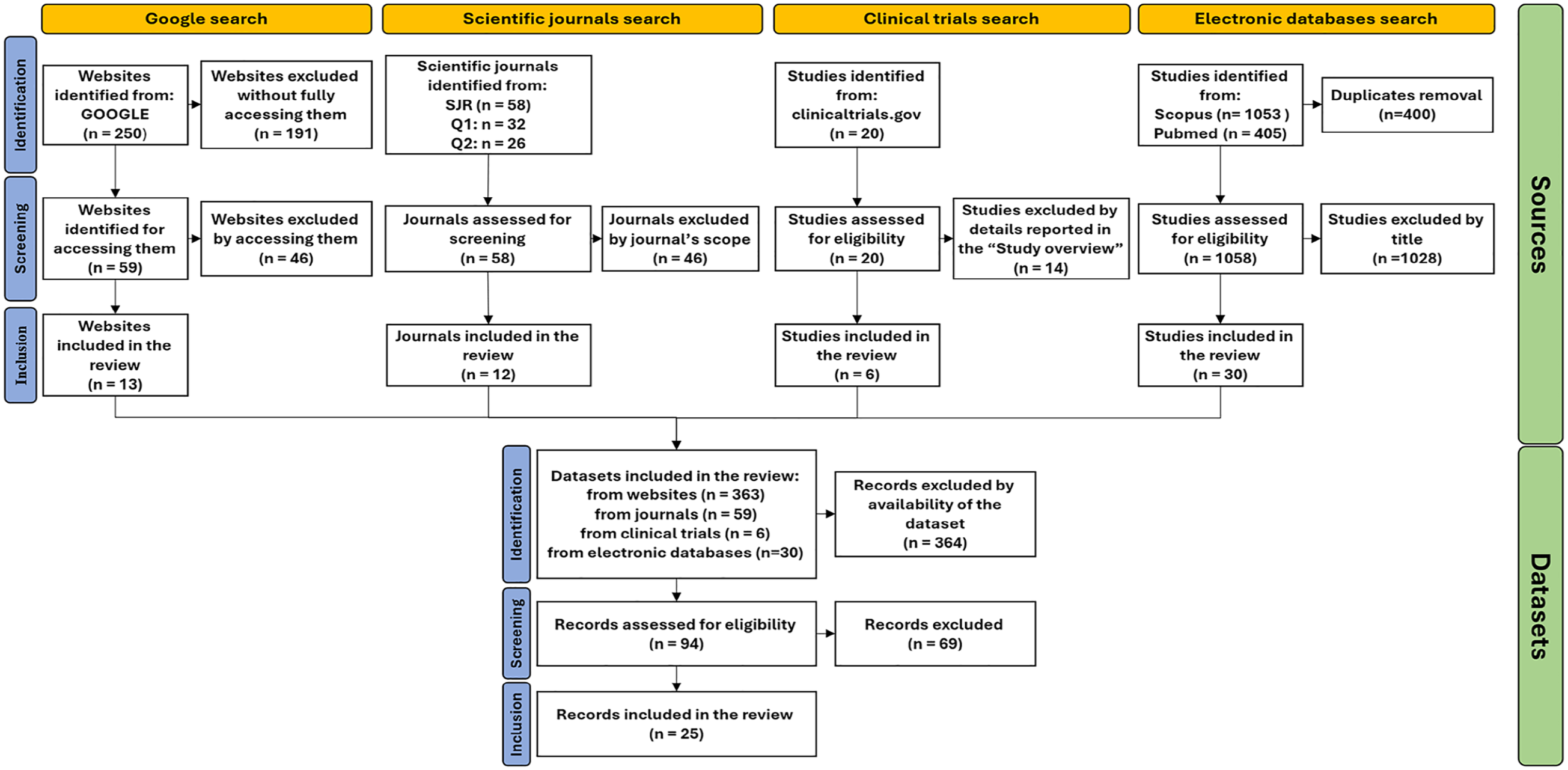
PRISMA flowchart.
List of the Portals in Which the Data Sets are Located.
Summary of the Data Sets Related to Challenge Tests.
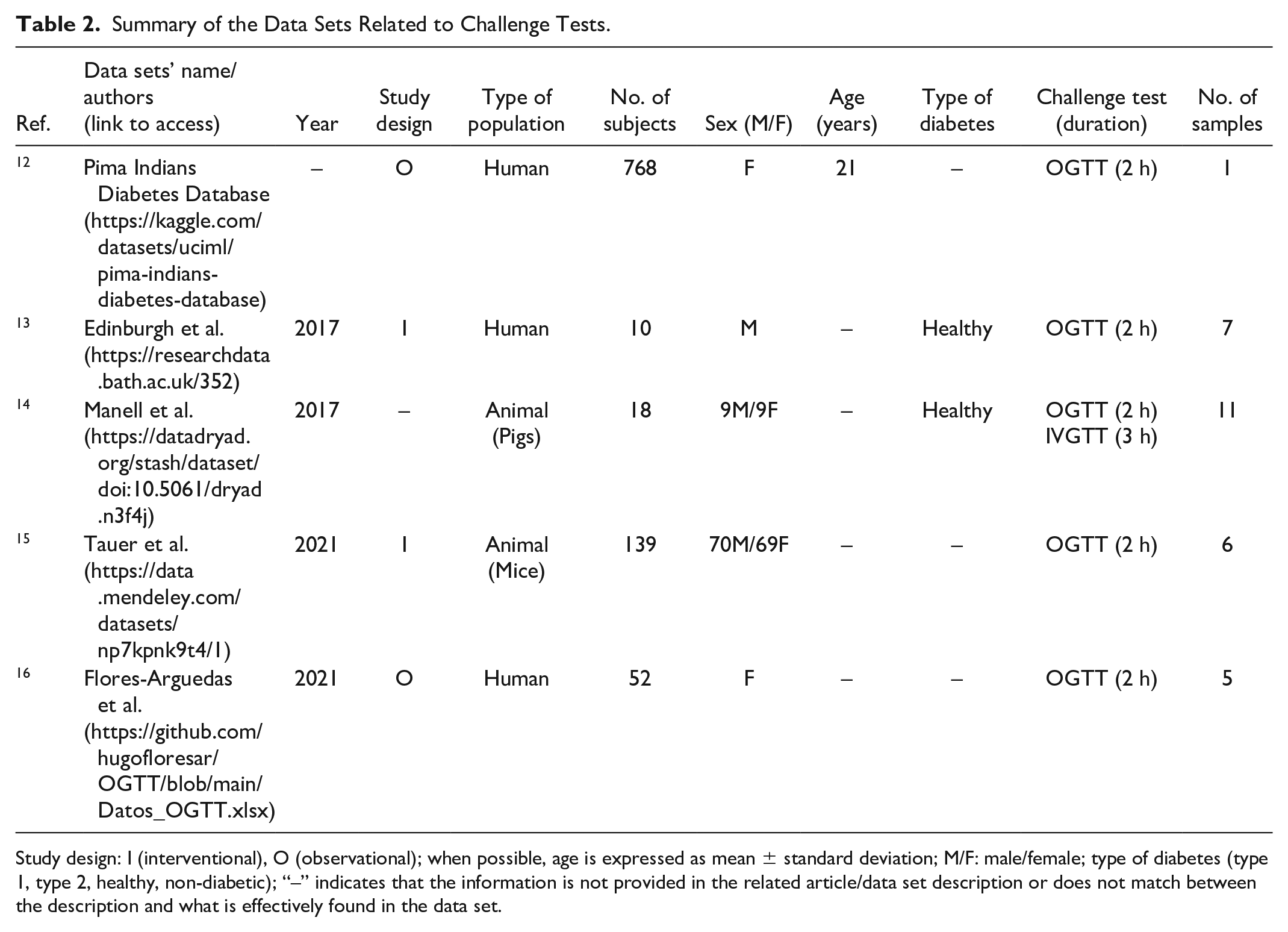
Study design: I (interventional), O (observational); when possible, age is expressed as mean ± standard deviation; M/F: male/female; type of diabetes (type 1, type 2, healthy, non-diabetic); “–” indicates that the information is not provided in the related article/data set description or does not match between the description and what is effectively found in the data set.
Variables Measured During the Challenge Test and Reported in the Data Sets.
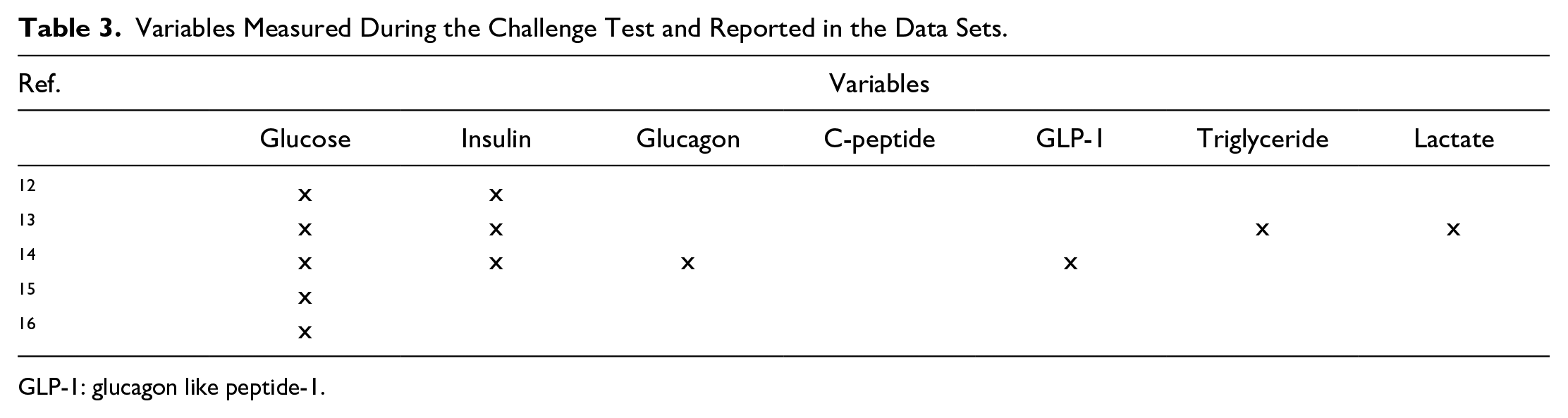
GLP-1: glucagon like peptide-1.
The second typology of data pertains to CGM, which represents a technique to monitor glucose every 1 to 5 minutes for 7 to 10 days or even more with a single glucose sensor. 37 As regards CGM data, it is necessary to distinguish between data sets that are freely available and directly downloadable from any user (detailed in Table 4) and data sets that are available under completion of a request form (detailed in Table 5). Such data sets are characterized by different duration, spanning from acquisitions lasting less than two weeks to acquisitions lasting more than nine weeks (Figure 5). Moreover, the number of subjects in the CGM data sets can vary, with the majority (11 of 20) characterized by more than 100 subjects (Figure 6).
Summary of the Freely Available and Downloadable Data Sets Related to Continuous Glucose Monitoring.
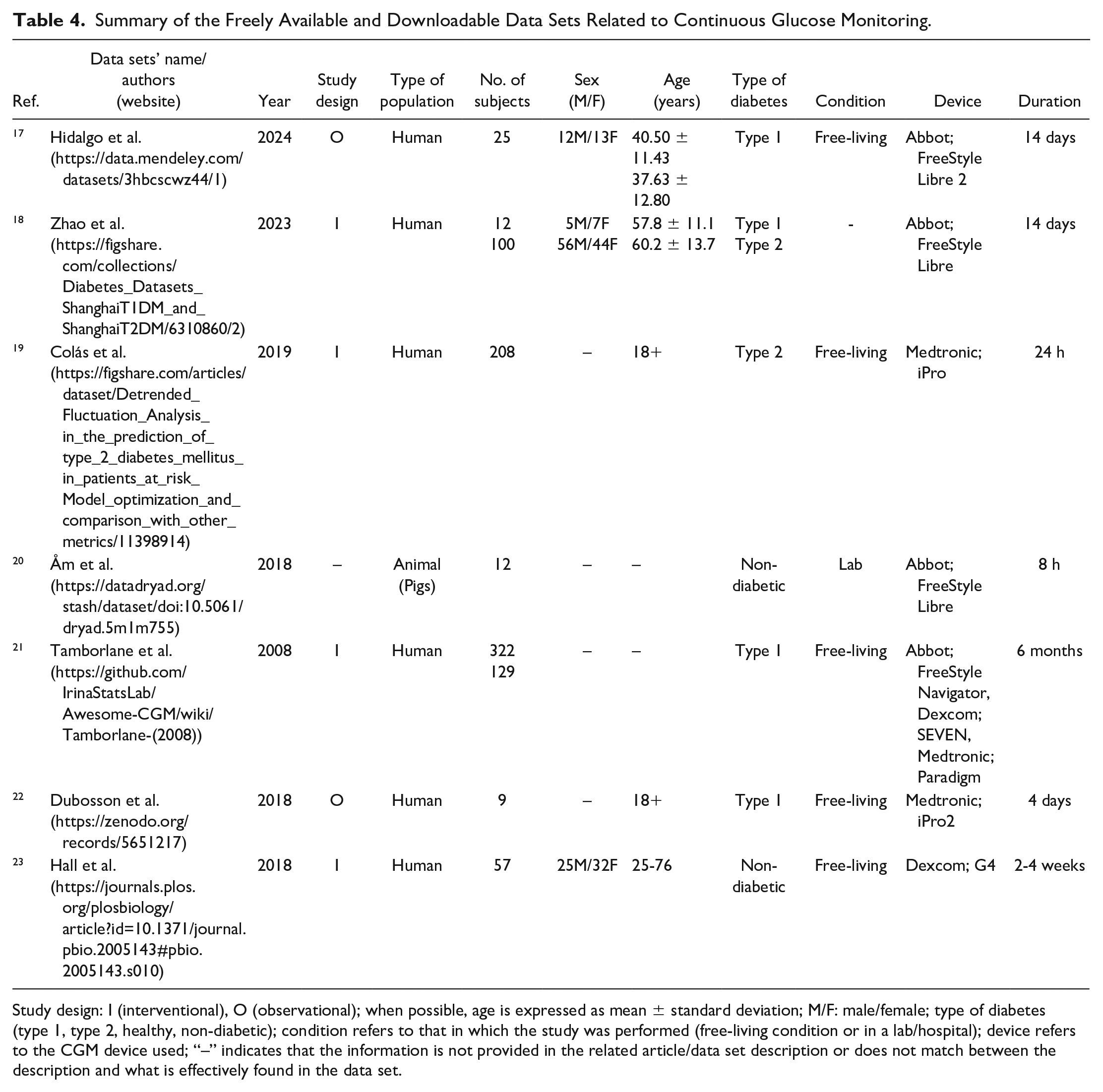
Study design: I (interventional), O (observational); when possible, age is expressed as mean ± standard deviation; M/F: male/female; type of diabetes (type 1, type 2, healthy, non-diabetic); condition refers to that in which the study was performed (free-living condition or in a lab/hospital); device refers to the CGM device used; “–” indicates that the information is not provided in the related article/data set description or does not match between the description and what is effectively found in the data set.
Summary of the Data Sets Related to Continuous Glucose Monitoring, Freely Available Upon Completion of a Request Form.
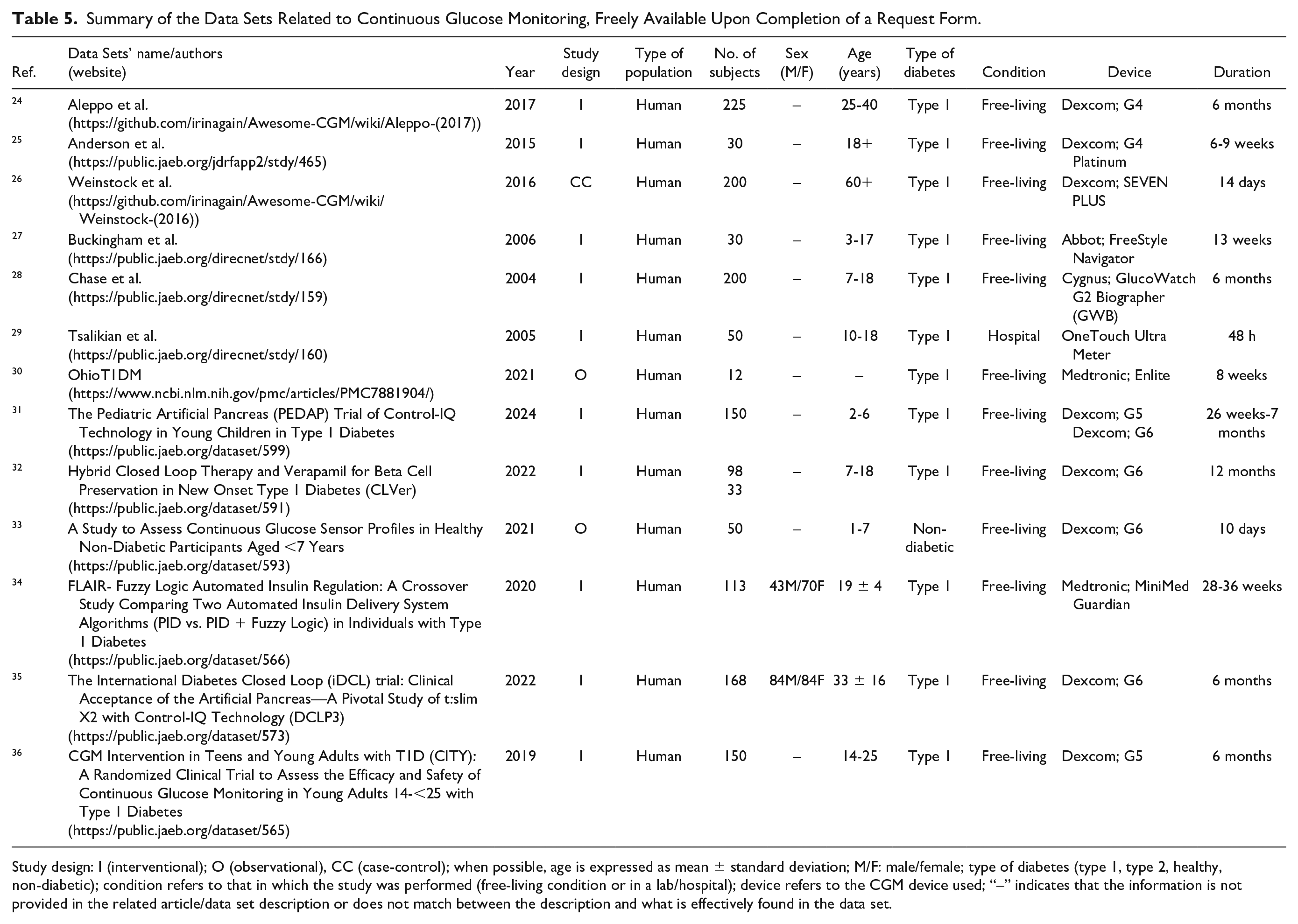
Study design: I (interventional); O (observational), CC (case-control); when possible, age is expressed as mean ± standard deviation; M/F: male/female; type of diabetes (type 1, type 2, healthy, non-diabetic); condition refers to that in which the study was performed (free-living condition or in a lab/hospital); device refers to the CGM device used; “–” indicates that the information is not provided in the related article/data set description or does not match between the description and what is effectively found in the data set.
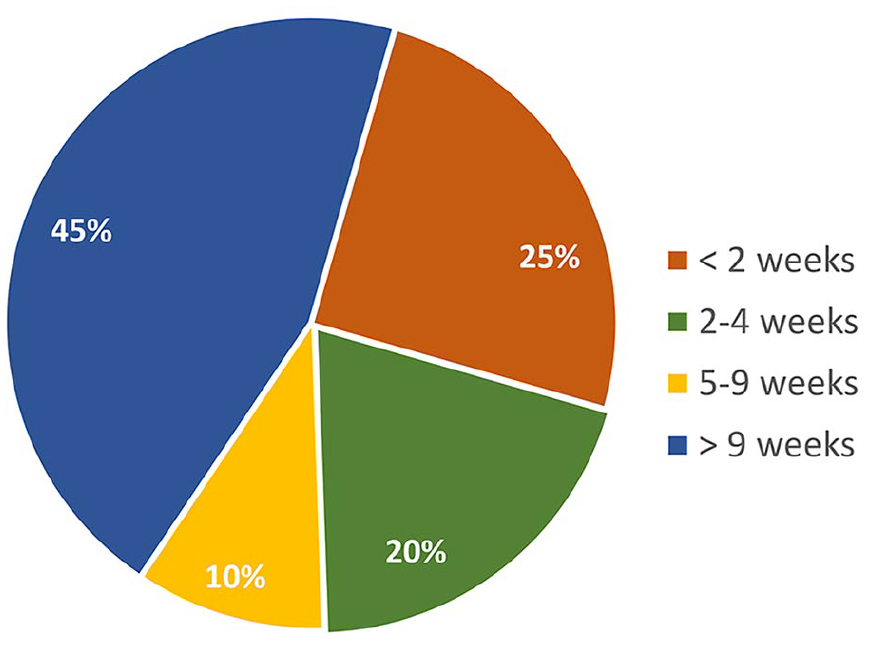
Duration of the continuous glucose monitoring (CGM) acquisitions across the reviewed data sets.
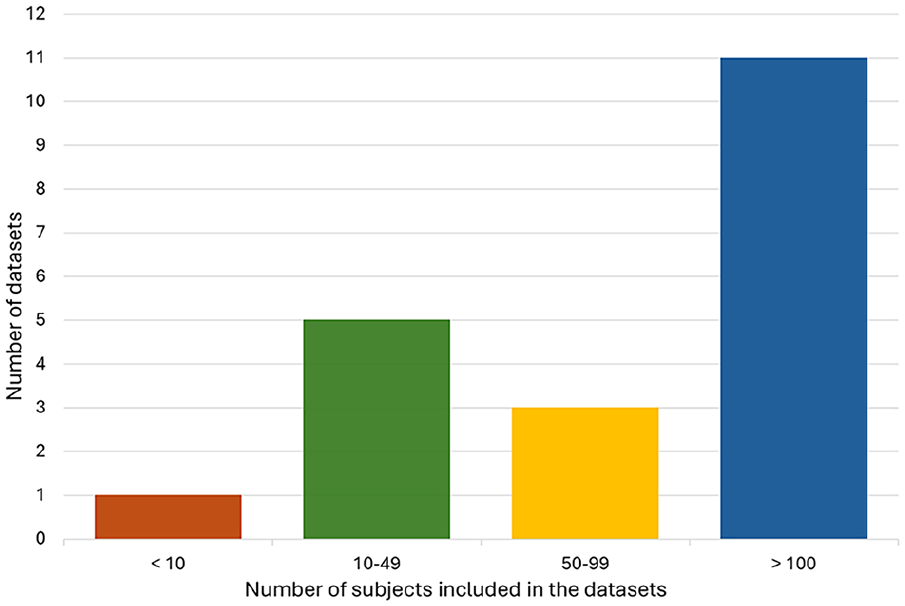
Number of subjects included in the continuous glucose monitoring (CGM) data sets.
Discussion
This review systematically analyzed the freely available data sets containing dynamic glycemic data that can be exploited in diabetes research. The 25 data sets included were retrieved from four different searches (ie, scientific journals, Google, clinical trials, and electronic databases), which made us confident that all potential sources of relevant information were covered. As regards the scientific journals search, a one-by-one screening was applied for scientific journals in the field, limiting the search to those in the top quartiles (Q1 and Q2, as per SJR). Although this choice may expose to the risk of not considering data sets potentially published in Q3-Q4 quartile journals, it was considered a reasonable choice since the most relevant quartiles were included. Moreover, a standard search in the databases of scientific literature (like SCOPUS and PubMed) was also included to complement the plethora of data papers or scientific papers with associated data available. In SCOPUS, the application of the filter “data paper” was necessary since the standard search was not viable due to the impossibility of applying a criterion to filter articles with associated available data, which is instead possible in PubMed. Of note, the strategy we applied for journal searching allows also to not filter a priori relevant results published as abstract only, if present in high-ranking scientific journals. The inclusion of Google search, as well as search in clinical trials, was motivated by the willingness to identify suitable websites and online sources where relevant data sets could potentially be located, even if they may not have an associated paper in a scientific journal. The exhaustive overview of the main websites and online sources provided by our search process is demonstrated by results reported in Table 1, which include, to the best of our knowledge, all the main portals devoted to data sharing. However, the implementation of a preliminary screening for the website search was necessary since Google is a vast search engine, housing an immense volume of data and, without some form of filtering, the provided results can become overwhelming and require substantial time for examination. As regards the clinical trials search, although different clinical trial registries could be considered, the search was limited to ClinicalTrials.gov; however, we did not consider this as a substantial limitation, as this registry is one of the largest and most used.
Overall, the outcomes derived from this analysis indicate that the available dynamic glycemic data can be classified into two main categories: challenge test and CGM data. As for the first category, all data from challenge tests pertain to OGTT. This result is not unexpected, being the 75-g OGTT one of the diagnostic screening tests for diabetes and prediabetes according to the American Diabetes Association (ADA) guidelines. 38 Concerning the variables provided within the OGTT data sets, the majority of the data sets reported the measurement of additional variables with respect to glycemia; in particular, insulin levels feature prominently in most of the data sets, whereas other information such as C-peptide, triglycerides, and glucagon-like peptide-1 (as a marker of incretin action) are rarely measured. It is also worth noting that all the OGTTs have standard duration (2 h). Conversely, with regard to the number of OGTT blood samples, it is noteworthy that the greater majority of the retrieved data sets provide a number of samples higher than that required for diabetes diagnosis (ie, two blood samples, at fasting and at 2 h during OGTT). Indeed, the collection of four or five samples at 30-minute intervals is confirmed as a common approach, 39 with some study protocols collecting even more samples. As regards IVGTT, only one of the data sets reported this type of data. Indeed, the IVGTT is a test typically used in the investigation of diabetes pathophysiology 40 and hence performed for research purposes rather than for direct clinical applications, thus usually resulting in data sets with a limited number of subjects. Of note, this review also targeted data from preclinical in vivo models, frequently studied with OGTT and IVGTT in diabetes pathophysiology research. 41 While accounting for a small percentage of data sets included in our review, it is worth noting that these data sets are characterized by a higher number of variables than those provided by studies on humans.
Applications exploiting this category of data may vary not only in relation to the number of subjects included in the data set but also to the protocol used for data acquisition. One of the most consolidated applications relies on the use of challenge test data jointly with mathematical (compartmental) models to extract parameters of clear physiological meaning (eg, insulin sensitivity, alpha and beta-cell sensitivity).42-47 It is worth noting that for this kind of application, the crucial factor is not represented by the number of subjects in the data set (being analysis performed on an individual basis) but rather by the characteristics of the protocol, which may lack suitability if the number of blood samples and/or variety of data do not match with the requirements in terms of model parameters to be estimated. An illustrative instance of this is the standard OGTT exclusively measuring glucose levels, which provides data not adequate for model-based approaches. Furthermore, other applications may rely on AI-based models, which have more urgent requirements in terms of the number of subjects included in the data set rather than protocol characteristics. Indeed, studies on open and proprietary data sets showed that, when a sufficiently large amount of data are exploitable (ie, in terms of number of subjects), even the basic challenge test data acquired in clinical practice, like the standard OGTT, may become useful to develop AI solutions with real applicability. 48 Also, when the exploitation of mathematical models is enabled by protocol characteristics, model-based features with clear physiological meaning can be extracted to power information gathered from the data and to feed AI models. 49
The CGM data sets cover a consistent percentage of the identified data sets (20 of 25). The use of CGM devices has become increasingly prevalent in recent years, primarily due to significant advancements in technology. By providing real-time and continuous monitoring of glucose levels (every 1-5 min), these devices are able to provide a large amount of data. Besides, an even more important advantage is their capability to monitor and describe the glucose fluctuations that take place in free-living conditions and in response to perturbations like meals and physical exercise. Information coming from CGM patterns is still widely unexplored, but efforts are in due course to provide standardization in their analysis in relation to the computable CGM metrics. 50 Thanks to the increasing availability of such kind of data sets and to their dimension (most of the reviewed data sets are characterized by more than 100 subjects), information from CGM devices could become crucial for the development of clinical decision support systems in diabetes field based on AI and machine learning approaches. In relation to this aspect and in consideration of the peculiarities of diabetes research field, best practices and pitfalls were described. 4 Finally, it should be acknowledged that the CGM data sets analyzed in this review are mainly related to T1D patients and less frequently to T2D (none in gestational diabetes mellitus [GDM]). On the contrary, this result was somehow expected, being CGM use still typically limited to patients with T1D, even though use in other populations is increasing. 51
One may argue on the usefulness of such a review study, given the widespread use of generative AI tools also in the field of literature search. However, when we attempt to use generative AI as a surrogate of the literature review here performed (question: “please search for open data sets containing glucose measurement data which are downloadable freely and put the results in a table”), only 4 of the 25 data sets were effectively retrieved, which are the most widely known, also easily identifiable as a simple Google search. This implies that, despite being very powerful tools in asking for more information regarding a specific data set, they are not able to replicate the manual effort done in the present study. A second element of criticism can be found in the choice of excluding data sets in which the data of interest are mainly expressed as a fasting glucose or glycated hemoglobin as a single value. The high number of data sets that may have this characteristic, however, would have not represented a real added value for this review; indeed, their potential for studies other than the ones for which they were acquired is usually limited with respect to dynamic data. Eventually, an additional element of criticism can be related to the obsolescence of results here reported, due to the possible availability of open data sets in the future. However, the methodology used in this study has to be considered as a search pipeline, which could be periodically updated to retrieve new data sets. This further emphasizes the usefulness of the present review.
Conclusions
This review identified a total of 25 data sets that can be freely downloaded. This number represents a small percentage with respect to the data sets initially anticipated from the four searches performed, proving that poor data accessibility still remains a limitation to overcome in this field. However, the possibility provided by this analysis to have an overview of the readily usable data sets and to easily locate them represents a step forward in fostering data access.
Supplemental Material
sj-docx-1-dst-10.1177_19322968251316896 – Supplemental material for Availability of Open Dynamic Glycemic Data in the Field of Diabetes Research: A Scoping Review
Supplemental material, sj-docx-1-dst-10.1177_19322968251316896 for Availability of Open Dynamic Glycemic Data in the Field of Diabetes Research: A Scoping Review by Libera Lucia Del Giudice, Agnese Piersanti, Christian Göbl, Laura Burattini, Andrea Tura and Micaela Morettini in Journal of Diabetes Science and Technology
Footnotes
Abbreviations
ADA, American Diabetes Association; AI, artificial intelligence; CGM, continues glucose monitoring; GDM, gestational diabetes mellitus; HbA1c, glycated hemoglobin; IVGTT, intravenous glucose tolerance test; OGTT, oral glucose tolerance test; PRISMA, Preferred Reporting Items for Systematic Review and Meta-Analysis; T1D, type 1 diabetes; T2D, type 2 diabetes.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
