Abstract
Background:
Continuous glucose monitors (CGMs) in research and clinical settings characterize glycemic profiles through repeated measurement of interstitial glucose levels on the order of minutes. Missing values from devices are unavoidable. Data from the Glycemic Observation and Metabolic Outcomes in Mothers and Offspring (GO MOMs) study were used to investigate the impact of missing data on CGM summary metrics. Several imputation techniques were evaluated by comparing mean relative bias (MRB) between true and imputed CGM data for the summary metrics.
Methods:
We used 105 CGM profiles with nine days of complete glucose measurements and introduced missing data strings using a zero-inflated negative binomial hurdle model. Overall missingness was introduced at 2% consistent with GO MOMs data and increased to 5%, 10%, and 20%. Imputation approaches included single, multiple, machine learning techniques, and hot-deck imputation, where missing values are replaced with the participant’s observed values. Removing missing values prior to analysis (complete case analysis) was also evaluated.
Results:
The MRB is minimal across most metrics and imputation methods at overall 2% missing data and increases with higher missing data frequency, with trends depending on metric and imputation method. Hot-deck imputation and complete case analysis show consistently low MRB.
Conclusions:
Missing CGM data are to be expected. For periods of wear with up to 20% missing data, hot-deck imputation and complete case analysis may be acceptable if data are missing completely at random. Explored imputation techniques are robust, but each has their own limitations, which should be considered if these techniques are implemented.
Keywords
Introduction
Continuous glucose monitoring (CGM) in clinical care and research provides an opportunity for characterization of glycemic profiles through rapid, repeated measurement of interstitial glucose levels. In the Glycemic Observation and Metabolic Outcomes in Mothers and Offspring (GO MOMs) study, 1 pregnant individuals wear blinded CGM for ten days at four different timepoints. These high-density data hold promise for novel insight into pregnancy glycemia.
Gaps in CGM readings lead to missing data. Pressure-induced sensor attenuation, leading to reduced blood and interstitial fluid exchange, may lead to inaccurate readings, commonly called “compression lows.” Compression lows are flanked by rapidly falling and rising glucose measurements (Figure 1), and it is generally assumed these are inaccurate readings. Missing values can also occur at random, possibly due to displacement of sensor filament or dropped transmission of data to a reader. These missing data strings vary in length and are flanked by values in a range typical of other readings for the individual (Figure 2).
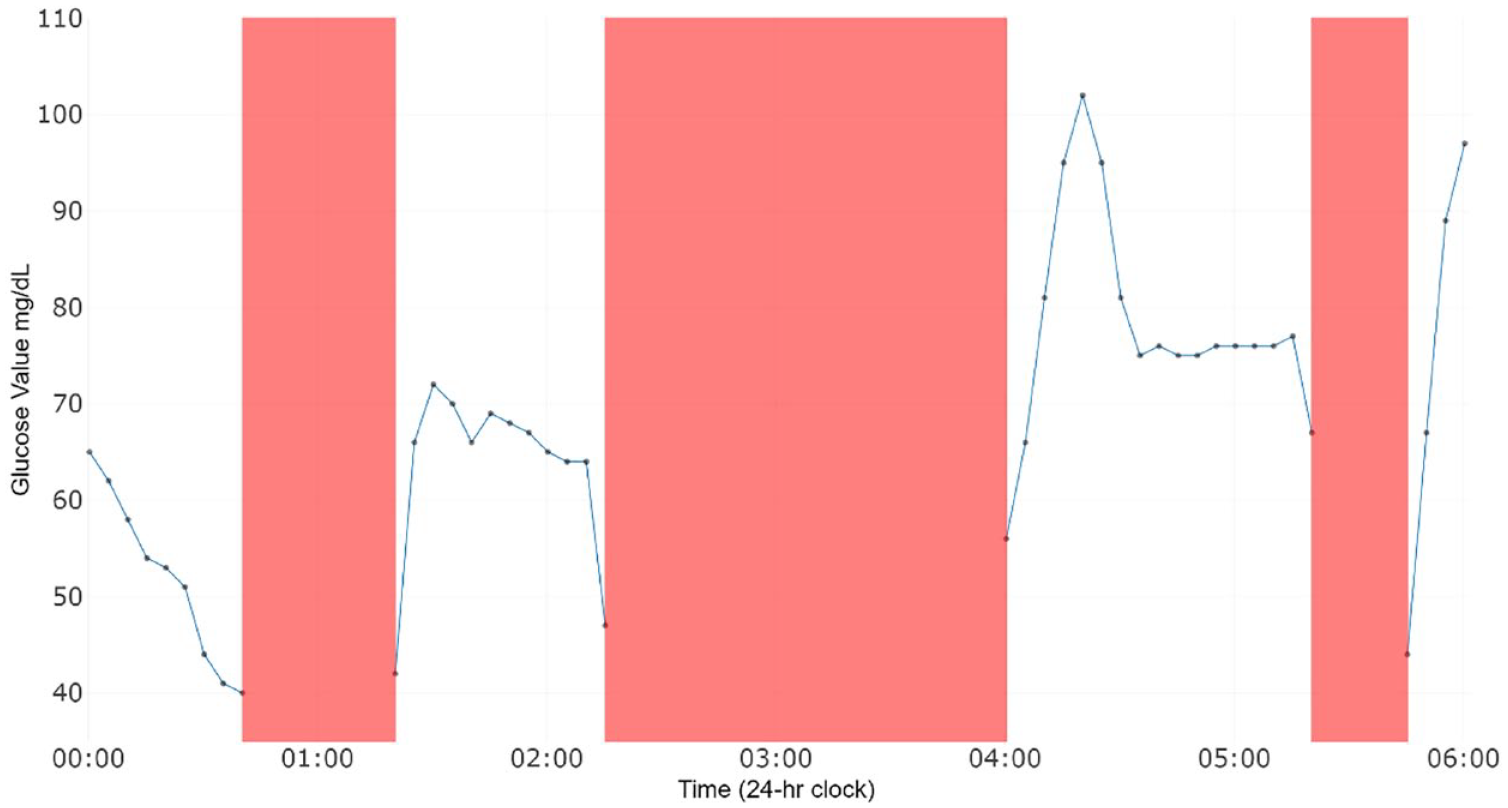
CGM profile from one day for a GO MOMs participant from midnight to 6:00 a.m. The x-axis measures time. The y-axis corresponds to glucose in mg/dL, which was measured in 5-minute increments. Gray dots are observed glucose measurements from the CGM device. Compression lows reported are marked in red. Compression lows are preceded by rapidly decreasing glucose measurements and then followed by rapidly increasing glucose values to a more typical range.
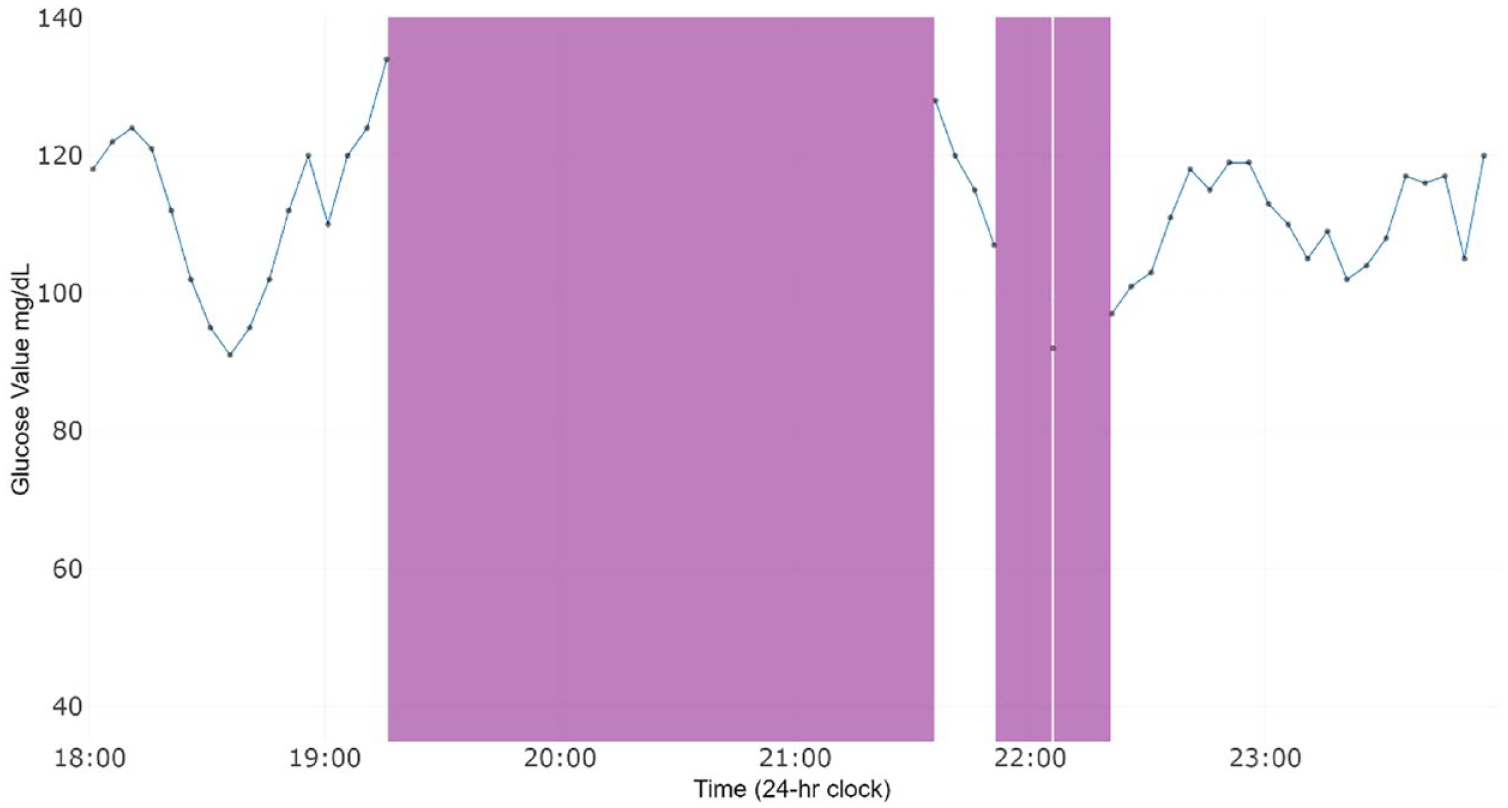
CGM profile from one day for a GO MOMs participant from 6:00 p.m. to midnight. The x-axis measures time. The y-axis corresponds to glucose in mg/dL, which was measured in 5-minute increments. Gray dots are observed glucose measurements from the CGM device. Unlike missing values due to compression lows, the strings of missing values marked in purple are apparently missing at random and are flanked by observed glucose values that are well above 40 mg/dL.
State-of-the-art CGM imputation techniques include interpolation, such as nearest neighbor imputation, where missing values are replaced with the nearest observed value 2 or linear interpolation based on observed data on either side of the missing values. 3 More sophisticated interpolation methods use cubic splines to model observations. 4 Millard et al 5 describe a hot-deck approach for replacing strings of missing values with observed values collected before and after the missing string. Drawbacks to these methods are that they are single imputations that make no distinction between imputed and observed values for analysis and do not make use of a participant’s CGM data collected on other days.
We sought to characterize the extent and patterns of missing data within CGM profiles for a subset of GO MOMs participants and used simulation to assess the impact of missing data on calculated CGM summary metrics. We explored a robust collection of imputation methods and identified resultant bias in calculated CGM metrics.
Methods
Glycemic Observation and Metabolic Outcomes in Mothers and Offspring Study
GO MOMs is a multicenter study to characterize longitudinal glycemia during pregnancy. 1 The primary objective is to use maternal glucose and clinical data at 10 to 14 weeks’ gestation to develop predictive criteria for gestational diabetes mellitus (GDM) at 24 to 28 weeks’ in mothers 6 and large-for-gestational age (LGA) delivery in newborns. 7 The GO MOMs participants provide informed consent.
The GO MOMs participants have study visits at 10 to 14 (visit 1), 16 to 20 (visit 2), 24 to 28 (visit 3), and 32 to 36 weeks’ (visit 4) gestation. At these visits, a blinded Dexcom G6 Pro CGM (Dexcom, Inc, San Diego, California) is inserted for ten days of wear as specified in the GO MOMs protocol. 1 The Dexcom G6 Pro measures interstitial glucose every 5 minutes for up to 288 measurements over a 24-hour period.
For this investigation, we compiled data from the first 15 participants who completed visit 1 CGM wear from each of the seven GO MOMs study sites and had nine days of complete observations starting at 00:00 (24-hour clock) on the day after device insertion through 23:55 nine days later (ie, 2592 glucose measurements). These 105 profiles provided reference data against which to evaluate imputation methods.
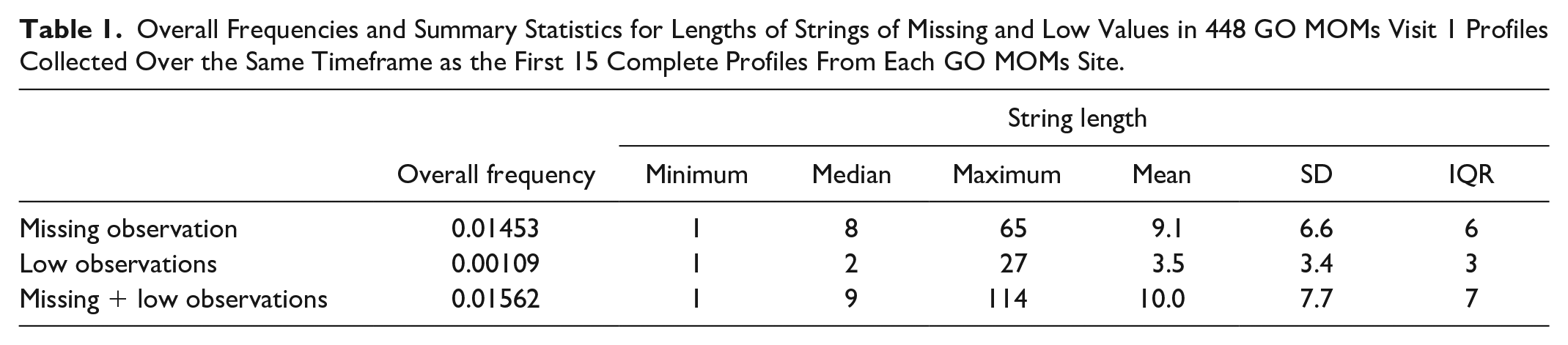
We additionally examined 448 profiles with at least one missing or “low” (below the threshold of detection) observation that were collected over the same timeframe as the 105 complete profiles. Table 1 summarizes overall missingness frequency and lengths of missing or “low” data strings. In GO MOMs, all values below the detection threshold and three flanking observations on either side of the “low” string that are <120 mg/dL are treated as missing.
Overall Frequencies and Summary Statistics for Lengths of Strings of Missing and Low Values in 448 GO MOMs Visit 1 Profiles Collected Over the Same Timeframe as the First 15 Complete Profiles From Each GO MOMs Site.

Methods
Simulation Study
Let
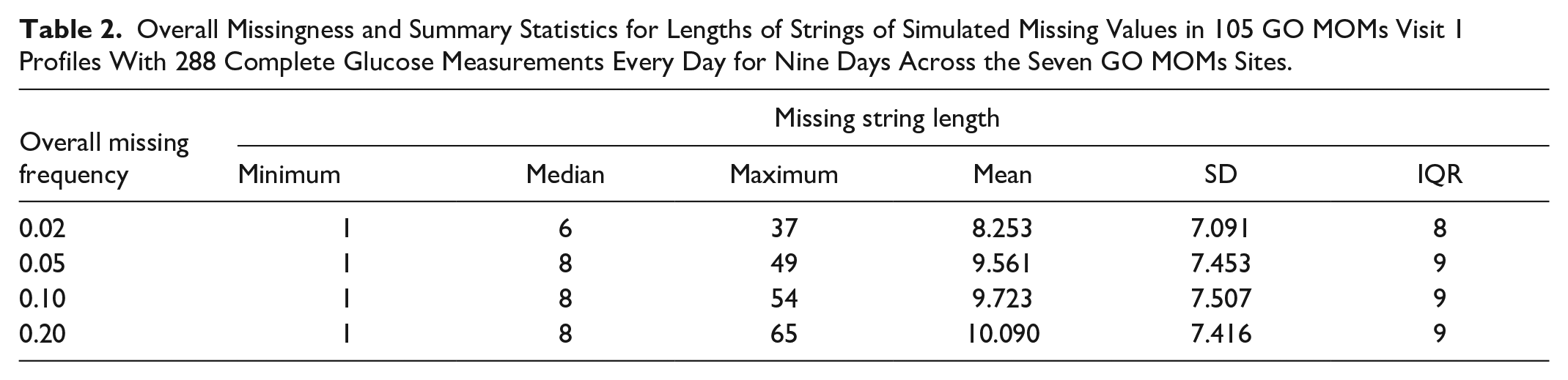
Overall Missingness and Summary Statistics for Lengths of Strings of Simulated Missing Values in 105 GO MOMs Visit 1 Profiles With 288 Complete Glucose Measurements Every Day for Nine Days Across the Seven GO MOMs Sites.

Imputation methods were applied after missing data were introduced. Let
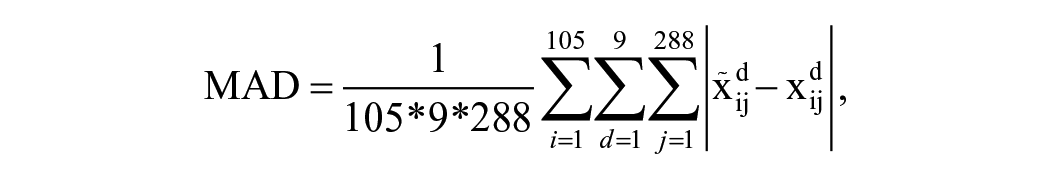
and
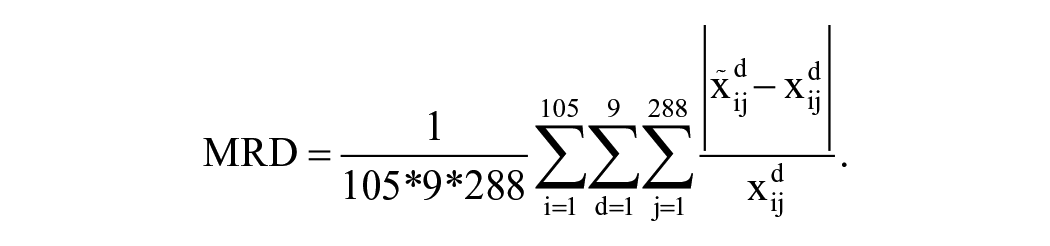
We calculated time-dependent (Supplemental Table 1) and time-independent (Supplemental Table 2) CGM summary metrics from the
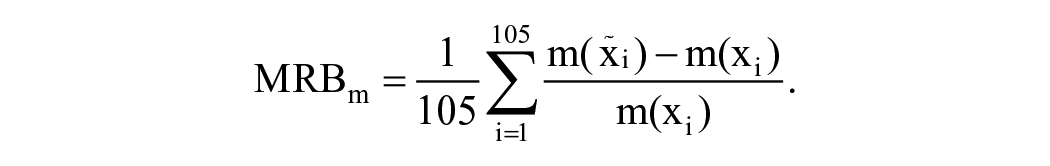
Imputation Methods
We examined single imputation, multiple imputation including hot-deck imputation (HDI) and chained equations and machine learning methods. Single and multiple imputation methods impute data for one participant at a time. Machine learning methods collectively utilize observed data for all individuals. We also evaluated complete case analysis in which missing values were omitted from all calculations. Examples of imputed data using all methods are illustrated in Figure 3.
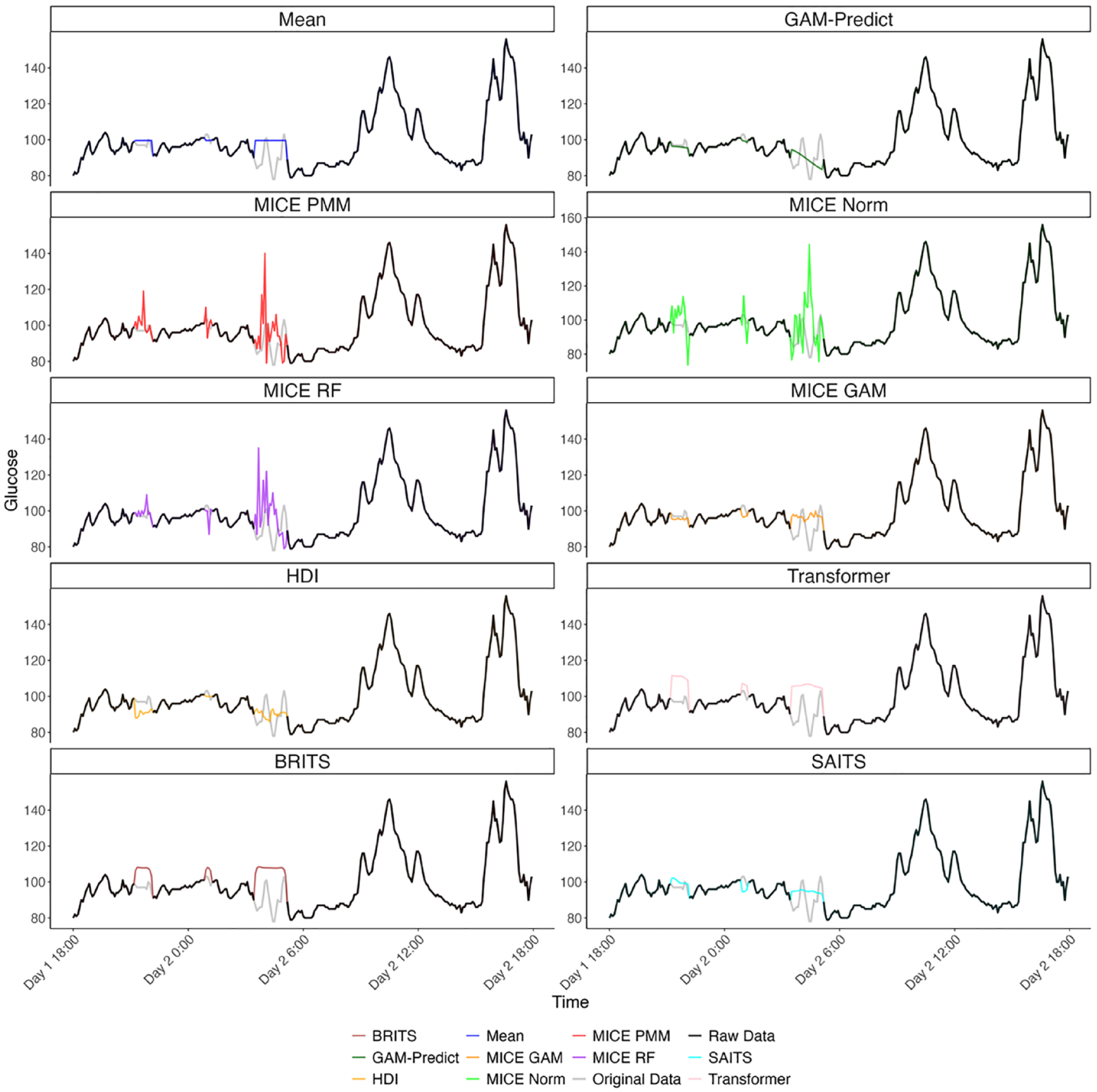
Examples of imputed data strings for all evaluated methods for 24 hours of data from one individual. Observed CGM data points that are analyzed as such are represented in black. Observed CGM data points that were treated as missing in simulations are represented in gray. Imputed values for the missing data points are represented in colors as indicated in the legend. For the multiple imputation methods, these figures illustrate one set of imputed values.
Single Imputation
Mean: Imputation of individual- and day-specific mean glucose values.
GAM-Predict: The
Multiple Imputation
HDI: HDI works by replacing missing values (recipients) with other observed values (donors). Since we have repeated days for each participant, we proposed donor cells be constructed using observed values from the same participant. First, for strings of missing data of six values or less, ie, up to 30 minutes, a donor cell is comprised of glucose measurements from the adjacent hour on either side of the missing data string. Values are sampled from this cell.
Second, for strings of missing data longer than six values, ie, more than 30 minutes, the donor cell is comprised of data strings from the other days for that individual at the same time of day. The string of data from the donor cell is selected using weighted sampling with weights determined by the mean of the two adjacent hours of CGM values on either side of the missing data string. The donor day is selected using the approach of Siddique and Belin, 11 with the donor randomly selected with probability inversely proportional to its distance from the recipient day. As donor strings may contain missing values, we iteratively repeat the two steps until no more missing values are present.
HDI details are described in full in Supplemental Material. This process is applied 10 times, and summary metrics are calculated over the multiple imputations.
Multiple imputation by chained equations (MICE): Given dense longitudinal observations,12
-15 we explored several MICE techniques. For each participant, columns in the data set were days, and each day was imputed by fitting a model conditioning on the other days. We first log-transformed the glucose values, performed 20 imputations, and then exponentiated to return values to their original scale. We employed four models as follows: MICE-PMM: Predictive mean matching (PMM) is an HDI approach. As implemented in MICE-Norm: As implemented in MICE-RF: Random-forest imputation can be effective in navigating complex, non-linear relationships.
17
Utilizing MICE-GAM: In addition to the single imputation GAM-Predict approach, we used GAMs in an MICE framework using
Machine Learning
Three deep learning techniques, Self-Attention Imputation Transformer with Self-Supervision (SAITS),
19
Bidirectional Recurrent Imputation for Time Series (BRITS),
20
and Transformer
21
were implemented using custom Python 3.9 scripts, leveraging
SAITS: SAITS employs a self-attention mechanism consisting of multiple layers to dynamically weigh observation importance and a feed-forward neural network.
BRITS: BRITS utilizes bidirectional recurrent neural networks to capture temporal dependencies in the data.
Transformer: Transformer consists of multiple encoder and decoder layers, each containing a self-attention mechanism and a feed-forward neural network.
The CGM data were normalized and reshaped. The data set was split into 80% training and 20% testing as required by these methods. After training, models were used to impute missing values in the validation data sets. Imputed values were then transformed back to their original scale.
Results
Mean Absolute Difference and Mean Relative Difference
The MAD and MRD were used to evaluate how closely each imputation method came to matching true individual glucose values, with lower values indicating concordance (Table 3). Across missingness frequencies, SAITS had the lowest MAD and MRD, outperforming other methods for recovering individual-level glucose values. The GAM-Predict had next best performance, followed by MICE-GAM. Transformer and other MICE methods demonstrated worse individual-level performance. For all methods, MAD and MRD were modest even at missingness frequencies as high as 20%, with an MAD of only 3.24 mg/dL (2.9%) for MICE-Norm, the worst-performing method according to these metrics.
MAD and MRD Values for the Investigated Imputation Methods Under a Range of Missing Value Frequencies.
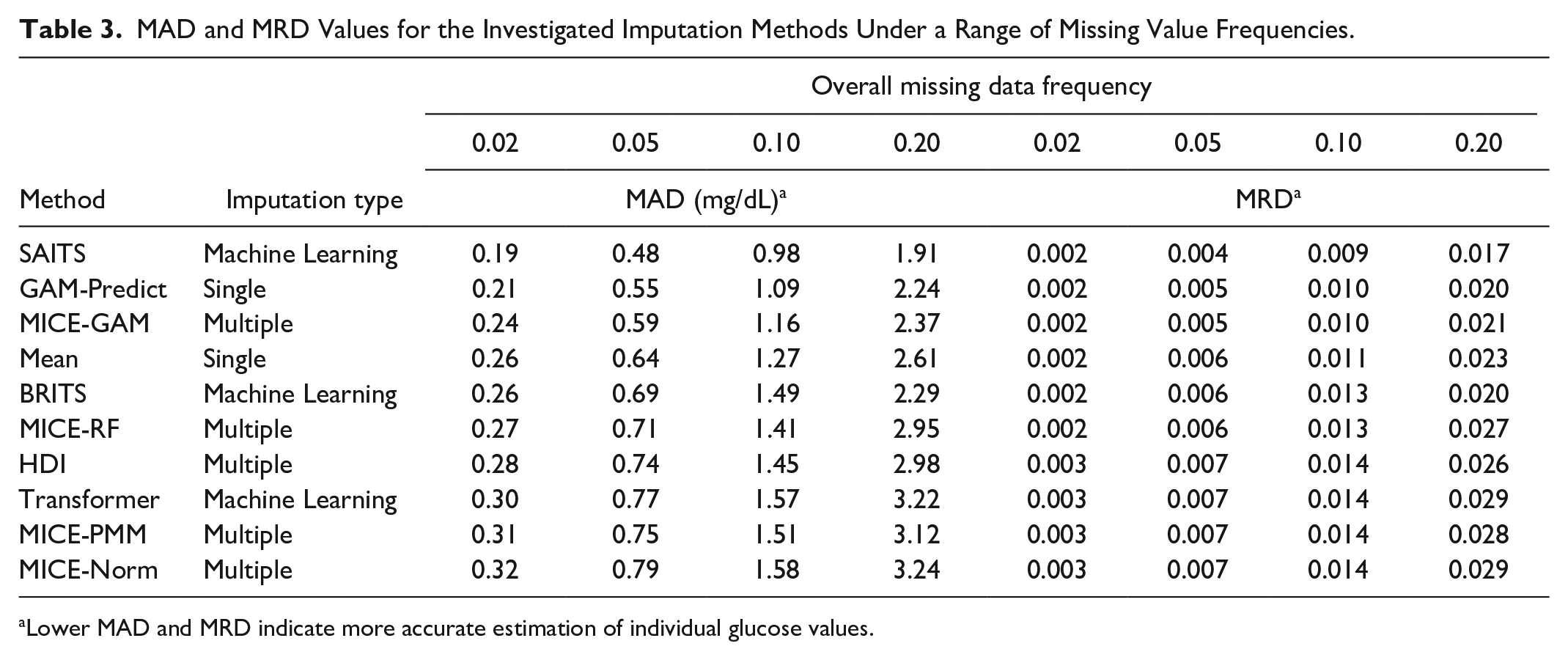
Lower MAD and MRD indicate more accurate estimation of individual glucose values.
Continuous Glucose Monitoring Summaries
Heatmaps demonstrating extent and direction of bias measured by MRB for time-independent and time-dependent CGM summary metrics are depicted in Figures 3 and 4, respectively, and in Supplemental Table 3. Results were excluded for GAM-Predict because over-estimation was so extreme that, when included in the heatmap, the results for the other metrics were visually indistinguishable.
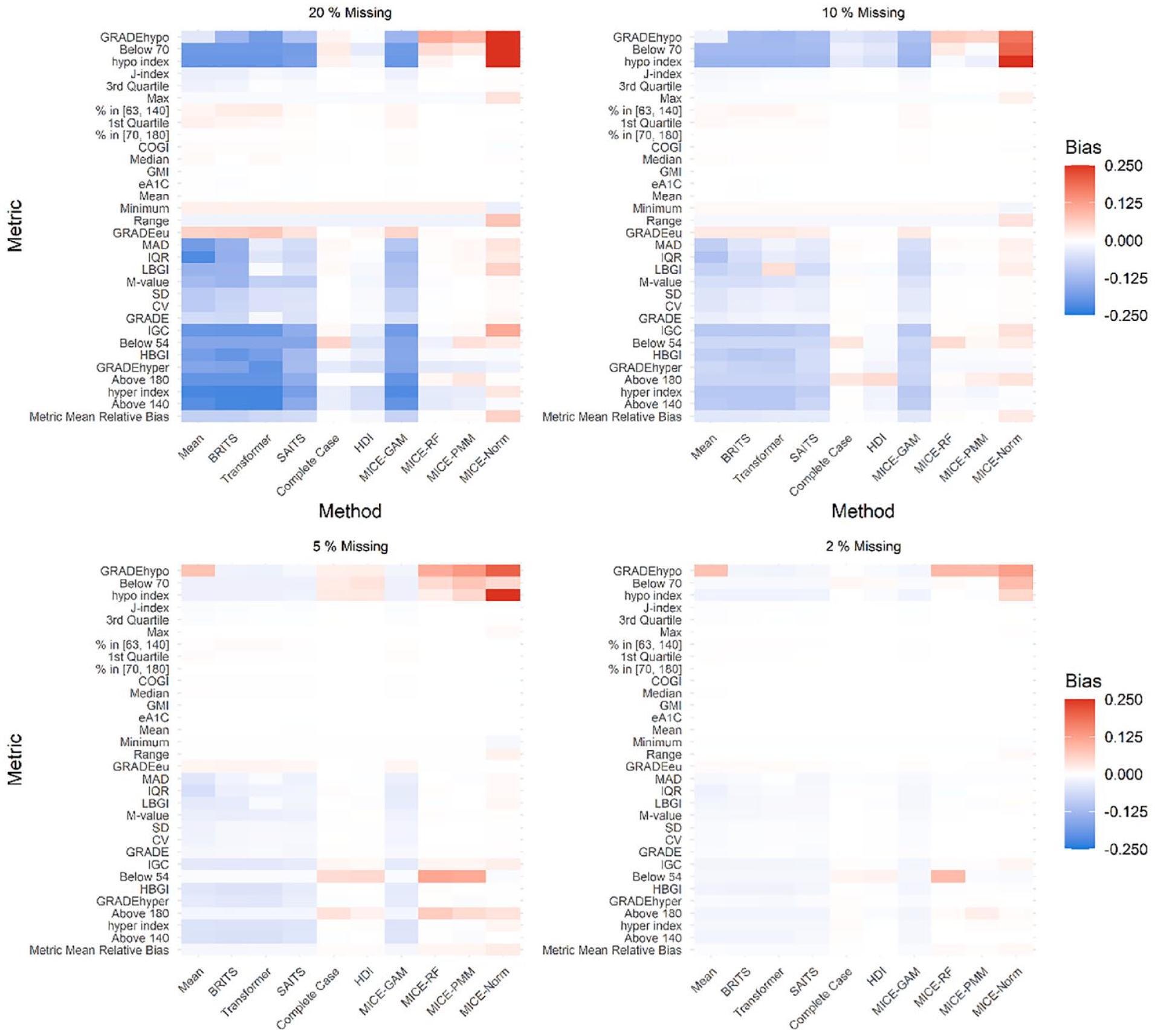
Heatmap of time-independent iglu metrics MRB across methods for the range of missing data percentages. Metrics (row labels) are defined in Supplemental Table 2.
For overall missingness of 2%, a frequency consistent with our 448 GO MOMs test cases with missing strings, MRB is minimal across all metrics and methods. Exceptions include the time-independent “GRADEhypo” metric for the mean method and three of the four MICE methods, where MRB is approximately 0.10. As expected, bias increases as missingness increases, with trends depending on metric and imputation method.
For time-independent metrics (Figure 4), as the percentage of overall missingness increases from 2% to 20% within a period of wear, trends emerge with mean, BRITS, Transformer, SAITS, and MICE-GAM methods demonstrating substantial negative MRB values indicating under-estimation for approximately half of the metrics. Performance of MICE-Norm is also worse as missingness increases, but with positive MRB values indicating over-estimation, with values approaching 0.25 for GRADEhypo, Below 70, and hypo index at 20% missing. The MICE-PMM and MICE-RF also tend toward over-estimation of these metrics. For central tendency metrics such as mean, median, J-index, continuous glucose monitoring index (COGI), glucose management indicator (GMI), and % time in target range (defined as 63-140 mg/dL for pregnancy or 70-180 mg/dL outside of pregnancy), all imputation methods perform comparably well with little bias at the highest levels of missingness. When viewed collectively according to mean MRB (bottom row of each heatmap), complete case, HDI, MICE-RF, and MICE-PMM demonstrate low MRB. For individual time-independent metrics, across all simulation settings, complete case analysis, where missing data were omitted and not imputed, and HDI maintain the smallest and best MRB.
For time-dependent metrics (Figure 5), trends emerge as missingness increases, with mean, BRITS, Transformer, SAITS, and MICE-GAM methods primarily demonstrating negative bias. Positive bias indicating over-estimation is increasingly evident for MICE-RF, MICE-PMM, and MICE-Norm for standard deviation of daily mean (SdDM), standard deviation of coefficient of variation (CVsd), standard deviation of rate of change (SdROC), glucose variability percentage (GVP), standard deviation within series (SdWSH), and glycemia risk index (GRI) metrics. Patterns for complete case analysis and HDI varied; according to mean MRB for all methods (bottom row of each heatmap), these approaches demonstrated the lowest collective bias. Of interest, area under the curve (AUC) demonstrated little bias across all methods and simulation settings. When limited to
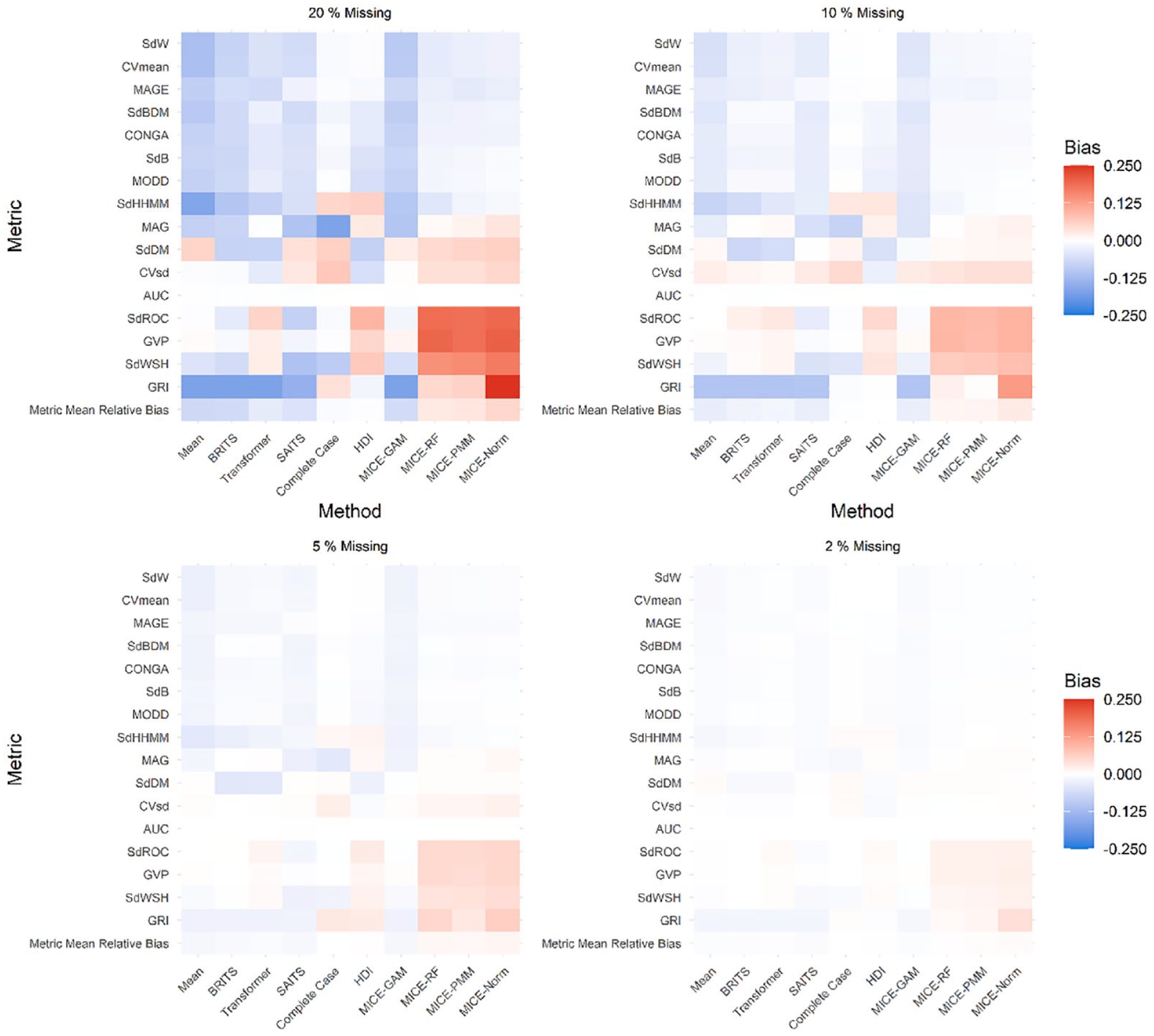
Heatmap of time-dependent iglu metrics MRB across methods for the range of missing data percentages. Metrics (row labels) are defined in Supplemental Table 1.
Discussion
Missing data are a documented phenomenon in CGM data. In GO MOMs, frequency of missing data is low, with 2% missing in the data set used for this analysis. In people with diabetes, CGM with only 70% to 80% completeness over 14 days is representative of 90 days of glycemia as reflected by hemoglobin A1c.22 -25 The goal of this investigation was to review bias across a range of CGM summaries for complete case analysis, where missing data were omitted, and several single, multiple, and machine learning imputation approaches. Our results indicate that complete case analysis and HDI demonstrate superior performance according to CGM metrics evaluated here.
At the level of individual data points, SAITS and GAM-Predict demonstrate best MAD and MRD. Imputation accuracy at the individual level for dense longitudinal data has been described for SAITS. 19 The accuracy of GAM-Predict at the individual level may be attributed to sensitivity to local trends when making predictions to estimate missing values. Worse MAD and MRD for MICE and certain machine learning methods may be attributable to variability that is inherent in multiple imputation methods.
While individual-level accuracy is a strength, analysis of individual CGM data points is not typically a primary goal. Owing to their longitudinal density, CGM data are generally summarized for statistical analysis. We evaluated a comprehensive set of metrics from the
Despite its strengths for individual-level imputation, SAITS, along with BRITS and Transformer machine learning methods, generally underestimated summary metrics, with bias growing worse as missingness increased. The MICE-GAM also demonstrated negative bias, whereas other MICE methods, MICE-RF, MICE-Norm, and MICE-PMM, showed varied patterns of bias depending on the metric.
In simulations, both complete case analysis and HDI demonstrated superior performance in terms of MRB across a variety of metrics and missing data scenarios. This finding suggests that, under certain conditions, HDI or avoiding imputation altogether might be preferred strategies, particularly when missing data are believed to be completely random. The HDI method, which we developed specifically for CGM data, has several features, which may explain its performance. First, it is a non-parametric method that only relies on identified good matches between donors and recipients. Second, HDI replaces
The CGM is considered the standard of care for individuals with diabetes on subcutaneous insulin regimens and is increasingly recognized for management in individuals with type 2 diabetes who are not on insulin therapy. 26 As use of these devices is on the rise in individuals who do not have diabetes, consideration of how to manage data will be critical. Summary metrics for CGM profiles are continually being developed. Strings of missing data, both at random and due to pressure-induced sensor attenuation, are to be expected. Based on our analysis of GO MOMs data under a range of simulated missing data frequencies and a large range of summary metrics, performing complete case analysis even for missing data percentages up to 20% may be acceptable if data are believably missing completely at random (MCAR) such that the probability a value is missing does not depend on observed or unobserved information. A value below the limit of detection may appear to violate the MCAR assumption since, by definition, it is missing due to the unobserved value itself. However, to the extent that these values are due to reduced exchange between blood and interstitial fluid rather than accurate measures of low interstitial glucose values, MCAR is satisfied, and for the observational GO MOMs study in which participants are blinded to their CGM values, an MCAR missing data mechanism is likely acceptable.
Conclusions
The choice of imputation technique must balance accuracy with practicality. If summary metrics are not amenable to missing observations, our results suggest HDI should demonstrate robust performance. If missing data can be accommodated, our simulations suggest that imputation is not necessary in many settings. This may ease computational burden and uncertainty around conclusions drawn from CGM data that include missing data strings. In the evolving field of CGM data analysis, it will be critical to continually refine and assess approaches to missingness and consider the use of imputation techniques. Future research should broaden these evaluations to varied contexts, delve into the effects of non-random missingness, and innovate imputation strategies specifically designed for CGM data.
Supplemental Material
sj-docx-1-dst-10.1177_19322968241308217 – Supplemental material for Imputation of Missing Continuous Glucose Monitor Data
Supplemental material, sj-docx-1-dst-10.1177_19322968241308217 for Imputation of Missing Continuous Glucose Monitor Data by Alan Kuang, Yuanzhi Yu, Juned Siddique and Denise Scholtens in Journal of Diabetes Science and Technology
Supplemental Material
sj-xlsx-1-dst-10.1177_19322968241308217 – Supplemental material for Imputation of Missing Continuous Glucose Monitor Data
Supplemental material, sj-xlsx-1-dst-10.1177_19322968241308217 for Imputation of Missing Continuous Glucose Monitor Data by Alan Kuang, Yuanzhi Yu, Juned Siddique and Denise Scholtens in Journal of Diabetes Science and Technology
Footnotes
Abbreviations
ARDL, Advanced Research and Diagnostic Laboratory; BRC, Biostatistics Research Center; BRITS, Bidirectional Recurrent Imputation for Time Series; CGM, continuous glucose monitoring; GAM, generalized additive models; GDM, gestational diabetes mellitus; GO MOMs, Glycemic Observation and Metabolic Outcomes in Mothers and Offspring; HDI, hot-deck imputation; LGA, large-for-gestational age; MAD, mean absolute difference; MCAR, missing completely at random; MICE, multiple imputation by chain equations; MRB, mean relative bias; MRD, mean relative difference; NUDACC, Northwestern University Data Analysis Center and Coordinating Center; PPM, predictive mean matching; SAITS, Self-Attention Imputation Transformer with Self-Supervision.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by grants from the National Institute of Diabetes and Digestive and Kidney Diseases: U01DK123795 to Massachusetts General Hospital; U01DK123791 to Kaiser Permanente; U01DK123759 and U01DK123745 to Northwestern University; U01DK123799 to Yale University; and U01DK123783 to Women & Infants Hospital of Rhode Island.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
