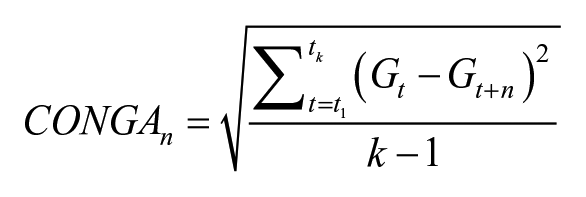Abstract
This report describes how a Conditional Generative Adversarial Network (CGAN) was used to synthesize realistic continuous glucose monitoring systems (CGM) from healthy individuals and individuals with type 1 diabetes over a range of different HbA1c levels. The results showed that even though the CGAN generated data, did not perfectly reflect real world CGM, many of the important features were captured and reflected in the synthetic signals. It is briefly discussed how heterogenous data sources constitutes a challenge for comparison of predictive CGM models. Therefore 40,000 CGM days were generated by the trained CGAN, equivalent to 940,000 hours of synthetic CGM measurements. These data have been made available in a public database, which can be used as a reference in future studies.
Introduction
The introduction of the continuous glucose monitoring systems (CGM) has proven to be a paradigm shift in both the management and understanding of diabetes. 1 CGM offers patients with diabetes and clinicians, the opportunity to improve the patient’s glycemic control by monitoring glucose levels on continuous basis. 2 This makes it possible to identify events of hypoglycemia, hyperglycemia and glycemic variability, which would not be detected with self-measurement of blood glucose (SMBG).3-5 As continuous blood glucose data become available, diabetes technologies such as an artificial pancreas/closed-loop system, personalized decision algorithms, and low/high blood glucose alarms has seen a significant increase in research interest and development. Methods such as predicting future blood glucose levels and modeling glucose dynamics are central to the development of these diabetes management technologies. Many studies have been published on these topics in recent years.6-10 However, the differences in reporting and data used for assessment, makes these systems hard to compare.6,11 There is a need for publicly available CGM databases, which could serve as common benchmark for these studies. 11 This report describes how a Generative Adversarial Network (GAN) was used to synthesize realistic CGM from healthy individuals and individuals with type 1 diabetes over a range of different HbA1c levels. Furthermore, a large database of synthetic CGM was made available, which can be used as a reference for future CGM modeling studies.
Methods
Modeling Approach
GAN is one of the most promising developments in deep learning and it can be used to produce synthetic data that resemble real data input. GAN was first introduced by Goodfellow et al. in 2014, 12 the publication describes the problem of unsupervised learning by training 2 deep (convolutional) neural networks, called generator and discriminator. The networks contest with each other in the form of a zero-sum game where the generator is trained to generate new data and the discriminator model tries to classify input as either real data or generated. This training procedure continues until the discriminator model is unable to distinguish between the real and the simulated data, which indicate that the generator model is generating a plausible signal.
Conditional GANs (CGANs) is an adjustment of regular GANs which can use data labels to generate data belonging to specific categories. In this study we trained a CGAN based on 24-hour CGM profiles from healthy individuals and individuals with type 1 diabetes. HbA1c levels, categorized into 4 labels: below 6.5% (healthy without diabetes), between 6.5% and <7%, between 7% and <8% and above 8%, were used as the condition. The architecture of the CGAN is illustrated in Figure 1.
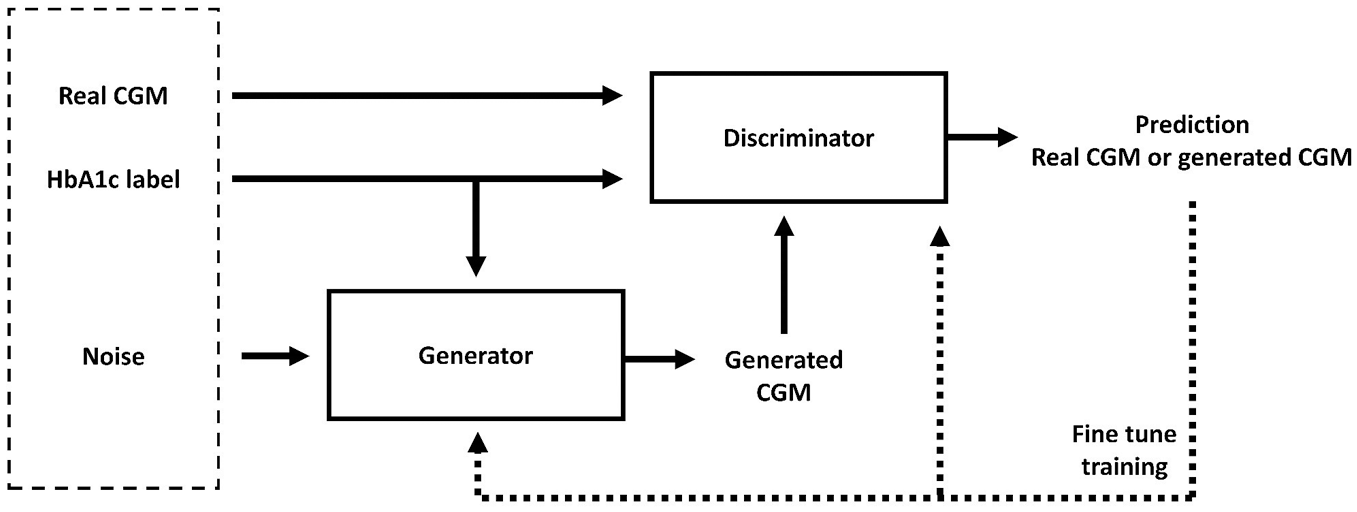
The architecture of the CGAN model.
Data Sources
The cohort for this study was combined from 2 studies, one on type 1 diabetes and one on healthy individuals - the T1D Exchange Severe Hypoglycemia in Older Adults with Type 1 Diabetes Study and a multicenter study on continuous glucose monitoring profiles in healthy participants without diabetes.13,14 A total of 786 full days of measurements were available from the 153 healthy individuals and a total of 1191 full days were available from the 200 individuals with type 1 diabetes. Among the participants with diabetes, 374 days were available from individuals with an HbA1c level of 6.5% to <7%, 421 days with an HbA1c level of 7% to <8% and 228 days with an HbA1c level above 8%. A single CGAN was trained on all available CGM data from the 2 studies.
Synthetic CGM Data Generation
Based on the trained CGAN, we generated 40,000 CGM days, equivalent to 940,000 hours of synthetic CGM values, which we made available as a public database. The CGAN was modeled and trained using Matlab R2020b (The Mathworks Inc., Natick, Massachusetts). The data are available at Mendeley data repository. 15
Statistical Comparison
To evaluate the similarities of the CGAN generated CGM measurements compared to the real CGM profiles we calculated several statistical measures often used to characterize glycemic control from CGM. 24-h (whole day) and night means were used to assess the general glucose level; this is highly correlated to HbA1c. 16 Time above 180 mg/dL and below 70 mg/dL were calculated to respectively assess hyperglycemia and hypoglycemia durations. Standard deviation of the 24-h mean (SD) and Continuous overlapping net glycemic action (CONGA) were used to assess the glycemic variability in the datasets.
CONGA
17
is calculated by determining the difference between values at different intervals.
Results and Comments
A total of 40,000 days of synthetic CGM measurements were successfully generated using the trained CGAN. Examples of CGAN generated CGM data are illustrated in Figure 2. The assessment of blood glucose characteristics for both the synthetic CGM and the real CGM datasets are presented in Table 1. The 24-h means for each of the 4 HbA1c groups were comparable between both datasets. The SD of the mean was lower in the synthetic data compared to the real CGM dataset, indicating that not all the 24-h variability was modeled by the CGAN. Time spend in hyperglycemia and hypoglycemia within each HbA1c group were also similar between the synthetic and real dataset. Both SD and CONGA were higher for each group in the synthetic dataset but the correlation between HbA1c level and increased glycemic variability was well captured by the modeling.
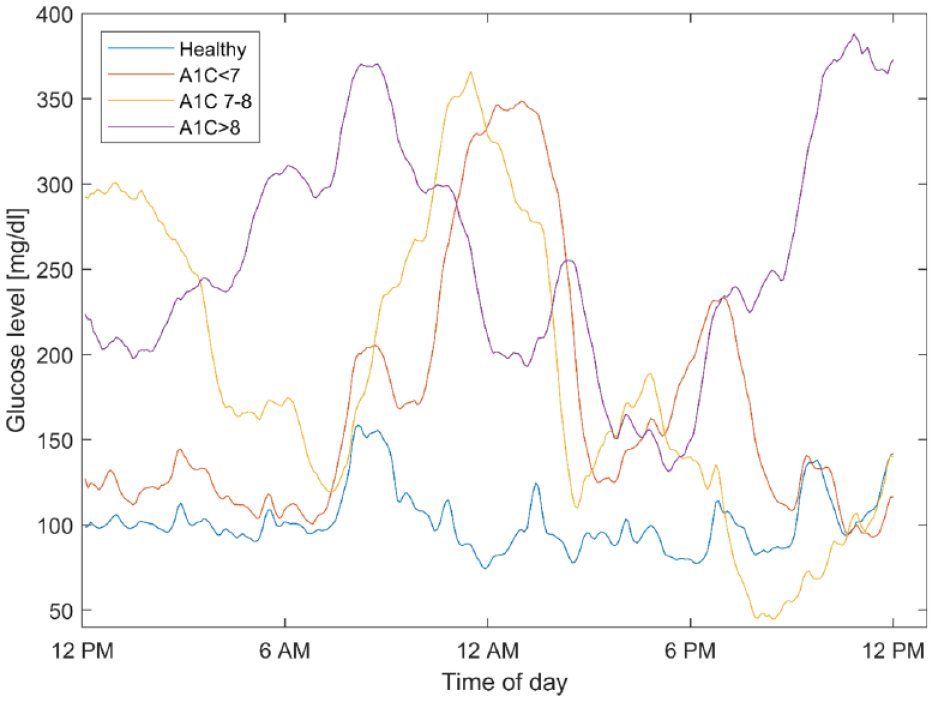
Examples of 4 synthetic generated CGM signals, one for each of the classes; healthy, diabetes with A1C <7%, diabetes with A1C 7-8%, and diabetes with A1C >8%.
Comparison of Characteristics Between Real CGM and Synthetic CGM Presented as Average (SD) or 50th (25th;75th) Percentile.
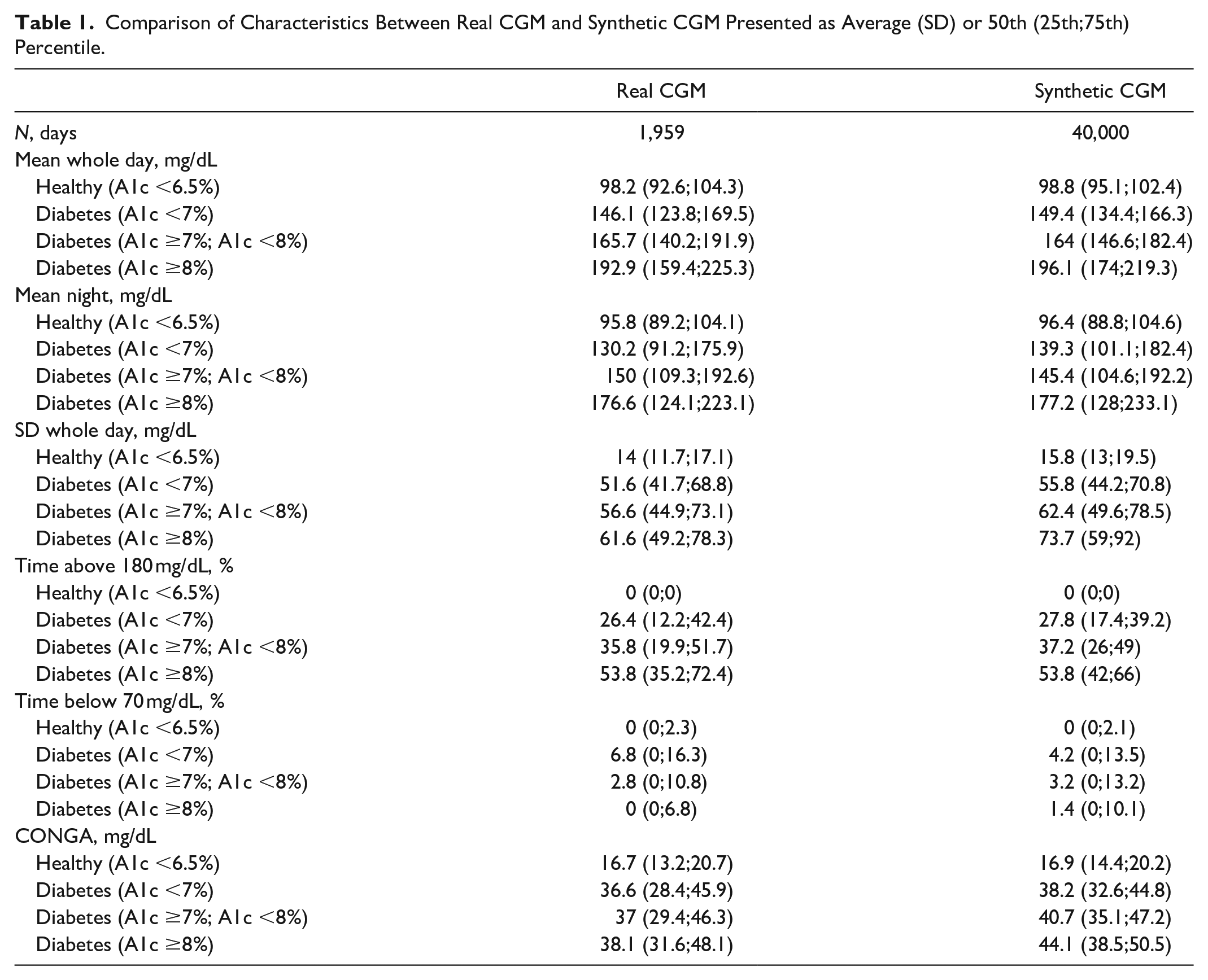
In summary, general increased mean levels of glucose, increased glycemic excursions, increased time spend in hyperglycemia and decreased time spend in hypoglycemia are known to be related to increased HbA1c in patients with diabetes. These characteristics were preserved in the synthetic CGM dataset.
Discussion
In this report, we propose a method to synthesize CGM signals, using a supervised machine learning approach, CGAN. The method was originally proposed as a method to generate synthetic images resembling real images. This study shows, how this method can be used to efficiently generate synthetic CGM signals that resemble real world data. While the CGAN generated data, does not perfectly reflect real world measurements, many of the important features from the individuals modeled are captured and present in the synthetic signals. The synthesized data is a readily accessible source of CGM measurements, that can be used as a common external comparator for predictive models developed on often heterogenous data sources. Predictive models based on CGM measurements could be a major advancement in technologies for the treatment of diabetes and have several scientific and clinical implications. The predictive performance of these models is often assessed by splitting the initial dataset into a training and a test set, with the test set thus considered an independent sample. This method is proven to be inefficient and risks incorrect assessment of actual performance. 18 Authors should consider using truly independent data for more correct assessment of potential overfitting as well as external validity. There is a need for an available database which can be used as a common benchmark in developing and assessing the performance of predictive models based on CGM. Furthermore, synthetic data could be used for clinical training of doctors and nurses working with diabetes patients. One of the potential use cases could be in a decision support system where the synthetic CGM is used as a reference for people being actively treated.
While most important characteristics are maintained in the proposed model, this type of model could be further developed, incorporating even more characteristics of the patients to accurately resemble specific patient characteristics and behaviors; for example, insulin dosages, meals and physical activity. This could have applications in many areas beyond predictive research. For example, in the pharmaceutical industry where large expensive cohort studies are often needed to validate findings, CGAN generated realistic data could be used as a part of in silico testing to reduce stress on patients and expensive need for in vivo testing.
Footnotes
Acknowledgements
N/A
Abbreviations
CGAN, Conditional Generative Adversarial Network; CGM, continuous glucose monitoring systems; CONGA, continuous overlapping net glycemic action; SD, standard deviation; SMBG, self-measurement of blood glucose.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Disclaimer
The source of the data is the T1D Exchange, but the analysis, content and conclusions presented herein are solely the responsibility of the authors and have not been reviewed or approved by the T1D Exchange.