Abstract
Purpose
Venous thromboembolism (VTE), including deep vein thrombosis and pulmonary embolism, is a leading cause of morbidity and mortality in cancer patients. Prostate cancer is associated with an elevated risk of VTE, yet the molecular drivers remain poorly defined.
Methods
In this study, we employed high-throughput proteomic profiling using the SomaLogic platform to analyze plasma from 85 prostate cancer patients, including 43 with and 42 without VTE. Samples were collected at cancer diagnosis, with VTE diagnosed at a mean of 96.8 months later.
Results
Principal component analysis showed modest proteomic separation between groups. Differential expression analysis identified enriched pathways in VTE patients, including hemostasis (TIMP1, JAM2, TMX3, F3, and ESAM), cell adhesion (CXCL12, CCL11, CCN5, COL18A1, and ADGRB1) and cell proliferation (TIMP1, REG1B, REG1A, CRLF2, and ALDH1A2). Receiver Operating Characteristic analysis using top fifteen proteins achieved an area under the curve of 0.859, indicating strong predictive value for VTE in this cohort.
Conclusions
We identified a specific cluster of circulating proteins associated with development of VTE in patients with prostate cancer. This work deepens understanding of systemic mediators of cancer-associated VTE and, pending validation in other cohorts, paves the way for improved risk stratification and long-term monitoring in this population.
Introduction
Venous thromboembolism (VTE), encompassing deep vein thrombosis (DVT) and pulmonary embolism (PE), is a serious complication in cancer patients. 1 VTE is a predominant cause of mortality in cancer patients, ranking second only to cancer-related fatalities. 2 Prostate cancer, one of the most prevalent malignancies in men, has been associated with an increased risk of thrombosis, contributing to morbidity and mortality.3,4 Multiple factors, including tumor-derived procoagulant activity, systemic inflammation, treatment-related effects, and patient-specific risks such as advanced age and comorbid conditions influence the hypercoagulable state in these patients. 5
Despite advances in understanding cancer-associated thrombosis, the precise molecular mechanisms remain unclear. Even though cancer is a local disease, it has systemic ramifications that are mediated by circulating factors that elicit distant effects. In the past, we demonstrated that small metabolites serve as systemic mediators of cancer-associated thrombosis. 6 The presence of cancer can also alter the proteomic composition from specific proteins secreted by cancer cells or proteins generated in other cell types in response to cancer. We, therefore, hypothesized that serum factors may transmit signals from the local cancer environment to distant organs, influencing deep vein thrombosis.
Emerging evidence suggests that sera proteins, such as tissue factor, C-reactive protein, soluble P-selectin and D-Dimer, play a critical role in modulating thrombotic risk, influencing coagulation, fibrinolysis, endothelial integrity, platelet activation, and immune responses.7,8 Identifying key plasma proteins associated with VTE in prostate cancer patients may provide valuable insights into disease pathophysiology and help establish novel biomarkers for risk prediction, early detection, and targeted prophylaxis thereby, optimizing the risk-benefit ratio. Therefore, we adopted an unbiased approach using Soma logic to understand how a systemic protein profile might be altered in the presence of cancer, and whether a set of proteins can be leveraged to predict venous thrombogenicity in patients with prostate cancer. Such a panel has the potential to improve patient stratification, guide personalized thromboprophylaxis, and enhance clinical management strategies to mitigate VTE-related complications in prostate cancer patients.
Methods
Data source and study population
We conducted a retrospective cohort study using data from patients enrolled in the Cooperative Studies Program (CSP) #719B between 2001 to 2006. 9 Clinical variables were obtained from two Veterans Affairs (VA) data systems: Corporate Data Warehouse (CDW) and the VA Cancer Registry System (VACRS). The CDW is a national repository that integrates clinical, administrative, and electronic health record (EHR) data from VA healthcare facilities across the United States. 10 VACRS is maintained by trained cancer registrars at each of the 132 VA Medical Centers that diagnose or treat Veterans with cancer, and includes detailed information on cancer diagnoses, staging, and treatments.11,12
Outcome definition
The primary outcome was the occurrence of venous thromboembolism (VTE) following prostate cancer diagnosis. VTE was defined as radiologically confirmed symptomatic or incidental pulmonary embolism (PE), lower extremity deep vein thrombosis (LE-DVT), and upper extremity DVT (UE-DVT). Occurrence of VTE was ascertained using a previously published algorithm that combines structured data (diagnosis codes and receipt of anticoagulation) and unstructured data (radiology reports parsed with a natural language processing method). 13 This method has been validated to have high sensitivity (96%) and positive predictive value (91%) in the VA database. 13
Covariates
Covariates were defined based on data recorded prior to prostate cancer diagnosis date. Age at the diagnosis was calculated using date of birth recorded in the CDW. Sex was recorded as per the CDW records. Race and ethnicity data was self-reported by patients and categorized in the EHR using standardized CDW-defined categories. The race categories included American Indian or Alaska Native, Asian, Black or African American, Declined to Answer, Native Hawaiian or Other Pacific Islander, Unknown by Patient, White, and White not of Hispanic Origin. The ethnicity categories included Hispanic or Latino, Not Hispanic or Latino, Unknown by Patient and Declined to Answer. For analysis, race and ethnicity were consolidated into four mutually exclusive categories: Hispanic, Non-Hispanic Black, Non-Hispanic White, or Other/Unknown. The Other/Unknown category encompassed responses from American Indian or Alaska Native, Asian, Native Hawaiian or Other Pacific Islander, Declined to Answer, and Unknown by Patient. Anticoagulation prescription data were extracted from the VA CDW in a window of 6 months before up to 30 days after prostate cancer diagnosis from outpatient settings. Pre-treatment Body Mass Index (BMI) was calculated using height and the nearest weight prior to prostate cancer diagnosis recorded within a 36-month lookback window prior to diagnosis, extracted from the VA vital status. Cancer-specific covariates included clinical stage and Gleason grade at diagnosis, both obtained from the VA Cancer Registry System. Cancer stage was classified according to the American Joint Committee on Cancer (AJCC) system (I, II, III, IV, or unknown); when multiple stages were recorded, the highest documented stage was used. Gleason grade at the time of prostate cancer diagnosis was extracted from the VA Cancer Registry. Platelet count data were extracted from the CDW laboratory files within a window of 365 days before to 365 days after prostate cancer diagnosis.
Statistical analysis
To evaluate differences in protein expression associated with VTE, patients diagnosed with VTE (cases) were matched to patients without VTE (controls) using propensity score matching. The propensity scores were estimated using regularized logistic regression with VTE occurrence as the outcome and the following covariates as predictors: age at diagnosis, race/ethnicity, body mass index (BMI), history of anticoagulation treatment, prostate cancer stage at diagnosis, and Gleason grade. A 1:1 nearest-neighbor matching algorithm with a caliper of 0.01 (1% of the propensity score distribution) was used to ensure balance between groups.
SomaScan proteomics
Plasma samples were collected in EDTA tubes from patients after exsanguination, prior to initiation of cancer therapy. After matching, samples underwent a SomaScan Assay v4.1 (SomaLogic, Boulder, CO, USA). This assay uses DNA-based aptamers rather than antibodies, which are modified at the nucleotide level to bind more tightly and with a slow off-rate. 14 These two factors confer a higher degree of specificity to protein targets. For each patient 100 µL plasma samples were sent to SomaLogic and analyzed for 7288 unique protein targets.
Downstream analysis to identify differentially expressed proteins
Processed proteomics data from the SomaScan assay were analyzed using R statistical software. Raw data were imported and converted into Expression Set objects using readat package (version 1.4.1).
15
Samples were categorized into two groups: control (no VTE) and cases (VTE). Principal component analysis (PCA) was performed using the base R function prcomp to assess sample clustering and detect batch effects. Due to limited variability in proteomic expression data, log-transformed expression values were filtered based on group-based signal-to-noise ratio (SNR), calculated as the difference in group means divided by the sum of standard deviations, to enrich for biologically relevant features. An absolute SNR > 0.2 cutoff was used to select 406 proteins for downstream differential expression analysis. Differentially expressed proteins (DEPs) were
Gene ontology and pathway analysis
Functional enrichment analysis of DEPs was conducted using the Database for Annotation, Visualization and Integrated Discovery (DAVID) platform. 17 Each list of DEPs was uploaded separately, and enrichment analyses for Gene Ontology (GO) biological processes and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathways were performed. Enrichment results were considered significant at p < 0.05. Bar graphs were generated to present the top enriched functions and pathways. Guided by results of global protein survey, proteins associated with key biological pathways-including hemostasis, cell adhesion and cell proliferation-were extracted for subsequent analysis. A heatmap was generated using these pathway specific proteins to depict the relative expression levels between the groups. These proteins were also annotated in volcano plots to highlight their statistical and biological relevance.
Linear regression of pathway proteins
To assess the predictive utility of proteins involved in thrombosis-associated biological pathways, generalized linear models (GLMs) were fitted using a binomial distribution, with VTE status (case/control) as the binary outcome. Four separate models were generated: one using the top five DEPs from all enriched pathways combined, and one each using the top five DEPs from the cell adhesion, proliferation, and hemostasis pathways individually. For each model, a receiver operating characteristic (ROC) curve was generated to assess model performance and the trade-off between true positive rate (sensitivity) and false positive rate (1- specificity). The area under the curve (AUC) was calculated for each model to quantify predictive accuracy and assess strength of pathway specific DEPs in discriminating VTE status. To estimate the variability in AUC associated with this model, we used non-parametric bootstrap resampling (n = 10,000) with replacement and calculated AUC scores for each iteration. The mean AUC and the 95% confidence interval (2.5th–97.5th percentile) were reported.
Results
A total of 85 human prostate cancer plasma samples were analyzed, comprising 43 samples derived from patients who developed venous thromboembolism (VTE) (Figure 1, Table 1, Supplementary Table 1) and 42 from patients who did not experience thrombotic complications. One patient was excluded due to insufficient specimen quality. There were no differences between the two groups in other parameters. All samples were collected subsequent to prostate cancer diagnosis, whereas VTE was diagnosed at a mean follow-up period of 96.8 months.
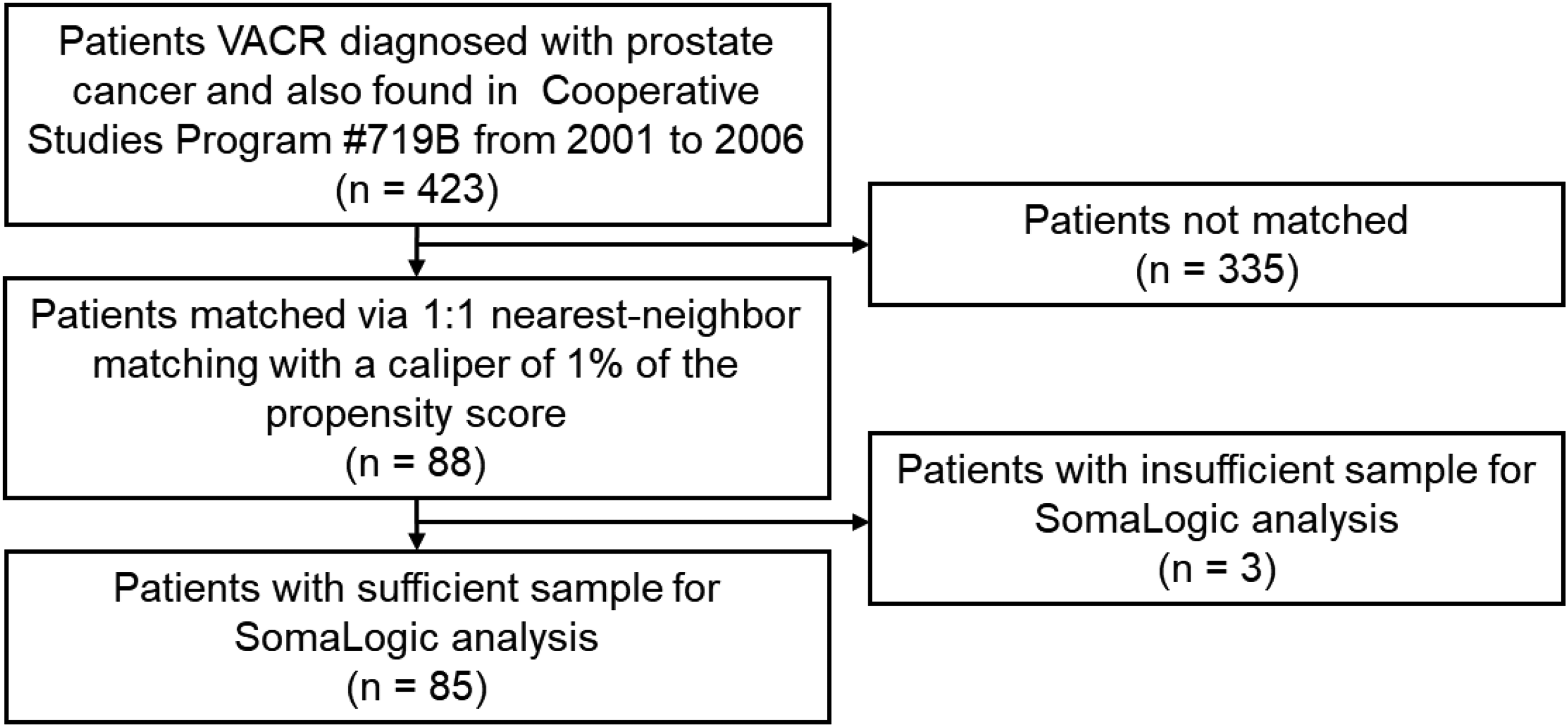
Patient flow diagram.
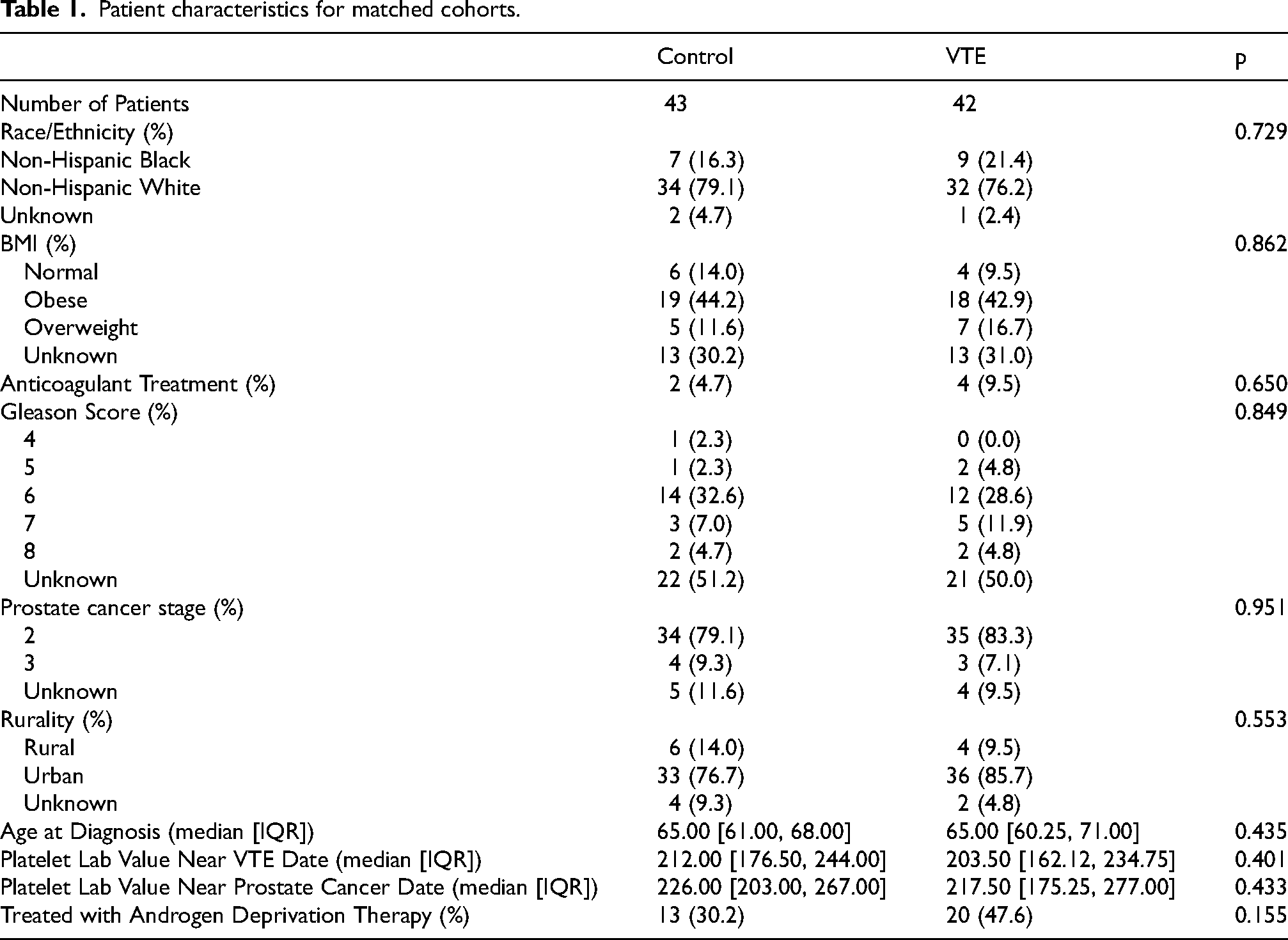
Patient characteristics for matched cohorts.
To identify potential circulating protein markers associated with thrombosis and/or cancer progression, we employed the SomaLogic platform for high-throughput proteomic profiling (see Methods). Principal component analysis (PCA) demonstrated a mild separation between the VTE and non-VTE groups (Figure 2A), suggesting distinct proteomic signatures. Assessment of clinic-pathological parameters revealed that Gleason scores were similarly distributed between the two groups when compared by age (<65 and >65 years) (Figure 2B and 2C), indicating that age and tumor grade were not significantly different between VTE and non-VTE patients. All variables that were adjusted for in the propensity score were similar between the two groups.
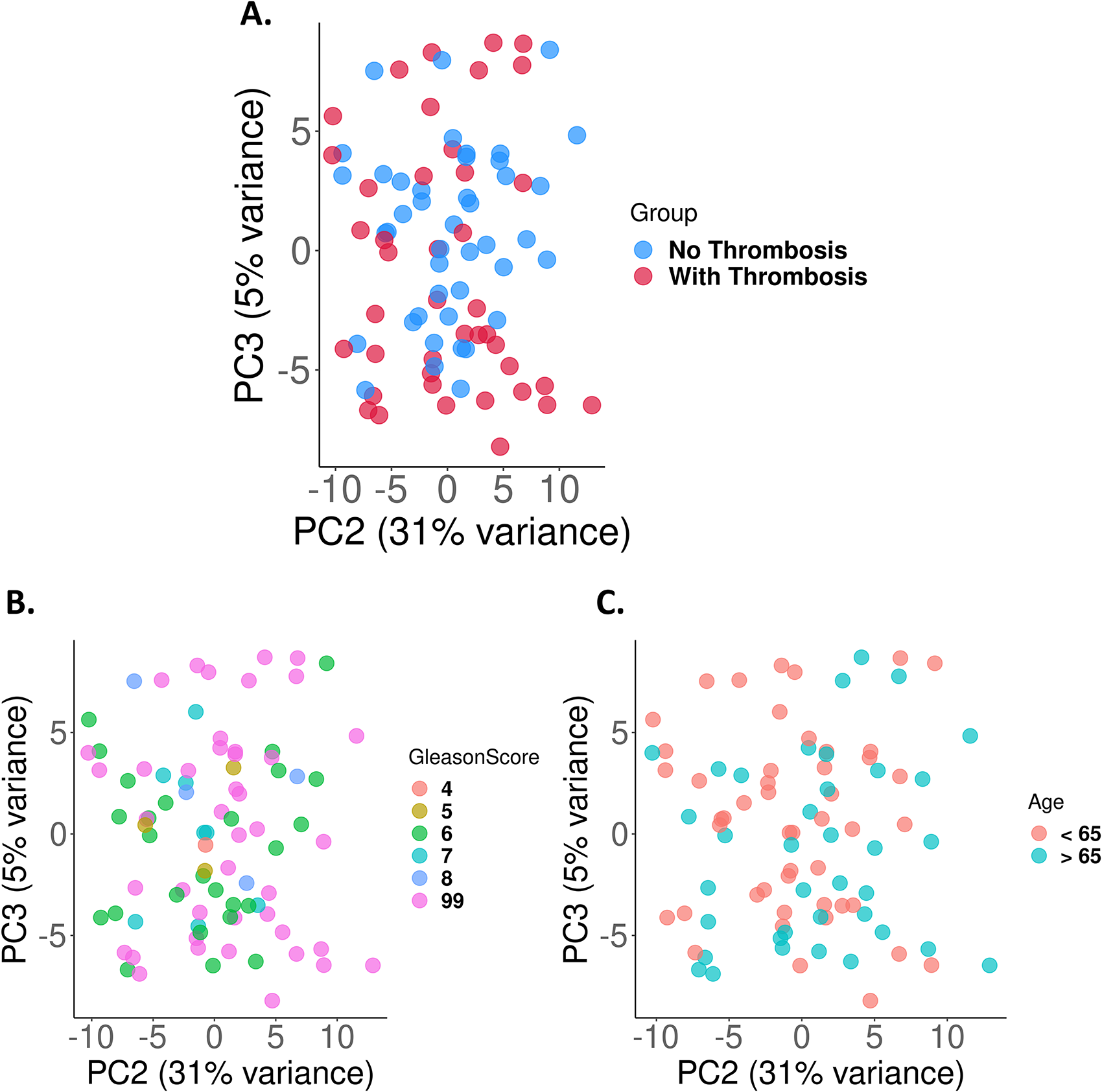
(A) Principal Component Analysis (PCA) plot of protein expression profiles for prostate cancer patient samples with and without thrombosis. PCA plot of protein expression profiles for prostate cancer patient samples grouped by (B) Gleason score and (C) age.
Differentially expressed proteins (DEPs) enriched in the thrombosis group were analyzed using the DAVID functional annotation tool. Functional enrichment revealed a significant association with pathways involved in cell adhesion, proliferation, and hemostasis (Figure 3A). The corresponding plasma proteins were notably elevated in patients who subsequently developed thrombotic complications (Figure 3B, Supplementary Table 1).
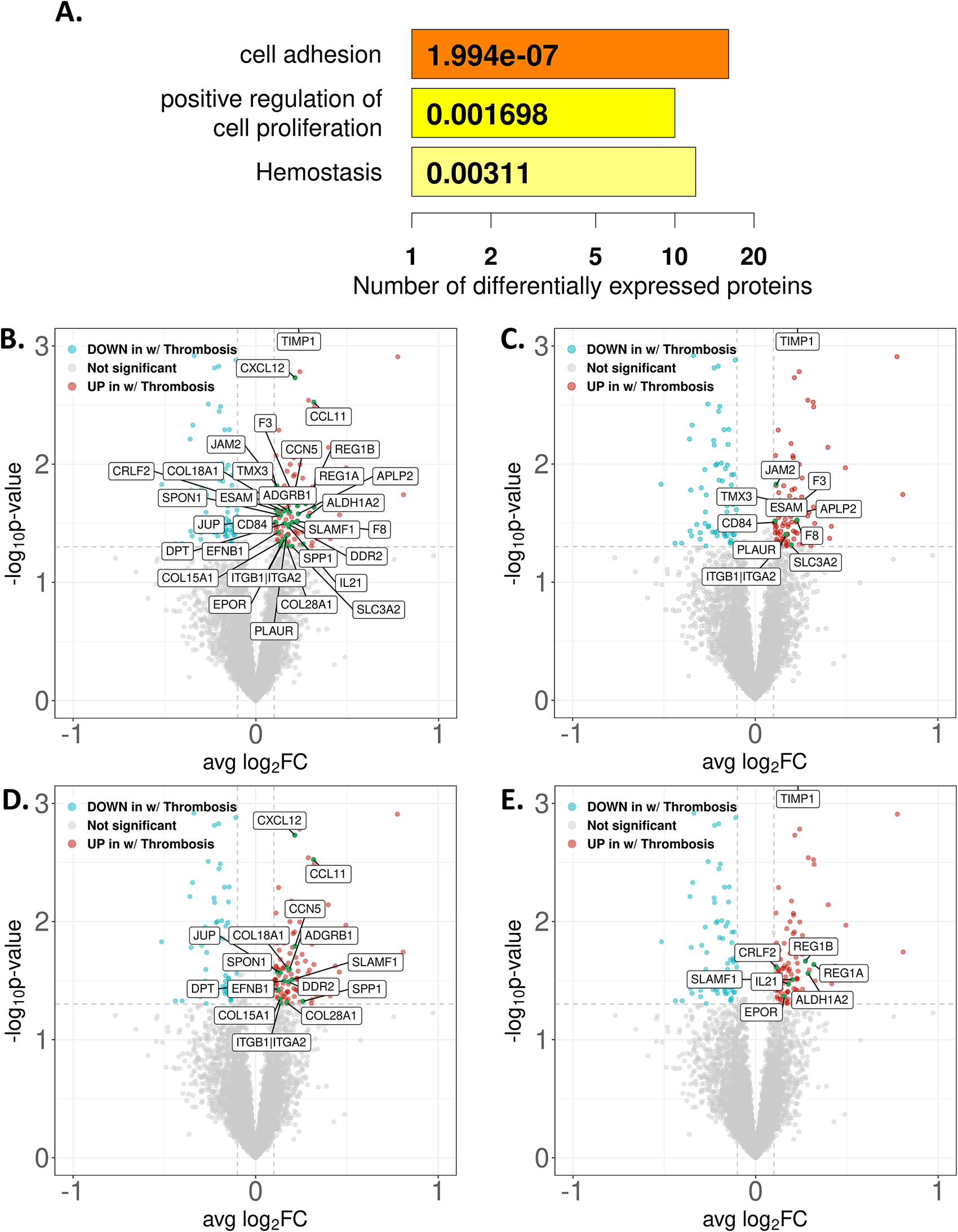
(A) Bar plot showing enriched pathways related to hemostasis, cell adhesion and cell proliferation identified from differentially upregulated proteins in thrombosis positive prostate cancer samples using the Database for Annotation, Visualization, and Integrated Discovery (DAVID) functional annotation tool. (B) Volcano plot illustrating differentially expressed proteins between prostate cancer samples with and without thrombosis. Individual volcano plots highlighting significantly regulated proteins associated with enriched pathways (C) hemostasis (D) cell adhesion, and (E) cell proliferation.
Among the DEPs, several proteins implicated in hemostasis, cellular adhesion, and proliferation- processes that contribute to both thrombosis and tumor progression- were significantly upregulated in the thrombosis group. These included hemostatic regulation proteins such as TIMP1, JAM2, TMX3, F3, ESAM, APLP2, F8, CD84, PLAUR, ITGB1,ITGA2, and SLC3A2 (Figure 3C). The proteins involved in cell adhesion that were significantly expressed in the thrombosis group included CXCL12, CCL11, CCN5, SLAMF1, SPP1, COL18A1, ADGRB1, SPON1, DPT, JUP, EFNB1, DDR2, COL15A1, ITGB1, ITGA2, and COL28A1 (Figure 3D). The proteins involved in cell proliferation that were significantly expressed in the thrombosis group included TIMP1, REG1B, REG1A, CRLF2, ALDH1A2, SLAMF1, IL21, and EPOR (Figure 3E). Collectively, these findings highlight candidate biomarkers and potential mechanistic links between thrombosis and prostate cancer progression.
To further investigate the prognostic value of the enriched pathways and their correlation with thrombosis in prostate cancer, we selected the top five statistically significant proteins associated with thrombosis from each of the three pathways. From the hemostasis pathway, TIMP1, JAM2, TMX3, F3, and ESAM were selected (Figure 4A); from the cell adhesion pathway, CXCL12, CCL11, CCN5, COL18A1, and ADGRB1 (Figure 4B); from the cell proliferation pathway, TIMP1, REG1B, REG1A, CRLF2, and ALDH1A2 (Figure 4C). Receiver operating characteristic (ROC) analysis was performed using a linear regression model to evaluate the predictive utility of these proteins for thrombotic risk in prostate cancer patients. Individual ROC curves were generated for each pathway as well as for a combined model integrating all 15 proteins. The combined model demonstrated strong predictive performance, yielding an area under curve (AUC) of 0.859, bootstrapped AUC of 0.917, and a 95% confidence interval (CI) of 0.841–0.984 (Figure 5A, Supplementary Table 2), indicating high discriminatory power for identifying patients at risk of thrombosis. Pathway-specific analyses revealed AUC values of 0.745 (bootstrapped AUC = 0.779, 95% CI: 0.674–0.873) for the hemostasis pathway (Figure 5B), 0.781 (bootstrapped AUC = 0.805, 95% CI: 0.704–0.894) for the cell adhesion pathway (Figure 5C), and 0.783 (bootstrapped AUC = 0.809, 95% CI: 0.712– 0.894) for the cell proliferation pathway (Figure 5D). While proteins involved in cell proliferation yielded the highest individual AUC, all three pathways contributed meaningfully to the overall predictive model. These results suggest that a multi-pathway protein panel could serve as a robust tool for predicting thrombotic complications in patients with prostate cancer.
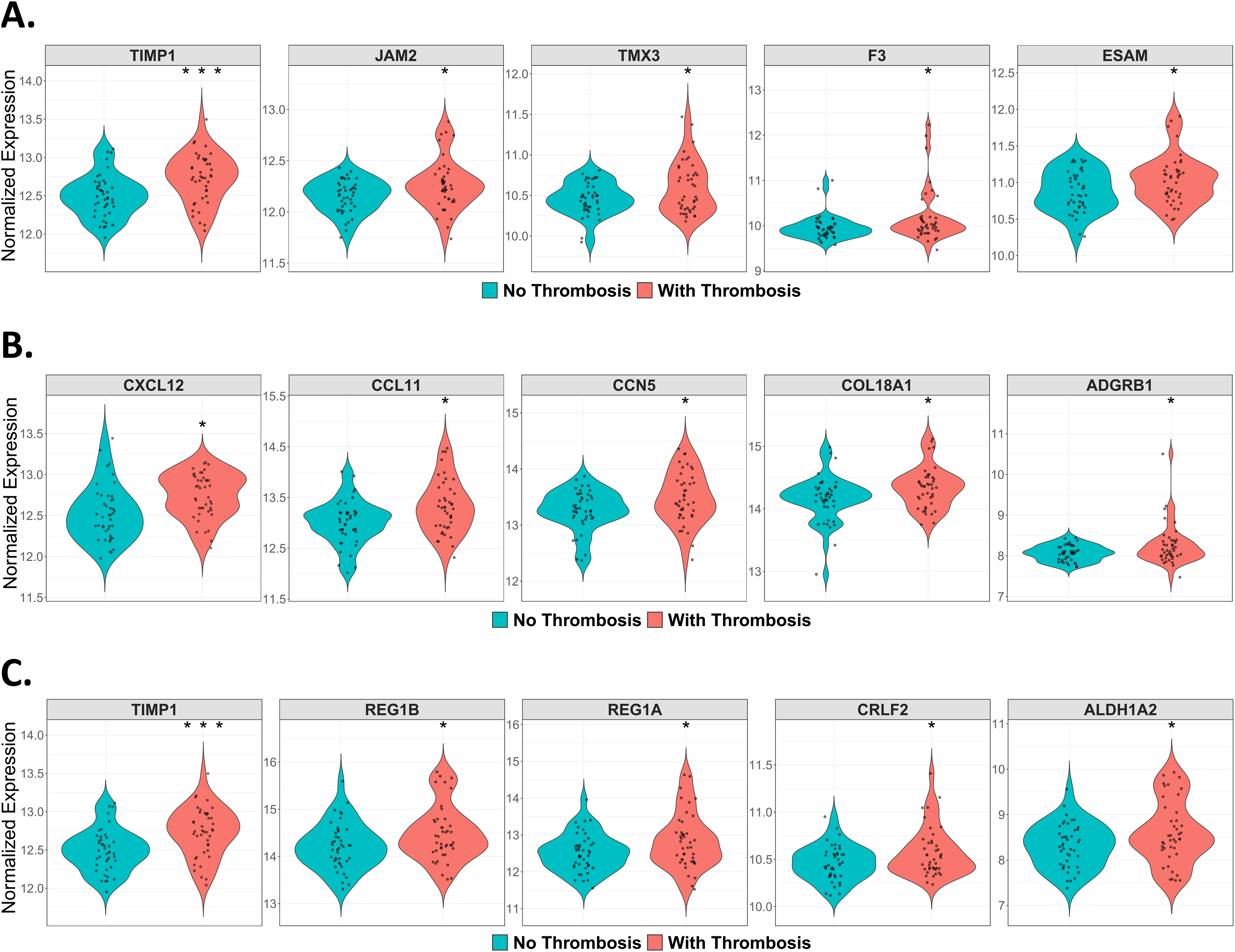
Violin plots displaying the distribution and variance of normalized expression levels for the top five differentially expressed proteins within each enriched pathway, including (A) hemostasis (B) cell adhesion, and (C) cell proliferation. Each plot represents protein expression across individual prostate cancer patient samples, highlighting differences in expression patterns related to thrombosis status.
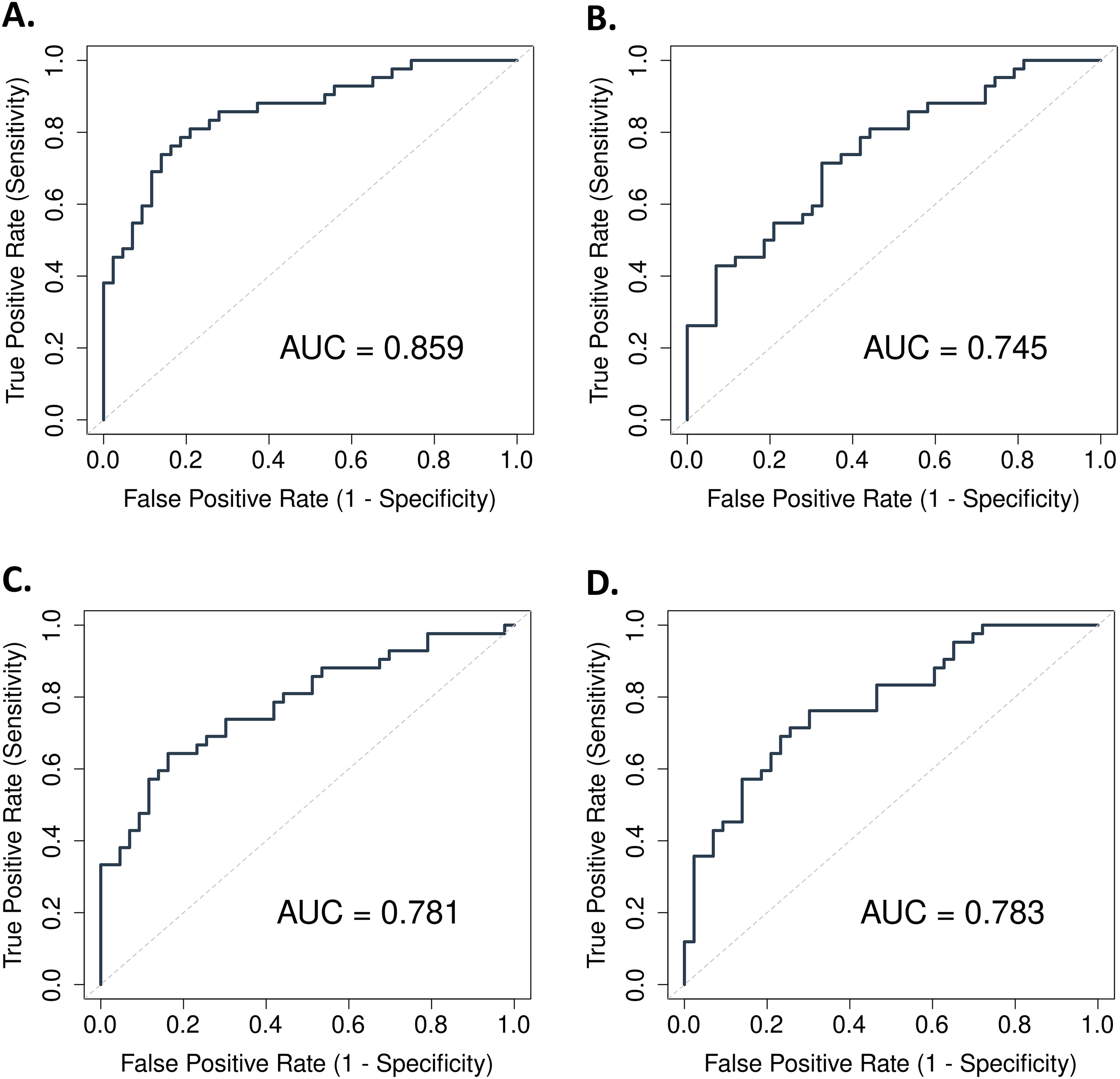
ROC curves and corresponding AUC scores of linear regression models trained to distinguish prostate cancer patients with and without thrombosis based on the top five differentially expressed proteins. (A) All three enriched pathways and each of the (B) hemostasis, (C) cell adhesion, and (D) cell proliferation pathways.
Discussion
In this study, we sought to identify soluble mediators associated with venous thromboembolism (VTE) in prostate cancer patients, using archived plasma samples from the VA CSP #719B cohort, which followed patients between 2001 and 2006. Importantly, samples were collected prior to the VTE. Through high-throughput proteomic profiling using SomaScan platform, we identified proteins in three pathways: hemostasis, cell proliferation and cell adhesion, which were significantly associated with VTE risk. A biomarker reduction strategy was employed to optimize clinical utility, and the resulting panel demonstrated high discriminatory power with an area under the ROC curve of 0.859, suggesting excellent predictive potential. These findings pave the foundation for future mechanistic exploration of the identified proteins and validation of their utility in an external cohort.
Proteomic profiling is a powerful tool for identifying predictive biomarkers across solid tumors using pre-treatment biospecimens. In breast and colorectal cancers, proteomic signatures have been used to guide targeted therapies and assess metastatic risk.18–20 In prostate cancer, studies have identified serum and tissue protein markers linked to disease progression, androgen deprivation therapy (ADT) resistance, and recurrence, with Clinical Proteomic Tumor Analysis Consortium (CPTAC) offering deeper proteogenomic insights. E26 transformation-specific (ETS) gene fusions, particularly involving ERG, human complement component 4a (C4a) exerted one of the most pronounced impacts on the proteomic landscape..21,22 Databases like Oncomine facilitate integration of these findings into clinical practice. 23 Cooperative oncology groups, such as Eastern Cooperative Oncology Group (ECOG) and Western Oncology Cooperative Group (WOCG), have been essential in validating biomarkers across populations. 24 Our study aligns with this framework, using pre-DVT sera in prostate cancer to develop a proteomic panel for thrombosis risk stratification.
Prostate cancer is associated with a moderate but clinically significant risk of VTE, with incidence estimates ranging from 2% to 10%, depending on disease stage and treatment modality.25,26 This risk increases notably in patients receiving ADT, such as luteinizing hormone-releasing hormone agonists (Leuprolide) or antagonists (Reluglix), which has been independently associated with a heightened incidence of deep vein thrombosis.27,28 ADT induces a hypogonadal state that may contribute to endothelial dysfunction, altered lipid profiles, and a procoagulant milieu—factors that collectively predispose to thrombosis. 29 Given that patients with prostate cancer often have favorable survival trajectories, they are at particular risk of experiencing thrombotic events during long-term follow-up. This underscores the importance of dissecting prostate cancer-specific biological pathways linked to thrombosis, which could reveal predictive biomarkers for targeted therapy.
The SomaScan 11k platform was selected for its broad assay coverage and high reproducibility, making it well-suited for large-scale population studies compared to Olink or mass spectrometry. However, as an aptamer-based platform lacking peptide-level resolution, its findings require quantative validation using complementary technologies.30–34 Among the top differentially expressed proteins identified in our study, several are involved in cancer and vascular biology. Tissue inhibitor of metalloproteinases 1 (TIMP1) was reported as biomarker for cardiovascular remodeling35,36 and is a regulator of the initiation and promotion of prostate cancer metastases. 37 Junctional adhesion molecule 2 (JAM2) is a member of immunoglobulin superfamily (IgSF) involved in cell proliferation, platelet activation, and tumor metastasis. 38 It also increases CXCL9/10 from tumor cells, which affects the immune microenvironment and can augment the thrombosis process. 39 TMX3, thioredoxin related transmembrane protein 3, plays an essential role in oxidation reduction system. TMX family genes are associated with tumor metastasis and immune checkpoints in pancreatic cancer. 40 Additionally, TMX3 has been detected in human megakaryocytes and platelets, and identified as a potential target for venous thrombosis in a Genome-Wide Association study for VT. 41
Tissue factor (TF) encoded by the gene F3 also known as thromboplastin or coagulation factor III, is a transmembrane glycoprotein responsible for coagulation cascade regulation and formation of blood clot. Elevated levels of TF-positive macrovesicles are considered as a mediator for cancer-associated thrombosis. 42 Circulating TF + MVs were detected in various cancers such as pancreatic, lung, gastric and breast43–45 and proposed as a useful biomarker of pancreatic cancer. Endothelial cell-selective adhesion molecule (ESAM), a member of the immunoglobulin receptor family, plays a role in endothelial cell migration and blood vessel formation.
C-X-C Motif Chemokine 12 (CXCL12), also known as stromal cell-derived factor 1 (SDF-1) acts as a chemokine by augmenting platelet aggregation and attracting inflammatory cells to vessels, which promotes arterial thrombosis. 46 CCL11 is a selective eosinophil chemoattractant, which regulates tumor growth by sending eosinophil cells to tumor cells, 47 and recruits myeloid-derived suppressor cells to the tumor to produce inflammatory mediators. 48 Cellular Communication Network Factor 5 (CCN5), also known as WNT1-inducible-signaling pathway protein 2 (WISP-2) was reported to antagonize CCN2's profibrotic actions and inhibit myocardial collagen deposition and fibrosis in chronic pressure overload of the heart. 49 Aldehyde Dehydrogenase 1 Family Member A2 (ALDH1A2), also known as Retinaldehyde Dehydrogenase 2 (RALDH2) was positively associated with immune cell infiltration, suggesting a protective role in prostate cancer progression. Cytokine receptor-like factor 2 (CRLF2), is a type I cytokine receptor that regulates cell proliferation and apoptosis of B cells and growth and inflammatory responses of T and dendritic cells. 50 Along with interleukin-7 receptor (IL7R), it activates STAT3&5 and JAK3 pathways involved in cell proliferation and hematopoietic system development. 51 The overexpression of CRLF2 is a biomarker of poor prognosis in T-cell acute lymphoblastic leukemia (T-ALL). 52 Our findings suggest a potential involvement of these proteins in cancer-associated thrombosis, in addition to their role in cancer growth and metastasis. These results strengthen possible links between coagulation activity, cancer cell proliferation and cancer progression.
This study, while highly informative, has some constrains. First, the use of archived samples allows for identifying strong associations but precludes causal inference. Future studies using experimental models could probe cause-and-effects. Second, the impact of biomarkers derived from archived serum samples may be attenuated in some patients due to the length of the interval between sample collection and VTE events. Future studies in cohorts with serial sampling are recommended to probe the role of the associations identified in our study in relation to timing of prostate cancer diagnosis, progression, and treatment. Third, the VA cohort, while nationally representative, may differ from the general population in demographics and unique military-related exposures. Our findings will benefit from confirmation in a civilian population. Fourth, certain clinical variables, including Gleason score and BMI, are missing in a number of patients due to the use of archived samples collected two decades ago. Of note, however, these variables are well balanced across cases and controls among those patients where they are measured. Fifth, while our study includes several cancer- and treatment-related variables (cancer stage, Gleason score, and whether androgen deprivation therapy was received), other variables are not included, including tumor subtype and the timing and duration of exposure to androgen deprivation therapy. Future studies validating our findings in the context of these additional variables in cohorts where they are available will be of value. Finally, while several new proteins emerged as differentially regulated between the two groups, some have been well characterized and others less studied. While direct links to thrombosis are not clear, our approach and study are hypothesis-generating, aiming at identifying potential protein panels that mark an/or contribute to thrombosis, warranting subsequent mechanistic analyses.
Conclusion
Our study identified a distinct plasma proteomic signature, combining 15 proteins from key biological pathways, that was highly predictive of future VTE events in patients with prostate cancer, underscoring the potential role of circulating proteins in mediating thrombotic risk. These results warrant further validation in larger, independent cohorts to examine the generalizability of the results, to confirm their predictive value and to advance the development of targeted strategies for early intervention and personalized thromboprophylaxis.
Supplemental Material
sj-csv-1-cbm-10.1177_18758592251390251 - Supplemental material for A panel of plasma proteins associated with venous thromboembolism in patients with prostate cancer
Supplemental material, sj-csv-1-cbm-10.1177_18758592251390251 for A panel of plasma proteins associated with venous thromboembolism in patients with prostate cancer by Asha Jose, Simon Lu, Chao Zhang, Jennifer La, Melissa Young, J Michael Gaziano, Nathanael R Fillmore, Katya Ravid and Vipul C Chitalia in Cancer Biomarkers
Supplemental Material
sj-xlsx-2-cbm-10.1177_18758592251390251 - Supplemental material for A panel of plasma proteins associated with venous thromboembolism in patients with prostate cancer
Supplemental material, sj-xlsx-2-cbm-10.1177_18758592251390251 for A panel of plasma proteins associated with venous thromboembolism in patients with prostate cancer by Asha Jose, Simon Lu, Chao Zhang, Jennifer La, Melissa Young, J Michael Gaziano, Nathanael R Fillmore, Katya Ravid and Vipul C Chitalia in Cancer Biomarkers
Footnotes
Acknowledgment
The views expressed are those of the authors and do not necessarily reflect the position or policy of the Department of Veterans Affairs or the U.S. government.
Ethical approval and informed consent statements
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The study was approved by the VA Boston Healthcare System Research and Development Committee as an exempt study prior to data collection and analysis, with a waiver of informed consent per the Common Rule due to use of existing data.
Authors contributions
VCC and KR developed the hypothesis and designed the research; AJ, SL, VCC, and KR reviewed and analyzed different parts of the data and wrote the manuscript; SL, CZ, JL, NF, MY, MG prepared the figures; All the authors contributed conceptually to different degrees and edited the manuscript.
Funding
This project was funded by the AHA Cardio-Oncology SFRN CAT-HD Center grant 857078 (AJ, JL, MG, NF, KR, and VCC); R01HL166608 (V.C.C and K.R.); Center of Cross Organ Vascular Pathology and by the Thrombosis and Hemostasis Affinity Research Collaborative (ARC) at Boston University Chobanian and Avedesian School of Medicine.
Declaration of conflicting interest
NF and JL have research funding from Bayer and Merck. MG also has research funding from Merck.
Data availability statement
VA data is available to qualified investigators with regulatory approval and consistent with VA policy.
Statements and declaration
Not applicable.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
