Abstract
Background
Gastric cancer is the fifth most common malignancy and third leading cause of cancer death in China, with advanced-stage five-year survival below 20%. βIII-tubulin (TUBB3) is overexpressed in cancers but its role in gastric cancer remains unclear.
Methods
TUBB3 expression was analyzed using TCGA data and clinical samples. Knockdown models assessed its effects on proliferation, migration, and invasion in vitro and in vivo.
Results
TUBB3 was significantly upregulated in gastric cancer tissues versus normal mucosa. High TUBB3 correlated with poorer disease-free and overall survival but not other clinicopathological features. Functionally, TUBB3 knockdown inhibited proliferation via G2/M arrest and reduced migration/invasion by disrupting invadopodia, without affecting apoptosis, EMT, or ECM degradation. In vivo, TUBB3 depletion suppressed tumor growth and metastasis. Mechanistically, TUBB3 promoted G2/M transition via p21/Cyclin B1 and enhanced invasiveness through Cortactin/JNK activation.
Conclusion
TUBB3 overexpression predicts poor prognosis in gastric cancer. It drives proliferation via cell cycle regulation and metastasis through invadopodia formation, suggesting its potential as a therapeutic target.
Introduction
Gastric cancer ranks fifth in incidence and third in mortality among malignant tumors in China. 1 Although the advent of targeted therapy and immunotherapy has transformed treatment paradigms and substantiallyprolonged patient survival, the overall five-year survival rate remains lamentably low at 35–40%, 1 with that of metastatic gastric cancer still plummeting below 20%.2–4 Consequently, further elucidation of gastric cancer's molecular underpinnings is imperative to identify novel therapeutic targets. In recent years, researchers have engineered an array of innovative small-molecule inhibitors targeting pivotal regulators of cell cycle progression and signal transduction pathways, such as cyclin-dependent kinases (CDKs), the epidermal growth factor receptor (EGF/Ras) pathway, and tubulin. These breakthroughs have fostered novel paradigms in anticancer drug discovery and development.
Microtubules, integral components of the cytoskeleton, comprise α-tubulin and β-tubulin and play pivotal roles in cellular morphology, intracellular trafficking, meiosis, and mitosis. 5 In cancer, aberrant tubulin function can precipitate chromosomal missegregation, aneuploidy, and tumor progression.6–8 Additionally, microtubules may facilitate metastasis by modulating the cytoskeletal framework. They also engage in key oncogenic signaling pathways, including MAPK and PI3 K/AKT, thereby influencing cell proliferation and survival. As such, microtubule-targeting agents (MTAs) have long been a mainstay in cancer treatment, 9 with nearly all MTAs exerting their therapeutic effects by binding to the β-tubulin subunit and disrupting microtubule dynamics. 8
Among the eight β-tubulin isotypes, βIII-tubulin (encoded by TUBB3) is predominantly expressed in neural and testicular tissues under physiological conditions. However, in a spectrum of epithelial-derived solid malignancies—including breast, colorectal, prostate, esophageal, and gastric cancers—βIII-tubulin is aberrantly overexpressed.10–15 Notably, its heightened expression correlates with significantly shorter progression-free survival (PFS) and overall survival (OS). 8 Intriguingly, in melanoma, reduced βIII-tubulin expression paradoxically serves as an independent prognostic indicator of poor outcomes. 16 Huang et al. conducted an analysis of βIII-tubulin expression in 146 cases of unresectable gastric cancer, revealing that patients with elevated expression exhibited markedly shorter PFS. 17 Furthermore, gastric cancer patients with βIII-tubulin overexpression demonstrated diminished responsiveness to taxane-based chemotherapy.18,19 Despite these findings, the precise mechanistic contributions of TUBB3 (βIII-tubulin) to gastric cancer progression remain incompletely understood.
In this study, we initially retrieved gastric cancer patient cohorts from the GEPIA2 and TCGA databases to investigate the relationship between TUBB3 mRNA expression, clinicopathological characteristics, and prognosis. We then validated our findings in a real-world cohort of gastric cancer patients treated at Peking Union Medical College Hospital (PUMCH), Chinese Academy of Medical Science. Having established the association between TUBB3 (βIII-tubulin) and poor prognosis in gastric cancer, we further explored its impact on tumor proliferation, metastasis, and regulatory mechanisms using gastric cancer cell lines and in vivo models.
Methods
Data acquisition and processing
The data (enclosing clinical, mRNA-seq, and miRNA-seq data) of 408 stomach adenocarcinoma (STAD) tissues and 211 adjacent normal controls were downloaded from the TCGA-STAD dataset in July 2024. After filtering out cases with incomplete tumor/node/metastasis (TNM) staging or ongoing data, 373 STAD patients were ultimately selected for inclusion in the analysis.
Gene expression analysis
TUBB3 mRNA expression profiles across diverse tumor types and their corresponding normal controls were obtained from GEPIA2 (Gene Expression Profiling Interactive Analysis 2) (http://gepia2.cancer-pku.cn), an intuitive and versatile tool for analyzing TCGA datasets. 20
Analysis of TUBB3 expression and its association with clinicopathological features and survival in gastric cancer patients using the TCGA online database
This study utilized RNA sequencing data (HTSeq-FPKM format) and corresponding clinical information from the TCGA-STAD database, retrieved via R(V4.0.2) and the TCGAbiolinks package. Data normalization was performed by converting FPKM to TPM format and applying log2 transformation using the ggplot2 package. The Wilcoxon rank-sum test was employed to assess differential TUBB3 expression between tumor and normal tissues, with bar plots visualizing the results. The optimal cutoff value for TUBB3 mRNA expression was determined using the surv_cutpoint function from the survminer package, stratifying patients into high- and low-expression groups. Associations between TUBB3 expression and clinicopathological characteristics, including gender and TNM stage (Chi-square test) as well as age (Wilcoxon rank-sum test), were analyzed, and the findings were systematically tabulated.
For survival analysis, the survminer and survival packages were utilized to construct Kaplan-Meier survival curves, with the log-rank test evaluating differences in OS and DFI between expression groups. Furthermore, univariate and multivariate Cox regression analyses were conducted to investigate the prognostic impact of TUBB3 mRNA expression alongside clinical variables (gender, age, TNM stage) on OS and DFI, providing insights into its potential as a prognostic biomarker in gastric cancer. OS is defined as the interval from initial diagnosis to all-cause mortality, whereas DFI is the time from diagnosis to the first documented recurrence of the disease.
Patient samples
Since September 2024, clinical data and characteristics of the 103 patients have been gathered. The collected information includes age, gender, surgical pathological staging, histological differentiation, surgical margin status, and prognostic outcomes, including OS and DFS. OS is defined as the interval from initial diagnosis to all-cause mortality, whereas DFS refers to the period after treatment during which a patient remains free from disease recurrence or progression. To evaluate the expression of βIII-tubulinin gastric cancer and adjacent normal tissues, a random selection of 20 patients who underwent surgery at PUMCH during the same period was made. Basic information of these patients was also collected, includes age, gender, surgical pathological staging, histological differentiation, surgical margin status.
These patients with GC who underwent surgical eradication at the PUMCH between November 2012 and October 2019. All cases were histopathologically confirmed as GC or other malignancies by board-certified pathologists at the PUMCH. Tumor staging was performed according to the 8th edition of the AJCC on Cancer TNM staging system. Patients were systematically followed-up via outpatient visits or structured telephones until March 4, 2025.
This study was approved by the Medical Ethics Committee of Peking Union Medical College Hospital. Written informed consent was obtained from all participants prior to enrollment.
TUBB3 expression, clinicopathological features, and survival in gastric cancer patients
Surgical pathological specimens from 103 gastric cancer patients underwent TUBB3 (βIII-tubulin) immunohistochemistry, and the expression level of TUBB3 (βIII-tubulin) was quantified using the Histochemistry score(H-Score). The median H-Score was used to classify the patients into low and high expression groups. Chi-square tests were applied to analyze the correlation between TUBB3 expression and clinicopathological factors (such as gender and TNM stage), while the Wilcoxon rank-sum test was used to assess its relationship with age.
Additionally, TUBB3 expression was evaluated in relation to patient survival, with Kaplan-Meier survival curves generated using the “survminer” and “survival” packages, and log-rank tests were used to assess the differences in OS and DFS.
Immunohistochemistry (IHC)
IHC staining was performed according to the standard protocol outlined in previous studies. 21 The pathological tissue samples were obtained from paraffin-embedded specimens preserved by the hospital's pathology department. The HE-stained slides of pre-cancerous and adjacent cancer tissues were confirmed by pathology before slicing. Paraffin sections were deparaffinized and placed in EDTA antigen retrieval buffer (pH 9.0), followed by antigen retrieval in a microwave. Endogenous peroxidase activity was blocked with hydrogen peroxide, and goat serum was used for blocking. Anti-βIII-tubulin antibody (1:200) was applied overnight at 4°C. After washing, a secondary antibody (1:100) was applied for 1 h at room temperature. DAB chromogenic reagent was added, followed by hematoxylin counterstaining. The slides were dehydrated, mounted, and examined under a microscope. The Aipathwell (Servicebio) software was used to analyze the tissue immunofluorescence staining. H-score was applied to convert the percentage of positive cells and their staining intensity in each slide into corresponding numerical values, allowing for a semi-quantitative analysis of both the staining intensity and the extent of positivity.
The H-Score formula is as follows:
H-Score (∑(pi × i) = (percentage of weak intensity cells ×1) + (percentage of moderate intensity cells ×2) + (percentage of strong intensity cells ×3)), where “i” represents the classification of positive cells: negative (no staining) = 0 points; weak positive (light yellow) = 1 point; moderate positive (brown yellow) = 2 points; strong positive (brown) = 3 points. “pi” represents the percentage of positive cells in each intensity category. The H-score ranges from 0 to 300, with higher values indicating stronger combined positivity in both staining intensity and extent.
Cells and treatment
MGC803 and HGC27 cell lines were procured from the Cell Resource Center of the Chinese Academy of Medical Sciences and authenticated to confirm their identity. MGC803 cells were cultured in Dulbecco's modified Eagle's medium (DMEM) (Gibco, Paisley, UK), whereas HGC27 cells were maintained in RPMI 1640 medium (Gibco, Paisley, USA). All culture media were enriched with 10% fetal bovine serum (FBS) (Gibco, Paisley, USA) supplemented with 100 U/mL penicillin and 100 μg/mL streptomycin. The cells were incubated under optimal physiological conditions at 37°C in a humidified atmosphere with 5% CO2 to ensure viability and proliferation.
Lentiviral particle transduction
Lentiviral particles carrying TUBB3-small interfering RNA(siRNA) (TUBB3 siRNA-1 sense 5’-UCUCUUCAGGCCUGACAAUTT-3,’ antisense 5’-AUUGUCAGGCCUGAAGAGATT-3’; TUBB3 siRNA-2 sense 5’-GACCUCAACCACCUGGUAUTT-3,’ antisense 5’-AUACCAGGUGGUUGAGGUCTT-3’; TUBB3 siRNA-3 sense 5’-GCACGUUGCUCAUCAGCAATT-3,’ antisense 5-‘UUGCUGAUGAGCAACGUGCTT-3’) and negative control siRNA (sense 5’-UUCUCCGAACGUGUCACGUTT-3,’ antisense 5’-ACGUGACACGUUCGGAGAATT-3’) were biochemically synthesized and integrated into the LV3 (H1/GFP&Puro) vector by GenePharma (Shanghai, China). For lentiviral transduction, MGC803 and HGC27 cells were exposed to lentiviral supernatants for > 24 h, followed by puromycin selection. The efficiency of selection was confirmed by fluorescence microscopy by detecting Enhanced Green Fluorescent Protein (EGFP), and transfection efficiency was further validated by qRT-PCR and western blotting.
Xenograft assay in nude mice
All animal experiments were conducted in accordance with the ARRIVE (Animal Research: Reporting of In Vivo Experiments) guidelines and approved by the Peking Union Medical College Hospital Application Form Welfare & Ethical Inspection in Animal Experimentation (Approval No: XHDW-2022-1004). In this study, BALB/c nude mice (6–8weeks old, male, weighing 18–22 g) were obtained from SPF Biotechnology(Beijing, China) and housed under specific pathogen-free conditions with a 12-h light/dark cycle, controlled temperature (22–25°C), and humidity (40–60%). The mice were provided with standard chow and water ad libitum.
To establish a subcutaneous gastric cancer xenograft model, si-TUBB3-MGC803 and NC-MGC803 cells (5 × 106 cells per mouse) were suspended in 50% Matrigel/PBS solution and injected subcutaneously into the right flank of each mouse. Tumor growth was monitored every 3 days using caliper measurements, and tumor volume was calculated using the following formula: V = (width)2 ×length/2. Mice were euthanized when the tumor size reached 1500 mm³ or when body weight loss exceeded 20%, following the institutional humane endpoint guidelines.
Tumor tissues, liver, and lung specimens were formalin fixed and paraffin embedded for histological and IHC examinations, as mentioned above. 22
All procedures were performed under isoflurane anesthesia to minimize animal suffering, and postoperative analgesia was administered as necessary. The study adhered to the 3R principles (Replacement, Reduction, Refinement) to ensure the ethical use of animals in research.
RNA extraction and qRT-PCR
This segment of the experiment adhered to the methodological framework established in the literature. 22 The primers used for real-time PCR are detailed in the Supplementary Table 1 (all custom-synthesized by Sangon Biotech, Shanghai, China).
Western blot
Western blot analysis was performed following established methodologies detailed in previous studies. 23 Primary antibodies used in WB, including those against GAPDH rabbit mAb (CST #5174, Danvers, MA, USA), TUBB3 (βIII-tubulin) rabbit mAb (CST #5568, Danvers, MA, USA), Cortactin rabbit mAb (CST #3503S, Danvers, MA, USA), β-catemin mouse mAb (Proteintech #66379-1-IG, Rosemont, Illinois, USA), ZO-1 rabbit mAb (Proteintech # 21773-1-AP, Rosemont, Illinois, USA), E-cadherin rabbit mAb (Proteintech #20874-1-AP, Rosemont, Illinois, USA), Vimentin rabbit mAb (CST #5741, Danvers, MA, USA), FN1 rabbit mAb (CST #7020, Danvers, MA, USA), MMP2 rabbit mAb (CST #40994, Danvers, MA, USA), LOX rabbit mAb (Proteintech # ET1602-23, Rosemont, Illinois, USA), FAK rabbit mAb (CST #13009, Danvers, MA, USA), p-FAK (CST #3824, Danvers, MA, USA), p21 mouse mAb (MCE #HY-10331, Monmouth Junction, NJ, USA), Cyclin B1 rabbit mAb (MCE #HY-13051, Monmouth Junction, NJ, USA), CDK1 rabbit mAb (MCE #HY-13716, Monmouth Junction, NJ, USA), MEK rabbit mAb (MCE #HY-12072, Monmouth Junction, NJ, USA), pMEK rabbit mAb (MCE #HY-15244, Monmouth Junction, NJ, USA), ERK1/2 rabbit mAb (MCE #HY-10369, Monmouth Junction, NJ, USA), pERK rabbit mAb (MCE #HY-10432, Monmouth Junction, NJ, USA), JNK rabbit mAb (MCE #HY-13634, Monmouth Junction, NJ, USA), pJNK rabbit mAb (MCE #HY-11960, Monmouth Junction, NJ, USA).
The secondary antibodies used in WB were goat anti-rabbit IgG (H + L) HRP (MultiSciences, China; 70-GAR0072) and goat anti-mouse IgG (H + L) HRP (MultiSciences, China; 70-GAM0072). Finally, chemiluminescence was monitored using chemiluminescence ECL detection reagent (Millipore, USA).
Cell proliferation assay
Cells (1.5 × 10³ cells/well) were inoculated in 96-well plates and inoculated for 4 days. Cell viability was assessed every 24 h by adding MTS reagent (10 µL/well) (Promega, Madison, USA), followed by incubation at 37°C for 1 h. Absorbance was quantified at 490 nm using a microplate reader (Molecular Devices). Each assay was performed in triplicates to ensure reliability and reproducibility.
Cell cycle detection
TUBB3 knockdown groups (MGC803-Lipo-siRNA-TUBB3 and HGC27-Lipo-siRNA-TUBB3) and control groups (MGC803-Lipo-siRNA-NC and HGC27-Lipo-siRNA-NC) gastric cancer cell lines were collected 48–72 h post-lipofection. Cells were fixed in 70% ice-cold ethanol overnight, centrifuged at 1500 rpm for 5 min, washed twice with PBS, and resuspended in PBS. RNAse A (100 μg/mL) was added and incubated at 37°C for 1 h. PI (50 μg/mL)(YEASEN, Shanghai, China) was then added, and cells were incubated in the dark at room temperature for 30 min. After filtering through a 40 μm cell strainer, flow cytometry was performed and cell cycle analysis was conducted using FlowJo software.
Apoptosis detection
TUBB3 knockdown groups (MGC803-Lipo-siRNA-TUBB3 and HGC27-Lipo-siRNA-TUBB3) and control groups (MGC803-Lipo-siRNA-NC and HGC27-Lipo-siRNA-NC) gastric cancer cell lines were collected 48–72 h post-lipofection. The pellets were resuspended in 100 μL PBS, followed by addition of 5 μL Annexin V-647 and 10 μL PI(Annexin V-YSFluorTM 647/PI Apoptosis Detection Kit, YEASEN, Shanghai, China). Cells were incubated at room temperature in the dark for 30 min. After filtering through a 40 μm cell strainer, flow cytometry was performed, and apoptosis was analyzed using FlowJo software.
In vitro assays for cell migration and invasion
This segment of the experiment adhered to the methodological framework established in the literature. 22 Transwell chambers (8 μm pore size; Costar, Kennebunk, ME, USA) were used for the in vitro migration and invasion assays. For migration assays, siTUBB3-MGC803 and siTUBB3-HGC27 cells and corresponding control, were seeded into the top chambers. For invasion assays, siTUBB3-MGC803 and siTUBB3-HGC27 cells, along with an equal number of corresponding NC-MGC803 and NC-HGC27 cells, were seeded into the top chambers, which were coated with Matrigel (30ug) (Corning, Corning, NY, USA). The cell seeding number was 4 × 105 cells, and the times for the migration and invasion assays were 24 h and 72 h, respectively. Migrating or invading cells were counted and photographed in five arbitrary 100× fields under a microscope. All measurements were performed in triplicate, and the results are presented as the mean ± SEM.
Invadopodia assay and immunofluorescence experiments
This portion of the analysis was implemented in conformity with the rigorously defined protocols described in preceding research. 24 The cells were fixed at room temperature in 4% paraformaldehyde for 10 min, permeabilized with cold acetone for 10 min, and stained with Alexa Fluor® 647-conjugated anti-cortactin antibody (1:200; Abcam, Cambridge, UK) at room temperature for 1 h. After washing with PBS, 100 μL/well (96-well plate) of 1× Fluorescein Phalloidin (#HY-K0902, New Jersey, USA) was added and incubated in the dark at room temperature for 20–90 min. After washing with PBS, cells were incubated with 0.1 μg/mL Hoechst 33342 (Thermo Fisher Scientific, Waltham, MA, USA) before being observed under a confocal microscope (Olympus, Tokyo, Japan).
Statistical analysis
Statistical analysis was performed using GraphPad Prism V8.0 software (GraphPad Software). All quantitative data are presented as mean ± SEM. Discrepancies between the two groups were evaluated using two-tailed Student's t-tests or the Mann–Whitney rank-sum test. Differences among three or more groups were compared using a two-tailed analysis of variance (ANOVA) followed by multiple comparison tests. Qualitative data were compared using the chi-squared test. Survival analyses were performed using Kaplan–Meier curves combined with the log-rank test. Statistical p was set than 0.05. Each experiment was performed in triplicate, unless otherwise specified.
Results
We initially utilized the GEPIA2 database to assess the expression of TUBB3 in tumor and adjacent normal tissues across various solid neoplasms. Analysis findings revealed that TUBB3 mRNA is upregulated in tumor tissues compared to adjacent non-tumor tissues across several solid malignancies, including esophageal carcinoma (ESCA), glioblastoma multiforme (GBM), ovarian serous cystadenocarcinoma (OV), skin cutaneous melanoma (SKCM), and STAD (Figure 1A). Notably, in the TCGA-STAD dataset, TUBB3 mRNA expression was markedly upregulated in gastric adenocarcinoma tissues (n = 373) compared to adjacent normal gastric tissues (n = 32) (p < 0.05) (Figure 1B).
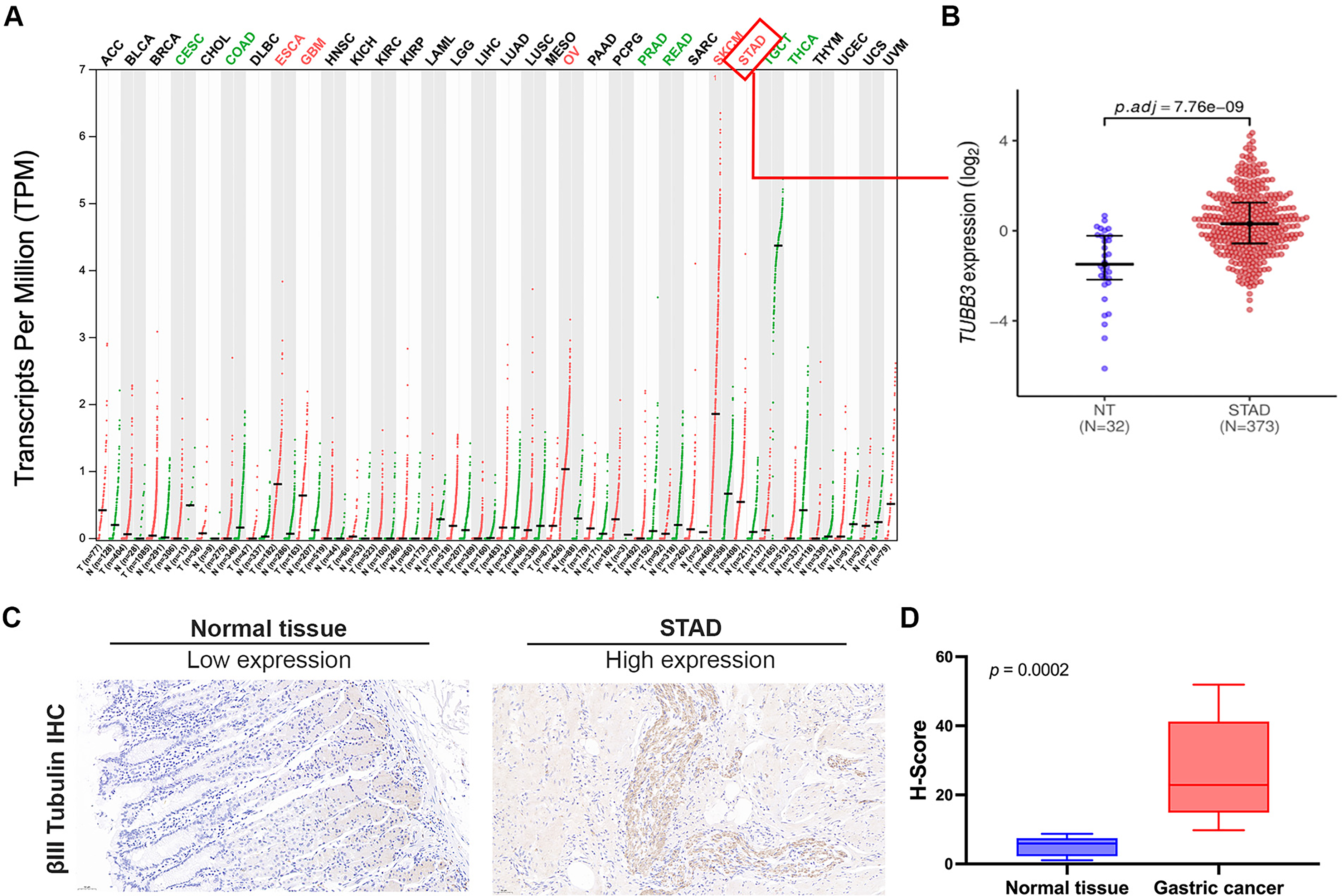
We subsequently evaluated the expression of βIII-tubulin protein in gastric cancer and adjacent non-cancerous tissue paraffin sections from 20 real-world patients (gastric cancer patients who underwent surgery at PUMCH)
The findings from both the TCGA-STAD gastric cancer cohort and the real-world gastric cancer cohort were consistent, with both demonstrating elevated TUBB3 expression in gastric cancer tissues.
2.
To explore the association between TUBB3 expression and the clinicopathological characteristics of gastric cancer, we extracted data from the TCGA database, including 383 gastric adenocarcinoma patients with available clinical information, among whom 338 had prognostic data. Utilizing the “surv_cutpoint” function from the “survminer” R package, we determined the optimal mRNA expression threshold for patients with survival data. Based on this threshold (12.93 TPM), patients were stratified into a high TUBB3 expression group (>12.93 TPM, n = 155) and a low TUBB3 expression group (≤12.93 TPM, n = 183). Comprehensive statistical analyses were performed to evaluate the correlation between TUBB3 mRNA expression and key clinical parameters, including age, sex, TNM stage, histological grade, and the presence of residual tumor post-surgery. The findings revealed no significant correlation between TUBB3 mRNA expression and these clinicopathological features (Supplementary Table 3).
To evaluate the clinical prognostic value of TUBB3, this study additionally enrolled a real-world cohort of postoperative gastric cancer patients (n = 103) (baseline characteristics are presented in Table 1). βIII-tubulin protein expression levels in surgical specimens were assessed by immunohistochemistry (IHC).Based on the median immunohistochemical score (H-Score = 19.07), patients were categorized into a high expression group (H-Score ≥19.07, n = 52) and a low expression group (H-Score <19.07, n = 51). Statistical analyses were performed to investigate the correlation between βIII-tubulin protein levels and clinical features such as age, sex, TNM stage, histological grade, and the presence of residual tumor after surgery. The results revealed no significant associations between βIII-tubulin protein expression and these clinicopathological characteristics (Table 1).
Correlation analysis of βIII-tubulin protein levels with clinicopathological characteristics of gastric cancer patients in the real world.
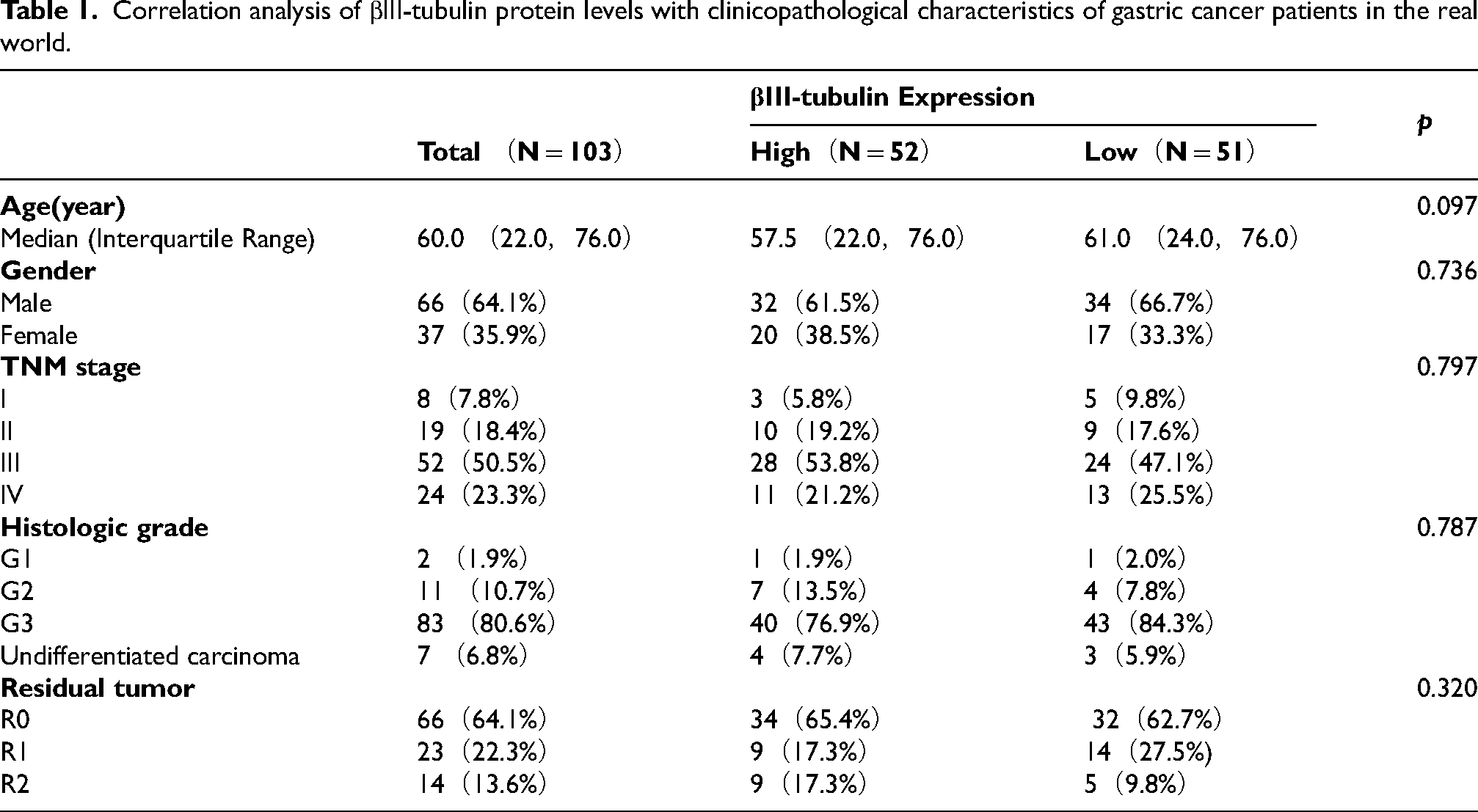
The findings from both the TCGA-STAD gastric cancer cohort and the real-world gastric cancer cohort were consistent, showing that TUBB3 (βIII-tubulin) expression in gastric cancer is not associated with clinicopathological features.
3.
In the TCGA database, analysis of 210 gastric cancer patients who underwent R0 resection with available follow-up data revealed that, based on the optimal TUBB3 mRNA cutoff, 21 out of 90 patients in the high-expression group experienced recurrence or progression, compared to 16 out of 120 in the low-expression group. While the median DFI was not reached in either group, high TUBB3 mRNA expression was associated with a significantly elevated risk of recurrence (HR = 2.43, p = 0.0063) (Figure 2A). Furthermore, among 338 STAD patients with available OS data, survival analysis demonstrated that in the low-expression group (n = 183), 60 patients died, with the median OS not reached (95% CI: 2.41 years – not reached), whereas in the high-expression group (n = 155), 74 patients died, with a median OS of 2.3 years (95% CI: 1.53–3.16 years). Notably, patients with high TUBB3 mRNA expression exhibited significantly poorer OS compared to those in the low-expression group (log-rank p = 0.00157), with a markedly increased risk of mortality (HR = 1.724, 95% CI: 1.225–2.426) (Figure 2B).
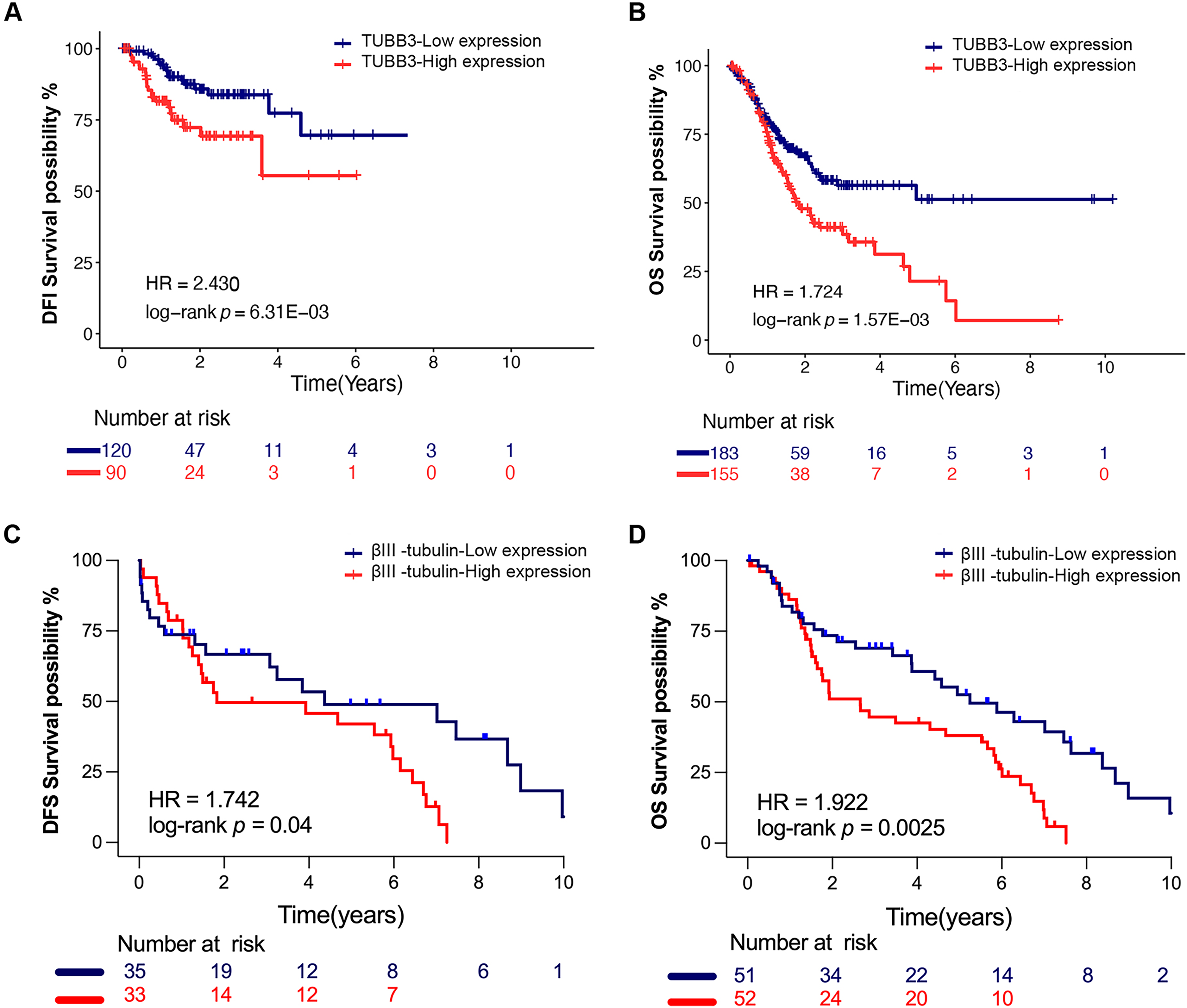
Using a Cox regression model, we evaluated the impact of TUBB3 expression, along with sex, age, TNM stage, and other clinical parameters, on OS and DFI in the TCGA gastric cancer cohort. Univariate Cox proportional hazards analysis indicated that high TUBB3 expression was significantly associated with an increased risk of shorter OS (HR = 1.15, 95% CI:1.02–1.30, p = 0.018) and a higher risk of shortened DFI (HR = 1.31, 95% CI:1.05–1.62, p = 0.016). After adjusting for age, TNM stage, tumor differentiation, surgical residual disease, and other potential confounders through multivariate Cox regression analysis, TUBB3 overexpression remained a robust independent predictor of poor prognosis, with an adjusted HR for OS was 1.14 (95% CI: 1.01–1.29, p = 0.035), and for DFI, it was 1.38 (95% CI: 1.11–1.72, p = 0.004). These findings underscore TUBB3 as an independent prognostic risk factor in gastric cancer (Supplementary Tables 4–5).
In a real-world cohort of 103 gastric cancer patients, we excluded those with pathological TNM stage (pTNM) of R2 or M1, leaving 68 patients with pTNM of R0/R1 and M0 for further analysis. Using the median βIII-tubulin expression level (H-Score = 19.07) as the optimal cutoff, we divided the patients into a high-expression group (H-Score ≥ 19.07, n = 33) and a low-expression group (H-Score < 19.07, n = 35). In the low-expression group, 21 patients experienced recurrence or progression, with a median DFS of 4.4 years (95% CI: 3.1 years – not reached). In contrast, among the high-expression group, 27 patients had recurrence or progression, with a median DFS of 1.9 years (95% CI: 1.4 years – 6.2 years). The high-expression group exhibited significantly lower DFS compared to the low-expression group (log-rank p = 0.04), with a 1.742-fold increased risk of recurrence or progression (HR = 1.742, 95% CI: 0.983–3.087) (Figure 2C). Further analysis of OS in the same cohort revealed that, using the median βIII-tubulin expression level (H-Score = 19.07) as the optimal cutoff, we divided patients into high-expression (H-Score ≥ 19.07, n = 52) and low-expression (H-Score < 19.07, n = 51) groups. In the low-expression group, 33 patients died, with a median OS of 5.3 years (95% CI: 3.9 years – 8.5 years). In the high-expression group, 43 patients died, with a median OS of 2.9 years (95% CI: 1.7 years – not reached). The high-expression group had significantly poorer OS than the low-expression group (log-rank p = 0.0025), with a 1.922-fold increased risk of mortality (HR = 1.922, 95% CI: 1.215–3.038) (Figure 2D).
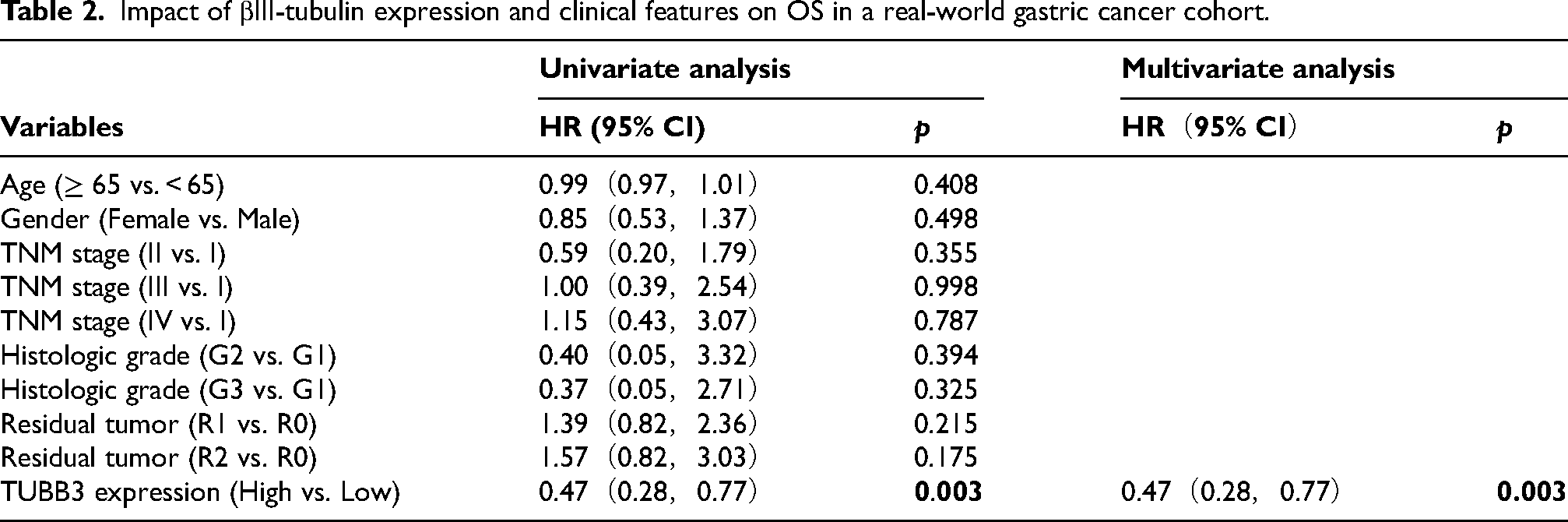
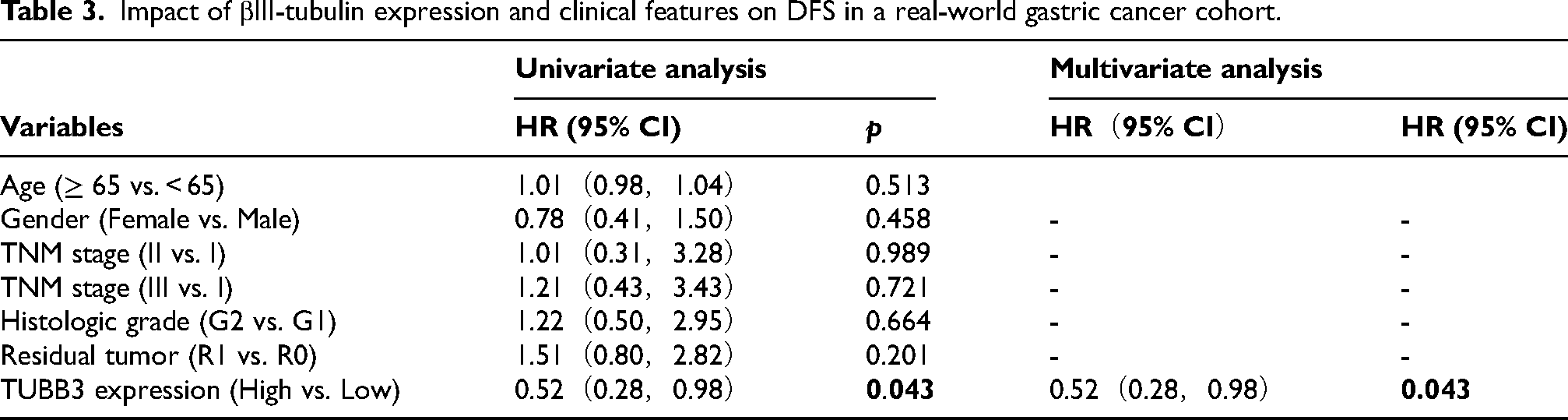
Using a Cox regression model, we analyzed the real-world gastric cancer cohort to assess the impact of βIII-tubulin levels, sex, age, and TNM stage on OS and DFS, calculating their HR. Both univariate and multivariate analyses revealed that high βIII-tubulin expression was significantly associated with shorter OS (HR = 0.47, 95% CI: 0.28–0.77, p = 0.003). After adjusting for age, TNM stage, tumor differentiation, and residual disease, high βIII-tubulin expression remained an independent prognostic factor for poor outcomes (Table 2). Similarly, both univariate and multivariate models showed that high βIII-tubulin expression correlated with shorter DFS (HR = 0.52, 95% CI: 0.28–0.98, p = 0.043). Even after adjusting for confounders, βIII-tubulin retained its role as an independent prognostic risk factor for gastric cancer (Table 3).
Impact of βIII-tubulin expression and clinical features on OS in a real-world gastric cancer cohort.
Impact of βIII-tubulin expression and clinical features on DFS in a real-world gastric cancer cohort.
Both the TCGA-STAD cohort (n = 338) and the real-world postoperative gastric cancer cohort (n = 103) indicate that high expression of TUBB3 (βIII-tubulin) is associated with poor prognosis in gastric cancer patients.
4.
We transfected MGC803 and HGC27 gastric cancer cell lines with lentiviral vectors encoding TUBB3 silencing (LV-shRNA-TUBB3) or control sequences (LV-shRNA-NC). Following transfection, green fluorescence was observed 48 h later due to the eGFP tag, confirming successful transduction (Figure 3A, Supplementary Figure 1A). Stable cell lines with reduced TUBB3 expression were selected using puromycin.
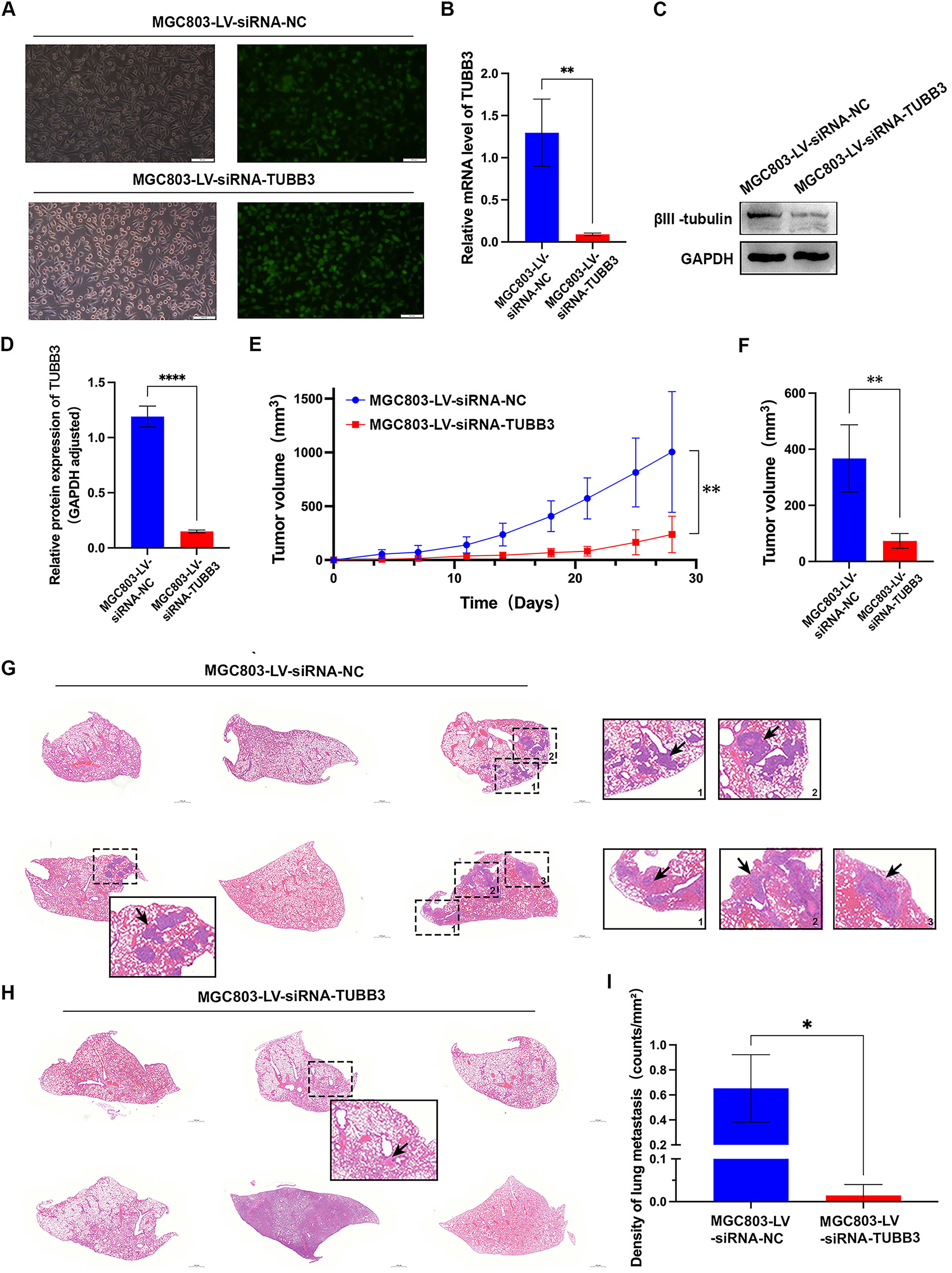
RT-qPCR and Western blot analysis demonstrated significant silencing efficiency. In the MGC803-LV-siRNA-TUBB3 group, TUBB3 mRNA expression was reduced to 12.56 ± 4.76% of the control (p < 0.01) (Figure 3B), and in the HGC27-LV-siRNA-TUBB3 group, it was reduced to 39.16 ± 9.26% (p < 0.05) (Supplementary Figure 1B). Corresponding reductions in βIII-tubulin protein expression were observed in the LV-shRNA-TUBB3 group (Figure 3C-D, Supplementary Figure 1C-D). Therefore, stable low TUBB3 expression MGC803 and HGC27 gastric cancer cell lines were successfully established, including the TUBB3 knockdown groups (MGC803-LV-siRNA-TUBB3 and HGC27-LV-siRNA-TUBB3) and the control groups (MGC803-LV-siRNA-NC and HGC27-LV-siRNA-NC) for subsequent experiments.
We subcutaneously implanted MGC803-LV-siRNA-TUBB3 cells (knockdown group) and MGC803-LV-siRNA-NC cells (control group) into nude mice and systematically monitored tumor growth and body weight. Throughout the observation period, tumors in the knockdown group (n = 6) remained significantly smaller than those in the control group (n = 6) (Figure 3E). At approximately three weeks post-implantation, tumor volume analysis revealed a statistically significant reduction in the knockdown group compared to the control (238.1 ± 169.29 mm³ vs. 1005.00 ± 562.11 mm³, p = 0.0095) (Figure 3F).
We subsequently assessed the impact of TUBB3 silencing on subcutaneous gastric cancer metastasis in nude mice. No hepatic metastases were detected in either the TUBB3 knockdown cohort or the control cohort. However, pulmonary metastases were observed in both groups.In the control cohort, 3 out of 6 mice developed lung metastases (Figure 3G), with metastatic foci numbering 20, 8, and 21, corresponding to metastatic densities of 0.80060 foci/mm², 0.34130 foci/mm², and 0.81573 foci/mm², respectively. Conversely, in the TUBB3 knockdown cohort, only 1 out of 6 mice exhibited lung metastases (Figure 3H), with a single metastatic focus and a markedly reduced metastatic density of 0.04409 foci/mm². Statistical analysis revealed a significant attenuation of metastatic burden in the TUBB3 knockdown cohort compared to the control cohort (n = 6, p < 0.05) (Figure 3I).
5.
We utilized the MTS assay to evaluate the effect of TUBB3 silencing on the proliferative capacity of gastric cancer cell lines MGC803 and HGC27. The findings demonstrated a marked suppression of cell proliferation in the TUBB3-silenced group compared to the control group (N = 6, p < 0.05) (Figure 4A). Consistently, a significant reduction in proliferation was also observed in the HGC27 cell line (N = 6, p < 0.05) (Figure 4B).
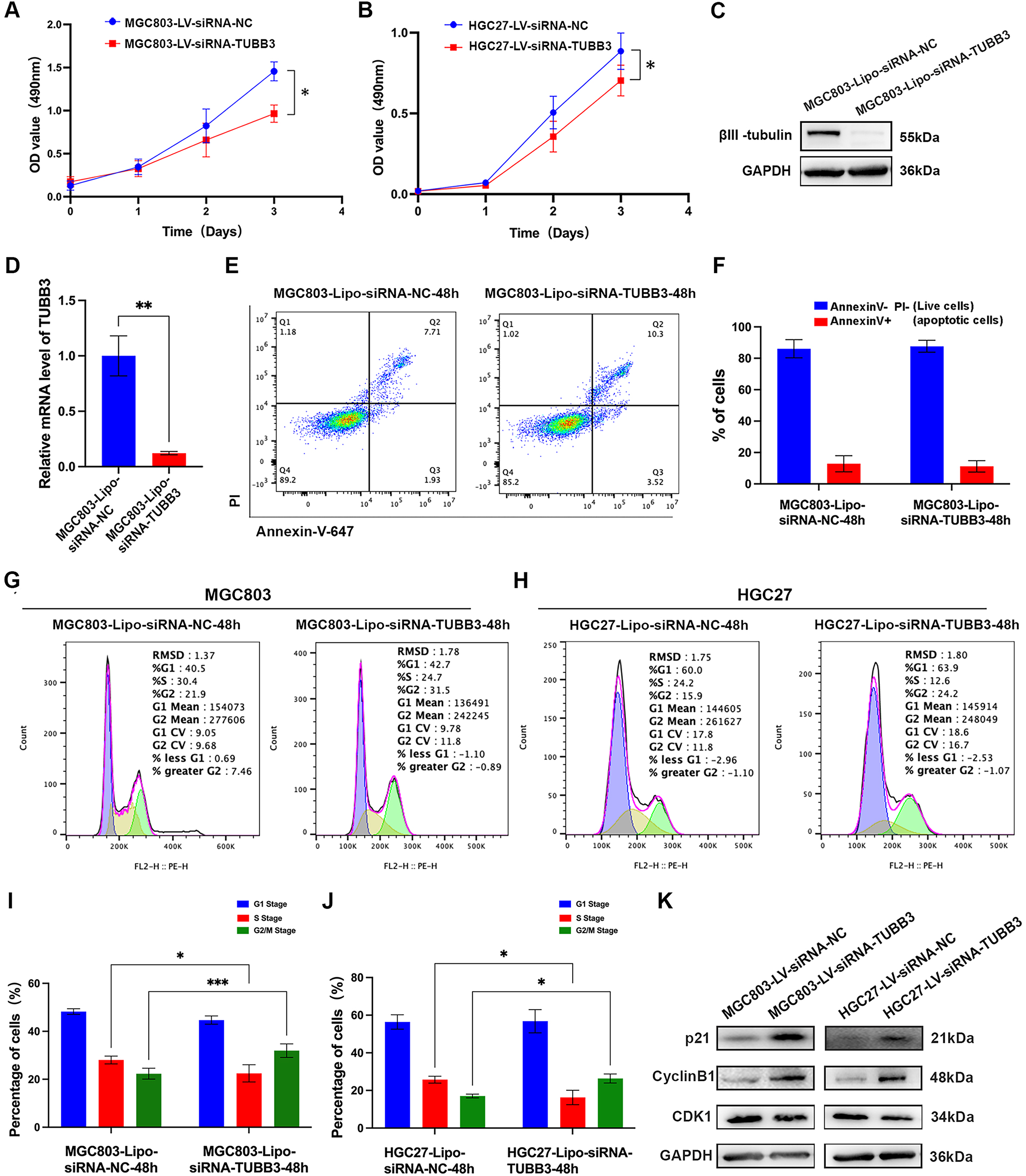
Suppression of TUBB3 expression profoundly curtailed cellular proliferation and elicited G2/M phase arrest during cell cycle progression, but exerted no discernible effect on apoptosis in gastric cancer cells in vitro. (A-B) The MTS test illuminated that the deletion of TUBB3 markedly inhibited the proliferation of MGC803 and HGC27 cells (N = 6). (C-D) Efficiency evaluation of liposome-mediated siRNA-TUBB3 in MGC803 cell lines (N = 3). (E-F) Knockdown of TUBB3 does not affect apoptosis in MGC803 cell lines (N = 3). (G-J) Cell cycle analysis in gastric cancer cell lines (MGC803 and HGC27) revealed a significant increase in the proportion of cells in the G2 phase and a corresponding decrease in the S phase in the siRNA-TUBB3 group compared to the control group (N = 3). (K) The effect of TUBB3 interference on G2/M transition-related proteins in gastric cancer cell lines. (ns, p ≥ 0.05; *, p < 0.05; **, p < 0.01; ***, p < 0.001).
As the lentiviral vector contained an eGFP-encoding sequence, the expression of green fluorescent protein in transfected gastric cancer cell lines interfered with flow cytometric analysis of apoptosis and cell cycle distribution. To overcome this limitation, we established an alternative gastric cancer cell model in which TUBB3 silencing was achieved via liposome-mediated transient transfection. Following transfection with either control shRNA-NC or shRNA-TUBB3 sequences for 48 h, total RNA and protein were extracted to evaluate TUBB3 expression at both the transcriptomic and proteomic levels in MGC803 and HGC27 cell lines. The results demonstrated a significant downregulation of TUBB3 mRNA expression in the siRNA-TUBB3-transfected group (MGC803-Lipo-siRNA-TUBB3; HGC27-Lipo-siRNA-TUBB3) compared to the control group (MGC803-Lipo-siRNA-NC; HGC27-Lipo-siRNA-NC) (N = 3, p < 0.05) (Figures 4D, Supplementary Figure 1E). Correspondingly, βIII-tubulin protein levels were markedly diminished in the siRNA-TUBB3 group (Figures 4C, Supplementary Figure 1F).
TUBB3 may modulate the proliferation of gastric cancer cell lines, potentially through its effects on apoptosis or cell cycle regulation. To further explore the impact of TUBB3 silencing on apoptosis, we performed flow cytometric analysis and observed no significant differences in the proportions of viable versus apoptotic cells between the siRNA-TUBB3-transfected group (MGC803-Lipo-siRNA-TUBB3; HGC27-Lipo-siRNA-TUBB3) and the control group (MGC803-Lipo-siRNA-NC; HGC27-Lipo-siRNA-NC) (N = 3, p > 0.05) (Figure 4E-F, Supplementary Figure 1G-H).
Cell cycle analysis demonstrated that, in the MGC803 cell line, the proportion of cells in the G2 phase was significantly increased in the siRNA-TUBB3 group (MGC803-Lipo-siRNA-TUBB3) compared to the control group (MGC803-Lipo-siRNA-NC) (31.97 ± 2.86% vs. 21.57 ± 1.60%, p < 0.001), while the proportion of cells in the S phase was significantly reduced (22.47 ± 3.49% vs. 29.03 ± 1.86%, p < 0.05) (Figure 4G, I). Likewise, in the HGC27 cell line, the siRNA-TUBB3 group (HGC27-Lipo-siRNA-TUBB3) exhibited a notable increase in the proportion of cells in the G2 phase (26.37 ± 2.36% vs. 17.03 ± 0.94%, p < 0.05), alongside a significant decrease in the S phase (16.27 ± 3.88% vs. 25.73 ± 1.80%, p < 0.05) when compared to the control group (HGC27-Lipo-siRNA-NC) (Figure 4H, J).
Following the observation that TUBB3 influences the G2/M phase transition in gastric cancer cell lines, we subsequently evaluated the expression of G2/M phase transition-associated proteins (Cyclin B1, p21, and CDK1) in both the TUBB3 interference group and the control group. The results revealed a significant upregulation of p21 and Cyclin B1 expression following TUBB3 interference, while no significant changes were detected in CDK1 expression (Figure 4K).
6.
In the migration assay, MGC803 cells in the TUBB3 interference group migrated 97.7 ± 28.9 cells/field, which was significantly fewer compared to the control group, with 200.7 ± 30.6 cells/field (N = 3, p < 0.05) (Figure 5A, E). In the invasion assay, the interference group showed 108.3 ± 12.3 cells/field, while the control group had 207.0 ± 20.5 cells/field, also demonstrating a significant reduction in the TUBB3 interference group (N = 3, p < 0.01) (Figure 5B, F). Similar results were obtained in the HGC27 cells. The TUBB3 interference group in HGC27 cells showed a migration of 98.0 ± 20.7 cells/field, significantly fewer than the control group, which migrated 229.5 ± 39.4 cells/field (N = 3, p < 0.05) (Figure 5C, G). The invasion assay in HGC27 cells revealed 41.3 ± 15.0 cells/field in the TUBB3 interference group, compared to 195.7 ± 16.6 cells/field in the control group (N = 3, p < 0.001) (Figure 5D, H).
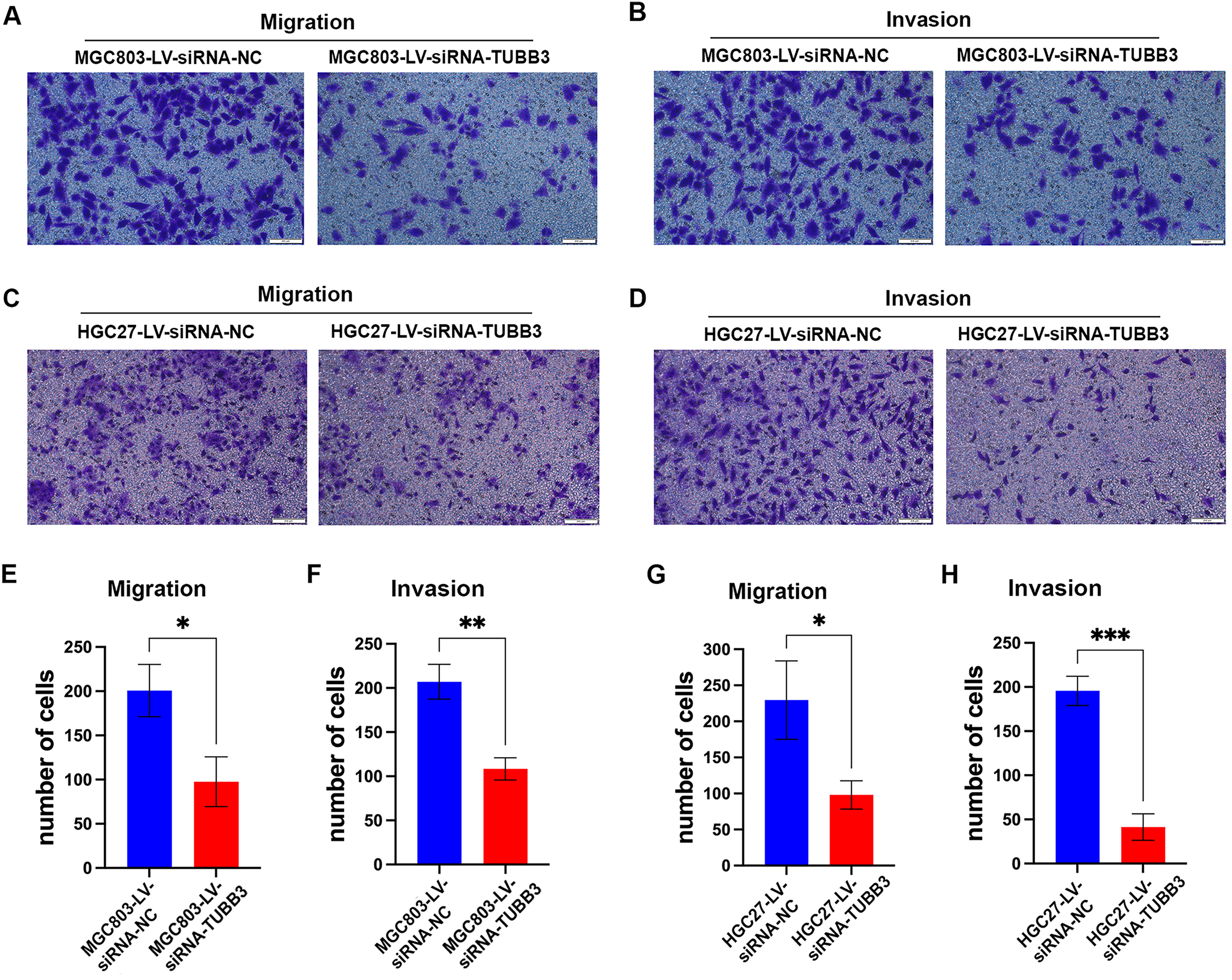
Previous studies have suggested that TUBB3 (βIII-tubulin) may regulate tumor cell migration and invasion through four primary mechanisms: promoting epithelial-mesenchymal transition (EMT),25,26 facilitating extracellular matrix (ECM) remodeling 27 and promoting anoikis resistance. 28
To investigate this, we first analyzed the expression of EMT-related proteins in the TUBB3 interference and control groups. In both MGC803 and HGC27 cells, TUBB3 interference did not lead to significant changes in the expression of epithelial markers, such as Zonula Occludens-1 (ZO-1) and E-cadherin, mesenchymal marker Vimentin, or key EMT-regulatory proteins like β-catenin, Focal Adhesion Kinase (FAK), and Phosphorylated Focal Adhesion Kinase (pFAK) (Supplementary Figure 2A). These results suggest that TUBB3 (βIII-tubulin) likely influences gastric cancer cell migration and invasion through mechanisms independent of EMT.
We then assessed the impact of TUBB3 on ECM remodeling-related proteins and found that TUBB3 interference had no effect on the expression of Matrix Metalloproteinase 2 (MMP2), an enzyme responsible for ECM degradation. Additionally, there was no significant impact on the expression of Fibronectin (FN), a key ECM component, or on Lysyl Oxidase (LOX), an enzyme that influences the crosslinking of collagen and elastin, thus affecting ECM remodeling (Supplementary Figure 2B). These findings suggest that TUBB3 may not play a role in ECM remodeling in gastric cancer.
Anoikis, a crucial barrier against tumor metastasis, prevents detached tumor cells from surviving and colonizing distant organs. Studies have shown that elevated TUBB3 expression enhances anoikis resistance, enabling tumor cells to persist and disseminate following detachment from the primary site. To assess the impact of TUBB3 knockdown on anoikis in gastric cancer cell lines, we performed flow cytometry analysis. The results demonstrated no significant differences in the proportions of viable and apoptotic cells under low-adhesion conditions between the TUBB3 knockdown group and the control group (N = 3, p > 0.05) (Supplementary Figure 2C-F), suggesting that TUBB3 depletion does not influence anoikis susceptibility in these cells.
Finally, many researches on other tumors has highlighted the role of invasive pseudopodia in tumor metastasis.29,30 we assessed the impact of TUBB3 interference on Cortactin, a crucial marker of invasive filopodia. 31 Our findings revealed a significant reduction in Cortactin protein expression in the TUBB3 interference groups compared to the control groups (Figure 6A).
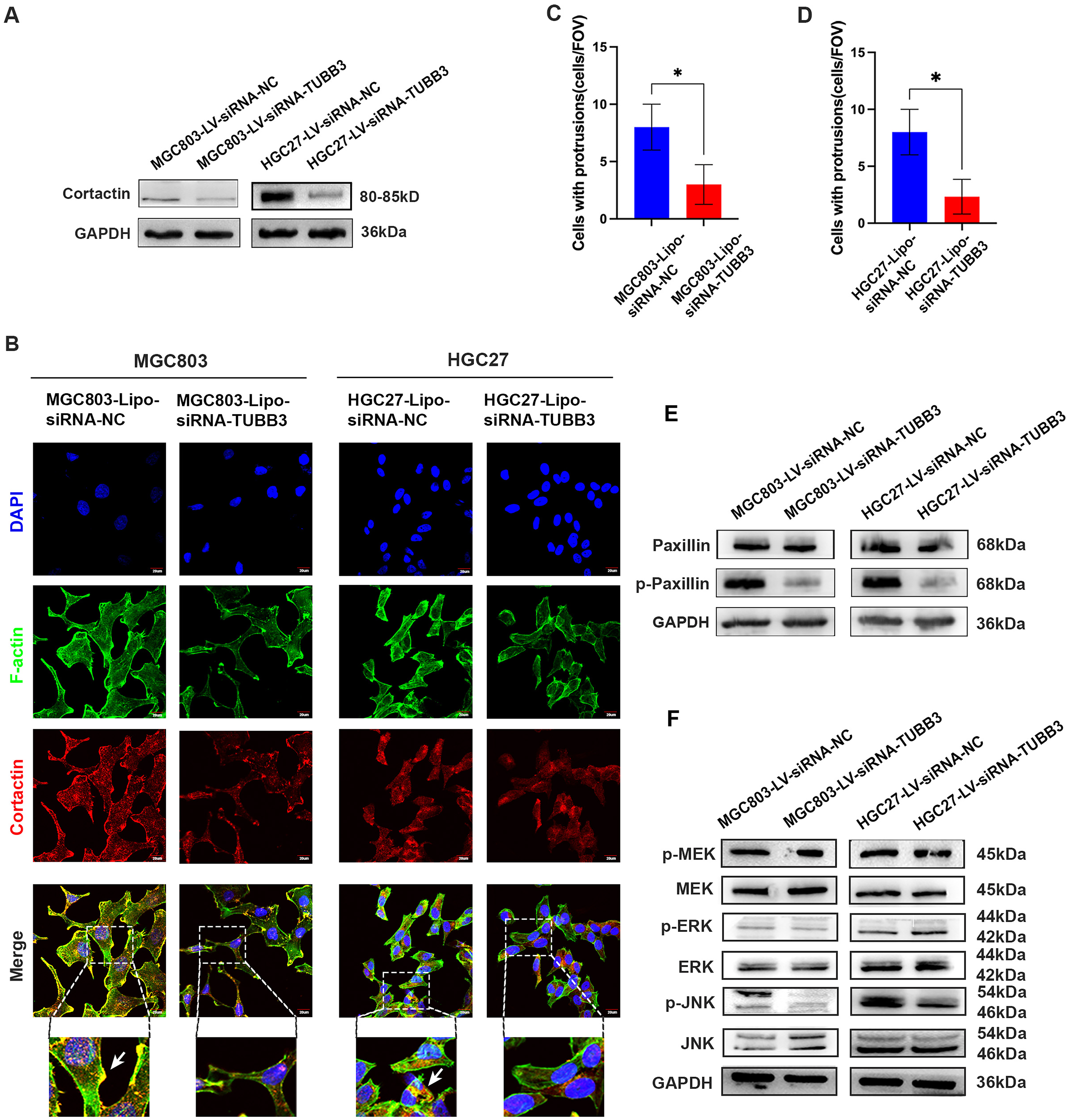
We then employed fluorescence confocal microscopy to assess the number of invasive filopodia in TUBB3-knockdown and control gastric cancer cell lines. The colocalization of F-actin and Cortactin serves as a hallmark of invasive pseudopodia formation. 24 The analysis revealed a significant reduction in the number of invasive filopodia—defined by the co-localization of Cortactin (red) and F-actin (green)—in the TUBB3-knockdown group (MGC803-Lipo-siRNA-TUBB3) compared to the control group (MGC803-Lipo-siRNA-NC) (N = 3, p < 0.05). Consistent findings were observed in the HGC27 cell line (N = 3, p < 0.05) (Figure 6B-D). These results indicate that TUBB3 suppression may inhibit gastric cancer cell migration and invasion by impairing invasive filopodia formation.
Paxillin is a key signaling adaptor protein localized at focal adhesions, where its phosphorylation plays a crucial role in regulating cell migration and cytoskeletal dynamics. Previous studies have demonstrated that phosphorylated paxillin facilitates the recruitment of actin-regulatory proteins to the leading edge, thereby promoting the formation of invasive protrusions.32,33 In the present study, TUBB3 knockdown resulted in a significant reduction in phosphorylated paxillin levels, while total paxillin expression remained largely unchanged (Figure 6E). These findings suggest that TUBB3 may modulate tumor cell migratory capacity by regulating the phosphorylation status of paxillin.
Previous studies have highlighted the involvement of the MAPK signaling pathway in invasive filopodia formation. 34 To further elucidate the impact of TUBB3 knockdown, we examined its effects on MAPK signaling proteins and their phosphorylation status in gastric cancer cell lines. Notably, compared to the control group, the TUBB3 knockdown group exhibited a significant reduction in JNK phosphorylation, suggesting a potential mechanistic link between TUBB3 and MAPK-driven invasion (Figure 6F).
Discussion
TUBB3 is located on chromosome 16q24.3, spanning approximately 4.5 kb and comprising four exons and three introns. It encodes βIII-tubulin, a key component of microtubules. 35 Initially identified as a neuronal marker, TUBB3 plays a crucial role in neurodevelopment, and germline mutations in TUBB3 have been associated with oculomotor nerve dysfunction. 36
TUBB3 is widely overexpressed in tumors; however, the mechanisms underlying its upregulation in cancer remain unclear. 35 First, treatment with chemotherapeutic agents such as vincristine and paclitaxel significantly increases TUBB3 expression, though the precise mechanisms remain elusive. This upregulation may result from direct induction by chemotherapy or from the selective elimination of TUBB3-low-expressing cell subpopulations, leading to an enrichment of TUBB3-high-expressing cells.37,38 Second, hypoxia has been implicated in TUBB3 upregulation, as hypoxia-inducible factor 1α (HIF-1α) may bind to hypoxia response elements within the 3’ untranslated region (3'-UTR) of TUBB3, promoting its expression. 39 Additionally, HIF-2α may regulate TUBB3 transcription via SOX9 (SRY-Box Transcription Factor 9). 40 Lastly, certain microRNAs, such as miR-200c, may post-transcriptionally regulate TUBB3 expression.41,42
Current research on TUBB3 and its encoded protein βIII-tubulin in cancer primarily focuses on three aspects: (i) its role as a prognostic marker, (ii) its involvement in tumor growth and metastasis, and (iii) its association with resistance to microtubule-targeting agents.
TUBB3 is aberrantly overexpressed in multiple solid tumors, including lung,43–46 colorectal, 47 breast, 48 ovarian,49,50 and prostate cancers, 51 where it generally correlates with poor prognosis, aggressive features, and chemotherapeutic resistance. Notably, in some malignancies such as melanoma, 16 TUBB3 overexpression is associated with improved survival, while in head and neck tumors its expression shows no clear correlation with tumor stage or patient outcome. 52 Functionally, TUBB3 can disrupt microtubule dynamics,53–59 attenuate drug-induced apoptosis,60,61 promote EMT,25,26,62,63 enhance metastatic potential,27,28,64 and modulate oxidative stress responses,65–67 though these effects are tumor-type specific. Consistent with this broader context, our study demonstrates that TUBB3 is highly expressed in gastric cancer and promotes cell proliferation and invasiveness, highlighting its potential as a therapeutic target in this malignancy.
In our study, we first analyzed online databases and found that TUBB3 expression was higher in gastric cancer tissues than in normal gastric mucosa. This was further validated in real-world patient cohorts, where βIII-tubulin expression was undetectable in normal gastric epithelial cells but significantly elevated in gastric cancer epithelial cells. Unlike in breast cancer, where TUBB3 expression correlates with lymph node metastasis and tumor stage, 68 our study found no significant association between TUBB3 (βIII-tubulin) expression and clinicopathological features such as age, sex, TNM stage, or histological differentiation in gastric cancer, consistent with findings from Höflmayer et al. in European gastric cancer patients. 69
Consistent with other solid tumors, high TUBB3 (βIII-tubulin) expression is associated with poor prognosis in gastric cancer. 8 Similar conclusions were drawn in studies by Huang and Höflmayer.17,69 In both the TCGA database and our real-world cohort of post-surgical gastric cancer patients, those with high TUBB3 expression exhibited shorter disease-free interval (DFI)/disease-free survival (DFS) and overall survival (OS). Notably, the TCGA cohort used DFI, which specifically measures the time from initial treatment to disease recurrence or progression, excluding non-cancer-related mortality. In contrast, our real-world clinical cohort utilized DFS, a more clinically relevant metric that includes both recurrence/progression events and cancer-related mortality, thus providing a more comprehensive assessment of prognosis. Although the definitions of DFI and DFS differ—DFI focusing on recurrence/progression and DFS encompassing broader clinical endpoints—both effectively evaluate the relationship between TUBB3 expression and tumor recurrence/progression. While the association between high TUBB3 expression and poor prognosis in gastric cancer is well established, the underlying mechanisms driving TUBB3-mediated tumor progression remain largely unexplored.
Immune checkpoint inhibitors have become a first-line treatment for advanced gastric cancer, prompting us to investigate the relationship between TUBB3 expression and the Tumor Immune Dysfunction and Exclusion (TIDE) score, a predictor of immunotherapy response. Zhao et al. reported that in lung cancer patients receiving immune checkpoint inhibitors, non-responders exhibited higher TUBB3 expression. In a lung cancer mouse model, combining a TUBB3 small-molecule inhibitor with PD-1 blockade showed synergistic effects, possibly due to TUBB3 inhibition suppressing PD-L1 expression via Snail. 70 However, our analysis of the TCGA gastric cancer cohort revealed no significant association between TUBB3 expression and the TIDE score, suggesting that targeting TUBB3 may not be an effective strategy for enhancing immunotherapy response in gastric cancer.
To further explore the mechanisms by which TUBB3 promotes gastric cancer progression, we silenced TUBB3 expression in two gastric cancer cell lines (MGC803 and HGC27) and observed a significant reduction in cell proliferation, migration, and invasion.
As a microtubule component, TUBB3 may regulate tumor cell proliferation by influencing mitotic processes such as spindle formation. Liu et al. found that TUBB3 promotes gallbladder cancer proliferation by suppressing p21 and cyclin B1 while upregulating cyclin D1. 71 Parker AL et al. reported that βIII-tubulin reduces glycolytic dependency in cancer cells, allowing them to survive under fluctuating nutrient conditions by rapidly activating AKT during glucose deprivation, thereby promoting non-small cell lung cancer survival and proliferation. 72 In our gastric cancer models, TUBB3 knockdown induced G2/M phase arrest, with upregulation of G2/M transition regulators p21 and cyclin B1, though CDK1 expression remained unchanged. These findings suggest that TUBB3 may promote gastric cancer proliferation by facilitating G2/M phase transition. However, unlike reports in breast and colorectal cancer where TUBB3 inhibition induces apoptosis,73,74 TUBB3 silencing did not affect apoptosis in gastric cancer cells.
The mechanisms by which TUBB3 enhances tumor metastasis appear to vary across different cancers. Previous studies suggest that TUBB3 (βIII-tubulin) promotes tumor migration and invasion via four main pathways: facilitating epithelial-mesenchymal transition (EMT), remodeling the extracellular matrix (ECM) and enhancing anoikis resistance. 28
First, high TUBB3 expression confers anoikis resistance, allowing tumor cells to survive and disseminate following detachment from the primary tumor. Dong et al. demonstrated that TUBB3 enhances anoikis resistance in prostate cancer by activating the αvβ3/FAK/Src signaling axis, 28 while McCarroll JA et al. found that TUBB3 promotes anoikis resistance in lung adenocarcinoma via PTEN-mediated AKT phosphorylation. 64 However, in our study, TUBB3 knockdown did not significantly increase apoptosis under low-adhesion conditions in gastric cancer cells.
Second, TUBB3 may enhance metastasis by promoting EMT. In thyroid cancer, TUBB3 knockdown downregulated EMT-related genes. 14 Additionally, βIII-tubulin interacts with EHD1 to drive EMT in non-small cell lung cancer, 75 while RPPH1 induces EMT via interactions with βIII-tubulin in colorectal cancer. 63 However, in our gastric cancer models, TUBB3 knockdown did not affect epithelial or mesenchymal markers or EMT-related signaling proteins.
Third, TUBB3 may facilitate tumor invasion by promoting ECM remodeling, primarily through MMP2 upregulation. However, in our study, TUBB3 knockdown did not alter MMP2 expression or affect ECM components such as fibronectin (FN) and LOX.
Our experimental investigations failed to demonstrate any significant positive findings. Consequently, we performed a comprehensive review of the existing literature. Notably, Arnold et al. have established that DCLK1-dependent modulation of invasive pseudopod dynamics significantly contributes to the aggressive progression of head and neck squamous cell carcinomas. 29 In parallel, Zhao et al. have documented the critical involvement of invadopodia formation in glioblastoma metastasis, a process mediated through the c-Met /LanCL2 /STAT3 /Cortactin signaling cascade. 30 In light of these compelling evidences, we subsequently examined the potential role of TUBB3 in gastric cancer metastasis, with particular focus on its regulatory effects on invasive pseudopodia formation.
As a microtubule component, TUBB3 may regulate cytoskeletal dynamics to promote invadopodia formation. Invadopodia are specialized actin-driven membrane protrusions that facilitate tumor cell invasion by degrading the basement membrane (BM). Cortactin is a key regulator of invadopodia formation and function. 76 We found that TUBB3 knockdown reduced cortactin expression, as confirmed by Western blot and immunofluorescence. Dual immunofluorescence staining of cortactin and F-actin further revealed a significant decrease in invadopodia formation in TUBB3-depleted gastric cancer cells. Given the reported involvement of the MAPK signaling pathway in invadopodia formation, we examined MAPK signaling alterations following TUBB3 knockdown and observed decreased JNK phosphorylation. 77 These findings suggest that TUBB3 may enhance tumor invasiveness through multiple pathways, including cytoskeletal remodeling and adhesion regulation. Further investigations will explore the role of focal adhesion proteins such as paxillin and other adhesion-related molecules.
Silencing of TUBB3 was found to induce a pronounced G2/M cell-cycle arrest, accompanied by upregulation of the cyclin-dependent kinase inhibitor p21 and downregulation of Cyclin B1, while concurrently attenuating invasive cellular phenotypes and diminishing cortactin/JNK signaling activity. The precise hierarchical positioning of TUBB3 within these pathways has yet to be delineated. As a structural constituent of βIII-tubulin, TUBB3 may modulate downstream signaling cascades and cell-cycle regulation by altering microtubule dynamics or through interactions with cytoskeletal regulatory proteins, thereby shaping malignant cellular behavior.
Although the present study does not yet provide direct molecular evidence, we intend to employ co-immunoprecipitation coupled with mass spectrometry, protein stability assays, and functional rescue experiments to substantiate these mechanistic interactions, thereby refining the mechanistic role of TUBB3 in gastric cancer and evaluating its candidacy as a therapeutic target.
TUBB3 represents a potential therapeutic target for gastric cancer. In other malignancies, Han et al. identified a novel shikonin derivative, PMMB276, which targets TUBB3 and suppresses triple-negative breast cancer growth in mice. 78 Kashyap VK et al. demonstrated that the small-molecule inhibitor VERU-111 suppresses TUBB3 expression via miR-200c, inducing G2/M arrest and apoptosis in pancreatic cancer cells through Bax, Bad, Bcl-2, and Bcl-xL regulation. 79 Our findings indicate that targeting TUBB3 may potentially suppress gastric cancer progression and could help overcome paclitaxel resistance. However, While our findings indicate that TUBB3 overexpression enhances proliferative and invasive phenotypes in gastric cancer, its direct clinical association with chemotherapeutic resistance remains to be fully elucidated. Considering that TUBB3 has been implicated in resistance to microtubule-targeting agents and other cytotoxic drugs across multiple malignancies, it is plausible that similar mechanisms may contribute to therapy resistance in gastric cancer. To address this, future investigations incorporating larger, well-annotated clinical cohorts with treatment-response data, alongside preclinical studies employing both pharmacological and genetic inhibition of TUBB3 in in vitro and in vivo models, will be essential. Such efforts are expected to clarify the role of TUBB3 in mediating chemoresistance and to further define its potential as a therapeutic target in gastric cancer.
In conclusion, this study highlights high expression of TUBB3 (βIII-tubulin) as a prognostic marker of poor outcomes in gastric cancer. TUBB3 may regulate the cell cycle by upregulating G2/M transition-associated proteins such as p21 and Cyclin B1. Furthermore, TUBB3 appears to promote invasive podosome formation and metastasis through the upregulation of Cortactin and phosphorylation of JNK. However, the mechanisms underlying TUBB3 upregulation in gastric cancer, as well as its role in G2/M phase transition and invasive podosome formation, warrant further investigation.
Conclusion
Elevated expression of TUBB3 (βIII-tubulin) serves as a prognostic indicator of poor outcomes in gastric cancer.
In gastric cancer, TUBB3 may modulate cell cycle progression by upregulating G2/M transition-associated regulators, including p21 and Cyclin B1.
TUBB3 may facilitate invadopodia formation and drive gastric cancer metastasis by enhancing Cortactin expression and promoting JNK phosphorylation.
Supplemental Material
sj-docx-1-cbm-10.1177_18758592251390145 - Supplemental material for TUBB3 (βIII-tubulin) drives gastric cancer progression and poor prognosis by regulating cell cycle and invadopodia formation
Supplemental material, sj-docx-1-cbm-10.1177_18758592251390145 for TUBB3 (βIII-tubulin) drives gastric cancer progression and poor prognosis by regulating cell cycle and invadopodia formation by Hui Ge, Hui Tang, Tingting You, Chunmei Bai, Zhao Sun, Qin Han and Robert Chunhua Zhao in Cancer Biomarkers
Supplemental Material
sj-xlsx-2-cbm-10.1177_18758592251390145 - Supplemental material for TUBB3 (βIII-tubulin) drives gastric cancer progression and poor prognosis by regulating cell cycle and invadopodia formation
Supplemental material, sj-xlsx-2-cbm-10.1177_18758592251390145 for TUBB3 (βIII-tubulin) drives gastric cancer progression and poor prognosis by regulating cell cycle and invadopodia formation by Hui Ge, Hui Tang, Tingting You, Chunmei Bai, Zhao Sun, Qin Han and Robert Chunhua Zhao in Cancer Biomarkers
Footnotes
Abbreviations
Acknowledgements
Thanks for the technical support by Experimental Medicine Center, Department of Medical Oncology, Peking Union Medical College Hospital, Chinese Academy of Medical Sciences & Peking Union Medical College, Beijing, China
Ethics approval and consent for publication
The study was conducted in accordance with the Declaration of Helsinki and approved by the Medical Ethics Committee of Peking Union Medical College Hospital (Approval No: ZS-2087). Written informed consent was obtained from all participants prior to their inclusion in the study.
All animal experiments were conducted in accordance with the ARRIVE (Animal Research: Reporting of In Vivo Experiments) guidelines and were approved by the Peking Union Medical College Hospital Application Form Welfare & Ethical Inspection in Animal Experimentation.
Consent for publication
Not applicable.
Author's contributions
HG, HT, and TY performed the experiments, gathered the data, and drafted the manuscript. CB, ZS, QH, and RC designed the experiments, supervised the research, and reviewed the manuscript. All authors contributed to the article and approved the final version for submission.
CRediT authorship contribution statement
Hui Ge: Writing – Initial draft, Verification, Analytical method. Hui Tang: Writing – Initial draft, Verification, Analytical method, Data curation. Tingting You: Data curation. Chunmei Bai: Writing – Evaluation & refinement, Assets, Project coordination, Inquiry, Financial support acquisition. Zhao Sun: Writing – Evaluation & refinement, Assets, Project coordination, Inquiry, Financial support acquisition. Qin Han: Writing – Evaluation & refinement, Assets, Project coordination, Inquiry, Financial support acquisition. Robert Chunhua Zhao: Writing – Evaluation & refinement, Assets, Project coordination, Inquiry, Financial support acquisition.
Funding
This study was sponsored by National High Level Hospital Clinical Research Funding (2022-PUMCH-D-001); Chinese Academy of Medical Sciences Innovation Fund for Medical Sciences (2024-I2M-ZD-001).
Competing interest
The authors certify that no known financial or personal conflicts of interest have influenced the integrity of the research presented in this study.
Availability of data and materials
The novel contributions of this study are presented within the article and supplementary materials. For additional information, please reach out to the corresponding author.
Supplemental material
Supplemental material for this article is available online.
Author Biographies
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
