Abstract
Background
Metastasis-associated in colon cancer-1 (MACC1) is a novel oncogene involved in the growth and metastasis of tumors, which is overexpression in various tumors. MACC1 could promote the growth, invasion, and metastasis of colorectal cancer (CRC) by activating the HGF/MET signaling pathway in vivo. It has been confirmed that MACC1 mainly promotes the Warburg effect in gastric cancer cells through the PI3 K/AKT signaling pathway.
Objective
Here, we mainly investigated the association between MACC1 and the glycolysis process in CRC cells.
Methods
The expression of MACC1 in CRC and its relationship with the patient's survival were analyzed by TCGA database. We used different concentrations of glucose medium to culture HT-29 and HCT-116 with or without siMACC1. Cell Counting Kit-8 assay was used to detect cell proliferation. The content of lactic acid and glucose in cells was examined by enzyme-linked immunosorbent assay, and extracellular acidification rate was also determined with Seahorse XFe96 Extracellular Flux Analyzer. Western blot was used to detect the protein expressions related to glycolysis. Immunofluorescence was conducted to observe the expression and distribution of GLUT4.
Results
In this study, we observed that MACC1 was highly expressed in CRC and negatively correlated with the survival of patients. The expression of glycolysis related enzymes was significantly increased under the stimulation of different concentrations of glucose in HT-29 and HCT-116 cells. However, MACC1 knockdown could significantly reduce high glucose-induced the expressions of glycolysis-related enzymes. Besides, MACC1 knockdown could decrease the content of glucose and lactate, and inhibit glycolytic function in HT-29 and HCT-116 cells. Moreover, the expression of GLUT4 was significantly decreased in the two cell lines treated by 4.5 g/L or 9.0 g/L glucose with MACC1 knockdown. Additionally, MACC1 knockdown could inhibit high glucose-induced membrane translocation of GLUT4.
Conclusions
MACC1 knockdown can attenuate glucose metabolism by inhibiting the membrane translocation of GLUT4 in CRC cells.
Introduction
Colorectal cancer (CRC) is one of the most common malignant tumors of digestive system. There are approximately 1.9 million people diagnosed with CRC in 2020, which is the third most commonly diagnosed form of cancer worldwide. 1 At present, it is generally believed that CRC is a disease with multi-factor action, multi-gene participation and multi-stage development. 1 With the development of tumor molecular biology, more and more genes have been found to be involved in the development of CRC.
In 2009, it was confirmed that metastasis-associated in colon cancer-1 (MACC1) was a gene closely related to CRC metastasis. 2 It is a novel oncogenic factor involved in tumor growth and metastasis, which is located on human chromosome (7p21.1). Studies have proved that MACC1 is aberrantly overexpressed in various types of tumor, including lung cancer, gastric cancer, CRC and breast cancer.3–7 It may be a novel marker of CRC prognosis due to the association with metastasis and prognosis of CRC.8,9 MACC1 could promote the growth, invasion and metastasis of CRC through activating HGF/MET signaling pathway. 6 It has been suggested that abnormally elevated MACC1 expression is associated with invasion, metastasis and poor prognosis in CRC.
There are great differences in glucose metabolism between tumor cells and normal cells. Normal cells metabolize glucose mainly through oxidative phosphorylation, while glycolytic pathway will be started under anoxic or anaerobic conditions with generation of a small amount of energy in the whole glucose metabolism process. However, even under the condition of sufficient oxygen, tumor cells still undergo the process of high anaerobic glycolysis, which is called aerobic glycolysis, also known as ‘Warburg effect’. 10 Previous studies have demonstrated that MACC1 promotes Warburg effect mainly through PI3 K/AKT signaling pathway in gastric cancer cells. 11 In hepatocellular carcinoma, MACC1 induced tumor glucose metabolism by upregulating hexokinase-2 (HK2) indirectly and causing high levels of proliferation. 12 All of these studies have proved that MACC1 are associated with tumor glucose metabolism. However, the relationship between MACC1 and glycolysis in CRC is still unclear. Therefore, the molecular mechanism that MACC1 leads to CRC progression by influencing glucose metabolism remains to be elucidated.
In this study, we first investigate the effects of different concentrations of glucose on MACC1 expression and glycolysis-related enzymes in CRC cells. In addition, we observe the effect of MACC1 on the expressions of glycolysis-related enzymes and the membrane translocation of GLUT4 in CRC cells. This study found the knockdown of MACC1 attenuated the expression of glycolysis-related enzymes through weakening GLUT4 membrane translocation.
Materials and methods
Cell lines and cell culture
Human CRC cell lines, HT-29 and HCT-116, were purchased from Procell Life Science&Technology Co.,Ltd Cels were cultured in McCoy's 5A medium(Viva Cell; LOT 2210062) supplemented with 10% FBS (Gibco; Thermo Fisher Scientific, Inc.) and 100 U/mL penicillin and 100 mg/mL streptomycin at 37 °C in a humidified atmosphere containing 5% CO2.
Gene expression profiling interactive analysis (GEPIA)
Utilize the GEPIA database (http://gepia2.cancer-pku.cn/) to analyze the expression of the MACC1 gene in cancer and its relationship with survival. The “Survival” module displays the relationship between the gene and overall survival rates in TCGA-COAD-READ, with survival analysis results shown through Kaplan-Meier curves indicating P-values obtained from the log-rank test. The criterion for statistical significance is established at a P-value of less than 0.05.
RNA isolation and real-time PCR
Total RNA was isolated according to the manufacturer's instructions using TRIzol reagent (Cat. no.9108, TaKaRa) and the concentration of RNA was determined on an ultraviolet spectrophotometer (Flash, China). Next, 1 μg RNA was reverse transcribed into cDNA using HiScript® III All-in-one RT SuperMix Perfect for qPCR (Cat. no.R333, Vazyme). Real-time PCR was performed using ChamQ Universal SYBR qPCR Master Mix (Cat. no. Q711, Vazyme) in a total volume of 20 μl on the X 960 Real-time PCR (Heal Force, China) as follows: 95 °C for 30 s, followed by 40 cycles of 95 °C for 5 s and 60 °C for 20 s. The relative mRNA levels of target genes were calculated using the 2−△△Ct method.
Cell immunofluorescence
Cells were seeded in a 6-well plate with acid-washed glass coverslips. The cells were washed with PBS after different treatments, then the cells were fixed with 4% paraformaldehyde for 30 min, washed five times with phosphate-buffered saline (PBS), and permeabilized with 0.1% Triton X-100 for 30 min at room temperature (RT). Following that, the cells were incubated with blocking buffer for 30 min at RT. Then, cells were incubated with anti-GLUT4 (1:400) at 4°C overnight. The next day, the second antibody was incubated after washed five times with PBS and stained with phalloidin at the same time. The cells were counterstained with DAPI for nuclear staining and the fluorescence was observed under a Zeiss confocal microscope. Images were analyzed and processed using ImageJ.
Small interfering (siRNA) synthesis and transfection
siRNA oligonucleotides targeting the human MACC1 gene (siMACC1) were designed and synthesized. A random sequence was used as negative control. The riboFECT TM CP Transfection Kit (Cat. no. C10511-05; Ribobio) was used to transfect siMACC1 (GCCCAUCAGAGUAUACAUATT) into HT-29 or HCT-116. In brief, 50 pmol siMACC1 with 1 × buffer and 12 µl Reagent were mixed for 15 min and co-cultured with HT-29 or HCT-116 for 24 h at 37˚C with 5% CO2. The time interval between transfection and the subsequent experiments was 24 h.
Cell Counting Kit-8 (CCK8) assay
Cells were seeded in a 96-well plate at a density of 5000 cells per well. When the cell density reached 60% confluence, the cells were subjected to transfection with either si-MACC1 or si-NC for 24 h. Subsequently, each well received 10 µL of CCK-8 reagent (Cat. C0038, Beyotime, Shanghai, China), followed by incubation of the plate in the dark at 37 °C for 2 h. The OD was measured at 450 nm using a microplate reader.
Western blot analysis
The harvested cells were lysed using RIPA Lysis Buffer (Cat. no. R0010, Solarbio). Equal quantities of the protein samples (30μg of total protein/sample) were separated by denaturing gel electrophoresis (Bio-Rad Laboratories, Inc.). Following transfer onto poly-vinylidene difluoride membranes (Cat. no. 03010040001; Roche). Blotted membranes were blocked in 5% non-fat milk for 1 h and then incubated overnight at 4°C with primary antibodies against MACC1(Cat. no. ER65531; HUABIO), Hexokinase II (Cat. no. DF6176; Affinity Biosciences), LDHA (Cat. no. DF6280; Affinity Biosciences), PFKM (Cat. No. CSB-PA017822LA01HU; CUSABIO), β-actin (Cat. no. WL01372; Wanleibio) and GAPDH (Cat. no. ET1601-4; HUABIO). The membranes were further incubated with horseradish peroxidase-conjugated secondary antibodies for 1 h, followed by signal visualization using an electrochemical luminescence reagent. Images were acquired using a chemiluminescence imaging system JP-K600 (Jiapeng, China).
Detection of intracellular glucose and lactate levels
Measure the glucose and lactate content inside cells using a reagent kit (Jiancheng, Nanjing, China). Add the sample and working solution separately into a 96 well plate according to the instructions, and set up a blank group and a standard group. Each group should have at least three parallel control wells. After gently shaking the plate, incubate it at 37 °C for 10 min, and measure the OD value of each well at a wavelength of 505 nm using an enzyme-linked immunosorbent assay (ELISA) reader.
Detection of cellular glucose metabolism function
Extracellular acidification rate (ECAR) was determined using a Seahorse XFe96 Extracellular Flux Analyzer (Agilent, CA, USA). Briefly, cells were seeded in XFe96 Cell Culture Microplates and incubated at 37 °C for 24 h. The culture medium was changed to XF assay medium supplemented with 1 mM glutamine for ECAR assay, and cells in the new medium were placed in a 37 °C incubator without CO2 for 1 h. ECAR was examined under basal conditions and during the sequential injection of 10 mM glucose, 1 µM oligomycin, and 50 mM 2-DG. After each assay, cells were lysed, and the protein concentration was measured to normalize the ECAR.
Statistical analysis
All data were statistically analyzed using the SPSS software (version 16.0; SPSS, Inc.). All data are presented as the mean ± standard deviation of the mean (SD) from at least three independent experiments (n ≥ 3), and each independent experiment was repeated three times to obtain the mean. The means derived from same treatment group were analyzed statistically. Student's t-test was used to evaluate the significance between two groups, while one-way analysis of variance followed by Tukey's test was used to estimate the difference among multiple groups. *P < 0.05 was considered to indicate a statistically significant difference.
Results
MACC1 was upregulated in CRC and correlated with poor prognosis in patients
To investigate the role of MACC1 in tumors, we initially employed the TCGA database to analyze the expression of MACC1 in various tumor tissues and adjacent normal tissues. The analysis revealed significant expression differences of MACC1 in multiple tumors (Figure 1A), suggesting its potential significance in the initiation and progression of cancer. We also found that MACC1 expression is higher in tumor tissues than adjacent normal tissues in CRC. Additionally, we performed survival analysis on CRC patients from TCGA and the findings indicated that patients with high MACC1 expression had significantly shorter overall survival (OS) and disease-free survival (DFS) compared to those with low expression (Figure 1B and 1C).
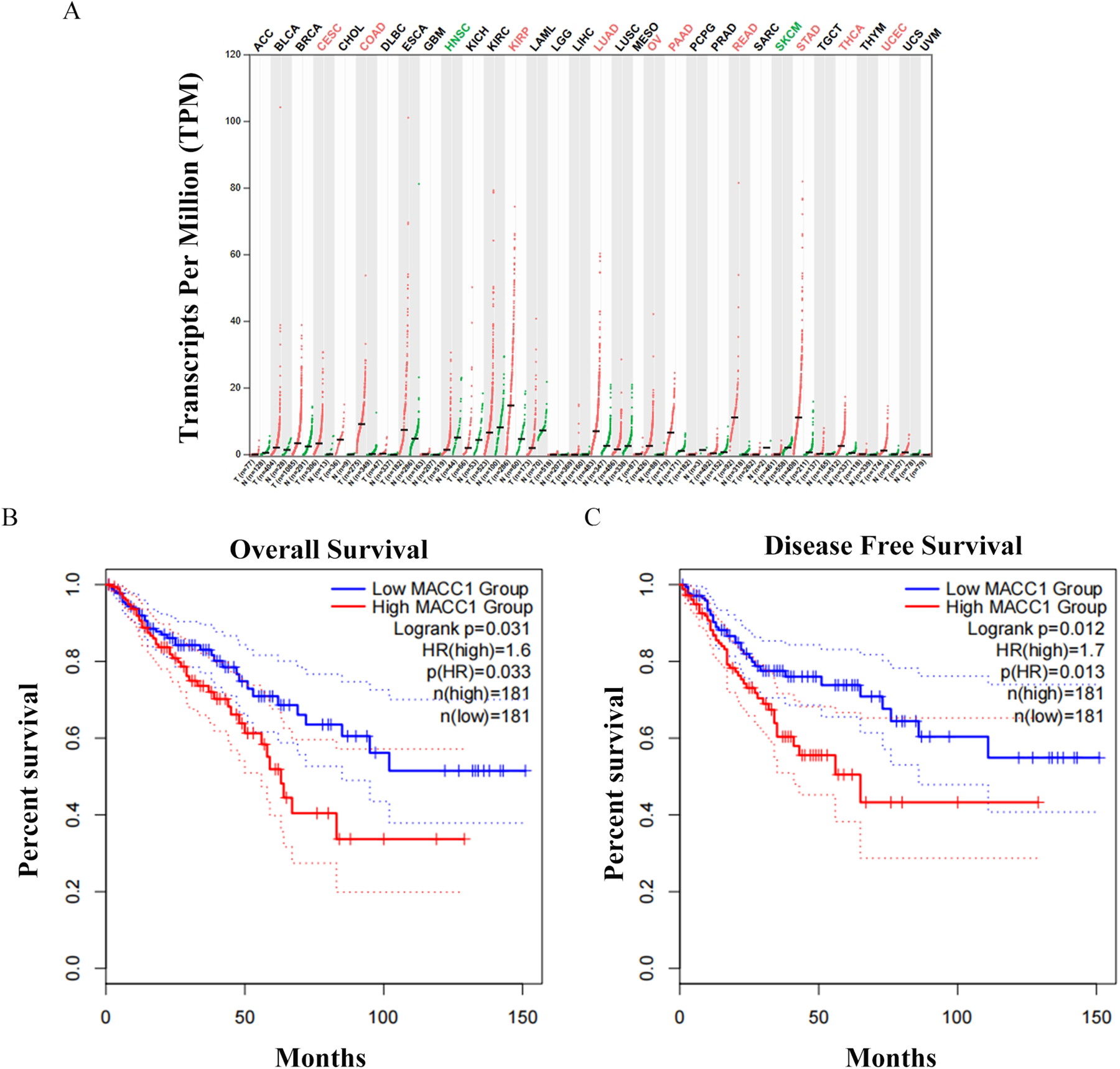
The expression of MACC1 in cancer and patient survival cycle. (A) MACC1 is highly expressed in cancer. (B) The relationship between the expression of MACC1 and the overall survival cycle of patients. (C) The relationship between the expression of MACC1 and the disease-free survival cycle of patients.
High glucose increased the expression of glycolytic related enzymes in CRC cells
To study the potential effect of glucose on the glycolysis process of CRC cells, the cells were treated with three concentrations of glucose medium (1.0 g/L(control), 4.5 g/L and 9.0 g/L). The protein expressions of glycolysis-related enzymes were detected by western blot to reflect the influence of glucose on glycolysis process. The 4.5 g/L and 9.0 g/L glucose medium could increase the protein expressions of LDHA, PFKM, HK2, GAPDH and MCCA1 in HT-29 cells (Figure 2A and 2B) and HCT-116 cells (Figure 2C and 2D). These results indicate that MACC1 may involve in glycolysis process.
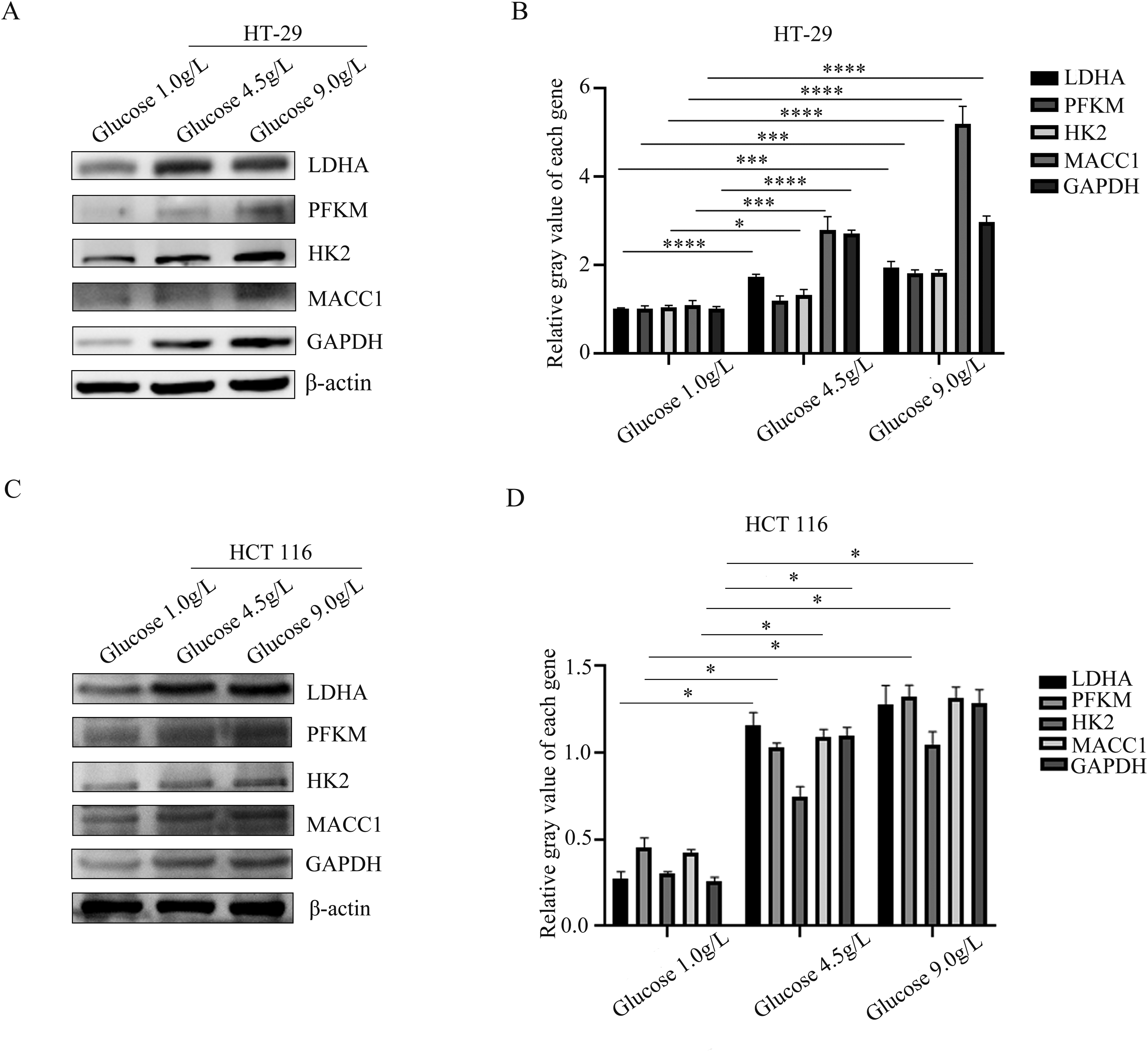
Protein expression of glycolytic related enzymes in different concentrations of glucose. The two cell lines were stimulated with glucose stimulation for 72 h. (A) The expressions of LDHA, PFKM, HK2 and MACC1 in HT-29 cells were detected by western blots. (B) The gray value of the strip was measured with ImageJ. (C) The expressions of LDHA, PFKM, HK2 and MACC1in HCT-116 cells were detected by western blots. (D) The gray value of the strip was measured with ImageJ. Data were expressed as mean ± standard deviation (SD), n = 3. *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001, compared with the glucose 1.0 g/L group.
The expressions of glycolysis related enzymes were decreased by knocking down MACC1 in CRC cells
According to the above results, we proved that the expression of MCCA1 was positive correlated with that of glycolysis-related enzymes. Therefore, to explore the effect of MACC1 on glycolysis of CRC cells, we first transfected the siRNA of MACC1 into HT-29 cells or HCT-116 cells with the culture medium of normal glucose concentration. The results showed that the knock-down efficiency of MACC1 reached 50% after 24 h of transfection with siMACC1 in both HT-29 cells (Figure 3A) and HCT-116 cells (Figure 3B).
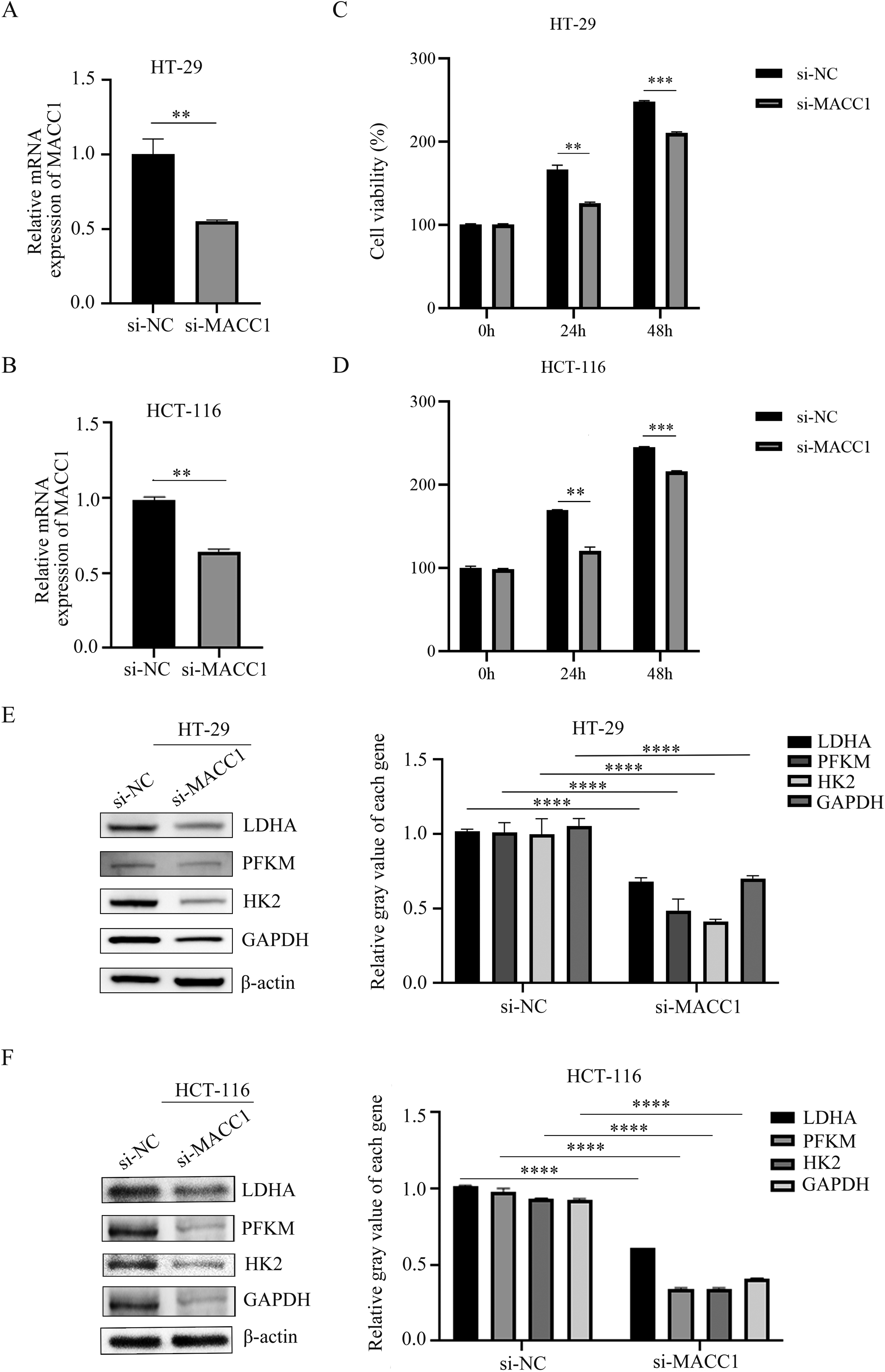
Effect of knocking down MACC1 on the expression of glycolytic related enzymes. (A-B) The expression of MACC1 at RNA level was detected after knocking down MACC1 in HT-29 (A) and HCT-116 (B) cells. (C-D) The effect of MACC1 knockdown on proliferation of HT-29 (C) and HCT-116 (D) cells. (E) The expressions of LDHA, PFKM, HK2 and MACC1 were detected by western blots in HT-29 cells. (F) The expressions of LDHA, PFKM, HK2 and MACC1 were detected by western blots in HCT-116 cells. Data were expressed as mean ± SD, n = 3. **P < 0.01,***P < 0.001, ****P < 0.0001, compared with si-NC group.
From the CCK-8 assays, we observed that knockdown of MACC1 inhibited the proliferation of HT-29 (Figure 3C) and HCT-116 cells (Figure 3D). Moreover, we detected the protein expressions of glycolysis-related enzymes by western blot. We found that MACC1 knockdown could decrease the protein expressions of LDHA, PFKM, HK2 and GAPDH in both HT-29 cells (Figure 3E) and HCT-116 cells (Figure 3F). These results indicate that MACC1 can affect the glycolysis process in CRC cells.
Knockdown of MACC1 inhibited glycolysis in CRC cells
Next, we investigated the effect of MACC1 knockdown on glycolytic levels in CRC cells, and some glycolysis indicators were tested. We found that si-MACC1 could decrease the glucose content, but it could increase the lactate level (Figure 4C and 4D) (P < 0.05). In addition, ECAR analysis showed that knocking down MACC1 reduced the levels of glycolysis in HT-29 cells (Figure 4E) and HCT-116 cells (Figure 4F). Taken together, these results indicate that knockdown of MACC1 can inhibit glycolysis in CRC cells.
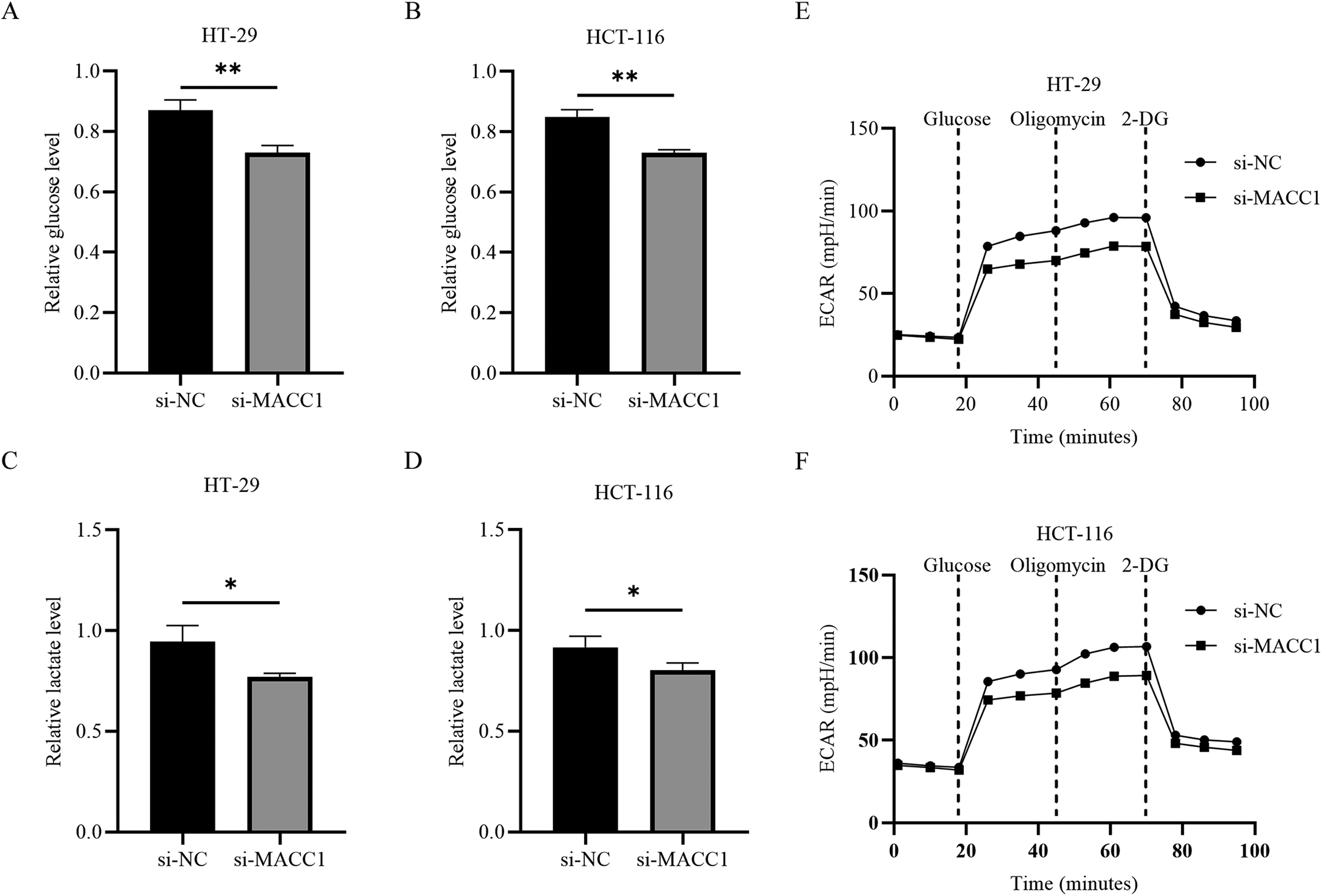
The effect of knocking down MACC1 on glycolysis function. (A-B) Glucose content was examined after MACC1 knockdown in HT-29 (A) and HCT-116 (B) cells. (C-D) Lactate content was examined after MACC1 knockdown in HT-29 (C) and HCT-116 (D) cells. (E-F) Glycolytic capacity of HT-29 (E) and HCT-116 (F) cells was measured by glycolytic rate assay. Data were expressed as mean ± standard deviation (SD), n = 3. **P < 0.01, compared with the si-NC.
The protein expressions of glycolytic related enzymes were inhibited with MACC1 knockdown in CRC cells treated with high glucose
To explore the effects MACC1 on the glycolysis process of CRC cells under different high glucose concentrations, we used western blot to determine the changes of glycolysis-related enzymes in CRC cells with or without MACC1 knockdown. After the cells were stimulated with 4.5 g/L or 9.0 g/L for 24 h under knocking down MACC1, the protein expressions of LDHA, PFKM, HK2, GAPDH and MACC1 were significantly decreased in both HT-29 cells (Figure 5A and 5B) and HCT-116 cells (Figure 5C and 5D).
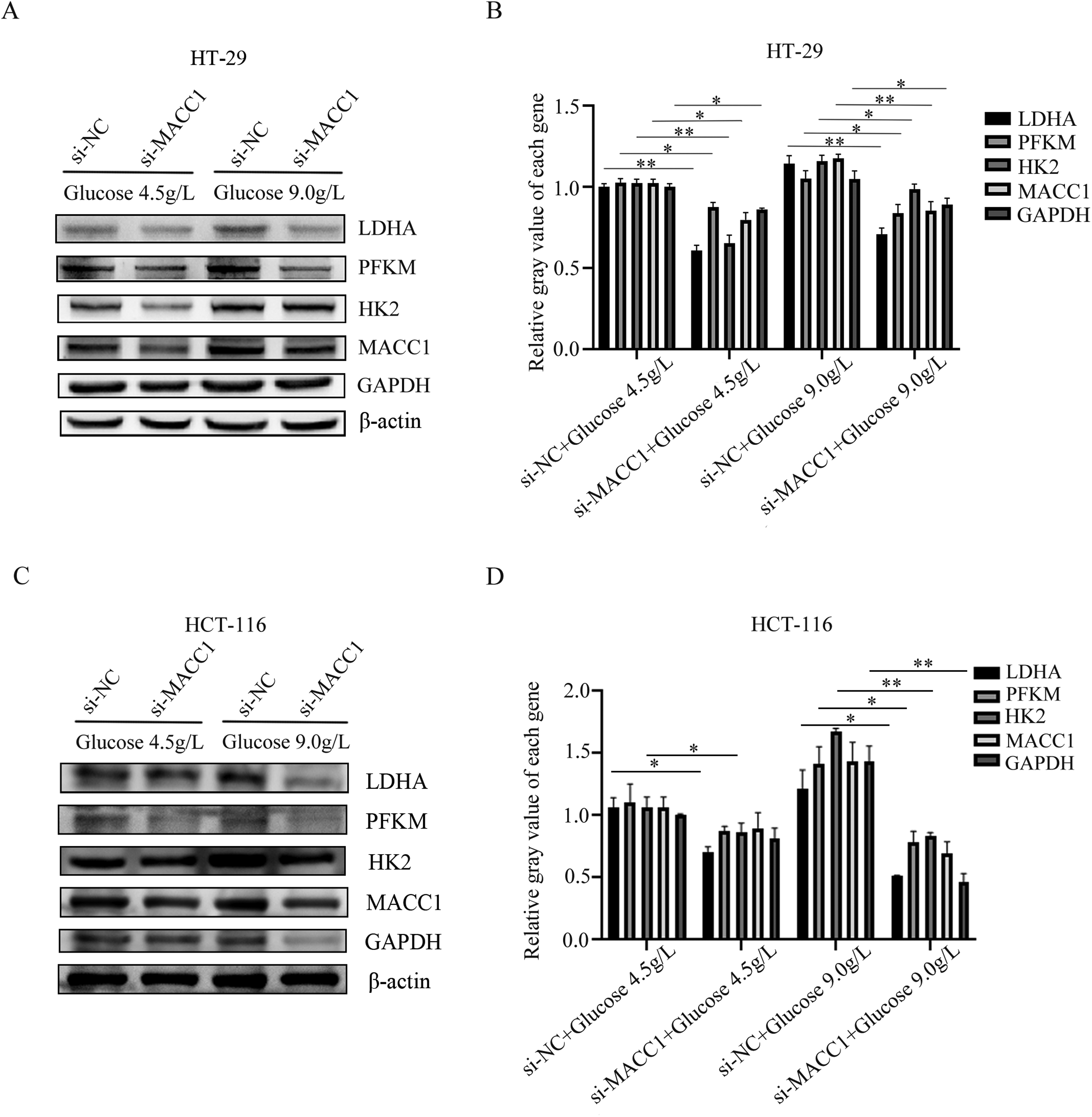
Under the stimulation of different concentrations of glucose, knocking down MACC1 will affect the protein expression of glycolytic related enzymes. (A-B) The protein expressions of LDHA, PFKM, HK2 and MACC1 were detected by western blot in HT-29 cells treated with 4.5 g/L or 9.0 g/L glucose with si-Con or si-MACC1. (C-D) The protein expressions of LDHA, PFKM, HK2 and MACC1 were detected by western blot in HCT-116 cells treated with 4.5 g/L or 9.0 g/L glucose with si-Con or si-MACC1. Data were expressed as mean ± SD, n = 3. *P < 0.05, **P < 0.01, compared with si-NC + glucose 4.5 g/L group or si-NC + glucose 9.0 g/L group.
Knocking down MACC1 could disturb the expression and distribution of GLUT4 under different concentrations of glucose in CRC cells
GLUT4 is one of glucose transporters, which participates in the rapid uptake of glucose in various cells, thus contributing to glucose homeostasis. After MACC1 knockdown, the expressions of glycolysis-related enzymes were inhibited under the stimulation of glucose with different concentration in CRC cells. Therefore, we suspected that MACC1 knockdown could affect the expression and distribution of GLUT4. Next, we used immunofluorescence to observe the expression and distribution of GLUT4 after MACC1 knockdown under glucose stimulation. The results showed that the expression of GLUT4 in siMACC1 group was significantly lower than that in siNC group. In addition, the distribution of GLUT4 to cell membrane was reduced after MACC1 knockdown. All these results indicated that the absorption of glucose by GLUT4 was weakened in both HT-29 cells (Figure 6A) and HCT-116 cells (Figure 6B) after MACC1 knockdown.
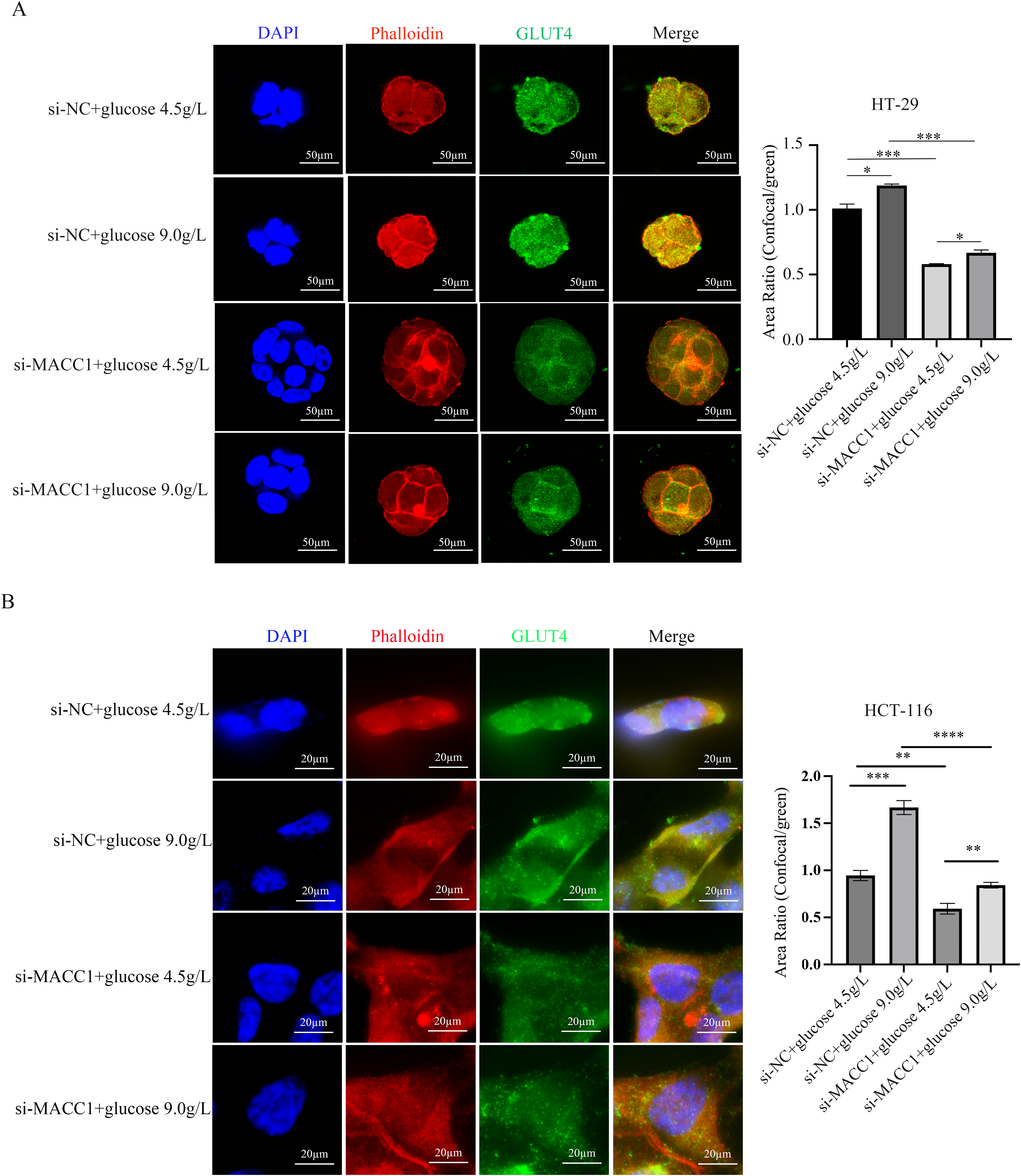
Under the stimulation of different concentrations of glucose, knocking down MACC1 will affect the expression and distribution of GLUT4. (A) Immunofluorescence staining determined the expression and distribution of GLUT4 in HT-29 cells treated with 4.5 g/L or 9.0 g/L glucose with si-Con or si-MACC1. Scale bar = 50 μm. (B) Immunofluorescence staining determined the expression and distribution of GLUT4 in HCT-116 cells treated with 4.5 g/L or 9.0 g/L glucose with si-Con or si-MACC1. Scale bar = 20 μm.
Besides, we also found that the morphology of cells was partially aggregated into balls with the loss of skeleton, the disorder of skeleton arrangement and the chaotic and fuzzy microfilament skeleton in CRC cells with MACC1 knockdown. However, the cytoskeleton structure showed that longer and more parallel F-actin stress fibers passed through the whole cell without knocking down the MACC1. The results showed that it could not only inhibit GLUT4, but also rearrange the cytoskeleton of cells in CRC cells with MACC1 knockdown.
Discussion
Unlike most normal somatic cells, cancer cells are metabolized mainly by glycolysis (Warburg effect), even under aerobic conditions. 13 Therefore, glycolysis process is essential for the study of tumor cells. The purpose of this function of tumor cells is to maintain their redox state and meet the energy and nutrition needed for their rapid proliferation. The glycolysis is regulated by a series of glycolytic enzymes, which is a multi-step reaction. 14 Once the glucose enters the cytoplasm, it will be phosphorylated into the membrane-impermeable glucose-6-phosphate by HKs. Glucose-6-phosphate is converted into pyruvate by a series of enzymes such as PFKs and pyruvate kinase.15,16 At this time, pyruvate is converted into acetyl coenzyme A under the catalysis of pyruvate dehydrogenase in the presence of oxygen, and into lactic acid under the action of LDHAs in the absence of oxygen.17,18 GAPDH is also an essential metabolic enzyme in glycolysis and gluconeogenesis. 19 MACC1 is a transcription factor which can promote tumor cell motility and invasion and cause metastasis in solid tumors, such as CRC, pancreatic cancer and breast cancer.7,20,21 Many studies proved that MACC1 were associated with tumor glucose metabolism. Therefore, we chose HK2, LDHA, GAPDH and PFKM as indicators to confirm whether MACC1 could affect the glycolysis process of CRC cells with high glucose culture. Our experimental results showed that the protein expressions of glycolysis-related enzymes and MACC1 were increased in a glucose concentration dependent manner in CRC cells. This result indicated that MACC1 was positively correlated with glycolysis process of CRC cells. Therefore, we suspected that MACC1 could affect the glycolysis process of CRC cells. To further verify our hypothesis, we detected the protein expressions of glycolysis-related enzymes in CRC cells with or without MACC1 knockdown. We found that knockdown of MACC1 could decrease the expression of glycolysis related enzymes and inhibit cell proliferation. This indicates that knockdown of MACC1 can restrain the proliferation of CRC cells by inhibiting glycolysis.
The increase of glucose metabolism and uptake are characteristic of many cancer cells. 14 In order to further prove that MACC1 is involved in the glycolysis of CRC cells, we detected the content of glucose and lactate, as well as the ECAR. The results showed that knockdown of MACC1 not only reduced glucose uptake and lactate production, but also inhibited the glycolytic function in CRC cells. Glucose uptake is the main process of glycolysis, while glucose enters the cytoplasm matrix through the cell membrane needing the help of GLUTs. 22 Studies have shown that the distribution and function of some types of GLUTs have tissue and cell specificity. 23 Different GLUTs located in different cells show different affinity for glucose, which adapts to their specific metabolic needs. 8 Among them, GLUT4 may be the most studied subtype of GLUTs, because it plays a major role in systemic glucose homeostasis and the pathogenesis of type II diabetes. 24 Therefore, we stimulated CRC cells with different concentrations of glucose after knocking down MACC1, and then observed the expression and distribution of GLUT4 by immunofluorescence staining. We found that MACC1 knockdown could affect the glycolysis process of CRC cells by disturbing GLUT4 location under high concentration of glucose treatment.
MACC1 is an important prognostic marker for metastases formation in CRC. In this study, we linked MACC1 with the glucose metabolism pathway of CRC cells. Our results proved that knockdown of MACC1 could suppress the proliferation by inhibiting the glucose metabolism of CRC cells. Besides, MACC1 could affect glucose metabolism and transport during glycolysis in CRC cells. These findings may provide a new strategy for the clinical treatment of colorectal cancer. However, although we have proved that knockdown of MACC1 could affect the expression and distribution of GLUT4, the underlying mechanism needs to be further explored.
Conclusion
In this study, we found that knocking down MACC1 could reduce the expression of glycolytic enzymes in cells. Moreover, MACC1 knockdown could reduce glucose uptake and decrease glycolysis rate by inhibiting GLUT4 membrane translocation, thereby suppressing cell proliferation.
Footnotes
Acknowledgements
This work was supported by Hebei Province Medical Science Research Plan Project (China) [grant number 20221808].
Author contributions/CRediT
Interpretation or analysis of data: Qingke Li, Lifang He, Zhiwu Wang
Preparation of the manuscript: Qingke Li, Lifang He, Zhiwu Wang
Revision for important intellectual content: Qingke Li, Lifang He, Yufeng Li and Jingwu Li
Supervision: Qingke Li and Jingwu Li
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
Conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
