Abstract
Central nervous system tumors, especially astrocytomas, are the solid neoplasms with the highest incidence and mortality rates in childhood. The diagnosis is based on histopathological characteristics, but molecular methods have been increasingly used. Translationally controlled tumor protein (TCTP) protein, encoded by the tumor protein, translationally controlled 1 (TPT1) gene, is a multifunctional protein with an important physiological role in the cell cycle. Expression of this protein has been associated with several neoplasms, including astrocytomas in adults. However, the role of this protein in pediatric astrocytomas is largely unknown. We aim to evaluate in cases of pediatric astrocytomas, the frequency of polymorphisms in the TPT1 gene and other genes associated with its molecular pathways, such as MTOR, MDM2, TP53, and CDKN1A, correlating it with protein expression and clinical variables, in formalin-fixed, paraffin-embedded (FFPE) samples. These samples were submitted to genotyping and immunohistochemistry analyses. The most revealing results refer to the MDM2 gene, rs117039649 [G/C], in which C polymorphic allele was observed only in the glioblastomas (p = .028). The CDKN1A gene, rs3176334 [T/C] presented a homozygous polymorphic genotype only in high-grade astrocytomas, when infiltrating tumors were compared (p = .039). The immunohistochemical expression of cytoplasmic MDM2 correlated with better survival rates in patients with glioblastoma (p = .018). The presence of polymorphisms in the MDM2 and CDKN1A genes, as well as a specific correlation between MDM2 expression, suggests a likely association with risk in pediatric astrocytomas. This study sought the probable role involved in the TCTP pathway, and associated proteins, in the tumorigenesis of pediatric astrocytomas, and some could have potential impact as prognostic markers in these patients.
Introduction
Translationally controlled tumor protein (TCTP) is a protein, highly conserved in eukaryotic cells, encoded by tumor protein, translationally controlled 1 (TPT1) gene (Acunzo et al., 2014; Bommer & Thilele, 2004). TCTP is usually expressed in mitotically active tissues, and it is involved physiologically in cell proliferation, protein synthesis, and immune response (Bommer & Thilele, 2004). Through interaction with other proteins, such as MTOR and MDM2, TCTP can affect the expression of p53 and p21 proteins, impacting apoptosis and cell proliferation (Acunzo et al., 2014; Amson et al., 2012; Bommer et al., 2015). All these proteins have physiological interactions but can also acquire pathological status, which has associated TCTP with several neoplasms, including gliomas (Amson et al., 2012; Chen et al., 2015; Gu et al., 2014; Miao et al., 2013; Rocca et al., 2015). Studies have shown that TCTP was more expressed in high-grade gliomas and indicated worse prognosis, in the adulthood, whereas its expression decreased in tumor reversion phenomena, a biological process in which neoplastic cells lose its malignant phenotype (Amson et al., 2013; Tuynder et al., 2004). Moreover, TCTP has been studied as a therapeutic target, and sertraline, an antidepressant, has been shown to inhibit TCTP levels, thus increasing p53 expression, and inducing tumor reversion (Amson et al., 2013; Baú-Carneiro et al., 2022; Boia-Ferreira et al., 2017).
Genetic variations are quite common throughout the human genome. The simplest form of variation is the exchange of a single nucleotide (single-nucleotide polymorphisms—SNPs). It is estimated that there are around 8 million SNPs with an allelic frequency greater than 1%. Although most SNPs do not have important functional associations, there are certain SNPs that can substantially alter the ultimate expression or function of a gene product (Chanock, 2001). These functional, or unfunctional, variants can affect the final observed phenotype and thus change the risk classification of the disease, contribute to the understanding of early diagnosis and the possible impacts on changing the response to treatment (Lehrnbecher et al., 2005; Pertovaara et al., 2006). Polymorphic genetic variants are the basis of genetic association studies in research on genetic risk factors for disease susceptibility. The expression of molecules from the transcription factor superfamily can be altered by the presence of a certain polymorphism, which makes this project attractive since it seeks for possible genetic markers that may be associated with worse prognosis in pediatric astrocytomas, the most common solid malignant neoplasia in children (Ostrom et al., 2020). Thus, the present study aims to investigate the association between SNPs in TPT1, MTOR, MDM2, TP53, and CDKN1A genes in pediatric astrocytoma samples, as well as protein expression, to try to evidence potential genetic biomarkers that would help predict the patients’ prognosis.
Methods
Study Population
This study was approved by the Human Research Ethics Committee of the Hospital Pequeno Príncipe, in Curitiba, Paraná (CAAE: 13240619.5.3001.0020/2019). This institution was founded in 1919 and today it is a renowned pediatric hospital in Brazil, a region with a particular colonization, compared to the other regions of our country. Most of its population descends from European immigrants, such as Italian, German, Polish, Ukrainian, among others. One hundred eighty-six primary pediatric astrocytomas diagnosed between 2003 and 2015 were selected, reviewed, and anonymized and with unrelated parentage.
The following variables were collected from the medical records: sex (male, female); age at diagnosis (years); outcome (death); and survival (months), and these subjects were grouped according to the World Health Organization (WHO) classification (Louis, 2016).
One hundred-one cases of the 186 cases previously reviewed still had remaining neoplastic tissue in their formalin fixed, paraffin-embedded (FFPE) samples. FFPE blocks were obtained from the HPP Biobank. The final sample consisted of 56 pilocytic astrocytomas (WHO grade 1), 23 diffuse astrocytomas (WHO grade 2), 3 anaplastic astrocytomas (WHO grade 3), and 19 glioblastomas (WHO grade 4).
Attempting to homogenize the phenotype to improve the analyses, groupings based on the four histological grades were used. This grouping was carried out based on morphological, molecular, and prognostic differences between the astrocytomas, and the samples were divided into circumscribed astrocytomas, which included only grade 1, and infiltrating astrocytomas, referring to grades 2, 3, and 4.
Seeking the most appropriate analyses from a clinical point of view, three forms of analysis were organized. The astrocytomas were compared in isolated histological groups (1, 2, 3, and 4) and in two different groupings: low-grade×high-grade (grades 1 + 2×grades 3 + 4), and low-grade×high-grade in infiltrating astrocytomas (grade 2×grades 3 + 4). Table 1 shows the baseline characteristics in the study population.
Baseline Characteristics in the Study Population.
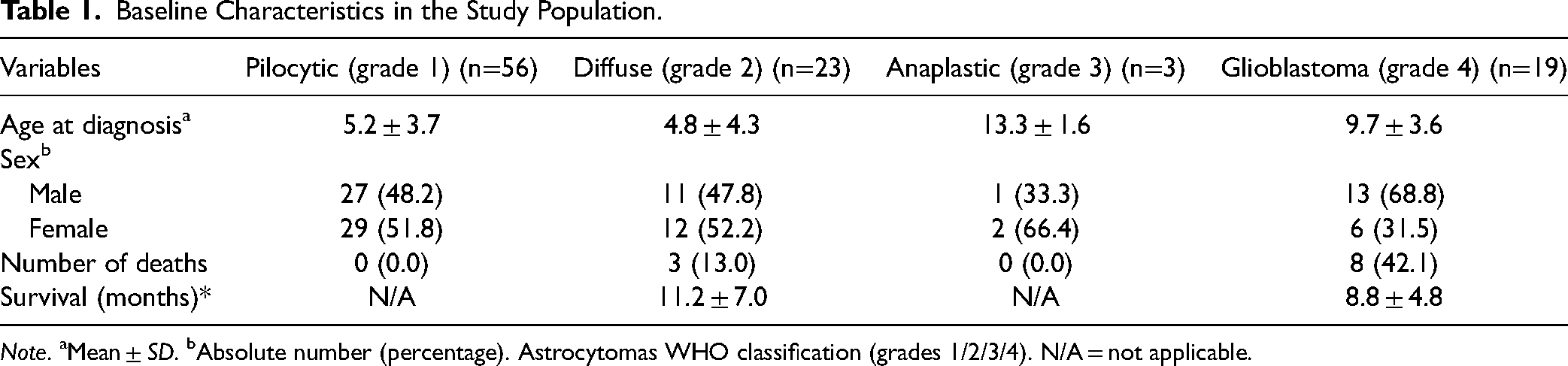

Note. aMean ± SD. bAbsolute number (percentage). Astrocytomas WHO classification (grades 1/2/3/4). N/A = not applicable.
DNA Extraction and Genotyping
The DNA was obtained from paraffinized cuts of the samples using a commercially available paraffin DNA extraction kit (Qiagen®). Markers for coverage (target SNP—tag SNP) of five genes’ loci (TPT1, MTOR, MDM2, TP53, and CDKN1A) were selected according to available information in the SNPinfo website (SNPinfo, 2022). After this search three tag SNPs for TPT1 gene (rs2234222 [G/C]; rs9595305 [T/C] and rs1062420 [T/C]), two for MTOR gene (rs1034528 [G/C]; rs12732063 [G/A]), two for MDM2 gene (rs2279744 [T/G]; rs117039649 [G/C]), one for TP53 gene (rs121912664 [C/T]), and two for CDKN1A gene (rs3176326 [G/A]; rs3176334 [T/C] were selected. The patients’ purified DNA was amplified by real-time PCR (Applied Biosystems 7500 Real-Time PCR System) using TaqMan® system for genotyping discrimination. Tag SNPs used in the study met the following two quality control criteria: overall call rate >90%, and Hardy–Weinberg Equilibrium (HWE).
Tissue Microarray (TMA) and Immunohistochemistry
From each of the 101 cases, hematoxylin-eosin slides were prepared. Two different neoplastic areas free from tissue processing artifacts or degeneration, such as necrosis or calcification, were selected for TMA block construction. Each TMA block included eight cases, with two samples from each case; each sample was 3 mm in diameter. The FFPE TMA blocks were cut into sections and placed on microscopic slides, which were submitted to immunohistochemistry following the procedure described by da Silva-Camargo et al. (2018). The TMA slides were subjected to antigen retrieval and incubated with the following monoclonal mouse antibodies: anti-TCTP antibody (1:100, Abcam®, Cambridge, MA, USA), anti-human MTOR antibody (1:100, Cloud®, Katy, TX, USA), anti-MDM2 antibody (1:200, Affinity®, Jiangsu, China), anti-human p53 protein antibody (1:200, Dako®, Glostrup, Denmark) and anti-human p21 protein antibody (1:400, Cloud®, Katy, TX, USA). The reveal HRP detection system from Spring Bioscience® (Fremont, CA, USA), which included the secondary antibody, was used. For visualization, slides were incubated with a diaminobenzidine complex and substrate and counterstained with Harris Hematoxylin. As positive controls, samples of tissues expressing each protein were used. As a negative control, the primary antibody was omitted, as shown in Supplementary Figure 1 (Silva-Camargo et al., 2018).
Morphological Analysis of Protein Expression
The evaluation of protein immunostaining was performed as follows. For quantitative analysis, TCTP and MTOR slides were digitized by Axio Scan.Z1® scanner (Carl Zeiss, Oberkochen, Germany), resulting in 300 to 500 images for each sample. Approximately 50 images from each sample were selected, avoiding necrotic areas or the presence of tissue processing artifacts. Then, these images were analyzed using Image Proplus® software, which performs morphometric analysis by color according to a previously established color reference. The software distinguishes the color brown, which is positive, from other colors and provides the mean tissue positive area in each sample in square micrometers (μm2) (Silva-Camargo et al., 2018). Semiquantitative morphological analyses of TCTP and MTOR, as well as MDM2, p53, and p21 proteins were also carried out. The Allred score, as described for hormone receptors immunostaining evaluation in breast cancer, was used (Allred et al., 1998). The advantage of the Allred score compared to the morphometric analyses was to evaluate the intensity of the immunostaining, as well as its subcellular location.
Statistical Analysis
Nominal variables (sex, number of deaths, survival, and genotype) were expressed by absolute and relative frequency and non-nominal (age at diagnostic and morphometric expression) by means and SD. Genetic distribution analysis was made by logistic regression p-value. HWE was observed when the p-value was not significant (p-value for HWE cut-off = .001). For the immunohistochemical study, Pearson correlation analyses between the proteins were carried out. Data were analyzed using the computer program by IBM® SPSS Statistics v.20.0 software and considered significant p ≤ .05. Haploview 4.2 was used to estimate the HWE.
Results
Polymorphisms in Astrocytomas for all Groups
The distribution of genotypic analysis for all tag SNPs in the four groups (n = 101) can be observed in Table 2.
Genotypic Frequencies Distribution and Analysis Between all WHO Groups.
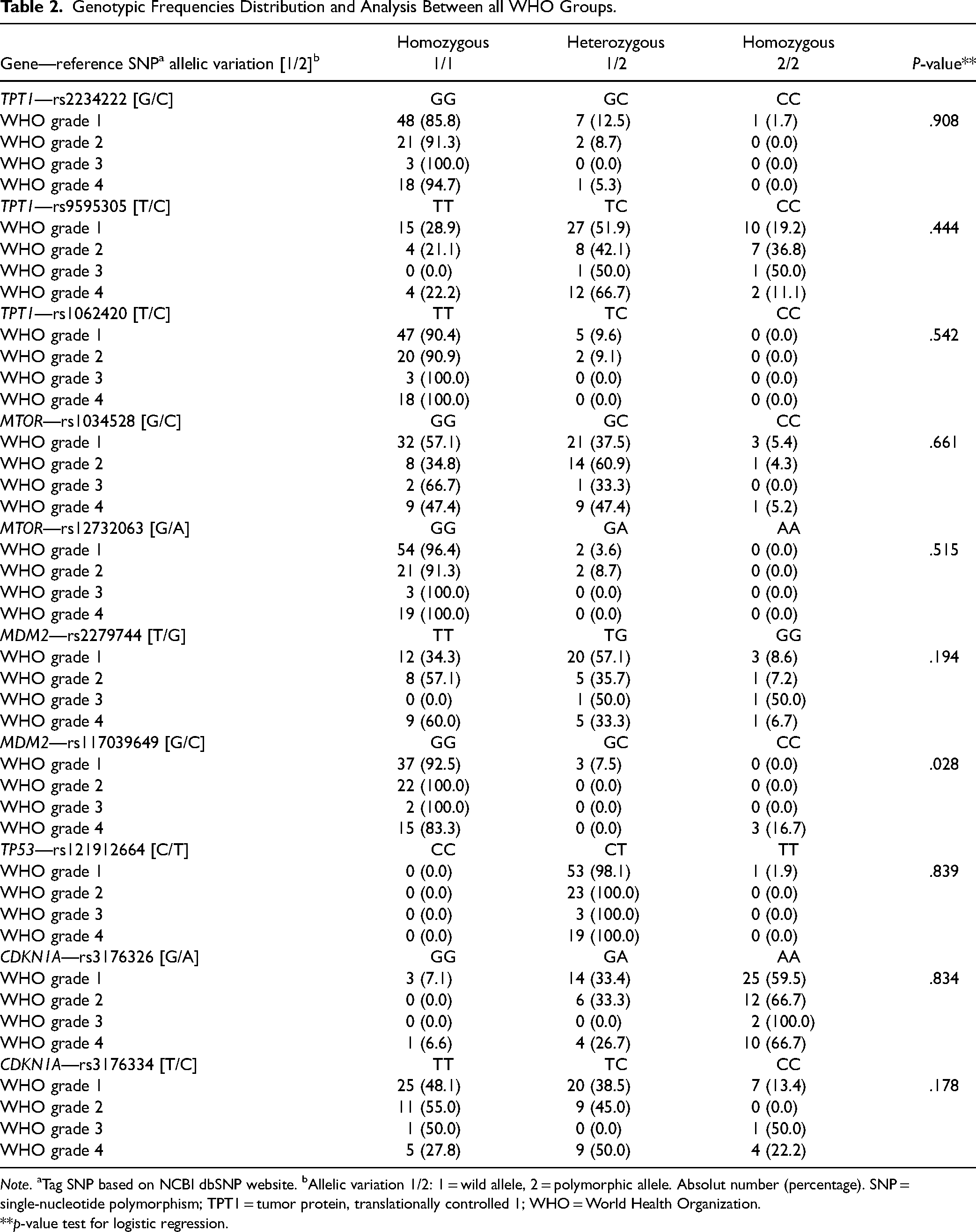
Note. aTag SNP based on NCBI dbSNP website. bAllelic variation 1/2: 1 = wild allele, 2 = polymorphic allele. Absolut number (percentage). SNP = single-nucleotide polymorphism; TPT1 = tumor protein, translationally controlled 1; WHO = World Health Organization.
**p-value test for logistic regression.
MDM2 rs117039649 [G/C] showed homozygosis (CC) for polymorphic allele only in glioblastomas (p = .028). An interesting observation can be made for the SNP of the TP53 gene, despite not having shown statistical difference. TP53 rs121912664 [C/T] showed no homozygosis for wild C allele. Almost all patients were heterozygous (CT), except for one patient homozygous for polymorphic T allele.
Polymorphisms in low-Grade Versus High-Grade Astrocytomas (Grades 1 + 2×grades 3 + 4)
In this analysis, the astrocytoma samples were separated into two groups (n = 101) following this grouping (Table 3): low grade (1 + 2) and high grade (3 + 4). The rs117039649 [G/C] for MDM2 showed statistically significant result (p = .005) like the ones described in Table 2.
Genotypic Frequencies Distribution and Analysis Between Astrocytomas (Grades 1 + 2×Grades 3 + 4).
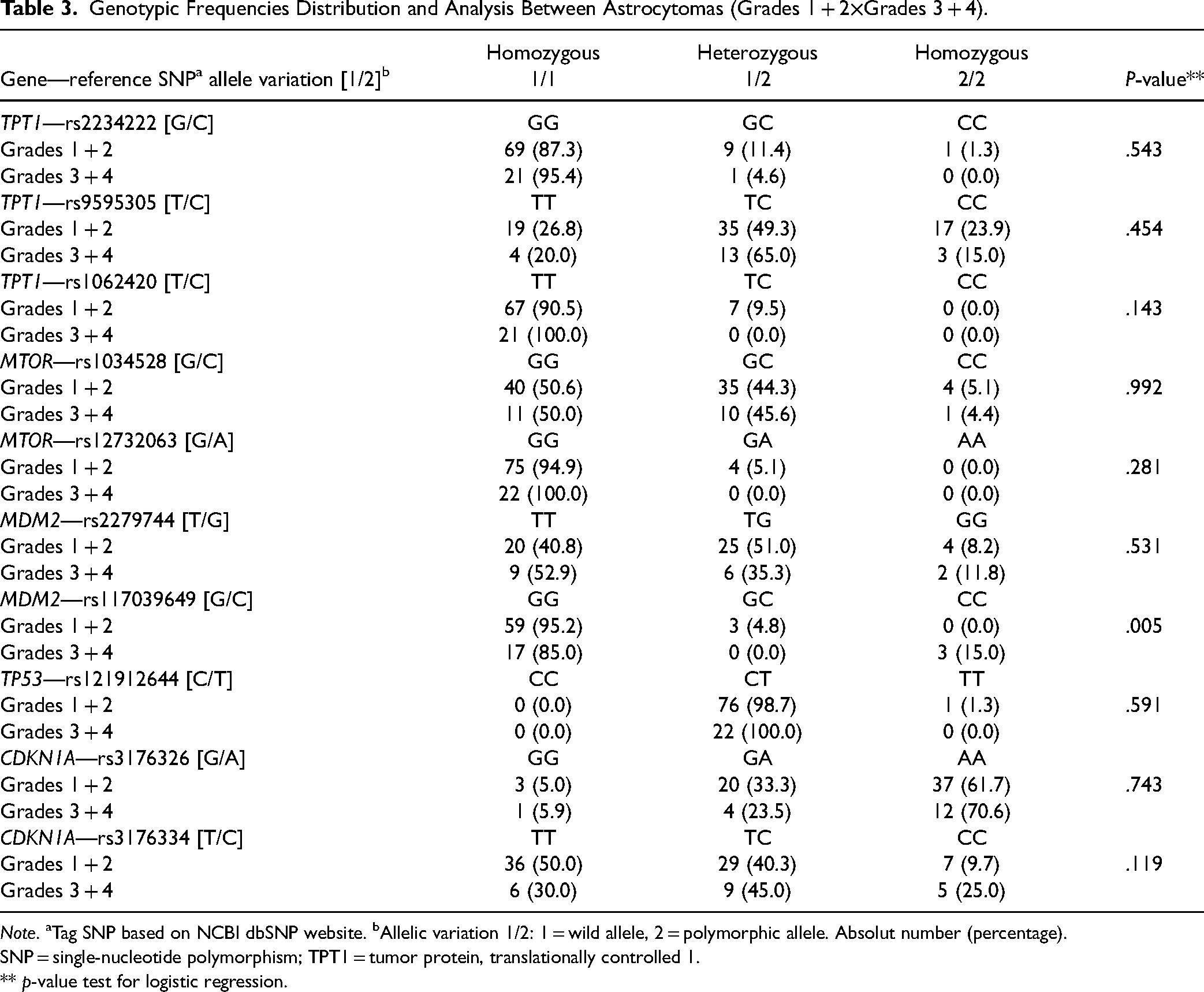
Note. aTag SNP based on NCBI dbSNP website. bAllelic variation 1/2: 1 = wild allele, 2 = polymorphic allele. Absolut number (percentage). SNP = single-nucleotide polymorphism; TPT1 = tumor protein, translationally controlled 1.
** p-value test for logistic regression.
Polymorphisms in Infiltrating Astrocytomas (Grade 2×grades 3 + 4)
We analyzed only infiltrating astrocytomas (n = 45), excluding grade 1 pilocytic astrocytomas, divided into low and high-grade tumors (grade 2×grades 3 and 4). The results are shown in Table 4. In this evaluation, CDKN1A rs3176334 [T/C] had a homozygous (CC) genotype for C polymorphic allele only in high-grade astrocytomas (p = .039). MDM2 rs117039649 [G/C] had borderline (marginal) statistical significance (p = .059).
Genotypic Frequencies Distribution and Analysis Between Astrocytomas (Grades 2 × 3 + 4).
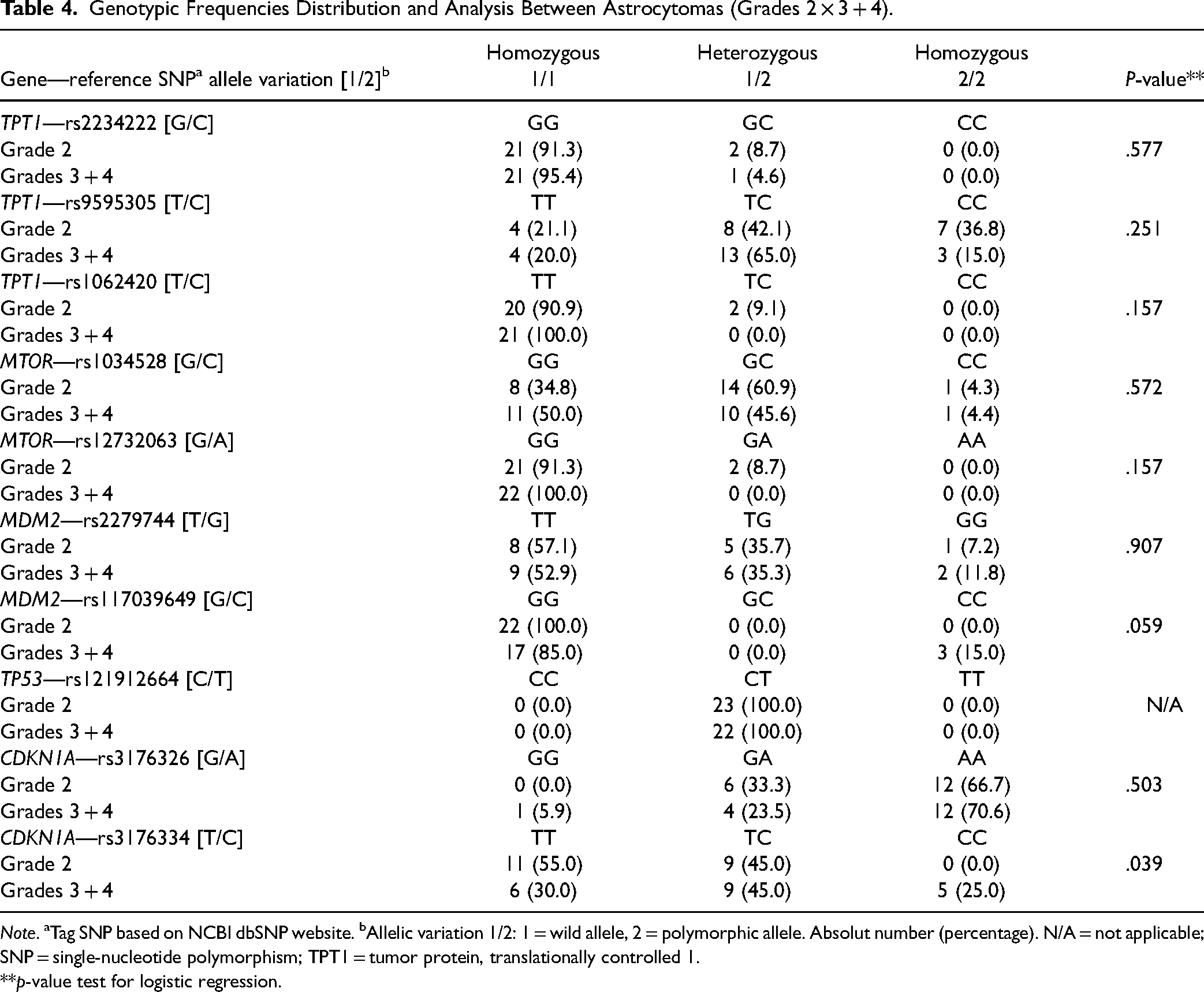
Note. aTag SNP based on NCBI dbSNP website. bAllelic variation 1/2: 1 = wild allele, 2 = polymorphic allele. Absolut number (percentage). N/A = not applicable; SNP = single-nucleotide polymorphism; TPT1 = tumor protein, translationally controlled 1.
**p-value test for logistic regression.
Immunohistochemistry in Astrocytomas
For the immunohistochemistry markers, the results were analyzed considering the four groups but Figure 1 only indicates groups 1 and 4. We found more TCTP morphometric expression in WHO grade 4 group (15,664,0μm2± 6418,9 μm2), but without statistical significance (Supplemental Table S1).
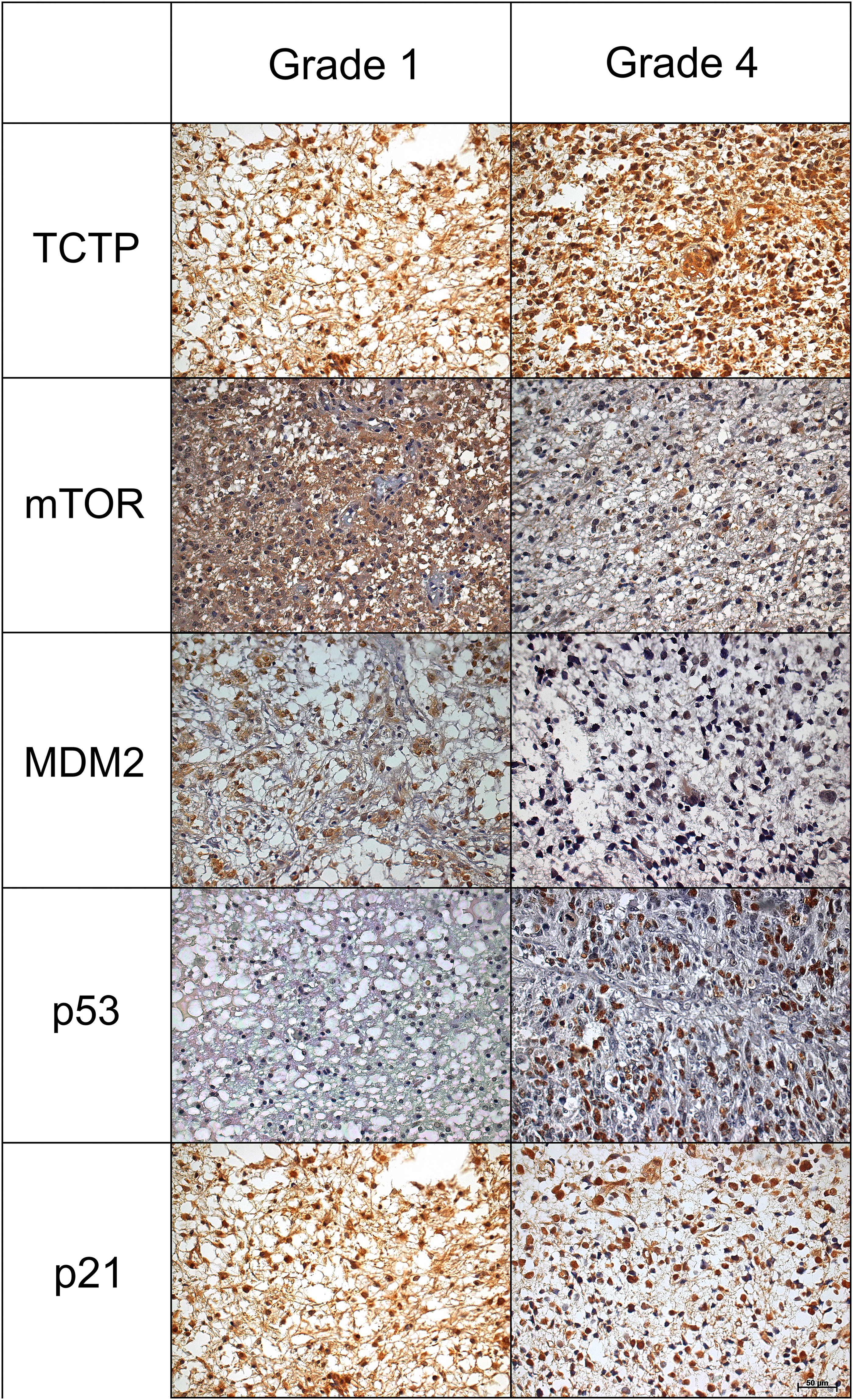
Immunoexpression for the five proteins TCTP, mTOR, MDM2, p53 and p21. Left column shows astrocytomas grade 1 and right column shows astrocytomas grade 4 (IHC, 200x).
Pearson correlation was made between TCTP and other molecules, and when we compared the expression of TCTP and p21 proteins, we have obtained interesting results. In grade 1 astrocytomas, there was a moderate positive Pearson correlation (r = 0.301; p = .025) between nuclear p21 and TCTP expression. As for grade 2 astrocytomas, we found a moderate positive Pearson correlation (r = 0.439; p = .036) between cytoplasmic p21 and TCTP expression.
Association Between Immunohistochemistry and Survival
This analysis was conducted between protein expression and survival of patients with glioblastoma (grade 4) who died. The mean survival of these patients was 8.8 (±4.8) months and, using this value as a cut-off of reference, the patients of group 4 were classified into two groups: below average survival (BAS) and above average survival (AAS). Four proteins had the potential to be analyzed considering Allred score and survival: TCTP, p53, p21, and MDM2 (Figure 2). mTOR protein was not included in this analysis, as their evaluation showed the same Allred score result in both groups, making it impossible to compare them.
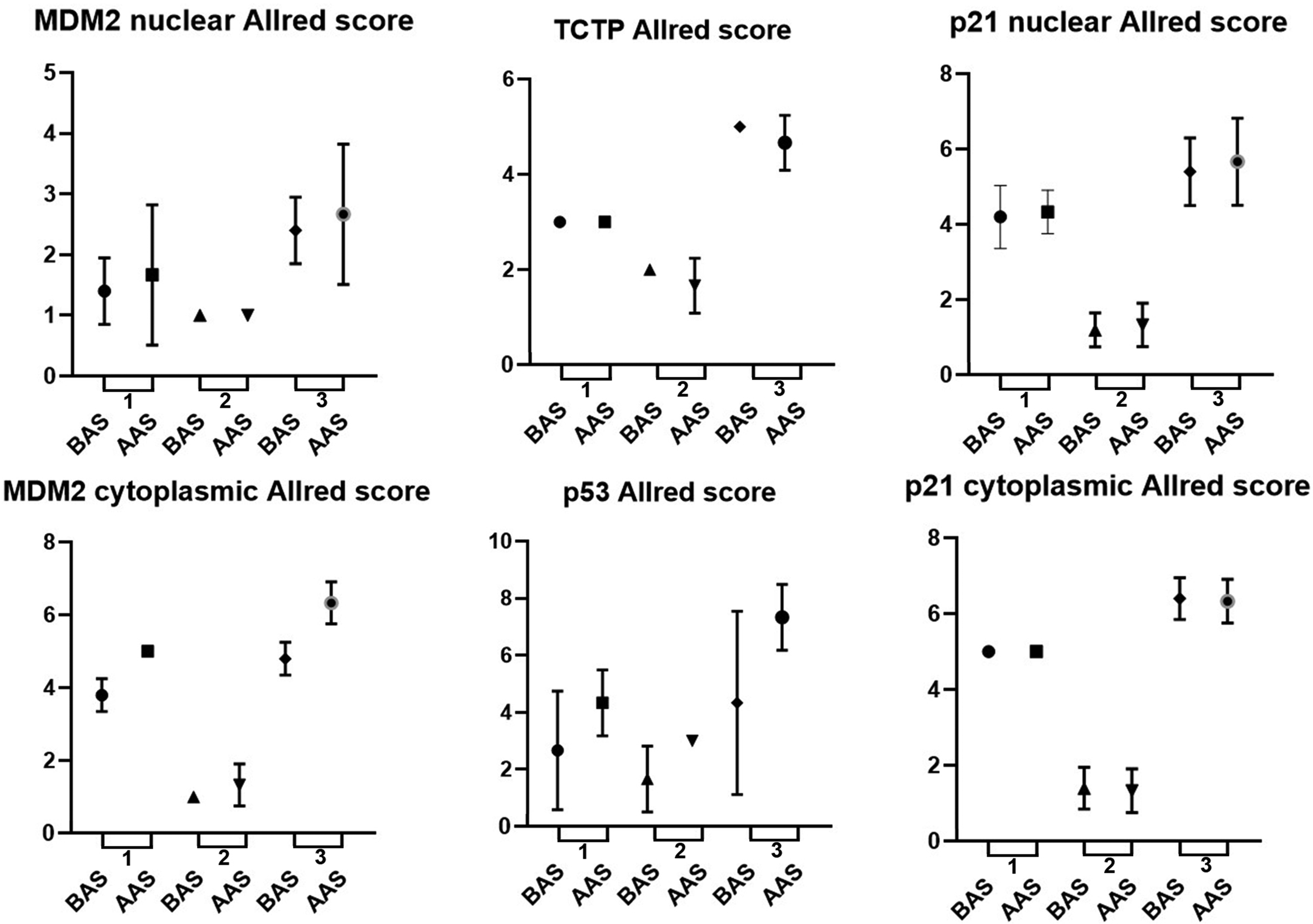
Survival and immunoexpression proteins (TCTP, MDM2, p53 and p21) distribution in glioblastomas (group 4). Survival was classified into two groups considering the mean of survival: below average survival (BAS) and above average survival (AAS). The proteins were evaluated according to the Allred value (distribution, intensity, Allred score). Each graphic has 3 analyses: 1 for distribution in below and above average survival; 2 for intensity in below and above average survival; 3 for Allred score in below and above average survival.
When we compared protein expression and survival rates in patients diagnosed with glioblastomas, considering the Allred score, we found that patients who survived longer had more expression of cytoplasmic MDM2 (p = .018) and p53 (p = .020).
Discussion
Tumors of the central nervous system (CNS) are the most common solid neoplasms in children, with the highest mortality rates and among the histological subtypes, astrocytomas are the most common (Ostrom et al., 2020). They are classified by the WHO in grades 1 to 4, which correlates directly to aggressiveness and poor prognosis. Grade 1 tumors are low-grade circumscribed astrocytomas, whereas grades 2, 3, and 4 are infiltrating astrocytomas (Louis, 2016). They can be subdivided into low-grade astrocytomas (grades 1 and 2) and high-grade astrocytomas (grades 3 and 4). These two groups of astrocytomas present different molecular pathways of gliomagenesis (Ryall et al., 2020). Several proteins have distinct roles in neoplasms, including astrocytomas, acting as either prognostic or therapeutic biomarkers. TCTP is one of these proteins, and through direct or indirect interaction with other proteins such as MTOR, MDM2, p53, and p21, it has been considered an independent prognostic factor in astrocytomas in adults (Gu et al., 2014). Our objective was to analyze SNPs in TPT1 and TCTP expression, as well as associated molecular pathways, in pediatric astrocytomas.
Regarding SNPs in TPT1, we did not observe an association with pediatric astrocytomas. Two genes showed significant results (MDM2 and CDKN1A) in all analyzes performed and all genes are in HWE (Supplemental Table S2) with the exception of MDM2 (rs117039649) and TP53 (rs121912664), and this aspect could be because the samples are from tumors.
MDM2 rs117039649 [G/C] showed homozygosis (CC) for polymorphic allele only in glioblastomas (WHO grade 4), both in isolated and low-grade×high-grade comparisons, which could indicate risk. Furthermore, in the specific analysis of the subgroup of infiltrative astrocytomas, this SNP showed a possible importance despite its marginal significance. MDM2 gene codifies MDM2 protein which inhibits p53 protein, impacting apoptosis (Moll & Petrenko, 2003). Perhaps, this polymorphism could provide a greater escape from apoptosis in glioblastomas. Previous studies have demonstrated an association between this SNP and neoplasms. In gynecological cancer, polymorphic C allele has shown a protective role (Gansmo et al., 2015; Roszak et al., 2015; Zhang et al., 2018). Different mechanisms of carcinogenesis, and different levels of gene expression, between gliomas and female genital tumors could explain this opposite role, compared to our results. When we compare with 1000 genome databases (NCBI, 2022), the frequency of the polymorphic C in this population is 1.3%, much lower than the frequency in our cohort, which was 5.5%. This could point to a risk factor of the polymorphic allele in pediatric astrocytomas.
TP53 rs121912664 [C/T] represents R337H mutation, which has high incidence in the south of Brazil, compared to other populations (Custodio et al., 2011; Ribeiro et al., 2001) perhaps due to the founder effect reported by the authors. We have found that almost every patient, except for one, showed heterozygosity for this SNP. Even though there was no statistical significance in this analysis, this result could indicate an important potential role for T polymorphic allele in pediatric astrocytomas. Furthermore, other studies have shown the impact of SNP in TP53 gene on prognosis of pediatric astrocytomas (Mascelli et al., 2016).
When we compared only infiltrating astrocytomas, we found that CDKN1A rs3176334 [T/C] presented a homozygous (CC) polymorphic genotype only in high-grade astrocytomas, which could also indicate risk. The 1000Genomes databases (NCBI, 2022) shows that the frequency of the polymorphic C in this population is 36.5%, a little higher than the frequency in our cohort, which was 33.7%. We have not found previous studies associating this SNP and gliomas or other neoplasms. But since CDKN1A is responsible for encoding p21 protein, which controls cell cycle, we could speculate that this polymorphism could enhance tumor cell proliferation.
Immunohistochemistry analysis did not show statistically significant results when we compared protein expression to WHO tumor grade. But when we correlated TCTP and other proteins, p21 showed positive Pearson correlation. Interestingly, p21 had nuclear expression in grade 1 astrocytomas, whereas grade 2 astrocytomas showed cytoplasmic expression. p21 protein subcellular location has been linked to different functions. When nuclear, it controls cell cycle; when cytoplasmic, it has been associated with a carcinogenic role (Kreis et al., 2019; Shamloo & Usluer, 2019). This could indicate that in low-grade astrocytomas, interaction between TCTP and p21 proteins is different: in grade 1, p21 protein develops its canonical function whereas in grade 2, it plays an oncogenic role.
Finally, we have found a correlation between cytoplasmic expression of MDM2 protein and better outcome in patients with glioblastomas. MDM2, when in the nucleus, is responsible for degrading the p53 protein and inhibiting apoptosis. But previous studies have observed a better outcome when cytoplasmic expression was assessed, as evidenced by Park et al. in patients with breast cancer (Park et al., 2013). Perhaps, cytoplasmic MDM2 could play a similar role in glioblastomas. The p53 protein also showed direct correlation between nuclear expression and survival. Mutations in TP53 gene indicate high-grade astrocytomas (Louis, 2016). However, the role of p53 protein expression regarding prognosis is still unclear. Studies have shown conflicting results, such as no association and association with poor prognosis (Chaurasia et al., 2016; Drach et al., 1996).
Our study has some limitations that should be highlighted in relation to the preliminary results. Even though the previous edition of WHO classification of tumors of the CNS has been used, the authors are aware that the new nomenclature distinguishes pediatric from adult-type gliomas. It also acknowledges the clinical and molecular differences between pediatric-type diffuse gliomas (Louis et al., 2021). Thus, molecular features, such as BRAF fusions and H3K27 and H3G34 mutations, should be included in future studies seeking a more accurate diagnosis, including morphological and molecular features. Our results mark an early study on pediatric astrocytomas and TPT1 and their associated molecular pathways. The authors suggest carrying out future studies that include larger and ethnically different populations. In addition to seeking to increase the sample number and carry out the classic case–control study design. Even though a new WHO classification of tumors of the CNS has been proposed in 2021, the previous classification was maintained in our study, considering that this was the reference at the time of the cohort's diagnosis. In addition, in order to update it, molecular testing (even though of extreme importance, currently unavailable in our institution), would be necessary.
Conclusions
To date, there are no studies evaluating the role of TPT1 and its molecular associations in pediatric astrocytomas, and in this article, two SNPs (MDM2 rs117039649 [G/C] and CDKN1A rs3176334 [T/C]) showed a suggestive risk association in glioblastomas. As for MDM2 protein expression, cytoplasmic expression indicated better overall survival. The results could suggest that this gene and its protein could become potential prognostic markers in pediatric astrocytomas.
Supplemental Material
sj-docx-1-asn-10.1177_17590914231153481 - Supplemental material for Polymorphisms in TPT1 Pathways in Pediatric Astrocytomas
Supplemental material, sj-docx-1-asn-10.1177_17590914231153481 for Polymorphisms in TPT1 Pathways in Pediatric Astrocytomas by Eduardo Morais de Castro, Leonardo Vinícius Barbosa, Aline Simoneti Fonseca, Seigo Nagashima, Caroline Busatta Vaz de Paula, Rafaela Zeni, Letícia Arianne Panini do Carmo, Luciane R Cavalli, Luiz Fernando Bleggi Torres, Andrea Senff Ribeiro, Lucia de Noronha and Cleber Machado-Souza in ASN Neuro
Supplemental Material
sj-docx-2-asn-10.1177_17590914231153481 - Supplemental material for Polymorphisms in TPT1 Pathways in Pediatric Astrocytomas
Supplemental material, sj-docx-2-asn-10.1177_17590914231153481 for Polymorphisms in TPT1 Pathways in Pediatric Astrocytomas by Eduardo Morais de Castro, Leonardo Vinícius Barbosa, Aline Simoneti Fonseca, Seigo Nagashima, Caroline Busatta Vaz de Paula, Rafaela Zeni, Letícia Arianne Panini do Carmo, Luciane R Cavalli, Luiz Fernando Bleggi Torres, Andrea Senff Ribeiro, Lucia de Noronha and Cleber Machado-Souza in ASN Neuro
Supplemental Material
sj-docx-3-asn-10.1177_17590914231153481 - Supplemental material for Polymorphisms in TPT1 Pathways in Pediatric Astrocytomas
Supplemental material, sj-docx-3-asn-10.1177_17590914231153481 for Polymorphisms in TPT1 Pathways in Pediatric Astrocytomas by Eduardo Morais de Castro, Leonardo Vinícius Barbosa, Aline Simoneti Fonseca, Seigo Nagashima, Caroline Busatta Vaz de Paula, Rafaela Zeni, Letícia Arianne Panini do Carmo, Luciane R Cavalli, Luiz Fernando Bleggi Torres, Andrea Senff Ribeiro, Lucia de Noronha and Cleber Machado-Souza in ASN Neuro
Footnotes
Acknowledgments
The authors are grateful to Pequeno Principe Complex for their support.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
