Abstract
We have previously shown that two anti-cancer drugs, CX-4945 and MS-275, protect and preserve white matter (WM) architecture and improve functional recovery in a model of WM ischemic injury. While both compounds promote recovery, CX-4945 is a selective Casein kinase 2 (CK2) inhibitor and MS-275 is a selective Class I histone deacetylase (HDAC) inhibitor. Alterations in microRNAs (miRNAs) mediate some of the protective actions of these drugs. In this study, we aimed to (1) identify miRNAs expressed in mouse optic nerves (MONs); (2) determine which miRNAs are regulated by oxygen glucose deprivation (OGD); and (3) determine the effects of CX-4945 and MS-275 treatment on miRNA expression. RNA isolated from MONs from control and OGD-treated animals with and without CX-4945 or MS-275 treatment were quantified using NanoString nCounter® miRNA expression profiling. Comparative analysis of experimental groups revealed that 12 miRNAs were expressed at high levels in MONs. OGD upregulated five miRNAs (miR-1959, miR-501-3p, miR-146b, miR-201, and miR-335-3p) and downregulated two miRNAs (miR-1937a and miR-1937b) compared to controls. OGD with CX-4945 upregulated miR-1937a and miR-1937b, and downregulated miR-501-3p, miR-200a, miR-1959, and miR-654-3p compared to OGD alone. OGD with MS-275 upregulated miR-2134, miR-2141, miR-2133, miR-34b-5p, miR-153, miR-487b, miR-376b, and downregulated miR-717, miR-190, miR-27a, miR-1959, miR-200a, miR-501-3p, and miR-200c compared to OGD alone. Interestingly, miR-501-3p and miR-1959 were the only miRNAs upregulated by OGD, and downregulated by OGD plus CX-4945 and MS-275. Therefore, we suggest that protective functions of CX-4945 or MS-275 against WM injury maybe mediated, in part, through miRNA expression.
Introduction
MicroRNAs (miRNAs) are short noncoding RNAs (∼22 nucleotides) that regulate the translation or stability of mRNA targets (Bushati & Cohen, 2007; Carthew & Sontheimer, 2009) and play an important role in regulating the cellular stress response. Deletion of many different miRNAs in mice does not produce a prominent phenotype unless a stressor is present (Mendell & Olson, 2012). For example, hearts of mice with deletion of miR-208 cannot handle pressure overload (Van Rooij et al., 2007), and miR-126 knockout mice have fragile, leaky vessels and impaired angiogenesis in response to injury (Kuhnert et al., 2008; Wang et al., 2008). Thus, miRNA mutant mice may appear to be normal, but exposure to a stressor may uncover severe phenotypic consequences.
In contrast, the impact of miRNAs in cancer cells is readily observed without the addition of stressors. For example, deletion of the miR-17-92 family of miRNA clusters induces apoptosis in lymphoma cells (Mu et al., 2009) and systemic delivery of let-7, miR-26a, miR-34a, or miR-143/145 suppresses tumorigenesis (Kota et al., 2009; Pramanik et al., 2011; Trang et al., 2011). In addition, the delivery of these miRNAs appears to selectively impact tumor cell growth and survival, while sparing normal cells. These findings suggest that cancer cells are more sensitive to the actions of miRNAs. It is possible that cancer cells experience chronic stress induced by DNA damage, aneuploidy, endoplasmic reticulum (ER) stress, protein misfolding/aggregation, metabolic reprogramming (Warburg effect), impaired mitochondrial dynamics, hypoxia (core), increased reactive oxygen species (ROS), and other toxic intermediates (Grandemange et al., 2009; Luo et al., 2009; Yadav et al., 2014). Cancer cells rely strongly on stress-induced signaling pathways to sustain activation of mitogenic and pro-survival signaling arising from oncogene activation or tumor suppressor loss, which distinguishes them from non-transformed cells and provides an opportunity for miRNAs to drive strong and selective phenotypic outputs in this pathologic setting.
Using mouse optic nerves (MONs), our studies have identified altered mitochondrial dynamics and metabolism, increased ER stress and glutamate-mediated excitotoxicity, impaired Ca2+ homeostasis, increased nitric oxide synthase (NOS) activity, and nitric oxide (NO) and ROS production with decreased glutathione levels (Baltan et al., 2011; Bastian et al., 2018; Tekkök et al., 2007) as underlying mechanisms of white matter (WM) injury. The optic nerve is the second cranial nerve and is a purely myelinated central nervous system (CNS) WM tract that preserves oligodendrocytes, astrocytes, microglia, oligodendrocyte precursor cells, axons, and myelin in their native three-dimensional cell-to-cell configuration. We and others have shown that the optic nerve offers several advantages to study the mechanisms of WM injury (Baltan, 2019; Bastian et al., 2020), including (a) MONs are sensitive to the aging process (Baltan et al., 2019; Cavallotti et al., 2002, 2003); (b) tissue isolation does not require extensive surgical interventions, so there is minimal preparation injury (Bastian et al., 2020); (c) the isolated MON is structurally and functionally stable in vitro for at least 18 h (Bastian et al., 2020); (d) there are no neurons, so there are no synapses or synaptic machinery to indirectly contribute to WM injury; (e) axonal function can be monitored by recording evoked compound action potentials (CAPs) (Stys et al., 1990); (f) cellular components can be identified through immunohistochemistry using cell-specific antibodies (Baltan et al., 2013); (g) cell-specific and/or inducible transgenic mice allow live imaging of individual glial cells, axons, or organelles such as mitochondria (Bastian et al., 2019a); (h) release of neurotransmitters such as glutamate can be quantitatively monitored (Baltan et al., 2013); (i) protein levels of interest can be quantified on western blots (Bastian et al., 2018); and (j) intravitreal injections enable a route of delivery of drugs to axons and glial cells (Chiha et al., 2020).
We were the first to report that MS-275, a Class I histone deacetylase (HDAC) inhibitor, promotes functional recovery in a model of WM ischemic injury (Baltan, 2012; Baltan et al., 2011, 2013, 2018; Bastian et al., 2018, 2019a; Murphy et al., 2014) when applied before or after ischemia by attenuating excitotoxicity, reducing oxidative stress, and preserving mitochondrial dynamics (Baltan et al., 2011; Murphy et al., 2014). In addition, we established a role of Casein kinase 2 (CK2) activation and its downstream signaling cascade of CDK-5, and AKT/GSK3β in mediating WM ischemic injury, and in the robust protection conferred by the CK2 inhibitor CX-4945 (Baltan et al., 2018). Interestingly, both the CK2 inhibitor CX-4945 and the HDAC inhibitor MS-275 are well-established FDA-approved anti-cancer drugs (Connolly et al., 2017; Pierre et al., 2011; Pili et al., 2017; Ryan et al., 2005; Zhong et al., 2020). MS-275 (Entinostat) is a potent and orally available inhibitor of HDACs currently in clinical trials for the treatment of several types of cancer (Ryan et al., 2005; Zhong et al., 2020). Similarly, CX-4945 exerts WM protection by inhibiting the PI3K/AKT/GSK3β pathway, a key pathway in multiple vital cellular processes of various cancers. CX-4945 (Silmitasertib), a potent and selective orally bioavailable small molecule inhibitor of CK2 for the treatment of cancer, is representative of a new class of CK2 inhibitors with K(i) values in the low nanomolar range and unprecedented selectivity versus other kinases (Battistutta et al., 2011; Siddiqui-Jain et al., 2010; Son et al., 2013). Furthermore, Silmitasertib is in Phase II clinical trials for the treatment of moderate COVID-19 patients (ClinicalTrials.gov Identifier: NCT04663737). Although these two drugs are equally effective when applied before or after ischemia across age groups, they mediate WM protection acting on entirely different mechanisms (Baltan et al., 2008; Bastian et al., 2018; Grandemange et al., 2009; Luo et al., 2009; Yadav et al., 2014).
In this study, we used a quantifiable miRNA profiling technique using nCounter® miRNA Expression Assay (NanoString Technologies, Seattle, WA) to test the hypothesis that the protective functions of CX-4945 and MS-275 may be mediated through differential regulation of WM miRNAs. We used an integrated bioinformatics analysis of miRNA expression profiles to develop a more comprehensive understanding of pathogenic mechanisms of WM ischemic injury. We report that oxygen glucose deprivation (OGD) altered miRNA expression levels in WM, and that CX-4945 and MS-275 differentially regulated miRNA levels in OGD MONs. We further confirmed miRNA expression across different conditions using in situ hybridization (ISH). To our knowledge, this is the first study that defines miRNAs that play a role in ischemic WM injury, adding a new perspective to our understanding of pathophysiological mechanisms and revealing new therapeutic targets for ischemic WM stroke. Repurposing of CX-4945 and MS-275 for stroke and identification of specific downstream effectors of these cancer drugs can provide further information regarding specific neuroprotection and neuro-restoration acting on more precise therapeutic targets.
Methods
Animals and Chemicals
All experimental procedures were approved by The Institutional Animal Care and Use Committee of the Cleveland Clinic and Oregon Health and Science University and were performed and reported in compliance with ARRIVE guidelines (Animal Research: Reporting In Vivo Experiments). C57BL/6J male mice (3 months of age) were purchased from Jackson Laboratory (Bar Harbor, ME). All mice were maintained on a 12 h light/dark cycle with food and water provided ad libitum. CX-4945 and MS-275 were purchased from Cayman Chemical (Ann Harbor, ME). The sources of other chemicals used were described previously (Baltan et al., 2008; Bastian et al., 2018; Stahon et al., 2016).
Optic Nerve Preparation, Electrophysiological Recordings, and OGD
Following CO2 asphyxiation, MONs were collected from randomly selected C57BL/6J mice at 3 months of age. MONs were gently cleared from the dural sheath and transferred to a Haas-top perfusion chamber (Harvard Apparatus, Holliston, MA) and superfused with artificial cerebrospinal fluid (ACSF) (pH = 7.4; 124 NaCl, 3 KCl, 2 CaCl2, 2 MgCl2, 1.25 NaH2PO4, 23 NaHCO3, and 10 glucose all in mM). MONs were allowed to equilibrate for at least 15 min in the recording chamber in normal ACSF bubbled with a 95% O2/5% CO2 mixture. All electrophysiological recordings were performed at 37°C. Suction electrodes back-filled with ACSF were used for stimulation and for recording CAPs. The stimulation electrode was connected to a stimulus isolation unit (A365R stimulus isolator; WPI, Sarasota, FL) and elicited CAPs every 30 s. Stimulus pulse (30 μs duration) strength was adjusted to evoke the maximum CAP possible, and then increased another 25% for supramaximal stimulation. The recording electrode was connected to an Axoclamp 900A amplifier (Molecular Devices, LCC, Sunnyvale, CA) and the signal was amplified 20 or 50 times, filtered at 3 kHz (SR560; Stanford Research Systems, Sunnyvale, CA), and acquired at 20–30 kHz. During experiments, the supramaximal CAP was elicited every 30 s. OGD was induced as previously described (Baltan et al., 2008, 2010, 2011; Bastian et al., 2018; Murphy et al., 2014; Stahon et al., 2016) by switching to glucose-free ACSF (replaced with equimolar sucrose to maintain osmolarity) and a gas mixture containing 95% N2/5% CO2. OGD was applied for 60 min, followed by restoration of glucose-containing ACSF and O2, and CAPs were recorded for up to 5 h post-OGD treatment. Experiments with CX-4945 and MS-275 were performed using ASCF containing the selected drug that was administered 30 min before, during, and 30 min after OGD. Electrophysiological experiments (Supplemental Figure 1) confirmed our previous findings that both CX-4945 and MS-275 protected and promoted axon function (Baltan et al., 2008, 2010, 2011; Bastian et al., 2018; Murphy et al., 2014; Stahon et al., 2016). All MON samples were collected at the end of the experimental protocol (see Supplemental Figure 1) and were used for the miRNA expression profiling experiments described below (n = 3 independent assays with each having a pool of 2 MONs; 6 MONs per group in total) and miRNA ISH experiments (ISH, n = 6 MONs from 3 mice per group).
miRNA Expression Profiling
For miRNA expression analysis, MONs were collected from C57Bl/6J mice exposed to control ACSF or OGD with or without CX-4945 or MS-275 as described above (n = 3 independent assays with each having a pool of 2 MONs; 6 MONs per group). MONs were collected at the end of experiments and preserved in RNAlater (#R0901-100ML, Millipore Sigma, St. Louis, MO) until RNA extraction. Tissues were homogenized and total RNA containing miRNAs were isolated following the protocol described in the miRNeasy Kit (#217004, Qiagen, Germantown, MD). RNA concentration and purity were determined using a NanoDrop 2000 Spectrophotometer (#ND-2000, Thermo Fisher Scientific, Waltham, MA). Unbiased miRNA profiling was performed using an nCounter® miRNA Expression Panel (mouse v1.5 miRNA assay (#CSO-MMIR15-212, NanoString Technologies) which detects the levels of 577 miRNAs, 33 murine-associated viral miRNAs, 4 mRNA probes, and 23 internal reference controls) and a master kit (#NAA-AKIT-012, NanoString Technologies). The hybridization cartridges were digitized with a GEN2 nCounter® Digital Analyzer instrument, the raw data (RCC files) were generated from the images with the nSolver™ 4.0 Analysis Software (NanoString Technologies), and miRNA levels were analyzed on a NanoString GEN2 analysis system according to the manufacturer's instructions (https://www.nanostring.com/wp-content/uploads/2020/12/Gene_Expression_Data_Analysis_Guidelines.pdf). The nCounter miRNA expression assay is designed to provide an ultra-sensitive (>1 copy per cell), reproducible, and highly multiplexed method for detecting miRNA in total RNA. The NanoString nCounter® technology uses a novel digital color-coded barcode, with each color-coded barcode attached to a single target-specific probe corresponding to each miRNA in the panel.
In Situ Hybridization
ISH was performed using a modified in situ protocol with locked nucleic acid-modified oligonucleotide probes (Exiqon, Denmark) as previously described (Stahon et al., 2016; Tripathi et al., 2019), with a few modifications. Briefly, well-characterized formalin-fixed, paraffin-embedded (FFPE) 7 µM thick sections were de-paraffinized and rehydrated followed by treatment with proteinase K (40 ng) at 37°C/30 min and then fixed in 4% paraformaldehyde. Next, sections were washed with phosphate-buffered saline (PBS) 3 times for 5 min each and then incubated in imidazole buffer, followed by incubation in EDC-Imidazole solution for 90 min at room temperature. Following 3 washes in PBS (3×PBS) for 5 min each, a DIG-labeled probe was hybridized to each section overnight (54°C). The next morning, sections were washed in 0.1× SSC, followed by endogenous peroxidase activity blocking with 3% H2O2/PBS. Sections were then placed in blocking solution (Roche) for 1 h and incubated in α-DIG-POD antibody (Roche) overnight at 4°C. The next morning, sections were washed (PBS/Tris–HCl/Triton-X100 buffer) and incubated with fluorescent-tagged TSA (Perkin Elmer, NEL741E001KT) to label the probe. After washing, sections were then washed in 3×PBS for 5 min, fixed in filtered auto-fluorescent eliminator regent (Millipore, 2160), and subjected to a series of 70% ethanol washes (6×), with a final wash in PBS. Sections were then mounted in prolong gold antifade reagent (Invitrogen, P36930) and micrographed under a fluorescent microscope (Leica DM5500 B).
KEGG Pathway Enrichment Analysis
For Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway annotation enrichment analysis, we used miRNAs that were differentially expressed and detected in OGD alone, OGD + CX-4945, and OGD + MS-275-treated MONs using DIANA-mirPath v.3.0 with DIANA-microT web server v.5.0 and gene union (Paraskevopoulou et al., 2013; Tripathi et al., 2019). MiRNAs were entered into a mouse KEGG analysis using the MIMAT accession number assigned to the NanoString probe, a false discovery rate correction was applied, and p <.01 and microT score of >0.8 were set as the thresholds.
Statistical Analysis
For electrophysiology experiments, each group consisted of six MONs (n = 6) obtained from three different mice (Supplemental Figure 1). Data are presented as mean ± SEM. Differences between three groups were tested using one-way ANOVAs followed by Bonferroni post-test. ** p <.01, *** p <.001 ****p <.0001. For NanoString experiments, each group consisted of three independent assays samples (n = 3, but a total of 6 MONs as each sample contained a pool of 2 MONs). Data were analyzed according to NanoString recommendations (https://www.nanostring.com/wp-content/uploads/2020/12/Gene_Expression_Data_Analysis_Guidelines.pdf. The background threshold was set at >22.4 average count based on the largest calculated (geometric mean) value of the negative control probes for each sample. MiRNAs included in the analysis were then normalized based on a top 100 miRNAs global normalization approach. Briefly, miRNAs were ordered from highest to lowest abundance based upon the average counts. For each sample, the geometric mean of the top 100 miRNAs was calculated. A sample-specific normalization factor was then calculated: normalization factor = (average of all geometric means)/(geometric mean of the sample). For each sample, the miRNA counts were multiplied by the sample's normalization factor. The normalized counts were log2-transformed and analyzed using a one-tailed heteroscedastic Welch's t-test, withp <.05 considered statistically significant. One-tailed tests were selected based upon the assumption that miRNAs will either increase or decrease with drug treatment. The –log10-transformed Welch's p-values and log2 fold change values were used to generate volcano plots, with thresholds set to p <.05 and 1.5-fold change. Non-parametric one-tailed Mann–Whitney U-tests were also calculated, and p-values reported in Supplementary Tables 1–6. Adjusted q-values for both Welch's t-test and the Mann–Whitney U-test were calculated using the Benjamini and Hochberg false discovery rate method, also reported in Supplementary Tables 1–6.
Results
The Most Abundant miRNAs in MONs Were Expressed in WM Astrocytes
MONs were obtained at the end of electrophysiological recordings that confirmed injury by OGD and axon function protection with CX-4945 or MS-275 against OGD (Supplemental Figure 1) as reported previously (Baltan et al., 2008, 2010, 2011; Bastian et al., 2018; Murphy et al., 2014; Stahon et al., 2016; Vlachos et al., 2015). A total of 577 mouse miRNAs and 33 murine-associated viral miRNAs were evaluated by the NanoString mouse miRNA expression analysis (Figure 1A). The array generated count-based expression levels and had intrinsic load and background controls. In MONs exposed to control ACSF, 277 miRNAs (average counts >22.4) (46%) were expressed above background levels. Figure 1A shows the distribution of average counts of all miRNAs that were expressed above background levels, and a representative list with highest expression are presented in Table 1.
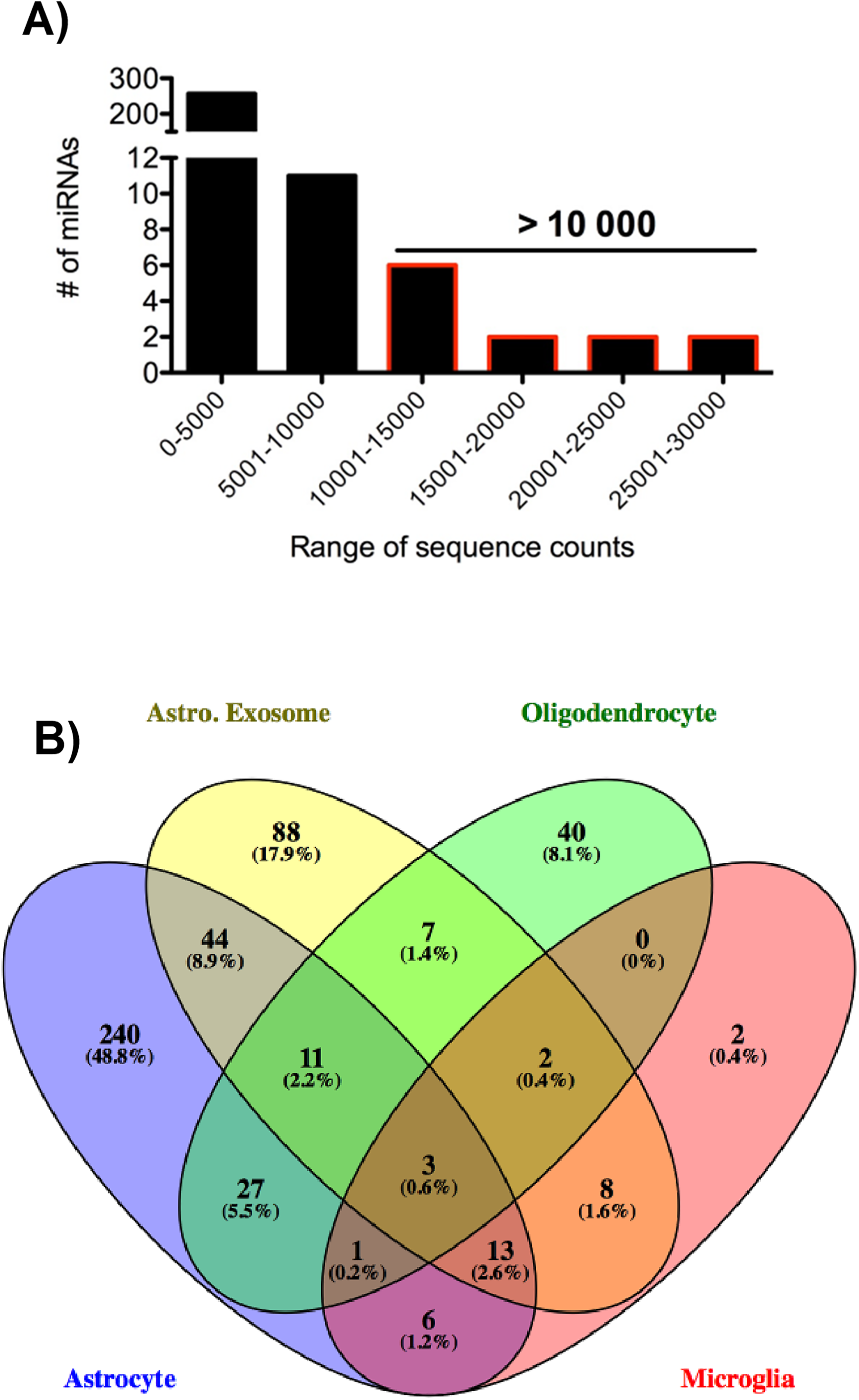
MiRNA expression in MONs. (A) Histograms of the average counts of the 277 miRNAs expressed above background levels in normal MONs. (B) Venn diagram analysis illustrates the cellular distribution of 492 previously identified glial miRNAs using cell-specific miRNA data (Butovsky et al., 2014; Fauré et al., 2006; Galloway & Moore, 2016; Jadhav et al., 2014; Lecca et al., 2016; Rao et al., 2016; Saika et al., 2017; Stary et al., 2016; Tang et al., 2017; Tu et al., 2017; Wang & Cambi, 2012; Wang et al., 2017; Zeng et al., 2017 ). This analysis shows that 240 miRNAs are selective to astrocytes, 88 miRNAs are selective to astrocyte exosomes, 40 miRNAs are selective to oligodendrocytes, and 2 miRNAs are selective to microglia. Note. miRNA = microRNA; MON = mouse optic nerve.
List of Highest-Expressed miRNAs in Mouse Optic Nerves (MONs) Showing Cellular Expression and Associated with a Neuronal Disease/Role/Disease Biomarker.
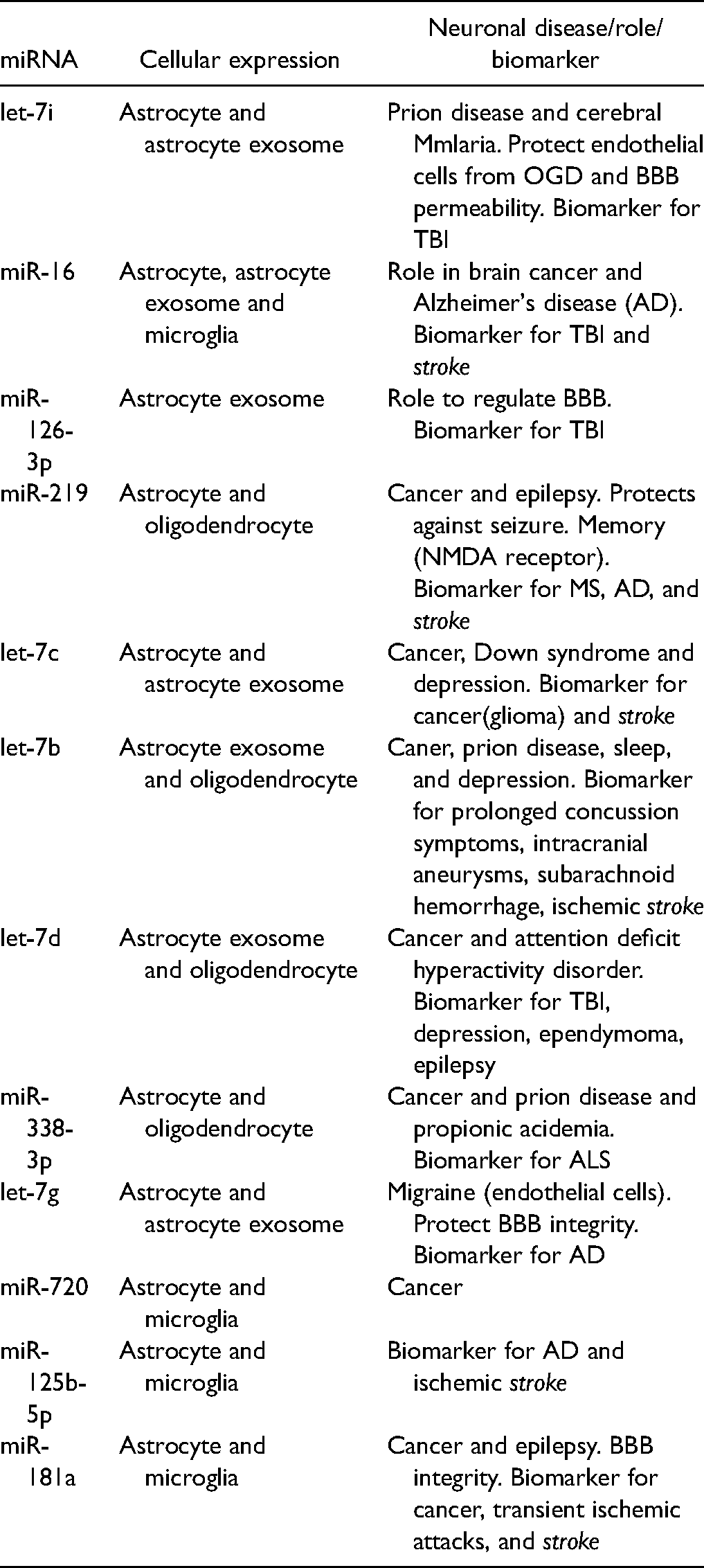
As MON tissue is a heterogeneous mixture of several cell types, we performed an analysis based on previously published literature to classify miRNAs from oligodendrocytes, astrocytes, astrocyte exosomes, and microglia. Venn diagram analysis shows localization of the miRNAs to oligodendrocytes, microglia, astrocytes, and astrocyte exosomes, with a vast majority being selectively expressed in astrocytes (240 miRNAs) and astrocyte exosomes (88 miRNAs). Forty miRNAs were expressed in oligodendrocytes, and only 2 miRNAs were selectively expressed in microglia (Figure 1B). Next, we used this literature-based analysis to identify the cellular localization of miRNAs in our MONs. The 12 miRNAs with greatest abundance in MONs were found to be associated with astrocytes (Table 1). Out of these, seven miRNAs were detected in astrocyte exosomes (58%), 4 were detected in microglia (33%), and 4 were detected in oligodendrocytes (33%) (Table 1). Only four of the most abundant miRNAs (let-7c, let-7i, let-7g, miR-126-3p) were selectively expressed only by astrocytes/astrocyte exosomes. Further data mining revealed that 6 of these 12 astrocytic miRNAs have been previously described to play a role in stroke in gray matter (GM; Table 1, italicized). Further, 8 out of these 12 miRNAs (67%) have also been shown to play a role in diseases in which WM has been shown to be involved in the etiology of the disease (Table 1).
OGD Differentially Modulates miRNA Expression in MONs
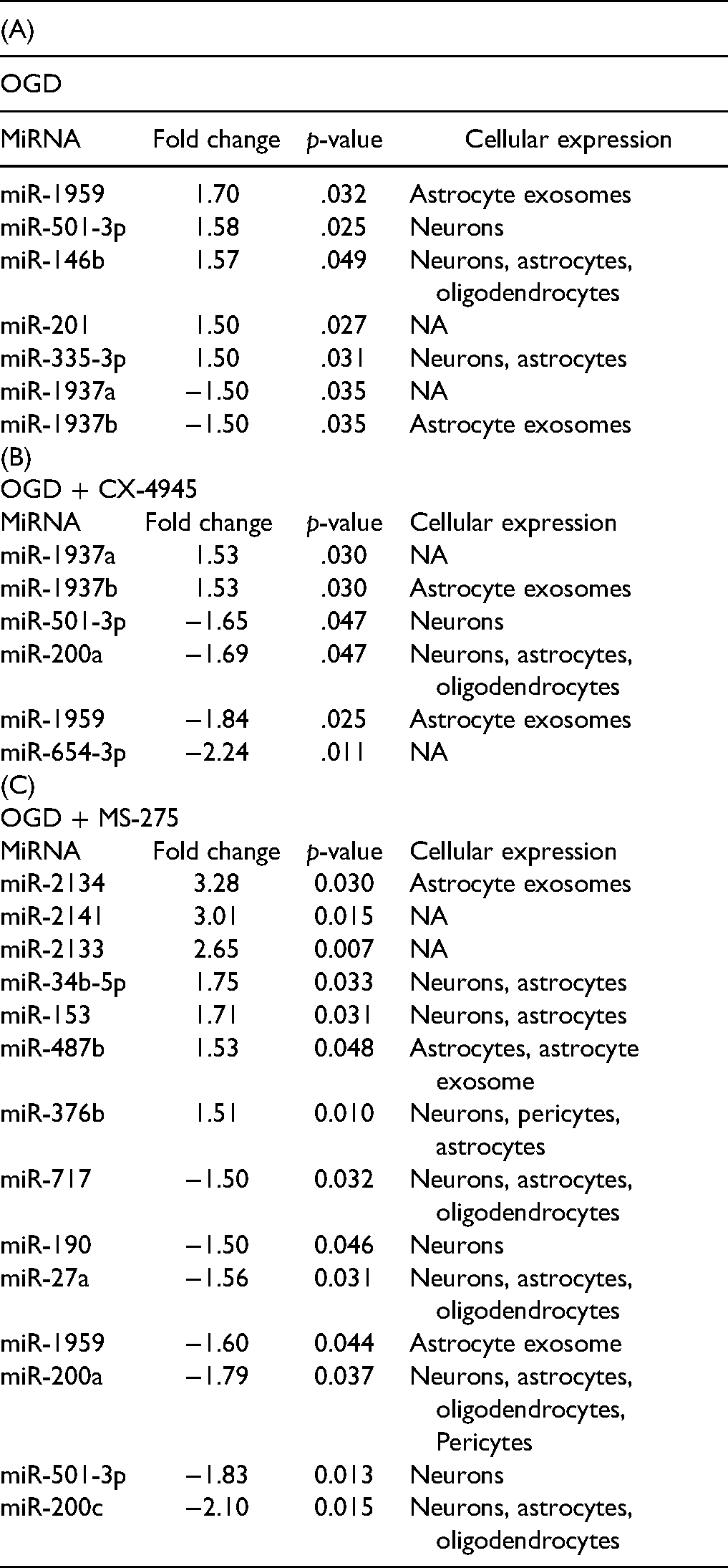
Differential expression analysis comparing control MONs and OGD-exposed MONs revealed that miRNA expression was either decreased (98 miRNAs) or increased (179 miRNAs) in treated MON samples (Figure 2A, Supplemental Table 4). Five miRNAs (miR-501-3p, miR-201, miR-1959, miR-146b, miR-335-3p) were significantly increased by fold change ≥1.5 in MONs exposed to OGD, while two miRNAs (miR-1937a, miR-1937b) were significantly decreased by a ≤−1.5-fold change (Figure 2A, Table 2A). We also assessed the effects of OGD on differential detection of miRNAs in MONs. Two miRNAs that had detectable expression levels in control MONs (above background threshold: >22.4) had undetectable levels in OGD MONs (Supplemental Table 7A), while eight miRNAs changed from undetectable expression levels in control MONs to detectable levels of expression in OGD-treated MONs (Supplemental Table 7A). Together, these results suggest that miRNAs expressed in MONs are differentially regulated by OGD and could contribute to the mechanisms by which ischemia leads to compromised recovery of axonal function.
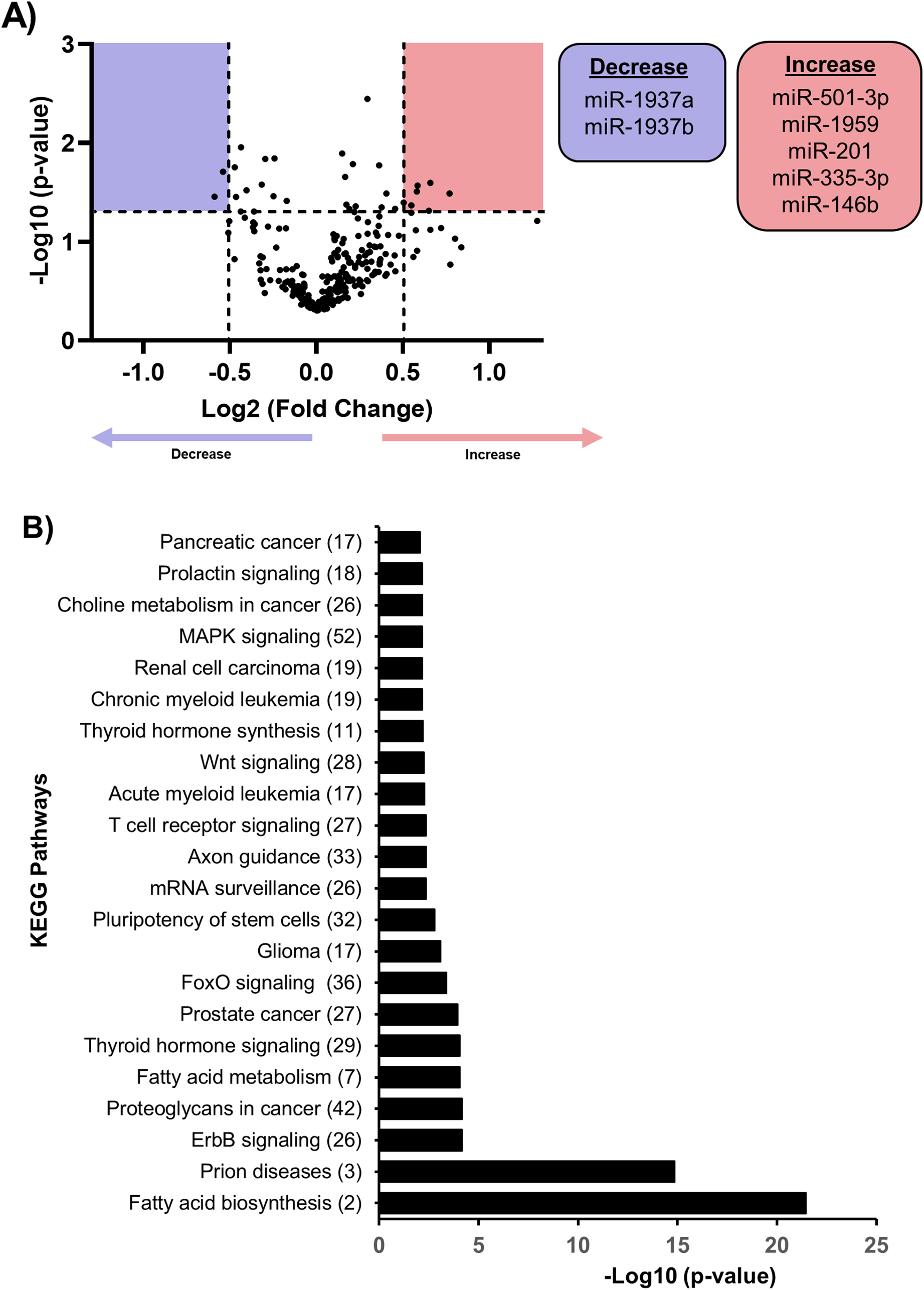
OGD regulated miRNA expression in MONs. (A) Volcano plot of the miRNA array analysis. The vertical dotted lines correspond to 1.5-fold change, and the horizontal dotted line represents a p-value of .05. The black circles in the red and blue areas represent miRNAs with statistically significant increased (list in red box) or decreased expression (list in blue box), respectively. (B) The 22 KEGG pathways were predicted to be targeted by miRNAs differentially regulated in OGD MONs. KEGG pathway analysis of was performed using MirPAth_v3 tool using MicroT-CDS and gene union option. The number of predicted miRNA targets is mentioned in the parenthesis along each category. A false discovery rate correction was included and p<.01 and a MicroT score >0.8 were set as thresholds for KEGG pathway enrichment analysis. Note. OGD = oxygen glucose deprivation; MON = mouse optic nerve; KEGG = Kyoto Encyclopedia of Genes and Genomes.
List of miRNA Shown in Volcano Plots with the Fold Changes and Welch's t-Test. (A) OGD Compared to Control. (B) OGD + CX-4945 Compared to OGD. (C) OGD + MS-275 Compared to OGD. Negative Sign Indicates Decrease.
To identify which signaling pathway(s) are targeted by the differentially expressed miRNAs (Table 2A) or that had undergone a change in detection threshold in OGD MONs (Supplemental Table 7A), we used the DIANA-mirPath v.3 (http://snf-515788.vm.okeanos.grnet.gr/). This analysis identified 22 KEGG pathways that are targets of the differentially regulated miRNAs in OGD MONs (Figure 2B), many of which are relevant to WM injury, including Wnt signaling (Back, 2017; Fancy et al., 2009; Ye et al., 2009), axon guidance (Hinman, 2014), FoxO signaling (Srivastava et al., 2018), fatty acid metabolism and biosynthesis (Gallucci et al., 2019; Jiang et al., 2016), and ErbB signaling (Li et al., 2020; Tang et al., 2017).
CX-4945 Treatment Differentially Regulates miRNA Expression in MONs Following OGD
We have previously shown that treatment with CX-4945 ameliorates the effects of OGD in MONs (Supplemental Figure 1, [Baltan et al., 2018; Bastian et al., 2018]). We therefore compared differential analyses of miRNA expression between MONs exposed to OGD and OGD + CX-4945. This comparative analysis revealed 149 miRNAs that were decreased and 128 miRNAs that were increased following CX-4945 treatment (Supplemental Table 5). Two miRNAs were significantly upregulated (≥1.5-fold, miR-1937a, miR-1937b, Table 2B) in MONs exposed to OGD + CX-4945, and four miRNAs were significantly decreased (≤−1.5-fold, miR-501-3p, miR-200a, miR-1959, miR-654-3p; Table 2B, Figure 3A). For differential detection analysis (above background threshold: >22.4), four miRNAs went from below background levels in OGD MONs to above background levels with OGD + CX-4945 treatment (Supplemental Table 7B). Conversely, five miRNAs went from above background levels with OGD treatment to below background levels in OGD + CX-4945-treated MONs (Supplemental Table 7B).
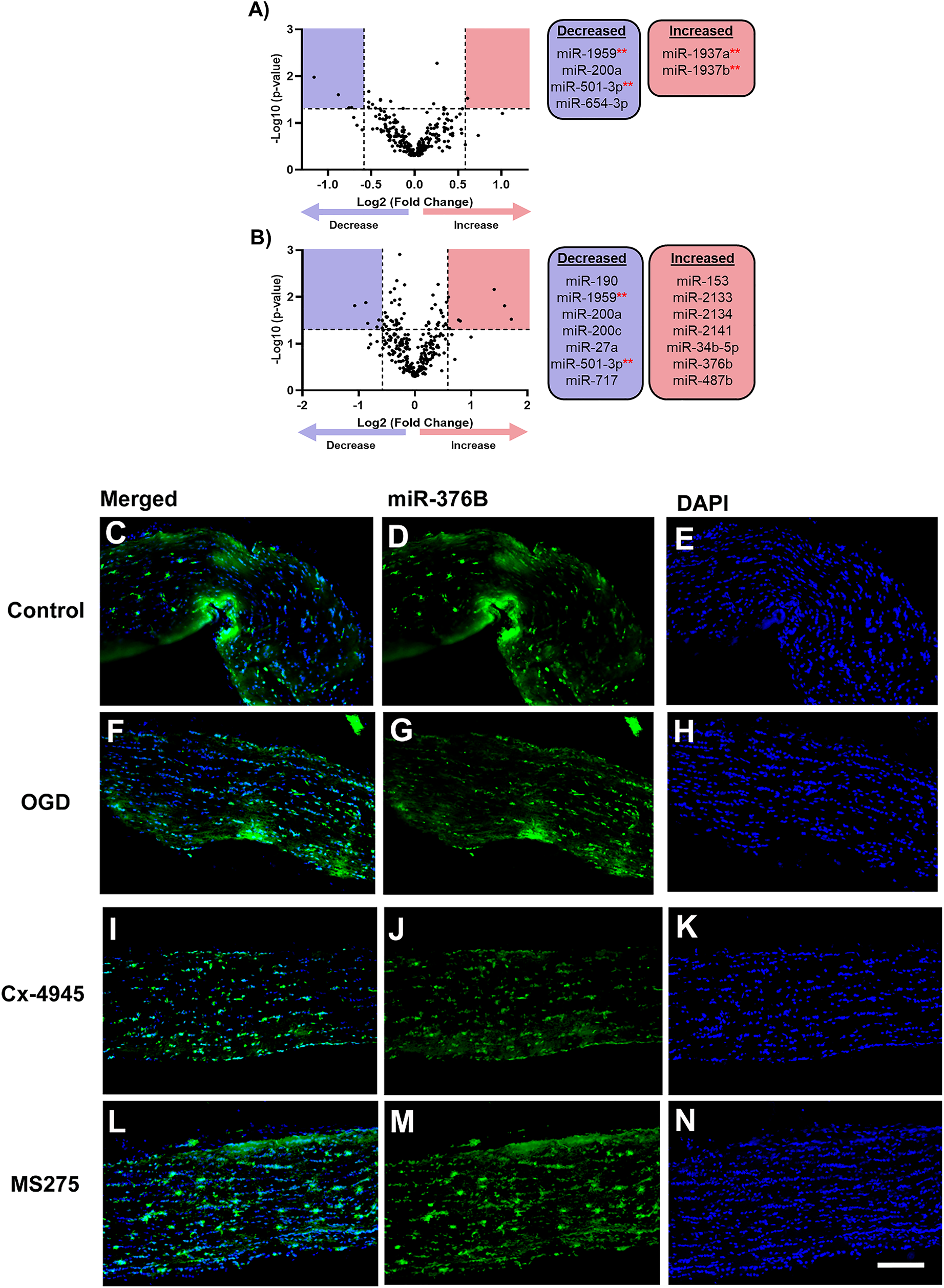
CX-4945 and MS-275 regulate miRNA expression of MONs with OGD. (A) Volcano plot showing miRNA expression analysis of MONs treated with OGD + CX-4945 (B) and OGD + MS-275. The vertical dotted lines correspond to a 1.5-fold change and the horizontal dotted line represents a p-value of .05. Significantly changed miRNAs are selected by the red and blue colored areas and listed in the box along the plot. **miRNAs that were also differentially regulated in OGD MONs (Figure 2A). Note both CX-4945 and MS-275 differentially regulated miR-501-3p and miR-1959 relative to OGD. (C–N) ISH showing expression of miR-376b in control, OGD, OGD + Cx4945 and OGD + MS275 MONs. miR-376b is labeled in Green (Panels C, D, F, G, I, J, L, M) while DAPI (nuclei) is in blue (Panels E, H, K, N). Scale bars = 50 μm. Note. OGD = oxygen glucose deprivation; MON = mouse optic nerve.
MS-275 Treatment Differentially Regulates miRNA Expression in MONs Following OGD
As MS-275 also renders protection following OGD, we performed differential analysis of miRNA expression between MONs exposed to OGD and OGD + MS-275. These results show that expression of 150 miRNAs were decreased and 127 miRNAs were increased following MS-275 treatment (Supplemental Table 6). Among these, seven miRNAs were significantly upregulated (≥1.5-fold, miR-376b, miR-2134, miR-2141, miR-2133, miR-34b-5p, miR-153, miR-487b) in MONs exposed to OGD + MS-275, while seven miRNAs were significantly decreased (≤−1.5-fold, miR-717, miR-190, miR-501-3p, miR-200a, miR-200c, miR-1959, miR-27a; Table 2C, Figure 3B). We selected miR-376b for additional validation (a common target for WM and GM ischemia) using ISH of MONs under conditions of OGD, OGD + CX-4945, and OGD + MS-275 (Figure 3C-N) using probes directed to detect miR-376b. These results showed increased expression of miR-376b following MS-275 treatment (Figure 3L-N) compared to control (Figure 3C-E) as well as OGD (Figure 3F-H). For differential detection analysis (above background threshold: >22.4), three miRNAs went from below background levels in OGD MONs to above background levels with OGD + MS-275 treatment. Conversely, 11 miRNAs went from above background levels with OGD treatment to below background levels in OGD-MS-275-treated MONs (Supplemental Table 7C).
CX-4945 and MS-275 Regulated miRNAs Target Multiple Pathways Relevant to Ischemia
KEGG pathway analysis identified 19 different biological processes predicted to be targets of differentially expressed miRNAs (Figure 3A; Supplemental Table 7B) in OGD + CX-4945 MONs
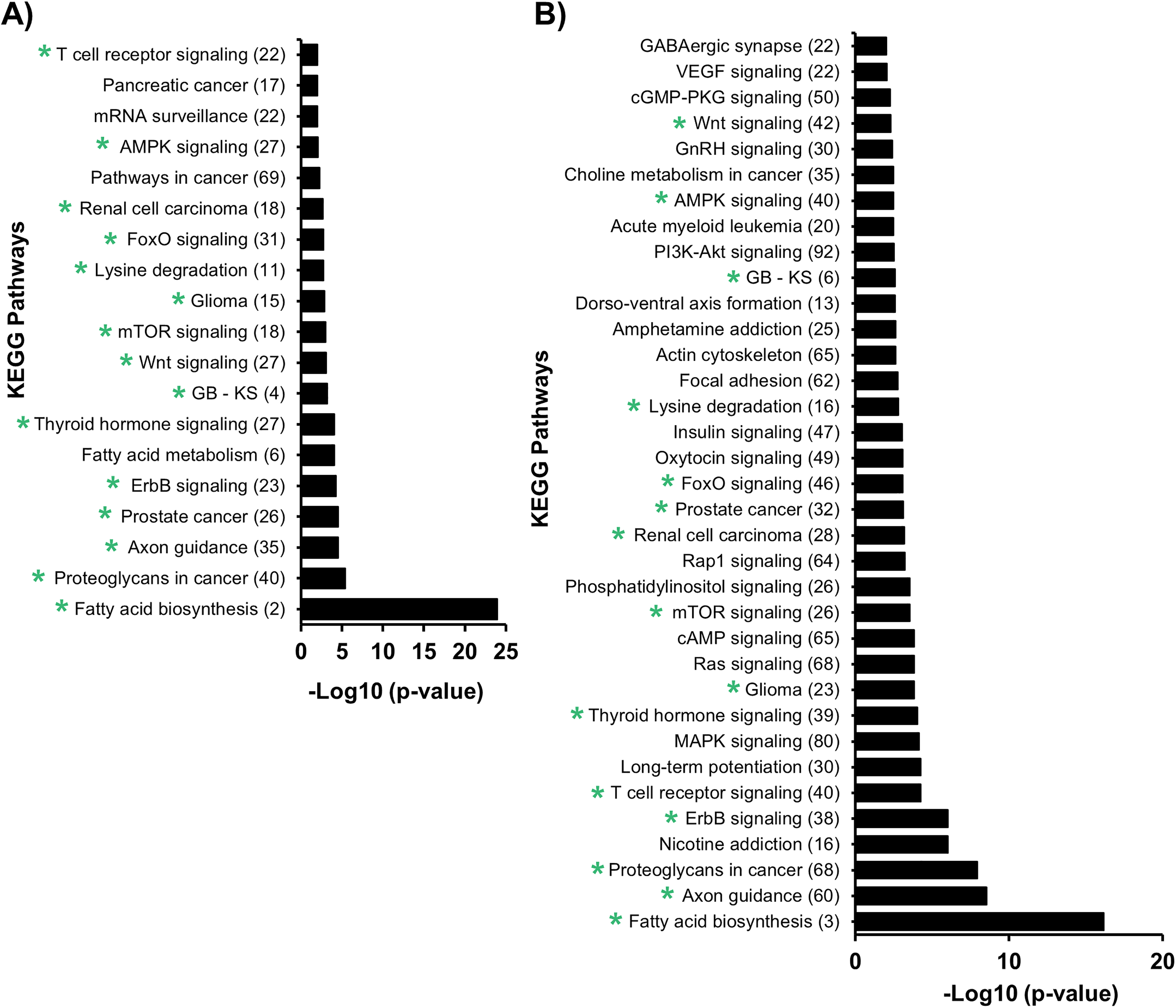
KEGG pathway analysis of miRNAs expressed in ischemic MONs treated with CX-4945 and MS-275. (A) The 19 KEGG pathways predicted to be regulated by miRNAs differentially regulated in OGD + CX-4945 MONs. (B) The 35 KEGG pathways predicted to be regulated by miRNAs differentially regulated in OGD + MS-275 MONs. Green* depicts the 15 KEGG pathways that are predicted to be regulated in both OGD + CX-4945 and OGD + MS-275-treated MONs. Analysis was performed using MirPAth_v3 tool using MicroT-CDS and gene union option. The number of predicted miRNA targets are mentioned in the parenthesis along each category. A false discovery rate correction was included and p<.01 and a MicroT score >.8 were set as thresholds for KEGG pathway enrichment analysis. Glycosaminoglycan biosynthesis–keratan sulfate (GB–KS). Note. KEGG = Kyoto Encyclopedia of Genes and Genomes; miRNA = microRNA; MON = mouse optic nerve; OGD = oxygen glucose deprivation.
Common miRNAs Between OGD, CX-4945, and MS-275 Treatment
While the mode of action of the two above-mentioned drugs differ, their final outcomes on enhancing WM function are similar. We therefore decided to determine the miRNAs that could be in common between the three experimental paradigms. Our results from Venn diagram analysis showed that OGD, OGD + CX-4945, and OGD + MS-275 regulate a series of common miRNAs, which are differentially regulated between OGD and OGD + drug treatments (Figure 5A). CX-4945 treatment relative to OGD differentially regulated miR-1959, miR-501-3p, miR-1937a, and miR-1937b (Table 2B). On the other hand, MS-275 differentially regulated miR-501-3p and miR-1959 compared to OGD (Table 2C). Note that miR-501-3p and miR-1959 were regulated by both CX-4945 and MS-275 (Figure 5A). We next validated the expression of miR-501-3p in MONs (Figure 5B-M), and the results showed increased expression of miR-501-3p in MONs undergoing OGD (Figure 5E-G) compared to controls (Figure 5B-D). Interestingly, treatment with both CX-4945 (Figure 5H-J) and MS-275 (Figure 5K-M) led to decreased levels of miR-501-3p expression. This analysis suggests that these differentially regulated miRNAs between OGD and drug treatments, and particularly miR-501-3p and miR-1959, may be important in the protective mechanisms of CX-4945 and MS-275 in promoting WM recovery after ischemic injury, although a causal role is yet to be established.
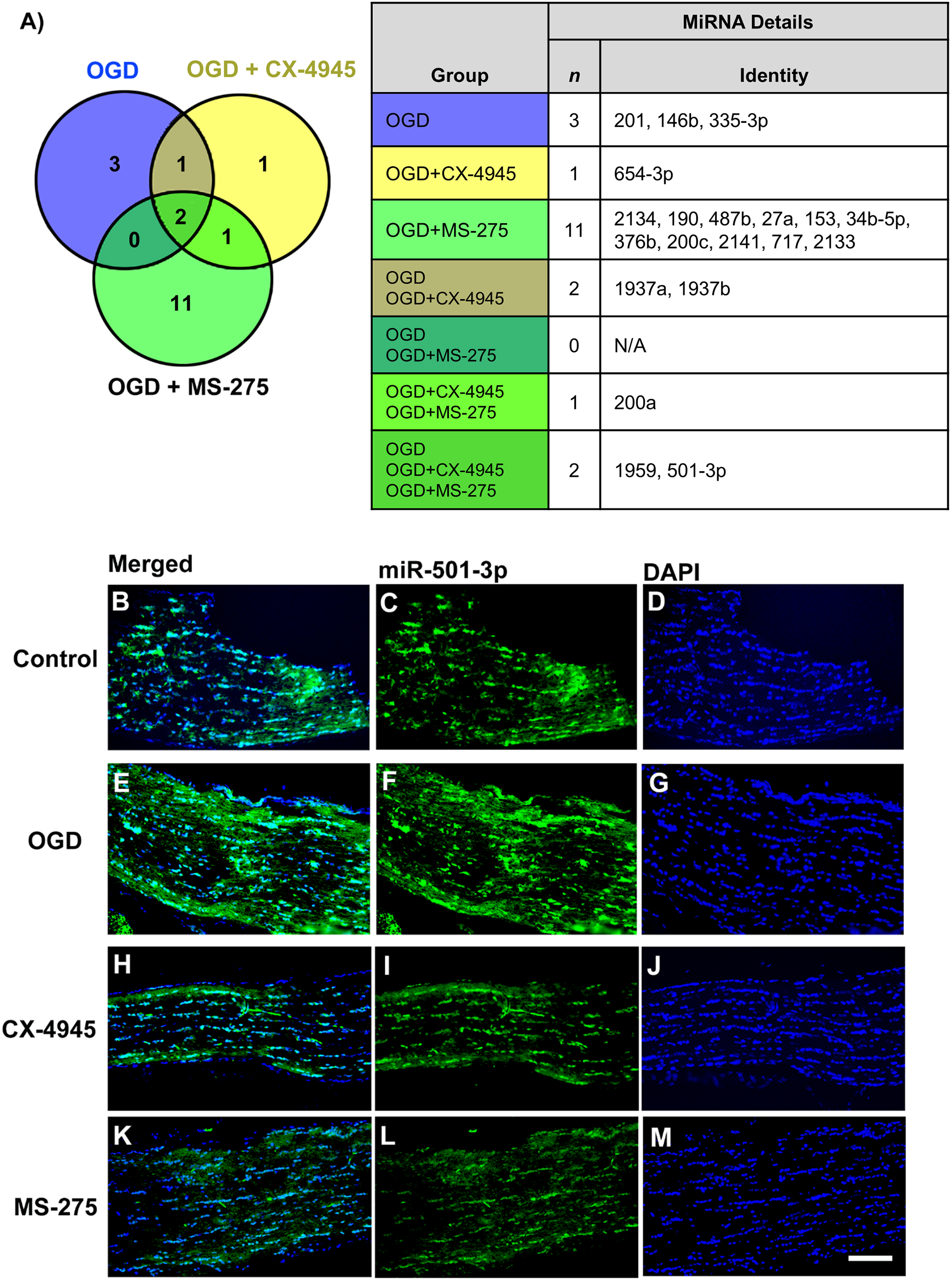
Two miRNAs were regulated by OGD, CX-4945, and MS-275. Venn diagram analysis shows combined miRNA data from OGD, OGD + CX-4945, and OGD + MS-275-treated MONs. Two miRNAs were regulated by all three conditions (miR-501-3p and miR-1959). The common miRNAs showed differential regulation relative to OGD. (B–M) ISH depicting expression of miR-501-3p in control, OGD, OGD + CX-4945, and OGD + MS-275 MONs. MiR-501-3p is labeled in green (Panels B, C, E, F, H, I, K, L) while DAPI (nuclei) is in blue (Panels D, G, J, M). Scale bars = 50 μm. Note. miRNA = microRNA; OGD = oxygen glucose deprivation; MON = mouse optic nerve.
miRNAs Expressed in Astrocytes Are Preferentially Regulated by OGD, CX-4945, or MS-275
Based upon the Venn diagram analyses to determine the expression of miRNAs in WM, based on literature, we determined the cell origin of the differentially expressed miRNAs with OGD, OGD + CX-4945, and OGD + MS-275. With OGD, the majority of the differentially regulated miRNAs were shown to originate from astrocytes and astrocyte exosomes (Table 2A, 57%). One miRNA has documented origin specifically from neurons (here axon in MONs), one from neurons and astrocytes, and one from neurons, astrocytes, and oligodendrocytes. Two of the miRNAs had no identified cellular origin and could not be determined (Table 2A). With CX-4945, the majority of the differentially regulated miRNAs originated from astrocytes or astrocyte exosomes (60%). Interestingly, only one miRNA originated from neurons, and one miRNA was found in neurons, astrocytes, and oligodendrocytes. Two miRNAs could not be identified to a cellular origin (Table 2B). Similar to OGD and OGD + CX-4945, the majority of the differentially regulated miRNAs by OGD + MS-275 were shown to originate from astrocytes and astrocyte exosomes (71%). Three of these 14 miRNAs (miR-2134, miR-487b, and miR-1959) were associated with only astrocytes or astrocyte exosomes. For the remaining, four originated from oligodendrocytes, astrocytes, neurons, and one of which was also from pericytes. There were two miRNAs (miR-190 and miR-501-3p) that were shown to only originate from neurons. The cellular origin of two miRNAs could not be identified (Table 2C). The miRNAs that both CX-4945 and MS-275 reciprocally regulated relative to OGD originated from astrocytic and neuronal origin. Each drug may target distinct additional miRNAs expressed in common glial cells (oligodendrocytes) as well as distinct glial cells, including pericytes (MS-275), to resist WM ischemic injury. For instance, miR-501-3p was shown to regulate AMPA receptor GluA1. Specifically, NMDA receptor GluN2A upregulates miR-501-3p to downregulate the expression of AMPA receptor GluA1 (Hu et al., 2017). These identified signaling pathways and receptors are shown to play major roles in WM function and injury (Baltan et al., 2008; Baltan Tekkök & Goldberg, 2001).
Discussion
WM injury is as important as GM injury in relation to functional recovery after an ischemic stroke in humans (Baltan, 2016; Baltan et al., 2013; Baltan Tekkök & Goldberg, 2001; Bastian et al., 2019a; Tekkök et al., 2007). WM lesion/damage has been found to be related to neurological deficits after a stroke (Sea Lee et al., 2005; Yamada et al., 2003). Because most current widely used stroke models (middle cerebral artery occlusion [MCAO] in rodents) only impact the cortex, the injury mechanisms gleaned from using such approaches would only apply to GM injury. This could be the reason why the majority of drugs developed using this approach have not successfully translated to stroke patients, since both GM and WM are damaged in most ischemic stokes. Recently, miRNAs have been implicated in rodent stroke model (GM) injury mechanisms; however, to date, no reports are available of miRNA profiling in ischemic WM, and the role of miRNAs in WM ischemic injury remains unexplored. Through a combination of a quantifiable miRNA profiling technique, ISH, and in silico KEGG pathway analysis, we have shown for the first time that (1) the most abundant miRNAs in MONs were expressed in astrocytes; (2) OGD differentially regulated miRNA expression in MONs; (3) CX-4945 or MS-275 treatment differentially altered miRNA expression in MONs following OGD; (4) miRNAs regulated by OGD, CX-4945, or MS-275 are predominantly expressed in astrocytes; and (5) OGD, CX-4945, and MS-275 affected some common KEGG signaling pathways. Overall, our results suggest that OGD regulates miRNAs to contribute to WM ischemic injury, and that the anti-cancer drugs CX-4945 and MS-275 modulate miRNAs to attenuate WM ischemic injury.
In this study, we measured miRNAs in a WM tract following ischemia using the NanoString nCounter® miRNA Expression Panel, an unbiased quantifiable profiling technology. Interestingly, miR-146b has been observed in conditions associated with WM injury (Li et al., 2018; Ma et al., 2014) and ischemic conditions like cardiac and renal ischemia (Kálmán et al., 2014). MiR-146b has also been observed in the plasma of ischemic stroke patients, suggesting that WM is injured in stroke clinically (Chen et al., 2018). Overall, our results combined with previous literature suggest that miRNA expression patterns following a stroke could provide signatures for WM and GM injury. These results could lead to the identification of WM injury biomarkers that could be measured in patients’ blood after a stroke. Thus, when combined with GM biomarkers, it would be possible to determine the extent of GM and WM involvement in individual stroke patients, and therefore, therapies could be uniquely tailored. In addition, the same biomarkers could be used to determine the success of therapeutic interventions and subsequently affect decisions to either continue the current treatment or to shift to an alternate therapy.
Our cell-specific miRNA query mapped the majority of altered miRNAs to astrocytes, although the cellular localization of miRNA expression might change with conditions. Astrocytes are increasingly being recognized to play important roles in the pathophysiology of different neurological diseases, and are being more frequently suggested as targets in new therapies (Pekny et al., 2016; Sofroniew & Vinters, 2010). Previously, we have shown that OGD activates astrocytes and reduces GLT-1 expression in WM (Saab et al., 2016). Class I HDAC inhibition protects WM integrity by attenuating astrocyte activation and upregulatingGLT-1 expression, together with conservation of mitochondria and ATP levels such that the subsequent glutamate release is substantially slower and smaller in amount (Baltan et al., 2010). Likewise, OGD increases eNOS activity in astrocytes, leading to axonal mitochondrial dysfunction and oligodendrocyte death in young and aging WM (Bastian et al., 2018). In contrast, astrocytes diligently store glucose as glycogen to derive lactate to support axon function during a metabolic challenge (Baltan, 2015; Bastian et al., 2019b; Brown et al., 2003; Tekkok et al., 2005). Therefore, the changes in astrocyte function through the action of miRNAs following an ischemic episode could dictate the functional status of the astrocyte being beneficial or detrimental for WM. Based upon our above analysis, it is difficult to determine whether astrocyte miRNAs regulated by ischemia are injurious or protective. Interestingly, some shared miRNAs with OGD were differentially regulated by the action of the two drugs studied, CX-4945 and MS-275. For example, CX-4945 differentially regulated miR-1937a, miR-1937b, miR-1959, miR-200a, miR-501-3p, and miR-654-3p, while MS-275 differentially regulated miR-2134, miR-2141, miR-2133, miR-34b-5p, miR-153, miR-487b, miR-376b, miR-717, miR-190, miR-27a, miR-200a, miR-501-3p, and miR-1959. miR-153 plays a role in mitochondrial ROS, and miR-1959 plays a role in the regulation of NF-kB (Zuo et al., 2016). CX-4945 and MS-275 both differentially regulated miR-1959 and miR-501-3p, which potentially implicates these two miRNAs in the protective actions of these two drugs. Interestingly, miR-501-3p was shown to mediate the activity-dependent regulation of AMPA receptor subunit GluA1 expression and was found to be associated with mitochondria (Sripada et al., 2012). Thus, miR-501-3p is an attractive target for further investigation as a potential therapeutic target. Overall, the identification of miRNAs that were differentially regulated by CX-4945 and MS-275, compounds which protect WM from ischemic injury, justifies the search for miRNAs that are important in the protection of WM from ischemic injury. Our prior studies reported that miR-200a and miR-27a levels were significantly upregulated by focal ischemia in both male and female mouse cortex (Lusardi et al., 2014). Here, we found that both CX-4945 and MS-275 decreased miR-200a levels, and that MS-275 decreased miR-27a levels, suggesting a potential mechanism whereby these drugs can act to reverse the expression of miRNAs increased by ischemia.
Our results suggest that each drug treatment regulated miRNAs distinctively from OGD, and in addition to two common miRNAs, miR-501-3p and miR-1959, each drug has its distinct set of miRNAs that may confer protection to WM against ischemic injury. It is of particular importance that miR-487b, miR-376b, and miR-27a, a set of miRNAs regulated by MS-275 in WM, had been previously reported to be regulated in GM ischemic stroke models (Alvarez et al., 2015; Feng et al., 2015; He et al., 2018; Jickling et al., 2014; Ryu et al., 2020). These miRNAs may be common targets to protect both GM and WM structure and function. Among those, miR-376b is regulated with hypoxia in pericytes, which may also target the blood–brain barrier (BBB) and the neurovascular unit (NVU) in addition to cellular protection. Because miRNA regulation is cell-specific and context-dependent, our ongoing research is evaluating whether the miRNAs that were regulated by CX-4945 and MS-275 have shared or merging signaling pathways to benefit WM function. Although the effects of the drugs on control conditions are not the focus of this study, we acknowledge the fact that the anti-cancer drugs most likely will regulate miRNAs in control tissue, and as such warrant further studies to examine this.
As a first pass to identify miRNA protein targets, we have performed an in silico prediction. We have annotated the identified targets into smaller and biologically meaningful groups using KEGG signaling pathways. Our analysis has identified common and specific KEGG signaling pathways between OGD, CX-4945, and MS-275 treatments. The importance of these pathways regulated by both drugs may emerge from their ability to repair and restore glia, myelin, and/or axon function by activating endogenous systems such as the ErbB signaling system, which is often referred to as an “endogenous protector” (Li et al., 2020). Any impairment of ErbB signaling may underlie WM defects by acting on oligodendrocytes, resulting in a defect in myelination and slower nerve transmission in the mouse brain (Carroll et al., 1997) . On the other hand, ErbB receptors are upregulated in neurons and macrophages/microglia following ischemic stroke and have been shown to be involved in neuroprotection and repair (Xu & Ford, 2005). Similarly, Wnt signaling is one of the most important pathways potentially targeting neurogenesis after cerebral ischemia, and various therapeutic approaches are being explored at preclinical stages to target endogenous neurogenesis induced by Wnt signaling after stroke onset (Qiu et al., 2019; Yu et al., 2018). The neuroprotective effect of mTOR inhibition on cerebral ischemia is another important protection step that acts on the cellular energy state during the ischemic period (Foster & Fingar, 2010; Koh, 2010; Koh et al., 2008; Sofer et al., 2005). Targeting axon guidance pathways is crucial in WM function, given that recovery is dependent upon axonal sprouting and reconnection (Hinman, 2014). Ischemia causes a major shift in WM myelin lipid composition, favoring the upregulated expression of diverse phospholipids and sphingolipids that are needed to support neuronal membrane, synaptic, metabolic, and cell signaling functions (de la Monte et al., 2020). Moreover, activation of fatty acid biosynthesis reduces lactoacidosis, supporting oxidation reactions including anaerobic glycolysis (Brose et al., 2016). Therefore, MS-275 and CX-4945 acting on these signaling pathways may at least in part mediate their protective roles in WM injury. Because we cannot determine whether the KEGG pathways that are in common or distinct are protective or injurious to WM ischemic injury due to the inconsistencies of in silico miRNA protein target prediction (Witkos et al., 2011), these results will need to be further validated experimentally.
In addition to success with cancer patients, accumulating evidence predicts long-term benefits of HDAC inhibition following acute CNS injury and neurodegenerative conditions (Denk et al., 2013; Formisano et al., 2015; Hahnen et al., 2006; Murphy et al., 2014; Pinho et al., 2016). Distinct effects of HDAC inhibitors may be attributed to their specific pattern of distribution among neurons and glial cells, and their ability to employ both transcriptional and non-transcriptional actions in axons and glial cell processes.
Overall, we have shown for the first time that miRNAs are regulated by OGD, and that the anti-cancer drugs CX-4945 and MS-275 alter miRNA profiles in WM. Many of the miRNAs that were regulated by OGD, CX-4945, and MS-275 were expressed by astrocytes and oligodendrocytes. We suggest that a combined WM and GM miRNA signature for ischemia could be used to identify the involvement of WM and GM injury following a stroke in individual patients. MiRNAs are readily detected in body fluids, therefore raising the interesting prospect of using these miRNAs as biomarkers for GM and WM injury as well as therapeutic interventions. Because the infarct following a stroke evolves, stroke patients could be monitored longitudinally, and the therapeutic intervention could be modified during the course of this disease. Also, because the drug treatment would have a distinctive miRNA signature, this signature could be used to determine if the therapeutic intervention is effective. In this study, we have also identified a number of miRNAs that were differentially regulated by CX-4945 and MS-275 compared to ischemia, which can help to focus the search for mechanisms of WM injury and to identify therapeutic targets to alleviate the burden of ischemic stroke that will ultimately improve the functional recovery of stroke patients. Note that ischemia and both drugs also regulated a group of murine-associated viral miRNAs (Supplemental Table 7D) which emerge as an interesting point in the light of CX-4945 being tested in clinical trials for Coronavirus disease in Covid19 patients (ClinicalTrials.gov Identifier: NCT04663737). Finally, our results may provide therapeutic targets for other diseases involving WM such as dementia, Alzheimer's disease, multiple sclerosis, periventricular leukomalacia, and Parkinson's disease.
Supplemental Material
sj-TIF-1-asn-10.1177_17590914211042220 - Supplemental material for Identification of miRNAs That Mediate Protective Functions of Anti-Cancer Drugs During White Matter Ischemic Injury
Supplemental material, sj-TIF-1-asn-10.1177_17590914211042220 for Identification of miRNAs That Mediate Protective Functions of Anti-Cancer Drugs During White Matter Ischemic Injury by Selva Baltan, Ursula S. Sandau, Sylvain Brunet, Chinthasagar Bastian, Ajai Tripathi, Hung Nguyen, Helen Liu, Julie A. Saugstad, Yalda Zarnegarnia and Ranjan Dutta in ASN Neuro
Supplemental Material
sj-docx-2-asn-10.1177_17590914211042220 - Supplemental material for Identification of miRNAs That Mediate Protective Functions of Anti-Cancer Drugs During White Matter Ischemic Injury
Supplemental material, sj-docx-2-asn-10.1177_17590914211042220 for Identification of miRNAs That Mediate Protective Functions of Anti-Cancer Drugs During White Matter Ischemic Injury by Selva Baltan, Ursula S. Sandau, Sylvain Brunet, Chinthasagar Bastian, Ajai Tripathi, Hung Nguyen, Helen Liu, Julie A. Saugstad, Yalda Zarnegarnia and Ranjan Dutta in ASN Neuro
Supplemental Material
sj-pdf-3-asn-10.1177_17590914211042220 - Supplemental material for Identification of miRNAs That Mediate Protective Functions of Anti-Cancer Drugs During White Matter Ischemic Injury
Supplemental material, sj-pdf-3-asn-10.1177_17590914211042220 for Identification of miRNAs That Mediate Protective Functions of Anti-Cancer Drugs During White Matter Ischemic Injury by Selva Baltan, Ursula S. Sandau, Sylvain Brunet, Chinthasagar Bastian, Ajai Tripathi, Hung Nguyen, Helen Liu, Julie A. Saugstad, Yalda Zarnegarnia and Ranjan Dutta in ASN Neuro
Footnotes
The authors thank Drs. C. Nelson for editorial assistance and R. Fairchild for access to Nano-string GEN2. This manuscript is dedicated to Dr. Sylvain Brunet (1968–2018).
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the National Institute of Aging (grant number AG033720, NS094881).
Supplemental material
Supplemental material for this article is available online.
