Abstract
Metabolic adaptations in the brain are critical to the establishment and maintenance of normal cellular functions and to the pathological responses to disease processes. Here, we have focused on specific metabolic pathways that are involved in immune-mediated neuronal processes in brain using isolated neurons derived from human autopsy brain sections of normal individuals and individuals diagnosed as Alzheimer’s disease (AD). Laser capture microscopy was used to select specific cell types in immune-stained thin brain sections followed by NanoString technology to identify and quantify differences in mRNA levels between age-matched control and AD neuronal samples. Comparisons were also made between neurons isolated from AD brain sections expressing pathogenic hyperphosphorylated AT8- positive (AT8+) tau and non-AT8+ AD neurons using double labeling techniques. The mRNA expression data showed unique patterns of metabolic pathway expression between the subtypes of captured neurons that involved membrane based solute transporters, redox factors, and arginine and methionine metabolic pathways. We also identified the expression levels of a novel metabolic gene, Radical-S-Adenosyl Domain1 (RSAD1) and its corresponding protein, Rsad1, that impact methionine usage and radical based reactions. Immunohistochemistry was used to identify specific protein expression levels and their cellular location in NeuN+ and AT8+ neurons. APOE4 vs APOE3 genotype-specific and sex-specific gene expression differences in these metabolic pathways were also observed when comparing neurons from individuals with AD to age-matched individuals.
Keywords
Genomic, metabolomic, and proteomic studies have provided important information on changes in the expression levels of genes, metabolites and proteins found in brains of humans with Alzheimer’s disease (AD) (Mihaescu et al., 2010; Kaddurah-Daouk et al., 2011, 2013; Darst et al., 2019; Huo et al., 2020; Mahajan et al., 2020). These techniques have revealed central components underlying the disease process. However, information on direct changes in critical cell-specific processes in AD has been lacking. Here, we have used immunohistochemically identified cell-types collected directly from brain sections of autopsied human patients diagnosed with AD using laser capture microscopy (LCM). This technique has been commonly used to study regional differences in brains from Down Syndrome (DS) patients and in studies of differences between AD brain and brains from normal non-diseased individuals (Alldred et al., 2015; Hondius et al., 2018). We have focused on differential gene expression between neurons that show hyperphosphorylated tau (AT8+) pathology and neurons from cognitively normal individuals. Equally importantly, we have compared AT8+ neurons that show hyperphosphorylated tau pathology to non-AT8+ neurons that do not demonstrate observable hyperphosphorylated tau pathology in humans diagnosed as AD. The genes studied were specifically chosen to reflect key interacting pathological processes associated with AD including immune-regulated metabolism, circadian rhythms and tau pathology. This cell- specific data has provided clues to the re-direction of metabolism that occurs in AD neurons.
Materials and Methods
Human Autopsy Samples
Frozen, de-identified human frontal cortex tissues were obtained from the Duke Kathleen Price Bryan Brain Bank and Biorepository (Duke BBB) at Duke University Medical Center and were kept frozen at −80°C until used. Information on age, sex, APOE genotype, and known behavioral and/or pathological diagnosis for each autopsy sample in the available pool was provided by the brain bank (Online Appendix Table 1).
Human Autopsy Sample RNA Quality Control
Prior to LCM, 50 mg of tissue from individual samples was assessed for RNA quality. Tissue samples were homogenized by sonication and RNA was extracted using the Zymo Research Direct-zol RNA mini-Prep Kit. All procedures were carried out under RNase-free conditions on ice and done as rapidly as possible. RNA quality was analyzed using the Bioanalyzer Pico Chip in a 2100 Agilent Bioanalyzer to determine fragment lengths. Samples with an adequate percentage of fragments between 50-300 nucleotides long were then chosen to undergo Nanostring mRNA analysis. We also obtained a general RNA Integrity Number (RIN) using the Bioanalyzer Pico Chip in a 2100 Agilent Bioanalyzer for each sample. However, RIN numbers were not used as an exclusive requirement for section inclusion in the Nanostring study. The nCounter platform provides high sensitivity detection for input levels of RNA in samples with low RIN numbers as discussed by Veldeman-Jones (Veldman-Jones et al., 2015). Nanostring technology permits robust detection sensitivity for low input RNA levels and does not require an amplification step. The RNA quality was analyzed and RNA Integrity Number (RIN) for each sample was calculated. Samples with an adequate percentage of fragments between 50-300 nucleotides long and a RIN greater than 2 were chosen to undergo Nanostring mRNA analysis. In addition, to limit decay and further loss of tissue slice RNA integrity during the LCM procedure, we collected no more than 400 cells per tissue sample. All procedures were carried out under RNase-free conditions on ice and done as rapidly as possible.
LCM: Preparation of Membrane Slides and Brain Sectioning
Nuclease-free membrane-coated slides (Carl Zeiss, NF 1.0 PEN, 415190-9081-000) were prepared for each sample by placing each slide in an individual airtight, RNase-free 50 mL tube. The tubes containing the slides were then placed under UV light at 254 nm for 30 minutes in a Laminar Flow hood to help overcome the hydrophobic nature of the membrane and aid in better adhesion of the tissue sections on the slides. Slides were stored as RNase-free until sample collection.
Frozen autopsied brain samples (frontal cortex) were sliced at a thickness of 8 microns using a Leica CM 1950 cryostat at -20˚C. Once sectioned, the tissue was placed immediately on the nuclease-free membrane-coated slides labeled with the sample number and AD status and kept on dry ice until staining. The brain sections then underwent immunohistochemical staining to visualize the cells of interest. All IHC staining for LCM analysis was performed on ice and all solutions used were RNase- and DNase-free.
LCM: Rapid IHC Staining
Prior to staining, the tissue sections were ethanol fixed onto the slides. For AD samples, a biotinylated PHF Tau AT8 antibody (ThermoScientific, MN1020B) was used to identify hyperphosphorylated tau-containing neurons in the tissue sections. For “non-AT8+ neurons” in AD brain sections or for neurons found in cognitively normal brain sections lacking AT8+ immunoreactivity, a mouse anti-NeuN primary antibody (Sigma-Aldrich MAB337) followed by a Biotinylated Goat-anti-Mouse IgG Fab2 secondary antibody (Vector BA-9200) was used to identify this type of neuron. The Avidin-Biotin complex method used the Vector Labs ABC kit (pk-6100). Visualization of staining was developed by using either- a) DAB plus nickel (Vector sk-4100) or b) NovaRed (Vector sk-4805). In select AD samples, the sections were double labeled and stained with antibodies for both AT8 and NeuN+ neurons in order to visualize both types of neurons in the same section of tissue. AT8 neurons were visualized with DAB plus nickel, then, normal neurons were visualized with NovaRed, applied sequentially, thus allowing distinction between the neuronal subtypes (Figure 1). Colorimetric development was stopped by washing with RNase DNase free water. (Antibody information including RRID numbers is found in Online Appendix Table 2).
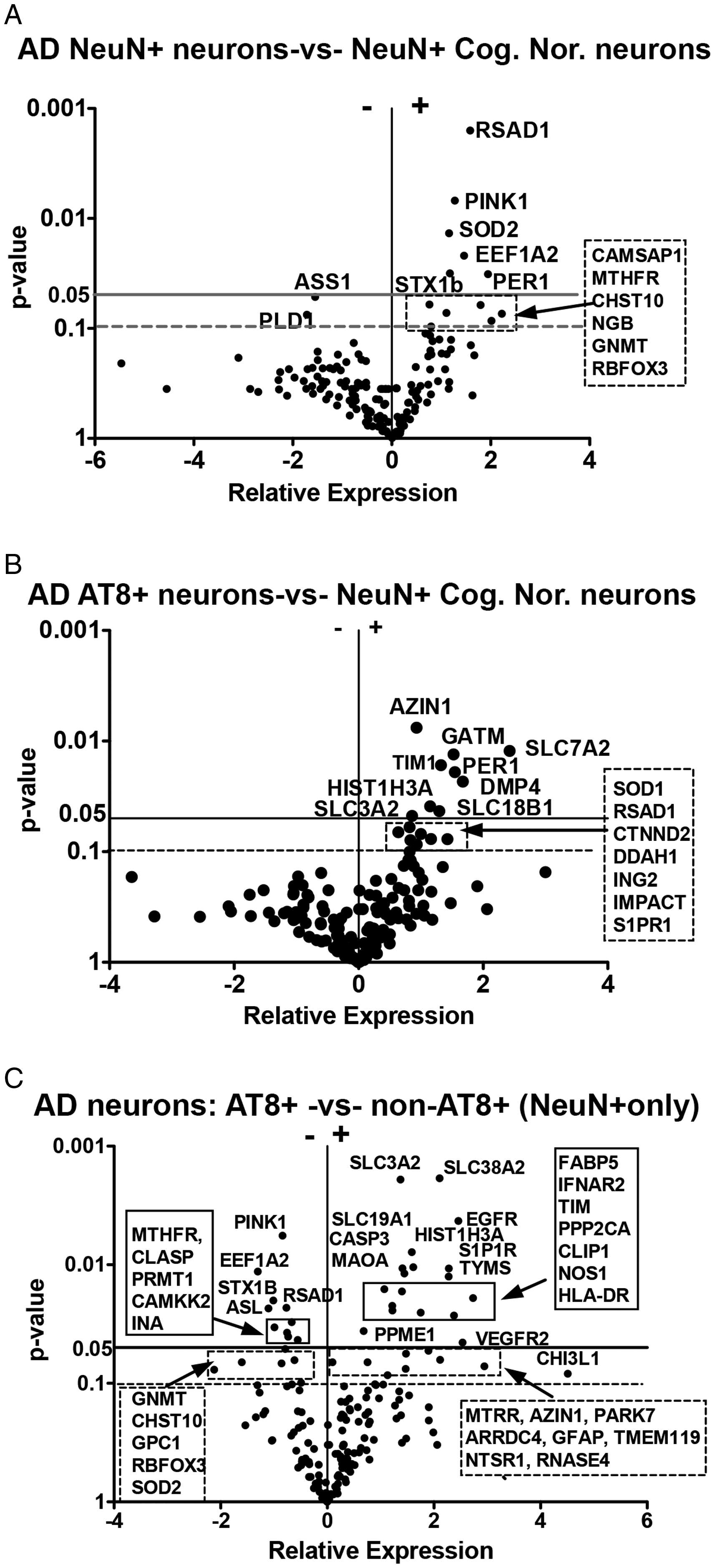
A: Differential mRNA expression between NeuN+ neurons captured using LCM from brain sections of individuals with AD compared to NeuN+ neurons captured from cognitively normal brain. Genes chosen for the study represented immune- modified metabolic pathways. Data are presented as a volcano plot where significantly increased mRNA expression levels at the p ≤ 0.05 level are shown in the upper right quadrant and genes with decreased mRNA levels at the p ≤ 0.05 level are shown in the upper left quadrant. MRNA level changes with a p-value of p < 0.1 are also indicated. Gene abbreviations are defined in Online Appendix Table 3. AD individuals: n = 20; Cognitively normal individuals: n = 11. B: Differential mRNA expression between LCM-captured AT8+ neurons from individuals with AD and NeuN+ neurons captured from cognitively normal brain. MRNA level changes with a p-value of p < 0.05 and p < 0.1 are noted. AD individuals: n = 20; Cognitively normal individuals: n = 11. C: Differential mRNA expression between AT8+ neurons from individuals with AD and NeuN+ neurons from the same AD brain. Cells were co-captured from an individual brain slice using double labeling techniques (n = 7). Genes with significantly (p ≤ 0.05) increased mRNA expression levels in AT8+ neurons compared to NeuN+ neurons in the same neuronal sample are shown in the upper right quadrant. Significantly decreased mRNA expression levels are shown for genes indicated in the left upper quadrant. MRNA level changes with a p-value of p < 0.05 and p < 0.1 are identified.
LCM: Cell Capture
After the brain samples were stained with the appropriate antibodies, they were rapidly transported on ice packs (−20°C) to the LCM facility. Stained cells on the LCM membrane slide were visualized using a 63X objective and collected using the Zeiss Palm Microbeam 4.2 LC-Microscope system with PALM Robomover Z software. Appropriate cells were identified and then catapulted via the hinge method into an individual opaque adhesive cap (Zeiss cat. no. 415190-9201-000). Approximately 250-300 neurons were collected rapidly from each slide, and required no more than 60 mins per slide. After cells from the sections on an individual slide were cut and catapulted, the adhesive material was immediately removed and placed into a 100 µl aliquot of chilled lysis solution and placed on ice until the RNA extraction was carried out.
LCM: Control for Cell Type
Cells captured for mRNA analysis were selected on the basis of cell-specific immunoreactivity. However, the capture technology, while accurate, can result in the collection of additional cells or pieces of cellular material from cells in the immediate vicinity. To better understand the degree of contamination resulting from capturing other cell types, particularly astrocytes, in our neuronal sample collections, 8-micron brain slices containing all cell types from multiple individual brains used in the study were processed for RNA using the same technique as used for the “captured” cell samples. RNA was extracted for each of these calibrator samples and NanoString gene expression analysis was carried out in the same manner as described. MRNA expression levels from each of the calibrator samples were measured for neuron-specific genes (Syntaxin1b; stathmin). Expression values for the neuronal genes were then averaged for each calibrator slice and divided by the corresponding average expression value for the glial acidic fibrillary protein (GFAP) gene from that calibrator sample (presented as the neuronal/GFAP ratio). GFAP levels were chosen to represent the most common contaminating cells (astrocytes) in the captured neuronal samples. Brain samples with a Syntaxin/GFAP ratio greater than 100 fold were chosen for use as described in detail by Tagliafierro et al. (Tagliafierro et al., 2016). It should be noted that GFAP expression levels were found to be higher in AT8+ cells when compared to NeuN-positive normal neurons but were not higher when compared to either NeuN+ neurons from AD brain or NeuN+ neurons from normal individuals. This implies the potential for increased contamination when isolating AT8+ cells or may represent a gene profile change in these specific neurons.
RNA Extraction From Isolated Cells
Total RNA was extracted from each 250-300 cell sample using the RNAqueous-Micro Total Isolation Kit (ThermoScientific AM-1931). RNA was eluted into 8 µl of Elution Solution, divided into 2 µl and 6 µl aliquots and stored at -80˚C until prepared for NanoString processing. RNA (6ul) was reverse-transcribed into complementary DNA by adding SuperScript VILO Master Mix (2ul, Invitrogen) as described (Tagliafierro et al., 2016). All samples also underwent target enrichment, and were stored at 4 °C within 24 hrs prior to using in the nCounter Single Cell Gene Expression assay (NanoString Technologies). NanoString assays were carried out by the Preclinical Genomic Pathology Laboratory at the Lineberger Center at the University of North Carolina School of Medicine, Chapel Hill NC.
Genes Assayed
Online Appendix Table 3 provides a list of the 197 genes and the HGNC identifiers provided by NanoString Technologies (Seattle, WA) used in the study. Genes were chosen for analysis based on their known involvement in arginine metabolism, immune function and cell pathophysiology.
Data Analysis
Gene expression data were analyzed using the nSolver Analysis Software 4.0 (NanoString Technologies). Cell sample data from individual brain used in the study were rigorously processed through multiple exclusion steps. These included background subtraction, positive control (geometric mean) normalization, and housekeeping gene normalization where ACTB and EIF4A2 were chosen as the appropriate housekeeping genes. A final calibration step as previously described was also included. Two types of neurons were distinguished, a) AT8-positive (AT8+) neurons and b) AT8- negative/NeuN+ neurons. NeuN+ cells were not further characterized for subtype of neuron. Volcano plots were used to evaluate the relative gene expression levels against the p value for each gene used.
Genes with p values less than 0.05 were judged as statistically significant. However, genes with p values less than or equal to 0.1 are also shown. The final numbers of samples used in the analyses are noted in the legends and were a limitation of this LCM, cell-based experiment. Gene selections were specifically designed to study metabolic and immune based pathways. However, the small sample size reduced the statistical power to detect significant differences after corrections for multiple comparisons (such as the False Discovery Rate (FDR) analysis) which are commonly used on large scale gene microarrays. Pawitan et al. (2005) and Jung and Sohn (2014) have discussed the use of False Discovery Rate (FDR) and False Negative Rate (FNR) (Anzulovich et al., 2006) in analyses required for small sample sizes and potential differences in the interpretation of a statistical value that arise for small sample sizes. Our study was based on specific interacting pathways that directed the choice of mRNAs to be analyzed and, thereby, was limited to a ‘small sample number’ category. Thus, the use of nominal p values in the framework of a Volcano plot permits identification of a small set of genes which can be further tested for replication in an independent sample.
AD Biomarkers
To determine AD status, soluble and insoluble amyloid beta 42 (Aβ42) and amyloid beta 40 (Aβ40) levels were measured using the Meso Scale Discovery assay system essentially as described by Weekman et al. (2014). The concentration of Aβ40 and Aβ42 was measured in both the soluble and insoluble fraction using the V-PLEX Aβ Peptide Panel 1 (4G8) Kit (No. K15199E), from Meso Scale Discovery (MSD; Mesoscale Diagnostics, LLC) and data were analyzed using the Discovery Workbench software from MSD. Data are shown in Online Appendix 4.
IHC on Formalin Fixed Hippocampal Brain Sections
To additionally confirm changed expression levels, brain sections from individuals diagnosed with AD or cognitively normal individuals were immuno-stained using standard techniques for selected proteins. The following proteins were identified using antibodies in situ; aggregated amyloid (4G8), hyperphosphorylated tau (AT8), Radical S-adenosyl methionine domain 1 (Rsad1); Radical S-adenosyl methionine domain 2 (Rsad2- also known as viperin) and methylene-tetra hydrofolate reductase (Mthfr). Antibody information can be found in Online Appendix Table 2. Microscope settings and information are detailed in Online Appendix 5.
Results
Neuronal Gene Expression Patterns for AD Compared to Cognitively Normal Individuals
Laser capture microdissection was used to isolate sub-types of neurons from brain sections of AD compared to age-matched cognitively normal individuals, with the goal of determining gene expression changes for specific cellular proteins within a neuronal subtype. The presence or absence of amyloid pathology was used to confirm AD-like pathology in each individual brain sample. Neurons in the brain sections were separately identified from other cell types (e.g., astrocytes, microglia) using the NeuN antibody (neuronal nuclear protein or NeuN, also known as Fox-3 (hexaribonucleotide binding protein 3). Neurons expressing hyperphosphorylated tau were distinguished using the AT8 antibody. In most cases, neuronal sections were double-labeled to specifically identify and separate NeuN+ neurons with hyperphosphorylated tau deposits from those NeuN+ neurons within the same section that did not show tau pathology. MRNAs selected for evaluation in the study were pre-selected to identify specific genes involved in key metabolic pathways, including arginine metabolism, polyamine production, the methionine and folate cycles, glutathione production and redox balance.
NanoString technology was used to obtain average mRNA levels for selected genes in the neuronal samples as described in the methods. Gene expression levels were then cross compared across subsets of neurons, and the volcano plot data of these comparisons were used to visualize differential gene expression data. The subsets used were classified as (1) NeuN+ neurons captured from individuals diagnosed at autopsy as cognitively normal, (2) NeuN+ neurons captured from individuals diagnosed at autopsy with AD and (3) AT8+ neurons captured from individuals diagnosed at autopsy with AD. AT8 immuno-stained neurons were not observed in sections from cognitively normal individuals. Cross comparisons between groups included (A) NeuN+ neurons from AD brain sections vs NeuN+ neurons from cognitively normal brains sections (Figure 1A and B) AT8+ neurons found in AD brains compared to NeuN+ neurons from cognitively normal brains sections (Figure 1B and C) AT8+ neurons found in AD brains (AD AT8+) compared to non-AT8, NeuN+ neurons also found in AD brains (AD NeuN+) using double labeling techniques on individual brain sections (Figure 1C).
Gene expression overlapped between the comparison groups. However, unique and informative mRNA expression differences were also found and are shown in Figure 1. In general, NeuN+ neurons collected from brain sections from individuals with AD demonstrated significantly higher levels of mRNA expression for genes associated with redox balance and mitochondrial redox properties when compared to NeuN+ neurons collected from cognitively normal individuals. The redox gene profile included Radical S-Adenosyl methionine Domain-containing 1 (RSAD1), PINK1 (PTEN induced putative kinase 1), and SOD2 (superoxide dismutase 2). It is worth noting that although PINK1 is more commonly associated with Parkinson’s disease, overlapping pathology with AD is clearly observed between neurodegenerative diseases (Monzio Compagnoni et al., 2020; Quinn et al., 2020). Decreased mRNA expression levels were observed for ASS1 (arginino-succinate synthase), a gene involved in citrulline production. Citrulline is a key amino acid used in NO production and in citrullination reactions such as histone regulation (Christophorou et al., 2014; Tessarz et al., 2014). Significantly increased mRNA expression levels were also observed between NeuN+ AD neurons and cognitively normal NeuN+ neurons for PER1 (Period 1), a clock gene, and EEF1α2 (Eukaryotic Elongation Factor 1α2), a critical translation factor expressed in neurons (Kenney et al., 2015, 2016). Increased gene expression levels for additional genes with p values at the p < 0.1 levels are also shown.
The gene expression profiles for AT8+ immuno-positive neurons from individuals with AD (Figure 1B) were also compared to the profiles from NeuN+ neurons collected from the same group of cognitively-normal individuals as above. Here, the relative mRNA expression levels detected in AT8+ neurons were increased for proteins associated with arginine/polyamine metabolism, including AZIN1 (antizyme inhibitor 1), GATM (glycine amidinotransferase), Golgi associated secretory pathway kinase (DMP4/FAM20C) and DDAH1 (dimethylarginine dimethylaminohydrolase 1). In addition, we found increased expression levels of critical amino acid transporter genes including SLC7A2 (also known as CAT2) and SLC3A2 (also known as LAT1) and the vesicular polyamine transporter, SLC18B1. Hist1H3A mRNA levels were increased, suggesting an underlying modification of histones in AT8+ neurons compared to normal, non-AD brain. Increased mRNA expression levels for circadian rhythm genes, that is, PERIOD 1 (PER 1) and TIMLESS (TIM), were also clearly observed in AT8+ AD neurons compared to cognitively normal NeuN+ neurons.
Differences between AT8+ neurons and NeuN+ neurons within an individual AD brain were further determined from double immune-labeling of single brain sections. Here, we compared gene expression for selected proteins between AT8+ neurons collected from the brain slice of an individual with AD to NeuN+
We also examined genes associated with arginine and methionine metabolism between AD AT8+ neurons with AD NeuN+ (non-AT8) neurons in the same brain slice. Genes that showed decreased expression levels in AD AT8+ neurons compared to the AD NeuN+ (non-AT8) neurons included methylene tetrahydrofolate reductase (MTHFR), protein methyl transferase-1 (PRMT1), RSAD1 and glycine N- methyltransferase-3 (GNMT-3), a primary methyl transferase that regulates S-adenosylmethionine (SAM) bioavailability (Hughey et al., 2018). TYMS (thymidylate synthase) which is involved in recycling of folic acid was increased in AT8+ neurons (Figure 1C). In addition, mRNAs for two key redox genes, superoxide dismutase 2 (SOD2) (critical to mitochondrial redox balance) and arginine-succinyl lyase (ASL) (involved in cycling arginine to nitric oxide) were reduced in AD AT8+ neurons when compared to AD NeuN+ (non-AT8) neurons. NOS1 (neuronal) mRNA expression levels were significantly higher in AT8+ AD neurons.
APOE Genotype-Based Effects on Gene Expression Levels
Autopsy brain samples were also analyzed based on APOE genotype. Figure 2A and B show gene expression data for individuals with an APOE4 genotype compared to individuals with an APOE3 genotype for NeuN-positive neurons captured from cognitively normal individuals (Figure 2A) and for AT8-positive neurons from individuals diagnosed with AD (Figure 2B). As shown in Figure 2A, no specific mRNA expression differences were observed between cognitively normal individuals with APOE4 versus APOE3 at the p < 0.05 level in the 195 genes analyzed in our directed mRNA study. Significant gene expression differences, however, were observed between AT8+ AD neurons expressing only APOE4 when compared to AT8+ AD neurons expressing only APOE3 (Figure 2B). In this case, increased mRNA expression levels for a number of genes were found and included PLD1 (phospholipase D1); BACE1 (beta secretase 1), IFNγR1 (interferon gamma receptor 1) and SPP1 (also known as osteopontin). The circadian gene, Timeless (TIM), was also significantly increased in APOE4 AT8+ neurons compared to APOE3 AT8+ neurons. A significant reduction in mRNA expression levels was found for RSAD1, eEF1a1 (Eukaryotic translation elongation factor 1a), and INA (Internexin neuronal intermediate filament protein alpha) in APOE4 AT8+ neurons compared to APOE3 AT8+ neurons.
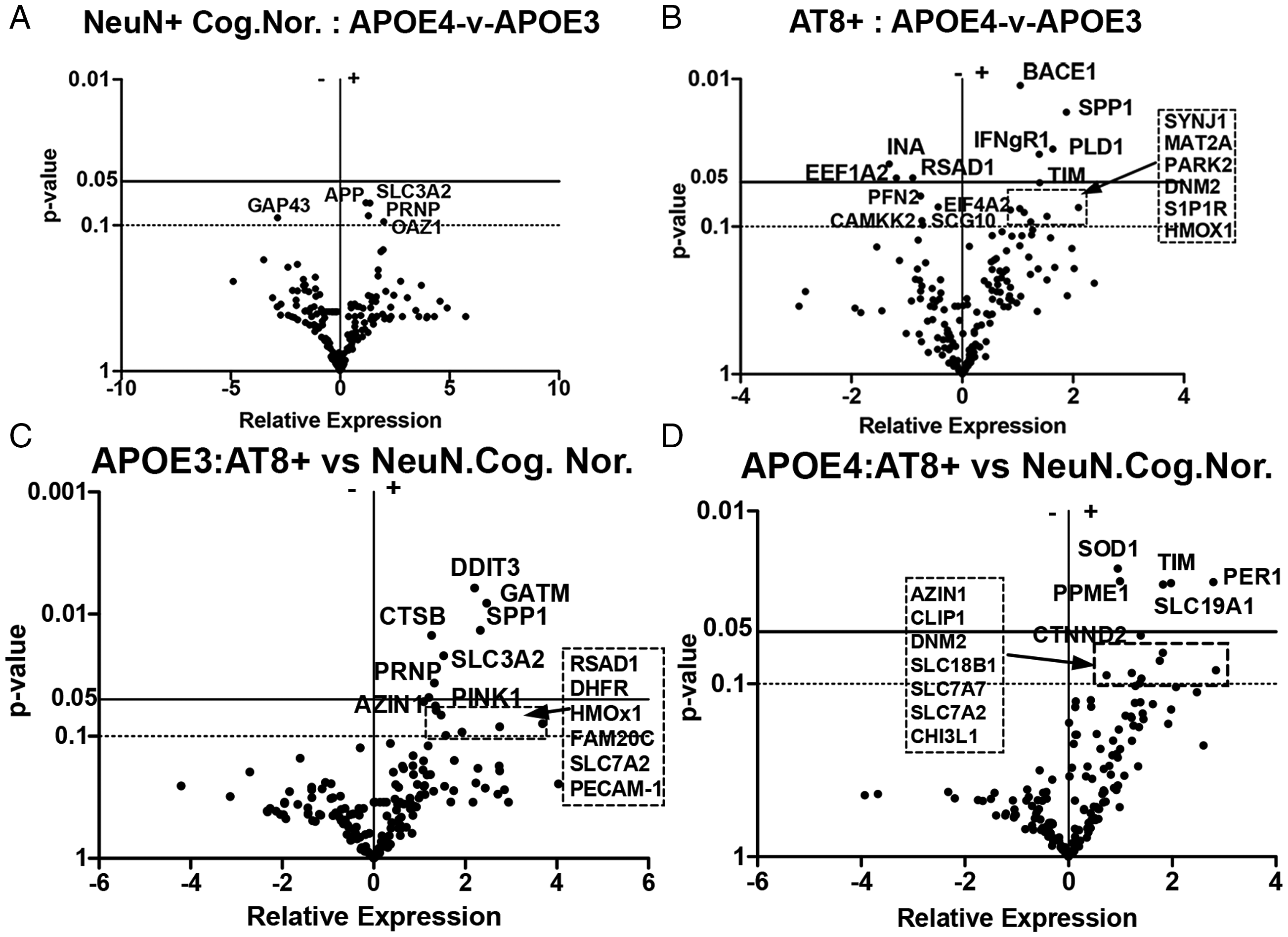
Impact of APOE4 v APOE3 Genotype on Gene Expression Patterns. Volcano plot analysis shows differential mRNA expression changes in (A) NeuN+ neurons collected from cognitively normal individuals that expressed an APOE4/4 genotype (n = 4) compared to NeuN+ neurons from cognitively normal individuals that expressed an APOE3/3 genotype (n = 6); (B) AT8+ neurons collected from individuals diagnosed with AD and expressing either an APOE4/4 genotype (n = 10) compared to AT8+ neurons with an APOE3/3 genotype (n = 9); (C) APOE3/3 AT8+ neurons collected from individuals diagnosed with AD (n = 9) compared to APOE3/3 NeuN+ neurons from individuals diagnosed as cognitively normal (n = 4); and (D) AT8+ neurons expressing an APOE4/4 or APOE4/3 genotype from individuals diagnosed with AD (n = 10).
The selective impact of APOE3 or APOE4 isoforms on neurons demonstrating AT8+ tau pathology compared to neurons from cognitively normal individuals was also investigated. Here, we compared gene expression for AT8+ neurons from AD brain to NeuN+ neurons captured from cognitively normal individuals expressing only an APOE3 genotype, or separately, expressing only an APOE4 genotype. APOE3- AT8+ neurons (Figure 2C) showed significantly increased levels of mRNA expression for multiple genes including prion protein (PRNP), osteopontin (SPP1), glycine amidinotransferase (GATM), cathepsin B (CTSB), antizyme inhibitor 1 (AZIN1), DNA damage inducible transcript 3 (DDIT3), PINK1 and others. The corresponding comparison of APOE4 AT8-positive brain samples (Figure 2D), however, showed a different profile, including increased expression of the genes that impact circadian rhythms such as Period 1 (PER1), delta-CTNND2, and Timeless (TIM) as well as altered mRNA expression for phosphoprotein methyl esterase- 1 (PPME1) and superoxide dismutase 1 (SOD1). Multiple membrane amino acid transporters were also increased, including transporters for reduced folate (SLC19A1), arginine (SLC7A7, SLC7A2) and SLC18B, a prominent polyamine transporter.
Sex Based Differences in Gene Expression
Expression levels of mRNA were compared based on sex in a similar manner as described above for APOE genotype. As shown in Figure 3A for individuals diagnosed as AD, mRNA levels for only 1 gene from our 198 gene profile was found to be significantly changed (decreased) at the p < 0.05 level in NeuN+ neurons from females with AD compared to males with AD. This gene coded for Dimethylaminohydrolase-1 (DDAH-1), the enzyme that degrades asymmetrical dimethyl arginine (ADMA). Reduced levels of ADMA have been previously observed in CSF of AD patients (Abe et al., 2001; Arlt et al., 2008).
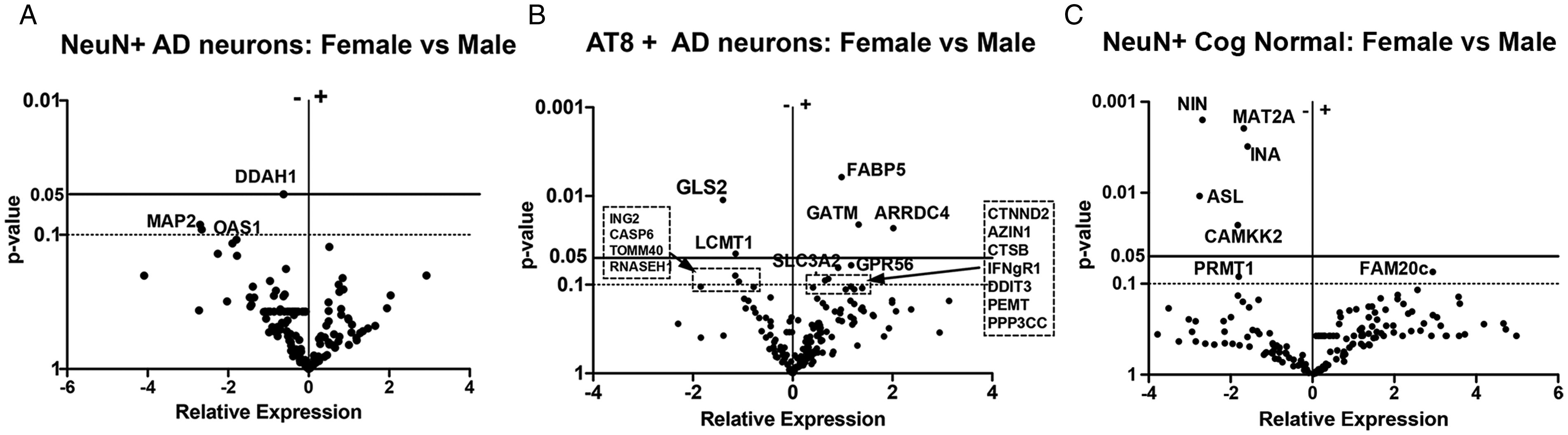
Differential Gene Expression Based on Sex. Volcano plot analysis shows differential mRNA expression levels in NeuN+ and AT8+ neurons between females vs males. A: NeuN+ neurons from individuals diagnosed with AD; n = 9 females; 10 males; (B) AT8+ neurons from individuals diagnosed with AD; n = 3 female 3; 5 males; and (C) mRNA expression levels from NeuN+ cognitively normal females vs male (5 female; 3 male).
A larger difference in mRNA profiles, however, was observed in AT8+ neurons from females with AD compared to AT8+ neurons from males with AD (Figure 3B). Here, mRNA levels were significantly increased in females compared to males and included genes encoding fatty acid binding protein 5 (FABP5), arrestin domain containing 4 (ARRDC4), known to be associated with voluntary motor activity (Kelly et al., 2012; Stockebrand et al., 2018) and glycine amidinotransferase (GATM, also known as AGAT) that alters mitochondrial energy regulation and ornithine recycling. Genes that showed significantly lower mRNA expression levels in females with AD compared to males with AD included glutamine synthetase 2 (GLS2) and leucine methyl transferase 1 (LCMT1) which is well known to play a key role in tau phosphorylation. While most commonly expressed within astrocytes, GLS2 is associated with oxidative damage of neurons in AD (Oliver et al., 1990; Robinson, 2001). Lower LCMT1 protein enzymatic activity can lead to demethylated PP2a and hence increased phospho-tau (Sontag and Sontag, 2014).
We also compared mRNA expression levels derived from brain tissue samples of cognitively normal females compared to cognitively normal males as shown in Figure 3C. Females showed a decrease in average mRNA expression levels for ninein (NIN) a cytoskeletal protein, methyl-adenotransferase 2 A (MAT2A), internexin neuronal intermediate filament protein alpha (INA), arginino-succinyl lyase (ASL) and Calcium/calmodulin-dependent protein kinase kinase 2 (CAMKK2). FAM20c was the only gene to show a slight increase in expression levels.
Protein Expression—Immunohistochemisty
Immunohistochemistry (IHC) on formalin fixed tissue was used to identify proteins corresponding to specific genes of interest that were significantly changed in our LCM study. Representative immuno-histological views from sections of control and AD brain tissues are shown in Figure 4 for cognitively normal individuals (Figure 4 left panels) and for individuals with AD (Figure 4 corresponding right panels). Based on the importance of methionine metabolism in AD, we focused on 3 specific proteins derived from the LCM based gene changes. These included MTHFR (Methylene-tetrahydrofolate Reductase), Radical S-Adenosy Methionine Domain-2 (RSAD2) and Radical S-Adenosyl Methionine Domain-1 (RSAD1). Typical patterns of immuno-positive protein expression for MTHFR, a well-known protein involved in methionine utilization pathways (Sadre-Marandi et al., 2018; Frohner et al., 2020), are shown in brain sections for age matched control individuals and for AD (Figure 4A and B). The protein immunostaining patterns supported the observed increase in MTHFR mRNA expression levels for AD brain compared to control brain sections. Both RSAD2 and RSAD1 are proteins involved in radical and redox processes and both showed increased protein expression in AD when compared to age matched control individuals. RSAD2 is a radical SAM domain protein that is now known to function as an anti-viral agent (Chin and Cresswell, 2001; Honarmand Ebrahimi, 2018; Kurokawa et al., 2019). The staining patterns of RSAD1, a closely related protein, however, were unique and in contrast to the RSAD2 pattern in the same brain sections. RSAD1 immunostaining was associated with a population of neurons that demonstrated tangle-like, structures (Figure 4F). Using double labeling for RSAD1 protein (Green, Figure 5A) and for neurons using NeuN (Red, Figure 5B) we further found that neurons immuno-positive for RSAD1 were associated specifically with fibrillar -like neuronal structures within AD brain (Figure 5C). Double-labeling techniques further confirmed that RSAD1 expression is primarily associated with neurons (Figure 5D-A) and is only secondarily associated with Amyloid plaques (Figure 5D-B). Western blot (Li-Cor Biosciences) was used to confirm a statistically relevant change in RSAD1 protein levels measured in brain slices from the same AD and control brains as used in the study (Figure 5E and Online Appendix 6). In contrast, we did not observe a significant difference between AD and normal cells using the control protein, vinculin.
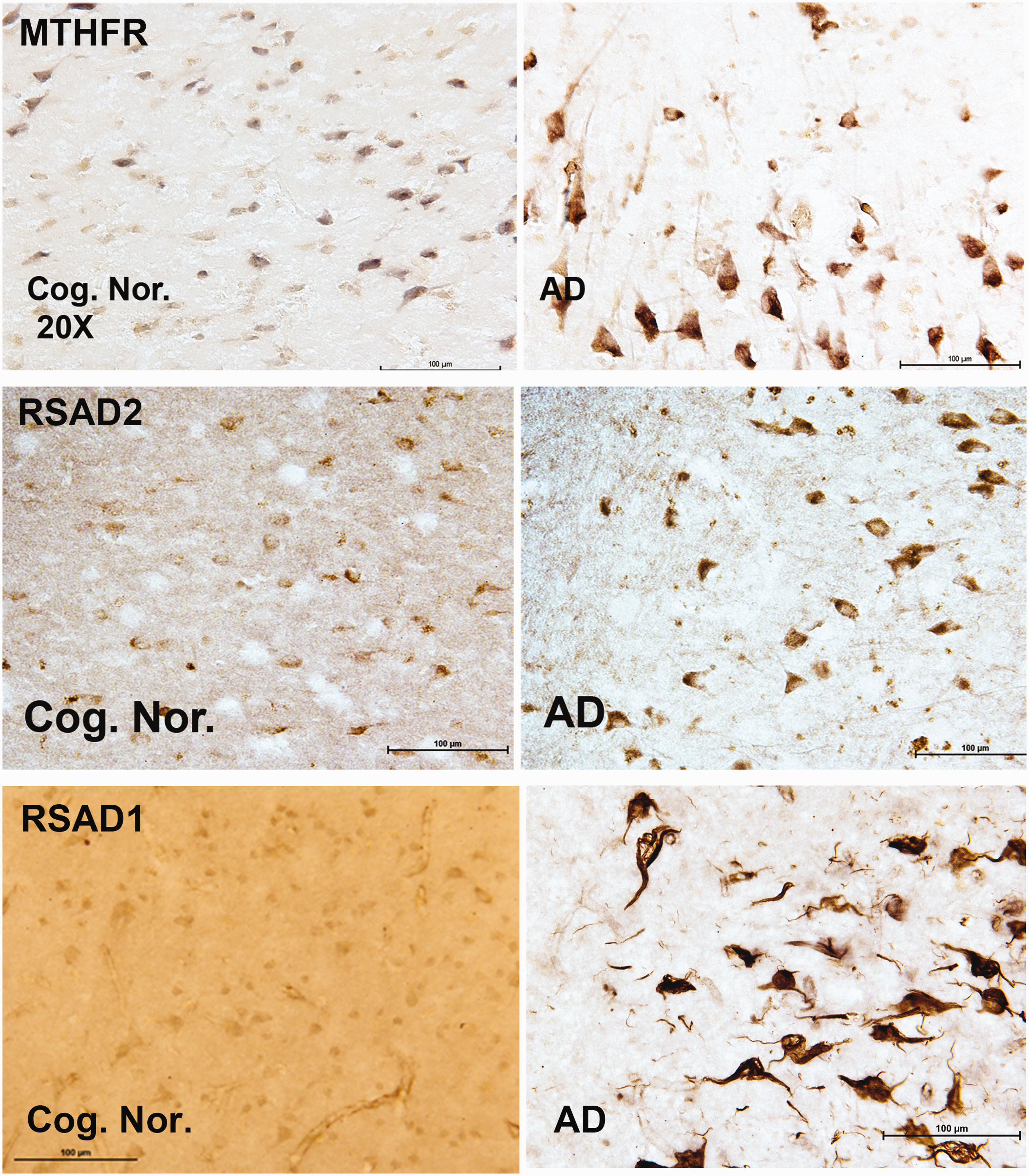
Immunocytochemical Differences Between Autopsy Brain Sections Derived From Individuals With AD Compared to Cognitively Normal Individuals. Typical protein expression levels for genes associated with methylation processes in sections from frontal cortex autopsy samples of cognitively normal individuals (left panels) compared to similar samples from individuals diagnosed with AD (right panels). A and B: Representative immunohistochemical profile for human autopsy brain sections stained for Methylenetetrahydrofolate Reductase (MTHFR) from a cognitively normal individual (A) compared to an individual diagnosed with AD (B). Left panel- Male; 86 years; cognitively normal; APOE 3/3; CERAD 1; B&B–stage 1. Right panel – Female; 86 years; AD, APOE3/4; B&B-stage 4; AA mild infarcts. C and D: Representative immunohistochemical profile for human autopsy brain sections stained for Radical S-Adenosyl methionine domain containing 2 (RSAD2- also known as viperin) from a cognitively normal individual (left panel) compared to an individual diagnosed with AD (right panel). Left panel- Female, 84 years, cognitively normal; APOE3/3, CERAD 1; Right panel- Female, 74 years, AD, APOE4/4, B&B-stage 5. E and F: Representative immunohistochemical profile for human brain sections stained for Radical S-Adenosyl methionine domain containing 1 (RSAD1) from a cognitively normal individual (Left Panel) compared to an individual diagnosed with AD (Right Panel). Left panel- Male; 86 years; cognitively normal; APOE 3/3; CERAD 1; B&B–stage 1; Right panel- Male, 89 years; AD, APOE3/3, B&B-stage 5; AA- moderate.
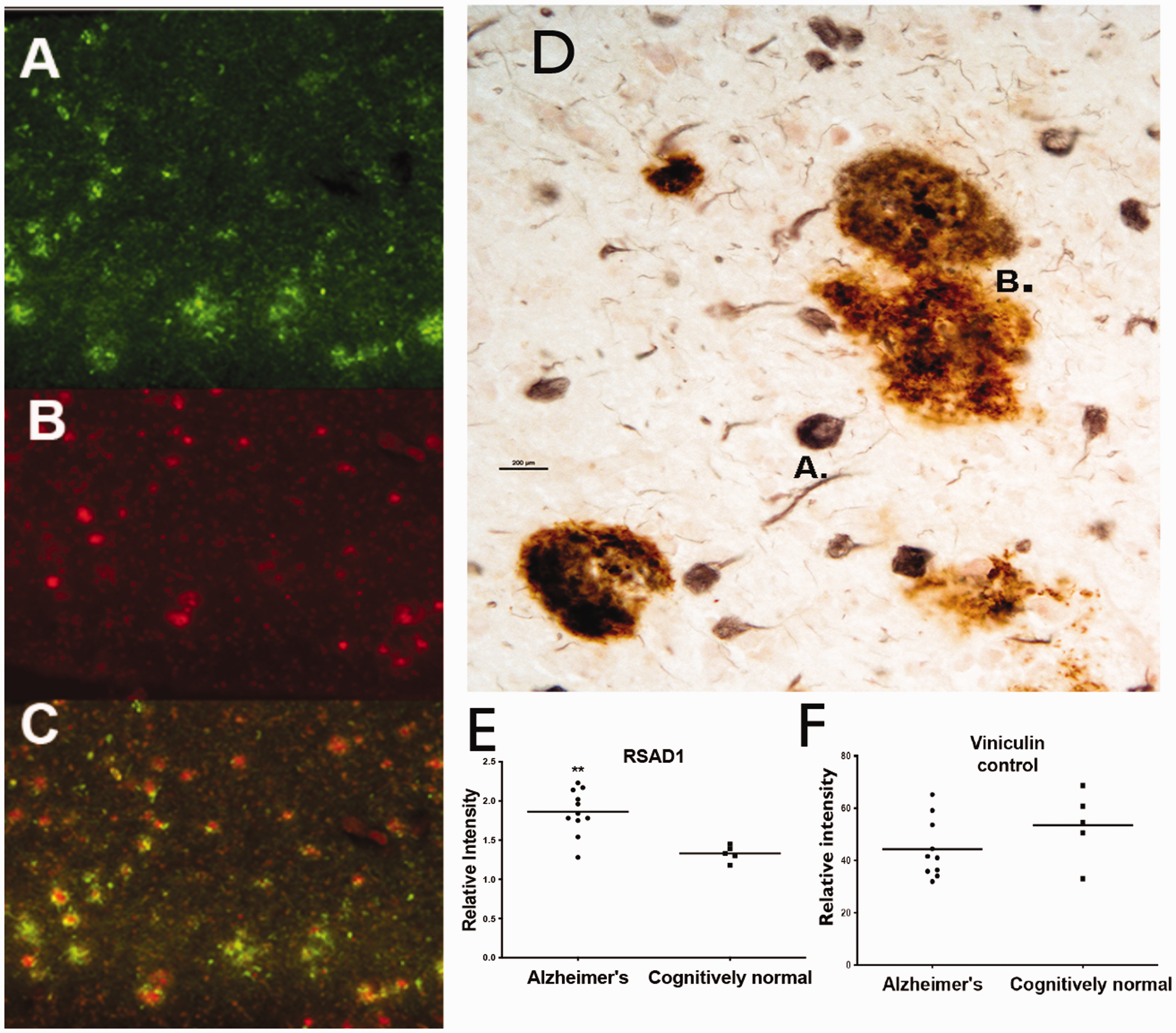
A to C: Co-localization of RSAD1 and NeuN immunoreactivities in a representative AD brain section. Fluorescent double labeling with RSAD1 and a neuron specific antibody (NeuN) was used to observe the overlap between RSAD1 and the population of neurons from a section of AD brain. A: Localization of neurons showing RSAD1 immunoreactivity (green). B: Localization of all neuronal sub types using immunoreactivity for the NeuN antibody (red). C: The presence of RSAD1 immunoreactivity co-localized with NeuN immnostaining on a sub-population of neurons (for example, (C) lower left). However, all NeuN+ neurons are not double labeled with RSAD1 immunostain (for example, (C) upper right). Pathology-Female; hippocampus, AD Brock Stage 5, 88 years. D: Double labeling was used to identify RSAD1+ cells (Black/gray stain) from 4G8+ amyloid containing neuritic plaques (Red/orange stain). E and F: Western blot was used to quantitate changes in RSAD1 protein levels (E) compared to a protein control (Vinculin, (F)). Samples were obtained from isolated brain sections from cognitively normal individuals compared to individuals with AD. ** p<0.01, n = 5 control (non-AD) and 11 AD brain samples.
Discussion
A primary goal of this study was to identify specific metabolic gene differences initiated by the presence of AT8+ hyperphosphorylated tau in neurons from individuals with AD when compared to neurons that did not demonstrate AT8+ pathology. To accomplish this goal, we used Laser Capture Microscopy (LCM) to identify and selectively isolate neurons from autopsied frozen brain tissues from individuals with AD or aged-matched normal individuals. MRNA data from captured AT8+ neurons were then compared to NeuN+ neurons captured in the same individual’s sample using double labeling techniques or to NeuN+ neurons captured from cognitively normal individuals. The neuronal specific nuclear protein marker, NeuN, represented a general post-mitotic neuronal population within the cortex (Dredge and Jensen, 2011). Three different between-group comparisons were made which included 1) NeuN+ neurons collected from AD brains compared to NeuN+ neurons from cognitively normal individuals, 2) AT8-positive neurons from individuals with AD compared to NeuN+ positive neurons from cognitively normal individuals, and 3) AT8-positive neurons versus NeuN+ (non-AT8+) neurons, both from AD brains. Our analysis focused on specific subsets of genes coding for critical proteins involved in metabolic processes that included energy utilization and redox balance, arginine usage, polyamine synthesis and methylation. These interrelated pathways form a critical and highly interactive component of the brain’s metabolic activity, are immune regulated and, when altered, are primary components of pathological changes. While the significance of these interactive pathways to human neurodegeneration may be partially obscured by death/autopsy-based factors, the comparisons between cognitively normal and individuals with AD have provided useful insights into disease- based differences. Our data support a pathophysiological metabolic switch in brain tissue of individuals with AD that directly affects both usage and cellular resupply processes for arginine.
Arginine-Polyamine-Methionine-Related Metabolic Pathways
Disruption of intracellular arginine levels are known to alter the physiological responses of both healthy and diseased cells and tissues (Kan et al., 2015; Bakshi and Morris, 2016; Morris, 2016). Loss of cellular arginine not only leads to induction of autophagy within individual cells, but also promotes compensatory pathways that redirect arginine usage to other metabolic pathways within a tissue (Liu et al., 2014; Kao et al., 2015; Morris, 2016). Expression and activity levels of arginase (AG1,2) and nitric oxide synthase 2 (NOS2) regulate arginine levels and are immune regulated, enabling changes in response to cellular and tissue conditions. Neither of these enzymes were upregulated in the pathological neurons in this study. However, we identified altered gene expression for two critical arginine-based metabolic enzymes, argininosuccinate synthase 1 (ASS1) and argininosuccinate lyase (ASL) that show altered mRNA expression levels in neurons from individuals with AD when compared to cognitively normal individuals. The sequential enzymatic action of ASS and ASL increases cellular/tissue arginine levels by converting citrulline to arginino-succinate, which in turn, is converted to arginine. Reduction in the activity levels of either ASS1 or ASL lowers cellular arginine levels (Nagamani et al., 1993). Our data show a differential expression pattern of these genes in both subtypes of AD neurons that indicates reduced cellular production of arginine from either low expression of ASS1 mRNA (as shown in AD NeuN+ neurons compared to NeuN+ neurons from cognitively normal brains) or from reduced levels of ASL mRNA expression (as observed in AD AT8+ neurons compared to AD NeuN+ neurons).
Ornithine Utilization/Polyamine Production
To gain further insight into potential downstream effects of altered arginine metabolism in the human AD brain samples, we analyzed gene changes associated with the production and use of ornithine (ORN), a key metabolite produced by the enzymatic activity of arginase 1 and 2 (Figure 6). Brain mRNA levels were measured for key enzymes that use ornithine. These included 1) ornithine amino transferase (OAT) that converts ornithine into pryolline-5 carboxylate to produce glutamate and proline, 2) ornithine transcarbamylase (OTC) that converts ornithine to citrulline and 3) ornithine decarboxylase (ODC) that forms putrescine, initiating polyamine synthesis. None of these enzymes demonstrated changes in mRNA expression when compared between subtypes of neurons. However, a significant increase in mRNA levels of Antizyme Inhibitor 1 (AZIN1) was observed in AT8+ AD neurons when compared to NeuN+ neurons from either cognitively normal individuals or NeuN+ (non-AT8+) neurons from individuals with AD. AZIN1 mRNA and the subsequent production of antizyme protein (AZ1) promote polyamine formation by blocking the inhibitory action of Ornithine Decarboxylase Antizyme 1 (OAZ1) on the enzymatic activity of ODC (Kanerva et al., 2008; Kahana, 2016; Ramos-Molina et al., 2018). The observed increase in AZIN1 mRNA levels in AT8+ neurons is permissive for active production of polyamines in these neurons (Inoue et al., 2013; Kan et al., 2015). Our data further showed elevated mRNA levels for the vesicular polyamine transporter, (SLC18B1) in AT8+ neurons in AD brain compared to NeuN+ neurons from cognitively normal brain sections. Changes in polyamine transport levels permit increased transcellular flux of polyamines, including acetylated polyamines, and are an integral process in cellular stress responses (Hoshino et al., 2005; Hiasa et al., 2014; Scalise et al., 2018; Fredriksson et al., 2019).
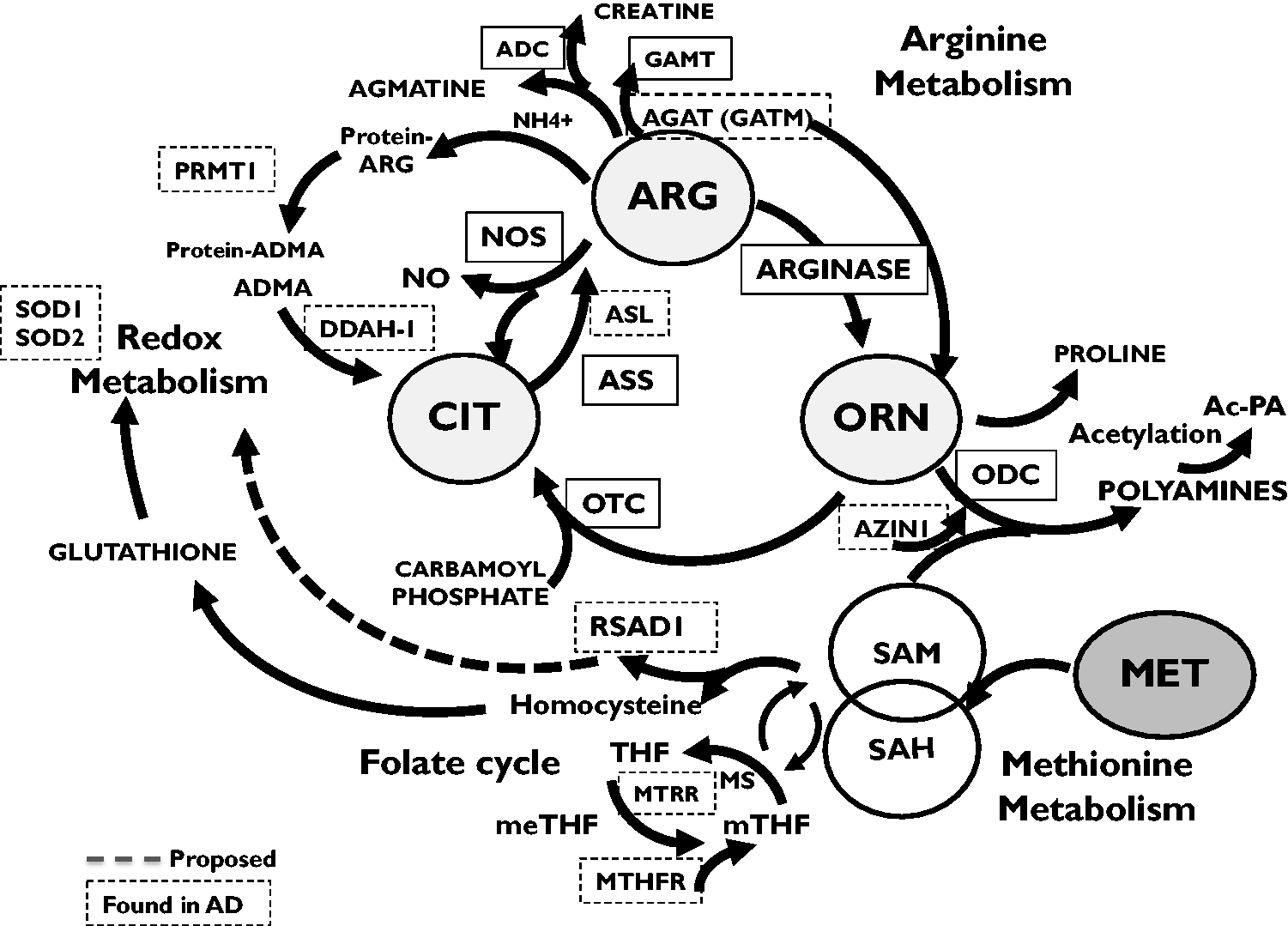
Representative Metabolic Pathway Interactions for Proteins Involved With Arginine, Ornithine, Methionine and Redox-Based Metabolic Pathways. Dashed box—Proteins affected in AT8+ neurons compared to non-AT8+/NeuN+ neurons. Dashed Line—RSAD1 activity in this pathway is based on its currently known cell-based effects and is estimated here.
Methionine Utilization
Methylation is a key process in brain metabolism and is altered in AD (Barodia et al., 2017). To better understand changes in methylation patterns between NeuN+ neurons from cognitively normal individuals compared to neurons expressing AD pathology, we measured changes in the mRNA expression levels of proteins associated with methionine usage in AD NeuN+ and AT8+ neurons compared to neurons from NeuN+ cognitively normal brains. The observed patterns of gene expression data showed increased mRNA levels in AT8+ neurons for MTHFR, PRMT1, GNMT, PPME1, RSAD1, and MTRR. We also used immuno-histochemistry to evaluate the corresponding protein expression levels to confirm specific gene changes. Prominent immunostaining for MTHFR was observed in brain sections from AD brain compared to brain sections from cognitively normal individuals. One of the most prominent and novel gene changes we observed involved a recently discovered class of enzymes that include radical SAM domain containing- 1 (RSAD1). RSAD1 mRNA was significantly elevated in NeuN+ (Non-AT8) AD neurons and in AT8+ AD neurons compared to cognitively normal neurons. Western blot confirmed an increased protein expression in related tissue sections. Furthermore, immunostaining using double-labeling techniques with specific antibodies provided a clear association of AT8+ and NeuN+ (Non-AT8) neurons with RSAD1 protein in cells from individuals with AD. Typical tangle like neurons were observed to be robustly stained with RSAD1 antibody and presented a unique staining pattern from either MTHFR or RSAD2, enzymes that are also shown to involve the use of SAM. Amyloid (Aβ+) deposits that lacked associated tau pathology were not stained in similar sections.
These novel data suggest a potential role for RSAD1 and its ability to undergo radical based reactions. RSAD1 and a related protein, RSAD2 (also known as viperin), are representatives of a superfamily of iron-sulfur cluster proteins, collectively called radical S-adenosylmethionine (rSAM) enzymes (Hutcheson and Broderick, 2012; Broderick et al., 2014; Bauerle et al., 2015; Horitani et al., 2015). The basic enzymatic activity of the rSAM enzyme superfamily results in electron transfer from a Fe-S cluster to SAM thereby initiating radical reactions by the generation of a 5′-deoxyadenosyl (5′-dAdo) radical. SAM is consumed in this reaction (Figure 6), potentially altering SAM/SAH ratios and methionine levels within neurons in AD brain. Importantly, RSAD1 activity has the potential to alter multiple targets within cells, including mitochondrial function (Layer et al., 2004; Bauerle et al., 2015; Latham et al., 2017). Although the exact mechanisms of RSAD1 activity in AD remain unknown, its direct association with AD neuronal pathology implies a critical and interesting role in the disease process.
While RSAD2 mRNA expression levels were not significantly increased, RSAD2 protein expression was clearly observed in AD neurons using immunohistochemistry. RSAD2 is linked to an anti-viral activity within neurons, most likely through a radical based mechanism (Chin and Cresswell, 2001; Honarmand Ebrahimi, 2018; Kurokawa et al., 2019).
Amino Acid/Metabolite Transporters
In addition to amino acids transporters that flux arginine and ornithine, solute carrier families transport multiple metabolites including creatine, carnitine and polyamines (Hiasa et al., 2014; Takeuchi et al., 2017; Scalise et al., 2018; Console et al., 2019; Hobbach and Closs, 2020). Here, we found increased mRNA expression levels in AT8+ neurons for a family of heterodimeric transporters that include the multifunctional SLC3A2 (4F2) subunits. These subunits associate with other transporter protein subunits to flux arginine, cystine and large branched chain and aromatic amino acids (Cantor and Ginsberg, 2012; Abdulhussein and Wallace, 2014; Cormerais et al., 2016). For example, SLC38A2 (a SNAT family member) that incorporates SLC3A2 was upregulated in AT8+ neurons and is one of a group of transporters associated with critical within-cell sensing and tissue signaling functions during amino acid starvation and autophagy in the brain (Hagglund et al., 2015; Nardi et al., 2015; Hellsten et al., 2018).
We also observed increased mRNA expression levels for the reduced folate transporter (SLC19A1) in AT8+ neurons compared to NeuN+ (non-AT8) neurons. Although initially discovered as a folate transporter, Ritchie et al. (2019) have recently identified SCL19A1 as a transporter for cyclic dinucleotides and in particular, 2’3’cyclic-GMP-AMP (cGAMP) into cells. cGAMP acts as a second messenger to initiate antiviral responses in cells. The selective association of SCL19A1 mRNA upregulation with AT8+ neurons and the presence of viperin (RSAD2) re-enforces the emerging evidence that viral-like responses may be observed in AD brain. Taken together, these specific changes in transporter mRNA expression levels imply down-stream regulation of differential activities in extended interconnected brain metabolic immune-regulated pathways.
Changes in Circadian Rhythm Genes
Disruption of sleep wake cycles is commonly associated with both preclinical symptoms and increased risk for dementia (Cammer and Tansey, 1987; Vitiello and Prinz, 1989; Lim et al., 2014; Musiek et al., 2018; Vanderheyden et al., 2018) and has also been linked to metabolic changes (Franceschi et al., 2018; Chan and Liu, 2021). Our data showed significantly increased mRNA expression of the circadian clock gene, Period1 (PER1) in both AT8+ and NeuN+ neuronal subtypes in AD brain samples compared to non-AD samples. Elevated PER1 mRNA has been observed previously using quantitative RT-PCR on regionally-matched brain tissue from aged-control subjects and from individuals diagnosed with AD (Cermakian et al., 2011). Our data also show increased expression of Timely (TIM) in AT8+ neurons captured from AD brain sections. TIM mRNA and protein are commonly found in embryonic neurons (Inaguma et al., 2015), but are also observed in humans with psychiatric disorders where circadian proteins contribute to the dysregulation of sleep patterns (Lamont et al., 2007; Terzibasi-Tozzini et al., 2017). Changes in ASS1 protein expression have also been reported to alter the activity of circadian proteins, potentially linking brain arginine metabolism to sleep-wake patterns and disruption of circadian rhythms.
The Impact of APOE Isoform Differences
We further stratified the gene expression data for this study in order to analyze specific gene expression differences based on APOE genotype. Comparing NeuN+ neurons from cognitively normal APOE4 individuals to NeuN+ neurons from cognitively normal APOE3 individuals showed no significant gene differences at the p < 0.05 level for the 198 genes included in our assay. Gene differences between APOE3 and APOE4 isoforms, however, were observed in neurons expressing AT8. For example, decreased mRNA levels for Neuronal Intermediate Filament Protein Alpha (as known as Internexin-Alpha, INA) in APOE4 brain sections were observed, suggesting disruption to neuronal filaments in APOE4 versus APOE3 brain. In addition, mRNA levels for BACE1, a critical enzyme for Aβ peptide formation, were increased in APOE4 AT8+ neurons compared to APOE3 AT8+ neurons. We also observed an increase in interferon gamma receptor-1 mRNA (IFNγR1) in APOE4 AT8+ neurons when compared to APOE3 AT8+ neurons. While IFNγ is normally associated with immune cell activation and may represent immune cell contamination of samples, Panagiotakopoulou et al. (2020) have recently identified IFNγ specific receptors on human iPSC-derived neurons.
Comparing AT8 positive neurons to NeuN neurons from cognitively normal individuals expressing either an APOE3 or APOE4 isoform also showed clear mRNA expression differences. For example, increased expression differences for APOE4 AT8+ neurons were prominently observed for genes associated with circadian rhythms (TIM; PER1; CTNND2), oxidation-based processes (SOD1; RSAD1; HMOX1), methylation pathways (PPME; AZIN1) and membrane-based transporters (SLC19A1; SLC18B1; SLC7A7). These gene changes predict differential circadian and metabolic outcomes between APOE3 and APOE4 AT8+ neurons.
The Impact of Sex on Gene Expression
Sex-based differences in mRNA expression levels were also observed for the selected gene profile used in this study. For NeuN+ neurons from cognitively normal individuals, females showed decreased expression levels in microtubule-related structural genes (NIN, INA) and in critical genes that participate in methylation and arginine based metabolic processes (ASL, MAT2A) when compared to brain samples from cognitively normal males. AD neurons showed a different sex-based gene profile which was dependent on the presence or absence of AT8 tau pathology. For NeuN+ AD neurons only DDAH1 mRNA levels showed a significant female to male difference. The reduction in DDAH1 mRNA levels would be consistent with disruption of citrulline to arginine recycling, thus impacting brain arginine levels differently in females vs males with AD. For AT8+ AD neurons a more diverse change in female to male gene profile was observed. These mRNA changes were consistent with an impact on tau pathology though changes in LCMT1 and on ornithine metabolism/mitochondrial function through increased mRNA expression of GATM (AGAT).
Summary
The data in this study have provided information on differential gene expression between individual neurons that show hyperphosphorylated tau (AT8+) pathology and individual neurons from cognitively normal individuals and, equally importantly, between neurons that selectively express AT8+ neurons in humans diagnosed as AD. The genes studied were specifically chosen to reflect key pathological aspects of brain metabolism, including the interaction of immune and metabolic pathways with sex and APOE genotype in brains of individuals with AD compared to unaffected, age matched controls. Pathophysiological processes affected included circadian rhythms, mitochondrial redox regulation, cellular methylation and cytoskeletal changes including hyperphosphorylated tau. This study further revealed the activity of a novel protein, RSAD1, that links radical based activity with methionine utilization and supports the hypothesis that immune regulated metabolic changes are a critical component of AD pathology. We also observed prominent sex and APOE differences as well as changes in circadian gene expression, further emphasizing the interconnection of metabolism with circadian regulation in AD. Overall, these guideposts allow us to discover and better investigate interacting pathways that set the stage for further examination of metabolic pathways in AD brain and how they are altered and/or activated by immune system input.
Summary Statement
Selective isolation of hyperphosphorylated (AT8+) neurons from frozen brain sections of post-mortem brain tissue from individuals diagnosed as AD and age matched control individuals was carried out using laser capture microscopy (LCM). Messenger RNA (mRNA) levels for genes representing specific metabolic-pathway proteins in the neuronal samples were then measured using NanoString technology and these data were then compared to non-AT8+ neurons from individuals with AD and from individuals diagnosed as non-AD. Data were also compared for the impact of APOE4 gene expression and for sex-based differences. These data provide information on interacting metabolic pathways that are abnormal in AD neurons and include the discovery of RSAD1, a new, but critical enzyme member of a unique radical SAM-based family that alters these pathways.
Supplemental Material
sj-xlsx-1-asn-10.1177_17590914211019443 - Supplemental material for Metabolism-Based Gene Differences in Neurons Expressing Hyperphosphorylated AT8− Positive (AT8+) Tau in Alzheimer’s Disease
Supplemental material, sj-xlsx-1-asn-10.1177_17590914211019443 for Metabolism-Based Gene Differences in Neurons Expressing Hyperphosphorylated AT8− Positive (AT8+) Tau in Alzheimer’s Disease by Audra York, Angela Everhart, Michael P. Vitek, Kirby W. Gottschalk and Carol A. Colton in ASN Neuro
Supplemental Material
sj-pdf-2-asn-10.1177_17590914211019443 - Supplemental material for Metabolism-Based Gene Differences in Neurons Expressing Hyperphosphorylated AT8− Positive (AT8+) Tau in Alzheimer’s Disease
Supplemental material, sj-pdf-2-asn-10.1177_17590914211019443 for Metabolism-Based Gene Differences in Neurons Expressing Hyperphosphorylated AT8− Positive (AT8+) Tau in Alzheimer’s Disease by Audra York, Angela Everhart, Michael P. Vitek, Kirby W. Gottschalk and Carol A. Colton in ASN Neuro
Footnotes
Acknowledgments
We would like to thank the Kathleen Price Bryan Brain Bank and Biorepository at Duke University Medical Center and, in particular John Ervin, Dr. Shih Hsiu “Jerry” Wang, MD and Dr. Diane Cruz for their outstanding work to preserve and archive valuable brain tissue for use for the study of Alzheimer’s disease.
Author Contributions
C. A. C. conceived the study, analyzed data and wrote the manuscript; K. W. G. and M. P. V. contributed to the planning of the study and proof reading of the manuscript; A. Y. and A. E. carried out Laser Capture Microscopy and other technical aspects of the study; M. P. V. and K. W. G. provided discussion and valuable insight into metabolic pathways.
Declaration of Conflicting Interests
The author(s) declared the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article: M. P. V.—Founder and share-holder in Cognosci, Inc., Durham NC.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the National Institutes of Health (NIH; R21AG051097) and the Alzheimer’s Association (AA; ZEN-16-363431); NIH AG 045422 and NIH R56AG057895. The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
