Abstract
Background:
Although various serum and tissue biomarkers have been investigated for glioma diagnosis, no gold standard has been identified. miRNA-21 was demonstrated to be a promising biomarker for the diagnosis of various brain tumors, whereas there remains uncertainty concerning whether miRNA-21 could be used as a good clinical diagnostic biomarker for glioma. The current meta-analysis aimed to evaluate the diagnostic accuracy of miRNA-21 as a potent biomarker in adults with suspected glioma.
Methods:
The Pubmed and Embase databases were searched systematically from inception to January 2020 to identify relevant research reports. Pooled sensitivity, specificity, positive likelihood ratio (PLR), negative likelihood ratio (NLR), and diagnostic odds ratio (DOR) were calculated. Summary receiver operating characteristic (SROC) curves were used to evaluate the overall diagnostic performance. Meta-regression and subgroup analyses were conducted to determine the source of heterogeneity and test the robustness of the results.
Results:
From 5394 citations with 997 subjects that met the inclusion criteria, 11 studies were selected. Summary estimates of the diagnostic performance of miRNA-21 were as follows: sensitivity, 0.83 [95% confidence interval (CI): 0.73–0.89]; specificity, 0.92 (95% CI: 0.85–0.96); PLR, 10.20 (95% CI: 5.10–20.30); NLR, 0.19 (95% CI: 0.12–0.31); and DOR, 54 (95% CI: 19–155). The area under the SROC curve was 0.94 (95% CI: 0.92–0.96). Deeks’s funnel plot revealed no evidence of publication bias (p = 0.59). Meta-regression analysis suggested that study publication year could attribute to the heterogeneity. Subgroup analysis found miRNA-21 had a constant high diagnostic accuracy across different ethnicity, glioma grade, sample source, and study region.
Conclusion:
This meta-analysis demonstrated that miRNA-21 has high diagnostic performance and could serve as a promising noninvasive diagnostic marker for glioma. Further large prospective studies are needed to validate its diagnostic value and its prognostic significance and therapeutic effects.
Introduction
The incidence of brain and central nervous system (CNS) neoplasms has been increasing rapidly for the past three decades, and gliomas account for more than half of all brain and CNS neoplasms. 1 According to the Cancer Statistics 2017 from the American Cancer Society, estimated numbers of new cases and deaths of brain and other nervous system tumors (NST) in the United States (US) are 23,800 and 16,700, respectively. 2 A status report of incidence and mortality worldwide for 36 cancers in 185 countries indicated that estimated numbers of new cases and deaths of brain and NST are 296,851 and 241,037, respectively. 3 Brain tumors are characterized by high morbidity and mortality owing to their localization and often locally invasive growth. Glioma, accounting for the majority of brain-cancer-related deaths, 4 are primary brain tumors that are thought to derive from neuroglial stem or progenitor cells. 5 Although surgery, radiotherapy, and alkylating agent chemotherapy are still the mainstay therapies, and all are applied in clinical treatment, the diagnosis and treatment of brain tumors, especially glioma, are the primary challenge for future neurologists, neurosurgeons, and oncologists. Therefor, there is an urgent need to explore novel molecular targets for better diagnosis of glioma.
MicroRNAs (miRNAs) are a class of small noncoding single-stranded RNA molecules that regulate RNA silencing and expression of certain genes. 6 They also play significant roles as oncogenes/tumor suppressors in various kinds of tumors.7–9 At the post-transcriptional level, miRNAs can regulate the expression of target genes by binding to their 3′-untranslated regions (3′-UTR) to participate in the regulation of life activities such as individual development, apoptosis, proliferation, and differentiation. 10 Accumulating evidence also demonstrates miRNAs are expressed abnormally in a series of diseases, especially in various tumors.11–13 Indeed, there are a large number of literature reports that miRNAs can serve as reliable biomarkers for the diagnosis and prognosis of human gliomas.14–16
miRNA-21 is one of the most studied types of miRNA. Previous studies have found that miRNA-21 is highly expressed in diverse cancer types and may serve as a biomarker for tumor diagnosis and prognosis.17–20 miRNA-21 is found not only in tissues, but also in a wide variety of extracellular fluid including cerebrospinal fluid (CSF), serum, plasma, saliva, and gastrointestinal fluids.21–25 Although numerous studies have proposed the diagnostic value of miRNA-21 in gliomas, results have been variable and inconclusive. The main purpose of this study was to investigate the diagnostic accuracy of miRNA-21 for detecting glioma, with the aim of determining whether miRNA-21 could be considered for use in screening patients with suspected glioma.
Methods
Protocol
This study was conducted following the Preferred Reporting Items for a Systematic Review and Meta-analysis of Diagnostic Test Accuracy Studies (PRISMA-DTA) guidelines (see Supplemental Table S1) and Assessing the methodological quality of systematic reviews (AMSTAR) Guidelines. 26 Neither ethics approval nor patient consent was required for this manuscript.
Search strategy
Before searching the literature, two reviewers (XZ and BL or ZM) searched the Cochrane Collaboration, PROSPERO, Joanna Briggs Institute (JBI), and INPLASY databases to avoid duplication. We then searched PubMed and Embase from inception to January 2020 using medical subject headings (MeSH), Emtree, and text word with no language limitations. Taking PubMed as an example, we combined the Medical Subject Heading (MeSH) terms and free-text words to implement search strategies including: (microRNA OR miRNA OR miR) AND (glioma* OR glioblastoma*) AND (“Sensitivity and Specificity” OR “diagnostic accuracy” OR “Predictive Value of Tests” OR “negative predictive value” OR “false positive” OR “false positive” OR “positive predictive value” OR diagnos*). Detailed search strategies are available in Supplemental Appendix 1. The literature search was also supplemented by a combined manual search in the reference lists from the related articles, reviews, and meta-analyses.
Two investigators (ZM and BL) independently and in duplicate carried out the initial research, importing the literature to EndNote X9.1 (Clarivate Analytics, Philadelphia, PA, USA), deleting duplicate records, excluding irrelevant literature, screening titles/abstracts, and enrolling studies with detailed classification. Eligibility was based on full text and supplement materials. Any inconsistencies were forwarded to a third reviewer (BY or TL) for a final decision.
Eligibility criteria
Included studies were required to meet all of the following criteria: (a) studies used the gold reference standard (histopathological examinations) to make definite diagnosis of brain tumors; (b) studies providing the diagnostic performance of miRNA-21 in blood (serum or plasma), tissue, and CSF for gliomas including World Health Organization (WHO) grade I–IV gliomas; (c) studies providing sufficient data for constructing the 2 × 2 contingency tables with true positive (TP), false positive (FP), true negative (TN), and false negative (FN) available.
Studies were excluded if they met one of the following criteria: (a) studies had insufficient data to yield diagnostic accuracy; (b) studies published in the form of letters, comments, reviews, or meta-analyses without original data; (c) studies belonging to basic research; (d) articles without peer-review or unpublished; (e) studies that were published repeatedly or had qualitative outcomes. Two independent authors assessed all of the studies for inclusion and exclusion. Any disagreements were discussed and resolved by consensus and involvement of a senior author if necessary.
Data extraction
Data from all included studies were abstracted independently by two authors using a standardized data collection form to address population features, reference standard, assay characteristics, methodological quality, study design, and diagnostic data, including study author, publication year, patient ethnicity and country, the number of cases and controls, glioma type or World Health Organization (WHO) grade, sample source, with or without reference gene, study design, and miRNA profiling, which were summarized in a standardized Excel (Microsoft Corporation, Redmond, WA, USA). All outcomes are dichotomous variables, including TP, FP, TN, and FN.
Cohen’s kappa coefficient
Cohen’s kappa coefficient (κ) – a statistical measure that permits investigation into study characteristics – was utilized to measure the inter-rater agreement of enrolled studies 27 ; Cohen’s κ measures the agreement between two raters who each classify N items into C mutually exclusive categories.
Risk of bias assessment
The risk of bias for the included studies was assessed using the Quality Assessment of Diagnostic Accuracy Studies-2 (QUADAS-2) tool. 28 The categories in the scoring system used for assessing risk of bias consisted of patient selection, index test, reference standard, flow, and timing. An answer of Yes, No, or Unclear was allocated to each assessed study. Only Yes answers were given a score. Risk of bias was performed independently by two authors and cross checked.
Statistical analysis
All meta-analyses were performed using the “Midas” module in the STATA 15.0 (Stata Corp LP, College Station, TX, USA). The summary sensitivity, specificity and receiver operating characteristic (SROC) curves were calculated and forest plots were generated to test diagnostic accuracy using the bivariate logistic regression model and hierarchical SROC model. In addition, a hierarchical SROC curve with a 95% prediction and confidence contours was plotted to present the results graphically.29,30 Spearman correlation test was used to assess the threshold effect. If no threshold effect was examined, the bivariate random-effect model was applied to conduct the meta-analysis. The I 2 statistic was calculated to assess inter-study heterogeneity. According to the Cochrane handbook, an I 2 statistic >50% was considered significant heterogeneity between studies. 31 Subgroup analysis was carried out to investigate the potential influential factors on the summary sensitivity and specificity, which included geographical area (US versus Europe versus Asia), study design (prospective studies versus retrospective studies), ethnicity (Caucasian versus Asian), publication year (before 2015 versus 2015 and after), glioma grade [mixed grade (WHO I/II–IV) versus WHO IV (glioblastoma) versus low grade (grade I–II) versus high grade (grade III–IV)], sample source (CSF versus blood versus tissue), sample size (<50 versus ⩾50), reference gene applied or not (yes versus no), miRNA profiling (simple versus multiple), and QUADAS-2 (high risk versus low risk). Deeks’ funnel plot was used to examine the possibility of publication bias with a p value <0.1 indicating existence of publication bias. 32 Fagan’s nomogram was also used to calculate pre-test probability and post-test probability to assess the diagnostic power of miRNA-21 in clinical practice. 33
Results
Literature search
The initial database search yielded 3168 citations after removing duplicates. Of these, 3143 irrelevant citations were removed after reviewing titles and abstracts, leaving 25 relevant studies for further consideration. We excluded another 14 articles after full-text reading based on the exclusion criteria (six without sufficient data for generating 2 × 2 contingency table; four letters, comments, reviews, or meta-analyses; two basic research studies; and two non-glioma brain tumors).34,35 Finally, 11 studies met the inclusion criteria and were retained in the meta-analysis (Figure 1).23,36–45 Cohen’s κ to measure the inter-rater agreement of enrolled studies was 0.85, indicating a favorable agreement.

Flow diagram of literature research.
Study characteristics
Baseline characteristics of the 11 included studies are displayed in Table 1. In summary, 11 studies with 19 sets of data enrolling 997 patients published between 2012 and 2017 were included in the meta-analysis. Four studies were conducted in Europe, two in the US, and five in Asia. Two studies had a prospective design, and the remaining nine studies had a retrospective design. In terms of ethnicity, six studies were conducted in Caucasian populations and five in Asian populations. The most common glioma investigated were mixed grade gliomas (WHO I/II–IV) in seven studies and WHO IV (glioblastoma) in five studies. Seven studies used CSF as samples, five used blood, and one used tissue. Seven studies were rated as high risk of bias and the other four were rated as low risk based on QUADAS-2.
Baseline features of included studies.

CSF, cerebrospinal fluid; GBM, glioblastoma multiforme; LG, low grade; HG, high grade; QUADAS, quality assessment of diagnostic accuracy studies; WHO, World Health Organization; US, United States.
Threshold effect
Spearman correlation coefficient of sensitivity and 1-specificity yielded −0.12 (p = 0.43), indicating no heterogeneity resulting from threshold effect.
Diagnostic performance
We used random effects model to estimate overall performance of miRNA-21 for diagnosis of glioma. Meta-analysis found that the summary sensitivity and specificity combining all included studies were 0.83 [95% confidence interval (CI): 0.73–0.89] and 0.92 (95% CI: 0.85–0.96), respectively (Figure 2). Significant inter-study heterogeneity was noted according to the Q test (sensitivity: Q = 93.26, p < 0.001; specificity: Q = 60.26, p < 0.001). The I 2 statistic also indicated substantial heterogeneity in terms of both sensitivity (I 2 = 80.7%) and specificity (I 2 = 70.1%). The summary diagnostic odds ratio (DOR) was 54 (95% CI: 19–155), suggesting the potential of a 54-fold higher level of miRNA-21 in subjects with positive glioma diagnosis compared with subjects with negative results, indicating a high diagnostic accuracy. The summary positive likelihood ratio (PLR) was 10.20 (95% CI: 5.10–20.30) and the summary negative likelihood ratio (NLR) was 0.19 (95% CI: 0.12–0.31). The area under the receiver operating characteristic (ROC) curve was 0.94 (95% CI, 0.92–0.96) (Figure 3). A Fagan nomogram was plotted to demonstrate the relations between pre-test and post-test probabilities and likelihood ratio (Figure 4). As noted in Figure 5, the summary PLR and NLR for miRNA-21 diagnosis of brain tumors were concentrated in the right lower quadrant, which indicated that PLR was <10 and NLR was >0.1.

Sensitivity and specificity of miRNA for diagnosis of gliomas.

SROC curve with 95% confidence contour and 95% prediction contour of miRNA-21 diagnostic value for gliomas.
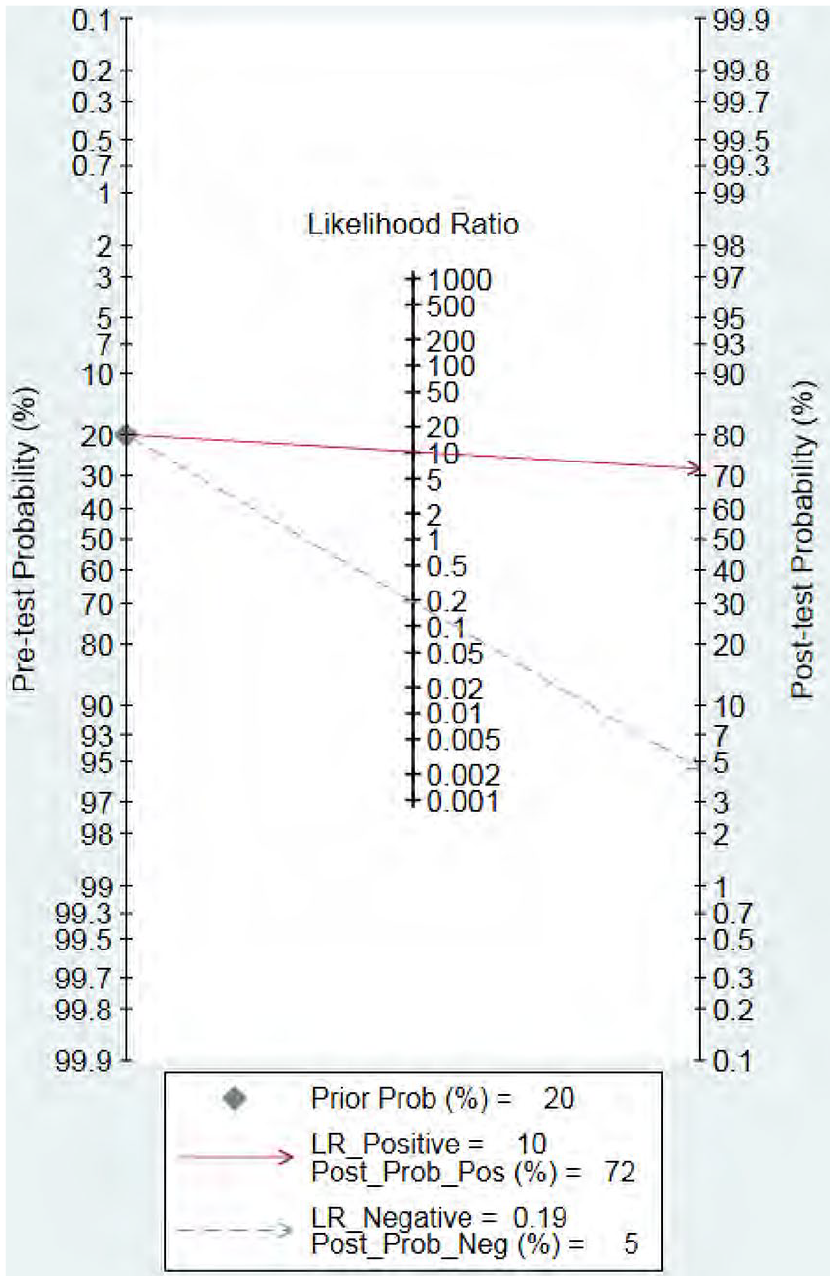
Fagan’s nomogram of miRNA-21 showing post-test probability with a fixed pre-test probability of 20% for diagnosis of gliomas.

Likelihood matrix indicates that summary PLR and NLR for miRNA-21 diagnosis of gliomas are concentrated on the RLQ.
Meta-regression and subgroup analysis
To further examine the potential sources of heterogeneity, we carried out a meta-regression analysis based on geographical area (US, Europe or Asia), study design (prospective studies or retrospective studies), ethnicity (Caucasian or Asian), publication year (before 2015 or 2015 and after), glioma grade [mixed grade (WHO I/II–IV) versus WHO IV (glioblastoma) versus low grade (WHO I–II) versus high grade (WHO III–IV)], sample source (cerebrospinal fluid, blood or tissue), sample size (<50 or ⩾50), reference gene applied or not (yes or no), miRNA profiling (simple or multiple), and QUADAS-2 (high risk or low risk). In the present meta-analysis, we found a significant effect on sensitivity (p = 0.03) for covariate of reference gene, and we did not find other covariates included in the meta-regression analysis to be the potential source of heterogeneity (all p > 0.05) (Figure 6). Results of subgroup analysis based on the above variables were consistent with the primary analyses in terms of sensitivity and specificity (Table 2).

Univariable meta-regression & subgroup meta-analysis
Subgroup analyses of the diagnostic accuracy of miRNA-21.

p < 0.1
CI, confidence interval; CSF, cerebrospinal fluid; miRNA, microRNA; QUADAS, quality assessment of diagnostic accuracy studies; WHO, World Health Organization; US, United States.
Sensitivity analysis
Sensitivity analysis was conducted by removing the included studies one by one and analyzing the SROC curve. As is shown in Figure 7, the results remained unchanged, suggesting that this meta-analysis was stable.

Sensitivity analysis of the included studies. (a) Goodness-of-fit, (b) bivariate normality, (c) influence analysis, and (d) outlier detection.
Publication bias analyses
Publication bias was examined using Deeks’ funnel plot test and visual inspection of funnel plot asymmetry. The shape of the funnel plot of the pooled DOR of miRNA-21 for the diagnosis of glioma revealed generally symmetry (Figure 8). Deeks’ asymmetry test also showed a statistically non-significant value (p = 0.59), further confirming no evident publication bias.

Deeks’ funnel plot asymmetry test for publication bias based on overall studies.
Discussion
Principal findings
This comprehensive review and meta-analysis is the first to explore the diagnostic value of miRNA-21 for glioma. Based on the findings of this study, we conclude that when miRNA is applied to the diagnosis of glioma, the rate of missed diagnosis (17%) and misdiagnosis (8%) will be quite low. The pooled sensitivity and pooled specificity reach 0.83 (95% CI: 0.73–0.89) and 0.92 (95% CI 0.85–0.96), respectively. And the diagnostic accuracy was high (DOR: 54, 95% CI: 19–155). This diagnostic accuracy was consistent when several subgroup analyses were performed (Table 2). In terms of these findings, miRNA-21, which can be detected in blood or CSF samples, has potential as a novel biological diagnostic tool for glioma.
Interpretation
Although histological diagnosis remains the gold standard for glioma, due to a type of highly heterogeneous tumors arising from brain parenchyma, molecular diagnostic markers such as isocitrate dehydrogenase (IDH) mutation status, chromosome 1p/19q status, copy number alterations of chromosome 7 and 10 and of telomerase reverse transcriptase (TERT) promoter, BRAF, and H3F3A mutations are of limited diagnostic value. Therefore, more and more studies began to focus on the integration of molecular aspects when diagnosing and managing gliomas, which included the use of miRNAs as diagnostic markers.40,46–48
Through bioinformatics analysis, it has been found that related miRNAs play a regulatory role in a variety of tumors, including CNS tumors. Previous studies have identified specific miRNAs associated with the diagnosis of gliomas. Recently, it has been confirmed that five combined miRNAs are involved in the alterations of MGMT, and that TP53 and is related to the progression of glioblastoma. 49 Among these miRNAs, miRNA-21 and miRNA-181d have been found to play a regulatory role in the carcinogenesis of glioblastoma, while miRNA-144 and miRNA-29a are related to the progression of glioblastoma. Although the diagnostic value of miRNA in glioma has been published in many different studies, further validation studies and comprehensive meta-analyses are needed with large cohort sample sizes to confirm the diagnostic performance of specific miRNA in glioma.
Imaging examinations such as magnetic resonance imaging (MRI) or positron emission tomography (PET) imaging are the first choice for the diagnosis of gliomas, but because of cost and low availability, they are not widely used, leading to delays in diagnosis. Therefore, there is an urgent need for a diagnostic method that is efficient, fast, and cost-effective. Blood and CSF samples are easier to obtain from patients and can be used to measure circulating miRNAs. Multiple studies have shown that various miRNAs (especially miRNA-21) in plasma and CSF of GBM patients are elevated significantly.
A study conducted by Santangelo et al. found that plasma exosomal miRNA had high sensitivity (0.81) and specificity (0.77) to high-level glioma, while high sensitivity was found at low-level glioma (0.75), low specificity (0.47). 40 A study by Qu et al. found that the CSF miRNA has both high sensitivity (0.88) and specificity (0.89) for the diagnosis of glioma. 50 Therefore, an updated and comprehensive synthesis of the evidence on the accuracy of miRNA-21 is warranted.
Moreover, blood or CSF sample measurement is convenient and less invasive, and qRT-PCR analysis techniques are already available in most routine laboratories. In addition, the results are readily interpretable by clinicians. Finally, this technique does not consume samples needed for other tests. Given the evidence from our findings, miRNA-21 has high sensitivity (0.83, 95% CI 0.73–0.89) and specificity (0.92, 95% CI 0.85–0.96) for the diagnosis of glioma, and the results are consistent in different WHO grades and other subgroups. Previous meta-analysis for the diagnostic accuracy of extracellular miRNA-21 for gliomas demonstrated an AUC of 0.94 in brain tumor and 0.95 in glioma, which is in line with values found from our study. 39 The results of meta-regression analysis demonstrated that the reference gene was a potentially significant factor influencing clinical heterogeneity in our study. Nevertheless, we cannot exclude the influence of some of the unmeasured causes of heterogeneity. Some other factors, such as patient-related factors including ethnicity, sample source, and tumor type, could also influence the results. Although significantly different effects were not found among subgroups, these factors should not be neglected due to the small sample size analyzed. The results, therefore, need to be interpreted with caution. The key findings of this study that were not observed in other similar studies lie in the fact that the diagnostic value of miRNA-21 was further validated in different subgroups, especially in different glioma grades including low (grade I/II) and high grade (grade III/IV), glioblastoma (grade IV), and mixed grade.
Strength and limitations
To the best of our knowledge, this study represents the largest and most comprehensive study on the assessment of the diagnostic accuracy of miRNA-21 in gliomas. The results regarding the diagnostic value of miRNA-21 in our meta-analysis were generally consistent with those from previous individual studies and systematic reviews,39,50–52 and further eligible studies were included and results were updated. Furthermore, we strictly followed the recent PRISMA-DTA guidelines for transparent and accuracy reporting to make research more credible and reproducible. One of important strengths of this study was that the retrieval strategy of this meta-analysis was formulated by a Cochrane Collaboration Member (TL). The strategy was thorough and specific, which aimed to enhance accuracy. MeSH Terms, Emtree, and free text-words were used in PubMed and Embase. The retrieval results included a wide variety of publications from inception to 2020, especially publications in the past 3 years that had not been enrolled by other meta-analyses. The included literature was searched by two reviewers independently, evaluated by Cohen’s κ. And the final decisions on literature screening and selection were sent to the senior authors (TL and ZM) for final decision, which contributed to a well-controlled review. Secondly, we did not limit publication date during the search of the major databases, making it unlikely to miss important publications. At least two independent authors were involved in the study screening, data extraction, and risk of bias assessment, and data were then cross checked for accuracy. Finally, we selected a random-effects model to generate a more conservative estimate. Compared with the previous meta-analyses, the present one included more additional studies and more thorough analyses were carried out, finding that the diagnostic value of miRNA-21 remained across various glioma grades, ethnicities, and sample sources.
There are several limitations to the present study. First, we did not include conference abstracts, unpublished gray literature, and studies not written in English, which may result in certain forms of bias of our findings. Furthermore, we omitted studies not indexed in our searched databases (e.g., Pubmed and Embase). Nevertheless, Deeks’ funnel plot suggested the absence of publication bias that could have influenced our results. Second, moderate-to-significant heterogeneity was indicated, which was expected partially because the studies included used different samples to detect miRNA-21, involved glioma patients with different WHO grades, had different study design, and other cases. Although we had conducted meta-regression analysis based on several factors for sensitivity and specificity (Table 2), this could only partly determine the potential source of heterogeneity. Third, according to QUDAS-2 tool, we found that most of the included studies did not report the assessment of blinding method, which showed a high or unclear risk of bias. These methodological issues should be avoided in future study design of diagnostic studies. Finally, although our results further demonstrate the diagnostic value of miRNA-21 for gliomas, the findings of the current study were based on aggregated, not individual, patient level data, which precluded adjustments for confounding factors and limited further analysis in certain groups of patients. And more than 50% (6/11) of the included studies had fewer than 50 patients for analysis. The small sample size did not allow for further subgroup analyses, although the consistent results were indicated in several subgroup analyses. We propose that future large well-designed studies to enroll a larger population with more homogeneous characteristics should be conducted, although this may result in selection bias. 53
Implications
Compared with several imaging diagnostic approaches such as MRI and PET imaging (the reported diagnostic sensitivity and specificity of 0.68–0.92 and 0.77–0.95 for MRI, respectively; and 0.88–0.90 and 0.73–0.75 for PET, respectively),54,55 blood miRNAs are considered as powerful cancer biomarkers with radiation-free and high diagnostic accuracy in various tumors. 56 The diagnostic sensitivity and specificity of miRNA-21 of our study yielded were 0.83 and 0.92, respectively, which was comparable with those of MRI and PET imaging.
The present study showed that miRNA-21 provided satisfactory diagnostic accuracy for glioma and can serve as a diagnostic marker comparable with conventional MRI and PET imaging, and which could complement the use of MRI and PET imaging. Whether the combined use of miRNA-21 and MRI or PET imaging can provide a more satisfactory diagnostic performance for the early diagnosis of glioma or not requires further investigation. Moreover, because glioma is a heterogenous disease, it may not be possible for miRNA-21 to serve as a gold standard biomarker. However, due to the inherent limitations of small sample size observational studies, we consider that only when further large-scale prospective, multicenter studies have validated the definite diagnostic value of miRNA-21 can the clinical application of miRNA-21 in the diagnosis of gliomas be applied routinely. Although currently the detection of miRNA-21 in the blood or CSF is not a gold standard biomarker for the diagnosis of glioma, it can provide meaningful reference information for clinicians that is not inferior to conventional MRI and PET imaging, while being more convenient and less expensive.
Conclusion
In conclusion, this meta-analysis indicates that miRNA-21 may be accurate enough to diagnose glioma. Subgroup analysis found miRNA-21 had a constant high diagnostic accuracy across different ethnicities, WHO grades, sample sources, and study regions. Although further large sample clinical studies are needed to establish the optimal approach to the application of miR-21, our evidence-based results clearly recommend, at least in part, the diagnostic value of miR-21 in glioma. Further large prospective studies are needed to validate its diagnostic value, its prognostic significance, and therapeutic effects.
Supplemental Material
sj-pdf-1-tam-10.1177_1758835920987650 – Supplemental material for miRNA-21 may serve as a promising noninvasive marker of glioma with a high diagnostic performance: a pooled analysis of 997 patients
Supplemental material, sj-pdf-1-tam-10.1177_1758835920987650 for miRNA-21 may serve as a promising noninvasive marker of glioma with a high diagnostic performance: a pooled analysis of 997 patients by Xinli Zhao, Zhihong Xiao, Bin Li, Hongwei Li, Bo Yang, Tian Li and Zubing Mei in Therapeutic Advances in Medical Oncology
Footnotes
Author contributions
Study concept and design (Bo Yang and Zubing Mei);
Acquisition of data (Xinli Zhao, Hongwei Li, Bin Li and Zubing Mei);
Analysis and interpretation of data (Xinli Zhao, Zhihong Xiao, Bo Yang, Bin Li and Zubing Mei);
Drafting of the manuscript (Xinli Zhao, Zubing Mei, Zhihong Xiao and Tian Li);
Critical revision of the manuscript for important intellectual content (all authors);
Study supervision (Bo Yang and Zubing Mei).
Conflict of interest statement
The authors declare that there is no conflict of interest.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the National Natural Science Foundation of China (grant no. 81774112) (to Z.M.) (grant no. 81401834) (to Z.X.), a grant from Siming Scholars from Shuguang Hospital (grant no. SGXZ-201913) (to ZM) and Health and Family Planning Commission Program of Wuhan City (grant no. WX18C29) (to Z.X.).
Data availability
All data generated or analyzed during this study are included in this published article and its supplementary information files.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
