Abstract
Background:
Acute ischemic stroke (AIS) imposes a major healthcare burden. It is hypothesized that bacterial infection could influence atherosclerosis and thrombus formation, potentially contributing to AIS.
Objectives:
We aim to systematically review all studies that have investigated the presence of bacterial signatures within thrombi retrieved following mechanical thrombectomy (MT) procedures in patients with AIS.
Design:
This systematic review is designed following the Preferred Reporting Items for Systematic Reviews and Meta-Analyses 2020 checklist.
Data sources and methods:
A comprehensive search was conducted in the Web of Sciences, PubMed, Scopus, and Embase databases to identify relevant studies.
Results:
The literature search and screening included 11 studies involving 674 patients, with 414 (61.4%) being male and 260 (38.6%) females. Among all the patients, 393 (58.3%) were positive for bacterial presence in their retrieved thrombi. The most utilized technique for bacterial signature detection was bacterial DNA extraction followed by polymerase chain reaction amplification of the 16S rRNA gene sequence. Staphylococcus aureus was the most studied bacteria among the studies analyzed.
Conclusion:
Bacterial infections and the presence of bacteria within thrombi may significantly contribute to AIS by initiating or exacerbating atherosclerosis or thrombosis. Understanding the mechanisms by which bacteria affect vascular health is crucial for developing effective preventive and therapeutic strategies for stroke patients.
Introduction
Stroke stands as a paramount public health challenge globally, ranking as the second leading cause of mortality and the third leading cause of morbidity according to the Global Burden of Disease 2019 report. 1 In the United States alone, the annual incidence of stroke is approximately 800,000 individuals, of whom over 600,000 encounter it for the first time. 2 The economic impact of stroke in the United States is profound, costing about 56.5 billion dollars during 2018–2019. 2 Notably, most strokes, around 87%, manifest as ischemic strokes characterized by diminished cerebral blood flow, predominantly attributable to large vessel occlusion (LVO) precipitated by atherosclerotic plaques or thromboembolic phenomena.1–3
Numerous comorbidities and risk factors contribute directly or indirectly to the development of atherosclerotic plaques, including diabetes mellitus (DM), hypertension (HTN), dyslipidemia, obesity, smoking, and inflammation. Recently it is hypothesized that bacterial inflammation may significantly influence the formation of atherosclerotic plaques or aggravating thromboembolic events on the pre-existing atherosclerotic plaque through dysregulation of coagulation or endothelial dysfunction.4–6 Exposure to microorganisms and bacterial infections may have a positive effect on the risk of cardiovascular disease, and it is mentioned that the risk of stroke may be tripled 3 days after bacterial infection.7,8 Some recent studies have indicated the presence of bacteria or their components within the atherosclerotic plaques and thrombi of patients with myocardial infarction or abdominal aorta plaques, suggesting their possible potential role in the pathophysiology of LVO.9–12
In this study, we aim to conduct a systematic review encompassing all studies that have investigated the presence of bacterial signatures within retrieved thrombi following mechanical thrombectomy (MT) in patients with acute ischemic stroke (AIS). A previous systematic review has studied the relationship of the microbiota of the body with AIS. 13 However, to the best of our knowledge, our study will represent the first systematic review to specifically assess the presence of any bacteria signature within thrombi obtained from individuals diagnosed with AIS.
Methods
Search strategy
This systematic review is conducted following the Preferred Reporting Items for Systematic Reviews and Meta-Analyses (PRISMA) 2020 checklist. 14 The literature search was performed by one author (A.Z.) on February 20, 2024, in four major databases including PubMed, Scopus, Embase, and Web of Science. The search strategy was a combination of Medical Subject Headings terms and keywords, including the following terms: “stroke,” “Ischemia,” “infarction,” “thrombectomy,” “endovascular therapy,” “bacteria,” “microbiota,” and “microorganism”; detailed search strategy is provided in the Supplemental File. In addition to the database search, we reviewed the bibliographies of included studies’ protocols and consulted field experts to identify any missed studies.
Screening and eligibility criteria
Nested Knowledge AutoLit software (version 1.79; Nested Knowledge, Saint Paul, MN, USA) was used for study screening and tagging. Two independent authors (A.Z. and S.G.) conducted the initial screening, assessing the relevance of articles based on their titles and abstracts. Subsequently, full-text articles were assessed to determine their eligibility for inclusion. The senior author (R.K.) resolved any inconsistencies between authors to reach a consensus. All studies that met the predetermined population, intervention, and outcome criteria were included in our analysis. The study population consisted of patients with AIS, and the intervention was thrombi retrieval by MT. The primary outcome of interest was the presence of bacteria or its component in the retrieved thrombi.
Exclusion criteria consisted of non-human studies, non-English studies, letters, abstracts, posters, case reports, review articles, and studies in which the thrombi were not retrieved.
Data extraction and synthesis
Data extraction was performed by one author (A.Z.) using a standardized data extraction form and Excel software. The variables extracted from each study included the title of the article, year of publication, first author name, study design, country, sample size, patients’ demographics, the National Institutes of Health Stroke Scale (NIHSS) score, stroke type, occluded vessel, intravenous thrombolysis (IVT) administration, comorbidities, exclusion criteria, most extracted bacteria, and bacteria detection method.
Assessing the quality of the included studies was performed independently by two authors (A.Z. and S.G.) using the JBI critical appraisal tool’s checklist for prevalence studies. 15 This checklist comprises nine questions surveying the quality of each section of the study, with possible answers including Yes, No, Unclear, or Not Applicable.
Categorical variables are shown as absolute numbers and percentages. Continuous variables are described by mean ± standard deviation (SD) or median and the interquartile range (IQR). Normally distributed values are shown as mean and SD, otherwise as median and the IQR.
Results
Literature search
The primary study search resulted in 1754 articles consisting of 477 articles in the PubMed database, 276 in the Scopus database, 178 in the Web of Science, and 823 articles in the Embase database. After excluding 449 duplicate articles, 1305 articles were assessed in title and abstract, and 1292 studies were excluded. The full text of 13 articles is database assessed for eligibility resulting in the remaining 11 studies. We also screened an additional 26 studies based on expert recommendations and previous resembling systematic reviews, of which none of them is included in our systematic review. Following the screening and exclusion processes, the literature review included a total of 11 studies. A PRISMA flow chart detailing the inclusion process is presented in Figure 1.

PRISMA chart.
Study characteristics
This systematic review comprised 11 studies, all investigating the retrieved thrombi from patients with AIS undergoing MT for the presence of bacterial signatures.16–26 These studies collectively involved 674 patients, with 414 (61.4%) being male and 260 (38.6%) females. It’s worth noting that the three studies conducted by Patrakka et al. at the Acute Stroke Unit of Tampere University Hospital, Tampere, Finland, appear to share the same patient sample and timeframe, constituting a subset of the larger Tampere Brain Microbes and Genetics study at the same institute. We did not exclude the duplicate studies because we are not conducting a meta-analysis. If we exclude the duplicated patients from the three studies by Patrakka et al., this systematic review will include 542 patients, with 321 (59.2%) being male and 221 (40.8%) females.
The anterior circulation is the most frequently affected site of LVO, with cardiac emboli and atherosclerotic stenosis identified as the primary causes of occlusion in these patients. In available data from the studies, there were 587 occluding thrombi in the anterior circulation and 63 thrombi in the posterior circulation. Of 585 patients with provided data regarding the reason for AIS, 311 (53.2%) patients had AIS due to cardioembolic events, and 274 (46.8%) patients had AIS due to atherosclerotic thrombosis. Thrombus retrieval in all studies was accomplished using aspiration, stent retriever, or a combination of both methods. Additionally, one study reported the necessity of employing a stent to achieve satisfactory reperfusion. Table 1 provides patient characteristics including first author name and year of study, study design and period in months, country, sample size, age, sex, pre-MT NIHSS, comorbidities, cause of stroke, site of LVO and IVT administration before MT.
Patients characteristics.
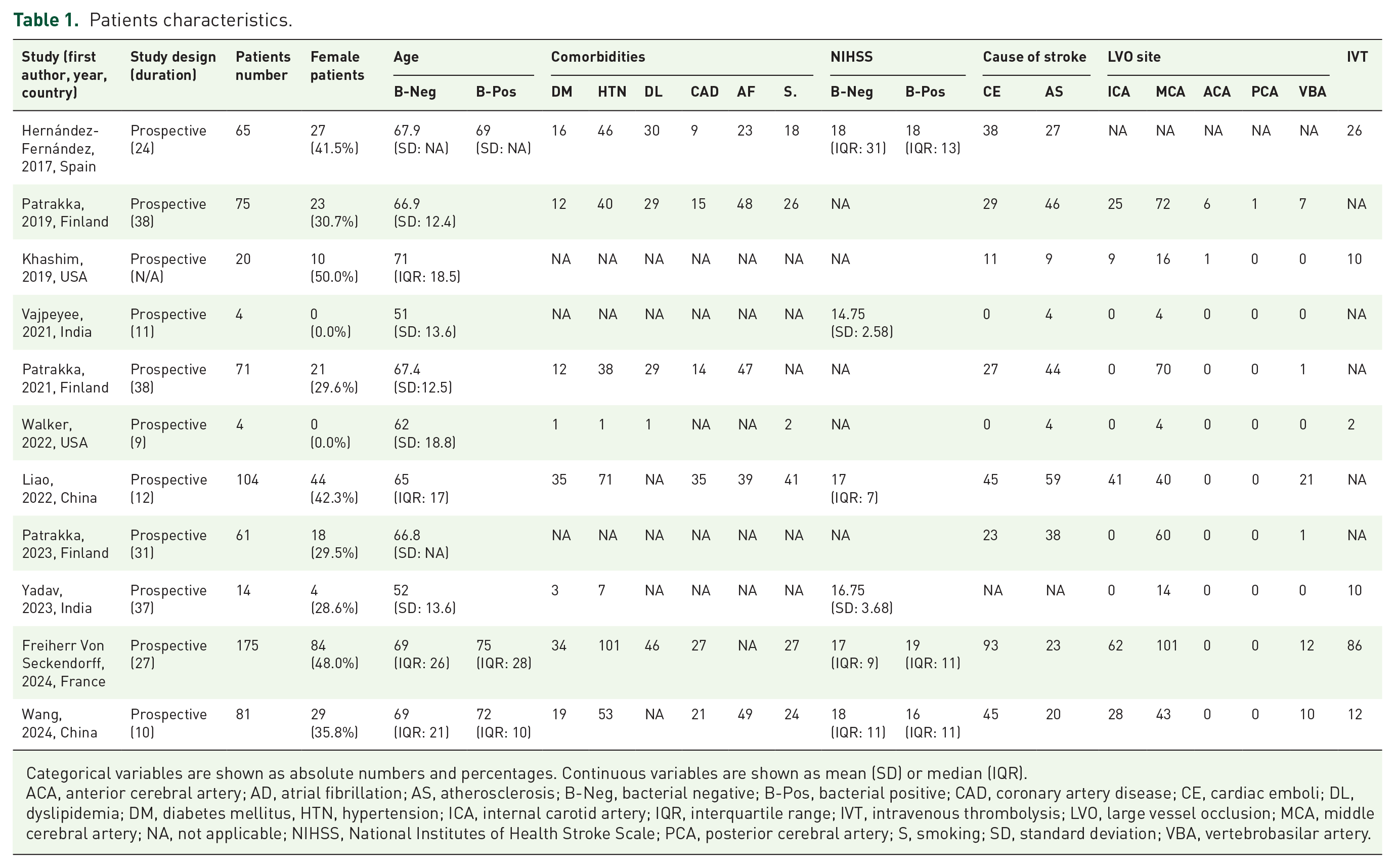
Categorical variables are shown as absolute numbers and percentages. Continuous variables are shown as mean (SD) or median (IQR).
ACA, anterior cerebral artery; AD, atrial fibrillation; AS, atherosclerosis; B-Neg, bacterial negative; B-Pos, bacterial positive; CAD, coronary artery disease; CE, cardiac emboli; DL, dyslipidemia; DM, diabetes mellitus, HTN, hypertension; ICA, internal carotid artery; IQR, interquartile range; IVT, intravenous thrombolysis; LVO, large vessel occlusion; MCA, middle cerebral artery; NA, not applicable; NIHSS, National Institutes of Health Stroke Scale; PCA, posterior cerebral artery; S, smoking; SD, standard deviation; VBA, vertebrobasilar artery.
Quality assessment
The quality assessment showed that the domains with highest concern in the studies were participant selection, the response rate, and the sample size, respectively. Figures 2 and 3 show the results of quality assessment of each study using checklist for prevalence studies of the JBI Critical Appraisal tool.

Quality assessment in each study.

Overall quality assessment in each domain.
Bacterial signature
Out of 674 patients across all 11 studies, bacterial signatures were detected in the retrieved thrombi of 393 cases (58.3%). However, after excluding the duplicated patients from the studies conducted by Patrakka et al., bacterial signatures were noted in 274 patients (50.6%) out of the total 542 patients. The prevalence of bacterial signatures in thrombi varied widely across studies, ranging from 0% to 100%. In one study comprising 20 patients, the authors did not detect any signs of bacteria in the retrieved thrombi. 18 Conversely, in three studies with smaller sample sizes, all patients had bacterial DNA present in the retrieved thrombi.19,21,24
Two studies solely employed staining and immunostaining for the detection of bacteria in the retrieved thrombus16,25; however, 9 out of 11 studies utilized extracted bacterial DNA for quantitative polymerase chain reaction (qPCR) amplification of the 16S rRNA gene. The 9 studies using DNA extraction for the detection of bacterial signature within thrombi consisted of 434 patients in which 330 (76%) samples were positive for bacterial signature; however, in the other 2 studies using other detection methods, there were 63 samples with positive bacterial signature from 240 patients (26.2%). It is noteworthy that the study in which no bacterial signatures were found in the retrieved thrombi of patients utilized both Gram staining and PCR amplification of the 16S rRNA gene for the detection of bacteria. 18 Table 2 shows the bacterial detection techniques and kits and the most prevalent bacterial species within the thrombi in each study.
Bacterial detection techniques and amplified bacterial species.

FISH, Fluorescent in situ hybridization; qPCR, quantitative polymerase chain reaction.
Discussion
Our systematic review, which examined the presence of bacteria or bacterial signatures in thrombi retrieved from patients with AIS, indicates that bacterial signatures were detected in the thrombi of more than 58% of AIS patients across the studies, albeit with a wide variation in the frequency of bacterial signature presence in thrombi, ranging from 0% to 100%. This wide range of bacterial signature detection could be due to various laboratory methods used for detection and also a small sample size of study; for example, in study of Khashim et al., in which the rate of bacterial signature presence in retrieved thrombi was 0%, they only studied 20 thrombi.
Bacterial infections are postulated to play a pivotal role in vascular diseases by either precipitating or exacerbating atherosclerosis and thrombosis processes.11,27 It is suspected that during the development of atherosclerotic plaque, the presence of bacteria within the plagues is responsible for the acceleration of the inflammatory process within the plaque leading to plaque rupture, thrombosis, and subsequent distal emboli.6–8 Moreover, some studies suggest that bacterial infections can induce thrombosis even in the absence of atherosclerotic plaques. 28
Several pathways and mechanisms are proposed to elucidate how the bacteria within the thrombus or infectious process can induce LVO whether by exacerbating LVO due to atherosclerosis or causing thrombosis and embolic process. For instance, it is shown that a cluster of Bacillus anthracis can directly activate prothrombin and factor X in human blood, leading to coagulation and thrombosis.28,29 Certain coagulation factors such as fibrin and fibrinogen matrix are associated with the antimicrobial defense system in the body, which will increase in blood during sepsis and result in thrombosis and LVO. 30 Staphylococcus aureus has been found to defend against the host’s immune system by secreting procoagulant factors and stimulating plasma clotting. 31 Some studies suggest that elevated levels of serum antibodies against bacteria in infectious diseases may increase the risk of stroke events in patients. 32 Gut microbial have also been implicated in promoting atherogenesis and thrombogenesis by activating Toll-like receptors. 33 Another proposed pathophysiology for the increase of cerebrovascular events during bacteremia is, that bacteria induce inflammation, are phagocytosed by neutrophils, and accelerate neutrophil extracellular trap secretion. This process can lead to bacterial colonization in atherosclerosis lesions, causing plaque instability, rupture, and subsequent platelet activation and thrombosis.34–37
Some studies have suggested a potential association between oral and endodontic infections and cardiovascular events. Pessi et al. 11 found a positive correlation between the presence of periapical abscess and the presence of Streptococci viridans DNA in the thrombi of patients with myocardial infarction. 38 Similarly, in three subset studies of Patrakka et al. 17 in Finland, results revealed a relationship between oral cavity infections and ischemic stroke. Their findings indicated that bacterial DNA was present in approximately 84% of thrombi retrieved from AIS patients. In which the most prevalent bacterium was the S. viridans mitis group.20,23 It is assumed that this bacterium can enter blood flow, adhere to vascular endothelium, induce platelet aggregation, and then form a biofilm that may serve as attachment points for other bacteria. In a cohort study surveying the presence of Porphyromonas gingivalis (Pg), a periodontal bacterium, in the thrombi of 175 stroke patients, authors detected this bacterium through immunostaining in 59 (33.7%) of patients. Considerably, the results showed that the near to complete reperfusion (Thrombolysis in Cerebral Infarction score of 2c/3) and 90 days favorable neurological outcomes were lower in patients with positive Pg in their aspirated thrombi. 25
Few studies have shown that the presence of bacterial signatures within thrombi can be linked to patients’ severity of disease, clinical outcomes, and functional independence following treatment. Therefore, detecting bacterial components after thrombus retrieval can provide insights into patients’ prognosis and assist physicians in initiating appropriate medical treatments. Additionally, identifying bacterial signatures in thromboembolic clots in AIS patients could help diagnose the source of the bacteria, such as in cases of infective endocarditis.11,22,39 Liao et al. in China investigated the microbial features of thrombi in stroke patients with LVO and their relationship with 90-day outcomes. They used DNA extraction, 16S rRNA gene sequencing, and Fluorescent in situ hybridization (FISH) for the detection of bacteria signatures in thrombus samples. Bacterial DNA was found in 96.2% of patients which mainly originated from plasma compared to microbiota of feces and oral samples. Interestingly, when evaluating the outcome of thrombectomy with bacterial DNA presence, they noticed that a higher concentration of Acinetobacter and Enterobacteriaceae DNAs were associated with a higher risk of perioperative adverse effects, and a higher abundance of Acinetobacter DNA was independently associated with a higher risk of 90-day mortality. 22
In each study included in this review different bacterial families’ signature was diagnosed within thrombi. As mentioned above, in three studies done by Patrakka et al.,17,20,23 the S. viridans mitis group signature had a higher concentration in the thrombi, which was attributed to oral infection. In Liao et al.’s study, the bacterial DNA with the highest average abundance were Acinetobacter (12.8%) and Stenotrophomonas (5.3%). Interestingly, the authors found that DNA of Lactobacillus and Chryseobacterium were significantly more abundant in thrombi from atherosclerotic stroke patients, while the DNA of Veillonellaceae family was more abundant in cardioembolic stroke patients. 22 The results of Wang et al.’s study suggests a distinct microbial profile within the thrombus compared to the circulating arterial blood. There was a higher abundance of Bacillus, Corynebacterium, Parabacteroides, Romboutsia, Roseburia, Prevotella, and Streptococcus species DNA in thrombi compared to arterial blood samples. 26 In a couple of studies in patients with myocardial infarction, the DNA of Chlamydia pneumoniae is extracted from the thrombi of coronary arteries.40–42 Although there is no definitive evidence linking specific bacterial species to different types of ischemic stroke (thromboembolic or atherosclerotic), some studies suggest trends. For instance, bacterial species found in patients with thromboembolic AIS are often those associated with septicemia or endocarditis. In contrast, the bacterial signatures observed in patients with atherosclerotic AIS tend to relate to body microbiota, such as those from the gut or oral cavity.
Studies in this systematic review have applied various laboratory techniques for the detection of bacteria in the thrombus. In this systematic review, the rate of detecting thrombi samples with positive bacterial signatures in the studies using DNA extraction and PCR was higher compared to two studies using other detection methods (76% vs 26%). Considering the available literature and studies, it can be concluded that DNA extraction followed by 16S rRNA gene sequencing and FISH techniques using universal and specific bacterial probes are the most effective methods for the detection of bacteria within the thrombus. Previously the culture and colony counting methods were the most common methods in the detection of pathogens. However, these methods have some limitations and can be slow, resource-intensive, and sometimes inaccurate; and they can assess live microbes or viable cells. In contrast, qPCR is faster, more specific, and more sensitive, detecting low concentrations of pathogens by targeting specific DNA sequences. Real-time PCR, in particular, offers superior detection limits compared to culture methods, which require live cells and have higher detection thresholds.43,44
Despite the insights provided by this systematic review, there are some limitations; the number of studies investigating bacterial signature in the thrombi of AIS patients is limited, however, there are plenty of studies evaluating the presence of bacteria in the thrombi of patients with myocardial infarction. Additionally, some studies concentrated solely on specific bacterial species. On the other hand, as noted in this systematic review, various techniques have been employed to detect bacterial signatures in retrieved thrombi; additionally, studies assessing bacterial DNA have utilized different bacterial 16S rRNA amplification methods, which limits the comparability of results across studies.
Conclusion
In conclusion, this systematic review showed that bacteria are present in more than half of retrieved thrombi from patients with AIS, which could raise the hypothesis of role of bacterial infections and the presence of bacteria within thrombus in cerebrovascular diseases by initiating or exacerbating atherosclerosis or thrombosis. Further research with larger sample sizes using precise laboratory methods such as PCR is recommended to explore bacterial signatures in the thrombi of ischemic stroke patients. Moreover, studies investigating the causality and the underlying pathophysiological mechanisms increasing the risk of cerebrovascular events associated with bacteria and studies investigating the associations between various bacteria and different stroke subtypes (cardioembolic, atherosclerotic, etc.) are warranted.
Supplemental Material
sj-docx-1-tan-10.1177_17562864241296713 – Supplemental material for Bacterial signature in retrieved thrombi of patients with acute ischemic stroke—a systematic review
Supplemental material, sj-docx-1-tan-10.1177_17562864241296713 for Bacterial signature in retrieved thrombi of patients with acute ischemic stroke—a systematic review by Armin Zarrintan, Sherief Ghozy, Kasthuri Thirupathi, Kalah Walden, Waleed Brinjikji, David F. Kallmes and Ramanathan Kadirvel in Therapeutic Advances in Neurological Disorders
Footnotes
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
