Abstract
Large language models (LLMs) transform healthcare by assisting clinicians with decision-making, research, and patient management. In gastroenterology, LLMs have shown potential in clinical decision support, data extraction, and patient education. However, challenges such as bias, hallucinations, integration with clinical workflows, and regulatory compliance must be addressed for safe and effective implementation. This manuscript presents a structured framework for integrating LLMs into gastroenterology, using Hepatitis C treatment as a real-world application. The framework outlines key steps to ensure accuracy, safety, and clinical relevance while mitigating risks associated with artificial intelligence (AI)-driven healthcare tools. The framework includes defining clinical goals, assembling a multidisciplinary team, data collection and preparation, model selection, fine-tuning, calibration, hallucination mitigation, user interface development, integration with electronic health records, real-world validation, and continuous improvement. Retrieval-augmented generation and fine-tuning approaches are evaluated for optimizing model adaptability. Bias detection, reinforcement learning from human feedback, and structured prompt engineering are incorporated to enhance reliability. Ethical and regulatory considerations, including the Health Insurance Portability and Accountability Act, General Data Protection Regulation, and AI-specific guidelines (DECIDE-AI, SPIRIT-AI, CONSORT-AI), are addressed to ensure responsible AI deployment. LLMs have the potential to enhance decision-making, research efficiency, and patient care in gastroenterology, but responsible deployment requires bias mitigation, transparency, and ongoing validation. Future research should focus on multi-institutional validation and AI-assisted clinical trials to establish LLMs as reliable tools in gastroenterology.
Plain language summary
Artificial intelligence (AI) is transforming healthcare by helping doctors make better decisions, analyze research faster, and improve patient care. Large language models (LLMs) are a type of AI that process and generate human-like text, making them useful in gastroenterology. This paper presents a structured framework for safely using LLMs in clinical practice, using Hepatitis C treatment as an example. The framework begins by setting clear goals, such as improving Hepatitis C treatment recommendations or making patient education easier to understand. A team of doctors, AI specialists, and data experts is assembled to ensure the model is medically accurate and practical. Next, relevant medical data from electronic health records (EHRs), clinical guidelines, and research studies is gathered and prepared to improve AI, ensuring it provides useful and fair recommendations. The right AI model is then chosen and improved to specialize in gastroenterology. To make sure the model is reliable and makes correct suggestions, its performance is checked and adjusted before use. A user-friendly interface is created so doctors can access AI-generated recommendations directly in EHRs and decision-support tools, making it easy to integrate into daily practice. Before full use, the AI is tested in real-world settings, where gastroenterologists review its recommendations for safety and accuracy. Once in use, ongoing updates based on doctor feedback help improve its performance. Ethical and legal safeguards, such as protecting patient privacy and ensuring fairness, guide its responsible use. Findings are then shared with the medical community, allowing for further testing and broader adoption. By following this framework, LLMs can help doctors make better decisions, personalize treatments, and improve efficiency, ultimately leading to better patient outcomes in gastroenterology.
Introduction
Artificial intelligence (AI) enables machines to simulate complex human cognitive functions in machines. 1 A key subset of AI, machine learning (ML), enables computers to learn from data and improve performance on specific tasks without explicit programming. Within ML, supervised maps input to outputs using labeled data, while unsupervised learning identifies hidden patterns within unlabeled data.2,3 Deep learning, a further subset of ML, employs neural networks inspired by the human brain function to analyze vast datasets and extract meaningful patterns without manual feature extraction. 4 Generative AI, advanced deep learning, can create new data—such as text, images, or audio—by learning patterns from large datasets and generating samples that align with the same underlying distribution.
Large language models (LLMs), a class of generative AI, have transformed natural language processing (NLP) in medicine, demonstrating success in clinical decision support, patient education, drug discovery, and biomedical research (Table 1).5–8
Applications of large language models in Gastroenterology.

AI, artificial intelligence; BDT, binary decision tree; CDS, clinical decision support; CoT-FSP, chain-of-thought-few-shot prompting; CPGs, clinical practice guidelines; IBD, inflammatory bowel disease; LLM, large language models; NLP, natural language processing; PAGC, program-aided graph construction; PROs, patient-reported outcomes; GI, gastroenterology.
Notably, MedPaLM 2 outperforms GPT-4 in USMLE-style medical questions and is currently being evaluated in real-world clinical interactions. 20 Similarly, BioGPT excels in biomedical literature mining, while UCSF-BERT has demonstrated >88% accuracy in detecting treatment-emergent adverse events from electronic health records (EHRs).21,22
In gastroenterology, LLMs have been leveraged for diverse applications, including generating patient education materials for bariatric surgery candidates, enhancing doctor–patient interactions in hepatocellular carcinoma and cirrhosis, responding to clinical queries, grading non-alcoholic fatty liver disease histology, and extracting data.9,13,20,23–30 An NIH-funded pilot study is currently evaluating LLMs for automating data extraction in hepatocellular carcinoma. 31 In addition, these models hold promise for improving telehealth scalability through patient triaging and serving as best practice tools for identifying patients eligible for colorectal cancer screening.32,33 LLMs also assist with research and data analysis. These models can streamline academic writing and accelerate labor-intensive tasks such as conducting literature reviews and meta-analyses.34,35
Despite their potential, LLM integration in research and clinical practice presents significant challenges. Concerns regarding data privacy, model bias, and the need for rigorous validation and transparency must be addressed. Overconfident errors or “hallucinations” pose safety risks, while explainability remains a key issue. Furthermore, real-world adoption is complicated by shifts in medical terminology, evolving clinical practices, and unpredictable factors such as disease outbreaks.36–38
To overcome these challenges, we propose a structured framework, illustrated in Figure 1, that integrates LLMs into research and clinical decision-making, using personalized Hepatitis C treatment model as an example. This framework incorporates fine-tuning versus retrieval-augmented generation (RAG), prompt optimization for structured outputs, active learning for continuous improvement, and methods to mitigate hallucinations and standardize outputs.
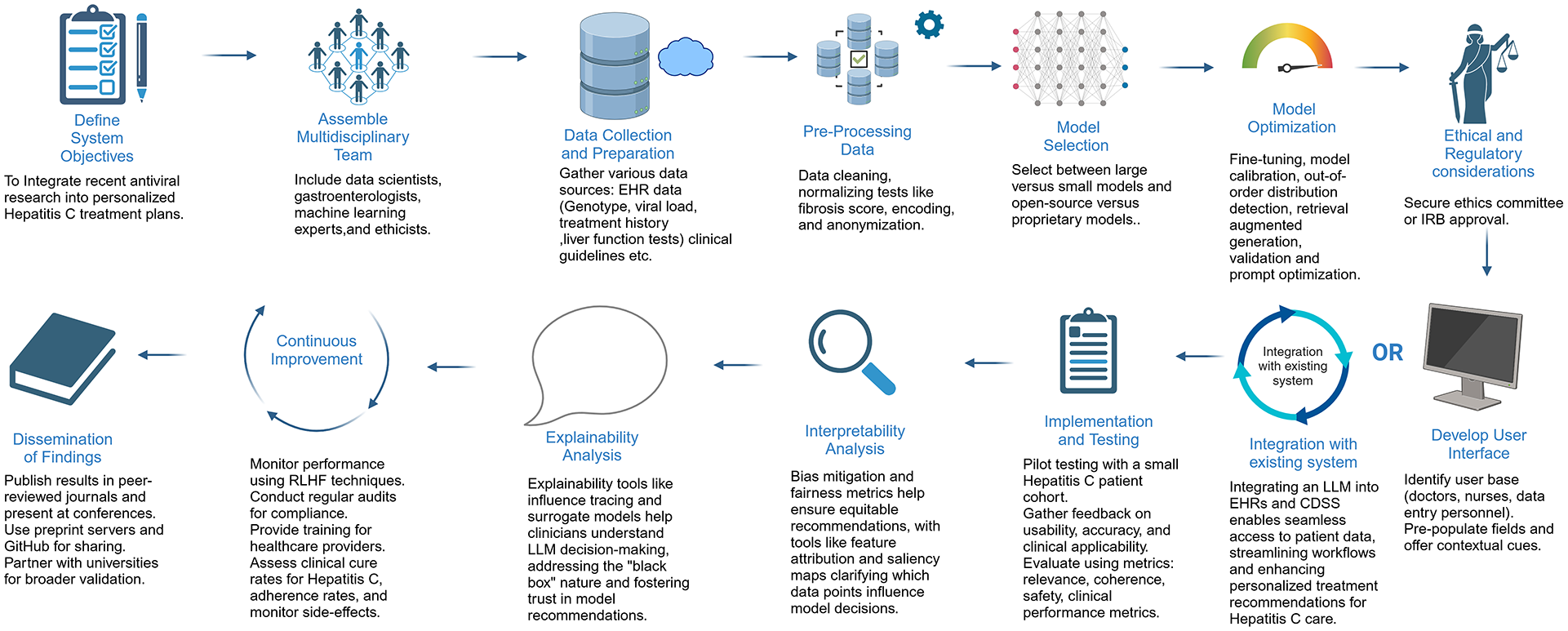
This figure illustrates a step-by-step framework for developing an LLM-powered clinical decision support system for Hepatitis C treatment. It outlines key stages, including ethical approvals, data collection, model selection, calibration, user interface development, integration with EHRs, and continuous improvement.
Framework
Define system objectives
Establishing clear goals for using an LLM model is essential. In our clinical scenario, the primary objective is to develop an AI-driven treatment recommendation system that generates personalized plans by incorporating genotype, viral load, fibrosis score, and prior treatment history. The model will integrate real-time clinical guidelines to ensure up-to-date recommendations while enhancing patient safety through risk stratification and optimized decision-making.
Assemble a multidisciplinary team
Developing an effective LLM for gastroenterology requires collaboration among AI specialists, data scientists, gastroenterologists, hepatologists, infectious disease experts, clinical informatics, quality assurance teams, bioethicists, and regulatory policymakers. 39 This team ensures the model is clinically accurate, seamlessly integrates into EHR systems, and complies with regulatory standards. 40
Data collection and preparation
Relevant data sources include EHR records (previous Hepatitis C treatments, patient demographics, alcohol use, diagnostic testing results, imaging, genotype data), clinical guidelines (e.g., American Association for the Study of Liver Diseases), medical literature, patient-reported outcomes, and regulatory and safety information (U.S. Food and Drug Administration approvals and adverse event reporting). Data must be adequate and representative of the intended population, as models fine-tuned on specific demographic groups may underperform in others. 38
Key preprocessing steps involve structured (e.g., HCV RNA levels, fibrosis scores) and unstructured data (e.g., clinical notes describing symptoms or treatment history) integration while ensuring Health Insurance Portability and Accountability Act (HIPAA)-compliant de-identification. Data cleaning is crucial to handle missing values, outliers, and inconsistencies. In pre-processing, structured data are converted to numerical forms, and unstructured data are processed using NLP techniques like tokenization and entity recognition to extract key medical concepts and map them to structured data, allowing the model to consider both empirical results and contextual patient history.
Following preprocessing, the dataset is divided into a training Set (70%–80%; for fine-tuning), a validation set (to prevent overfitting, which occurs when the model becomes overly specialized to the fine-tuning dataset, learning specific patterns that do not generalize well to new, unseen data), and test set (10%–15%).
Model selection
LLMs can be general purpose (GPT-4 and GPT-3.5), which are versatile but require domain adaptation and fine-tuned (BioGPT, UCSF-BERT, BioBert, PubMedBERT, etc.) which are tailored to specific applications41–44 (Table 2). In choosing between smaller models like Bidirectional Encoder Representations from Transformers (BERT) and large models like GPT-4, BERT should be preferred for real-time performance scenarios like bedside diagnostics due to faster inference times and lower communication costs.42,45 GPT-4 should be preferred for generating human-like text. For Hepatitis C, BERT can classify patient records and extract structured clinical information, while GPT-4 can generate personalized treatment recommendations based on multiple factors such as genotype, comorbidities, and prior treatments. 46 In our use case, a large model will be more suitable.
Pre-trained large language models.
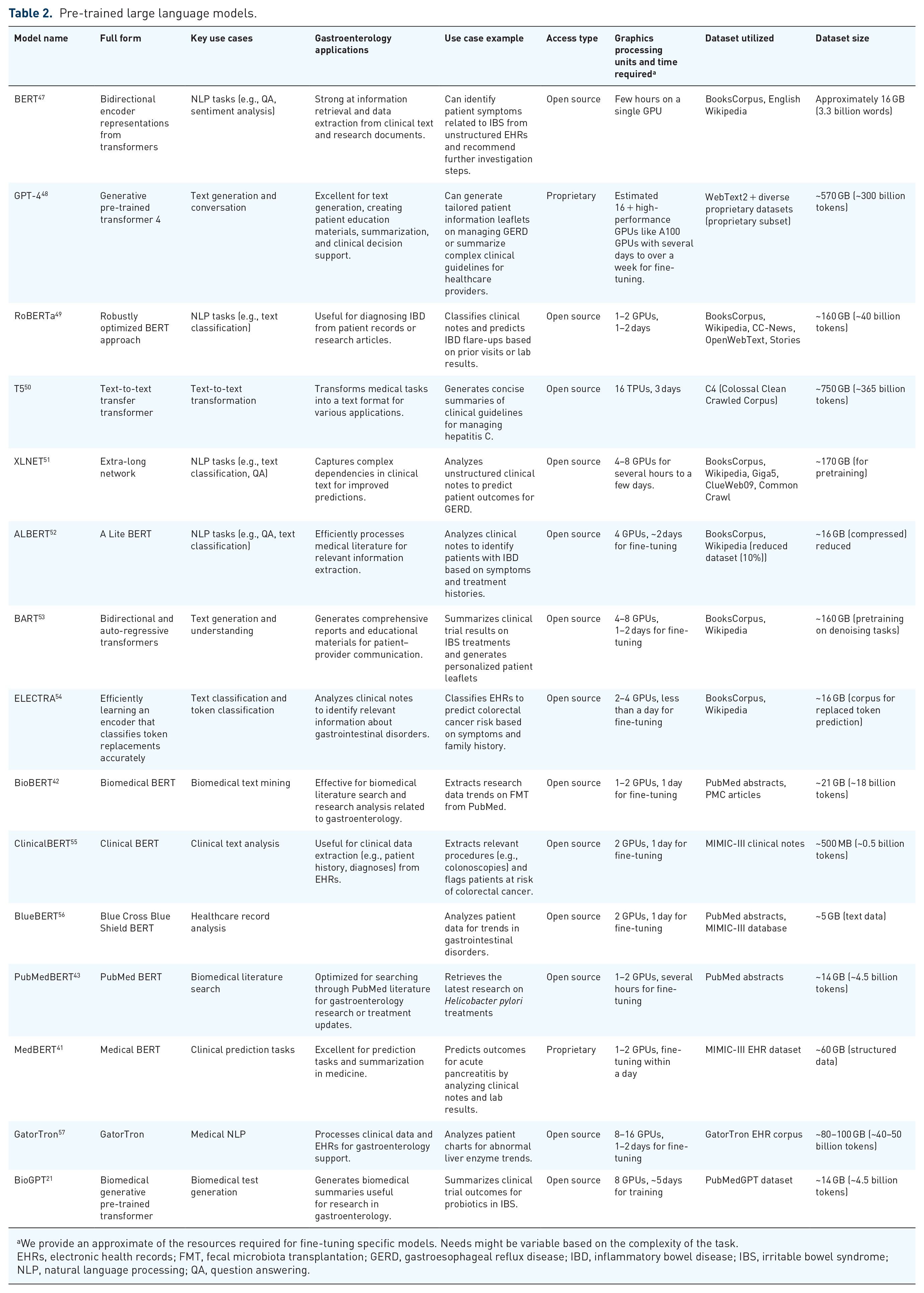
We provide an approximate of the resources required for fine-tuning specific models. Needs might be variable based on the complexity of the task.
EHRs, electronic health records; FMT, fecal microbiota transplantation; GERD, gastroesophageal reflux disease; IBD, inflammatory bowel disease; IBS, irritable bowel syndrome; NLP, natural language processing; QA, question answering.
Open-source models like BERT offer free access and customizability, allowing fine-tuning with medical data for tailored applications while ensuring data privacy when run locally, making them suitable for HIPAA compliance. However, they require significant computational resources and specialized expertise for setup and maintenance. By contrast, proprietary models like GPT-4 deliver state-of-the-art performance with minimal fine-tuning and seamless cloud-based scalability, making them ideal for healthcare deployment. 58 They benefit from continuous updates and easy API integration but come with high costs, limited control over model internals, and potential data privacy concerns when using cloud services. 59
Model optimization
Mitigating hallucinations and output variability
Hallucination occurs when an LLM generates incorrect or misleading information due to incomplete training data, overfitting, or lack of fact-checking. In clinical settings, this can lead to unsafe recommendations, such as incorrect Hepatitis C treatment guidance. 60 Cross-referencing outputs from trusted guidelines and developing internal layers that detect unsupported outputs can prevent hallucinations. Below, we discuss a few hallucination and output variability mitigation strategies.
Fine-tuning
Pretrained models learn from large, general datasets, making them broadly knowledgeable but lacking medical specificity. Fine-tuning adapts these models to clinical needs by training them on specialized datasets. 61 For Hepatitis C, fine-tuning ensures the model correctly interprets genotypes, fibrosis scores, and viral load and recommends AASLD-based treatment plans.
Fine-tuned models improve over time through active learning, where clinician feedback is used to enhance accuracy. This involves reviewing incorrect outputs, reinforcing correct responses, and periodically retraining the model with updated guidelines. Clinicians can actively flag incorrect recommendations, compare LLM outputs with expert decisions, and integrate the latest antiviral studies into the model’s learning process.
Model calibration and out-of-distribution detection
For gastroenterologists using LLMs in Hepatitis C treatment, ensuring that the model’s confidence aligns with real-world accuracy is critical. Model calibration adjusts confidence scores to prevent overconfidence in uncertain cases, helping clinicians gauge trust in recommendations. If an LLM suggests glecaprevir/pibrentasvir with 85% confidence, a well-calibrated model ensures this reflects an actual 85% likelihood of correctness. 62 Temperature scaling is a key calibration method that refines confidence levels by correcting systematic overconfidence. 63
Out-of-distribution (OOD) detection ensures the model recognizes when a case falls outside its training data, such as rare genotypic resistance mutations or Hepatitis C co-infections. 64 If the model flags a recommendation as uncertain or detects an OOD scenario, clinicians should seek additional expert input, refer to updated guidelines, or consider alternative antiviral options. By integrating calibrated confidence scores and OOD detection, LLMs become safer, more reliable tools for Hepatitis C decision-making.
Retrieval augmented generation
Unlike fine-tuning, which relies on static training data, RAG allows models to pull in real-time medical guidelines and research. This ensures treatment recommendations are based on the latest evidence without requiring constant retraining. 65 In Hepatitis C management, RAG is preferable over fine-tuning when real-time adaptability is needed, such as incorporating the latest AASLD/EASL guidelines, drug availability, insurance policies, and patient-specific factors from external sources. It allows the model to dynamically retrieve updated recommendations without retraining, making it ideal for rapidly evolving treatment landscapes and multi-center adaptability. 24 Fine-tuning is better suited for static, rule-based decision-making, such as structured genotype-based treatment selection. 66
Prompt optimization
Prompt optimization plays a crucial role in improving LLM responses without additional training. 67 Zero-shot prompts rely on the model’s general knowledge, such as asking, “What is the standard treatment for Hepatitis C genotype 1?” One-shot prompts provide an example before the query, improving contextual accuracy. Few-shot prompts offer multiple examples, refining model responses by demonstrating structured reasoning. Structuring prompts effectively—for instance, specifying “List first-line and second-line treatments for Hepatitis C based on fibrosis stage” rather than asking an open-ended question—reduces variability and enhances precision. To maintain consistency, techniques like controlled vocabulary use (ensures standardized medical terminology, such as using “F3 fibrosis stage” instead of the ambiguous “moderate liver scarring”), response formatting constraints (structure outputs predictably, like listing “Genotype → Preferred regimen → Treatment duration” for Hepatitis C therapy), and iterative prompt refinement (involves adjusting prompts to improve accuracy and completeness of responses) help reduce prompt sensitivity, ensuring reliable clinical decision support. 68
Validation
LLM variability should be validated through inter-clinician agreement studies and clinical decision support systems (CDSS) comparisons to ensure consistency with expert decisions and established protocols, ensuring stable and reliable clinical use. 69
Developing user interface/integrating with existing system
Designing an LLM interface for gastroenterology requires tailoring it to gastroenterologists, nurses, and administrative staff to ensure the interface aligns with clinical workflows and enhances usability.70,71
A structured input system with fields for patient ID, date of birth, and prior procedures (e.g., endoscopies, colonoscopies) ensures seamless data entry and accurate patient identification.
Pre-populating fields with recent lab results, imaging, and treatment history streamlines workflows, reducing administrative burden and minimizing errors. Key features for Hepatitis C decision support are outlined in Supplemental Table 1, with a wireframe of the interface in Figure 2.
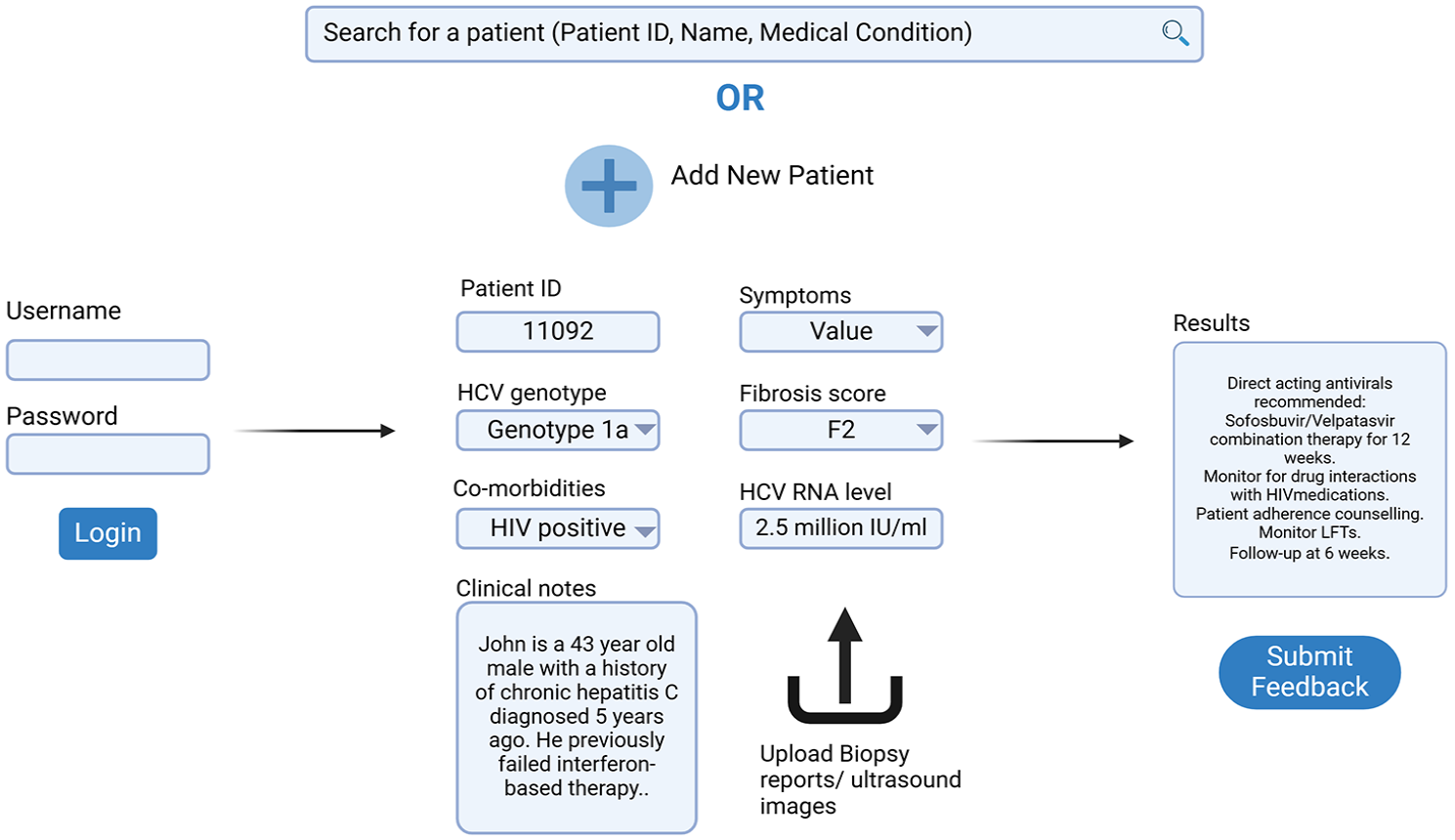
This figure presents a wireframe of the LLM-driven clinical decision support interface for Hepatitis C management. It demonstrates structured data entry fields for patient ID, genotype, fibrosis score, co-morbidities, and HCV RNA levels. The interface pre-populates critical fields, integrates biopsy/imaging uploads, and generates personalized treatment recommendations based on clinical guidelines. A feedback loop allows clinicians to refine model outputs, enhancing reliability and usability.
Integrating an LLM into existing clinical systems enhances efficiency and patient care by embedding AI-driven recommendations within EHRs, CDSS, and hospital management software. 72 Real-time EHR integration allows the model to analyze genotype, viral load, fibrosis scores, and treatment history without redundant data entry. When a clinician accesses a Hepatitis C patient’s record, the model provides tailored treatment suggestions directly within the EHR interface, minimizing disruptions and centralizing decision-making.
When embedded in a CDSS, the LLM serves as an advanced support layer, refining alerts and recommendations with context-aware insights.
Ethical and regulatory considerations
Approval from the ethics committee or institutional review board must be secured to ensure compliance with ethical standards. Adherence to the HIPAA in the United States and the General Data Protection Regulation (GDPR) in Europe is essential. 73 Patients do not typically provide consent waivers for using their de-identified data in model development.
Implementation and testing
After training the LLM on refined data and developing the user interface/integration with EHR, pilot testing with a small patient cohort under clinical oversight should be initiated. This stage is critical for gathering feedback from both patients and healthcare professionals, helping identify any challenges in usability, accuracy, and clinical applicability. Continuous feedback during pilot testing will help refine the model for real-world implementation.
Traditional evaluation metrics like accuracy, recall, precision, F1 score, and AUC-ROC are commonly used for predictive models; however, for LLMs generating personalized treatment plans, these metrics may not fully capture the model’s performance. Instead, assessment should focus on relevance (alignment with clinical expectations), coherence (logical consistency in recommendations), and safety (adherence to clinical guidelines). In addition, clinical performance metrics such as cure rates, treatment adherence, and incidence of side effects should be monitored to ensure the model provides actionable insights that improve patient outcomes.73,74
Interpretability and explainability analysis
Ensuring equitable recommendations is critical in gastroenterology, as Hepatitis C outcomes vary based on patient demographics such as age, sex, race, and socioeconomic status. Clinicians should be aware that LLMs can inherit biases from training data, leading to disparities in treatment recommendations.75–79 Bias detection tools, such as demographic parity analysis, can identify if certain patient groups receive different treatment suggestions. To mitigate bias, strategies like re-weighting underrepresented patient populations in training data or supplementing the model with diverse clinical trial data can help ensure fairer recommendations. For example, if a model consistently under-recommends direct-acting antivirals for certain racial groups due to historical underrepresentation in trials, re-weighting the data can correct this imbalance.80–82 As highlighted in recent GI literature on algorithmic bias, disparities in clinical trial participation and the unavailability of representative data can exacerbate these biases. 83
Transparency and explainability remain significant challenges for LLMs in healthcare, as LLMs do not provide traditional, step-by-step reasoning like clinical algorithms. 84 Feature attribution techniques can help clarify why the model suggested a particular regimen by highlighting key factors such as genotype, fibrosis score, or past treatment failures. 85 Saliency maps visually emphasize influential data points, such as “genotype 1a” or “prior failure with sofosbuvir,” making the rationale behind recommendations more intuitive. 86
Counterfactual explanations, on the other hand, enable clinicians to see how slight changes in input data, like assuming the patient was treatment-naïve, would alter the model’s output. 87 This helps clinicians understand the model’s decision boundaries and evaluate its relevance to the patient’s specific circumstances.
Understanding how the model was trained is equally important. If most recommendations favor sofosbuvir-based regimens, it may be because training data were heavily drawn from clinical trials emphasizing these therapies.
In cases where interpretability remains a challenge, simplified surrogate models can approximate the decision-making process of complex LLMs, offering clinicians a more intuitive pathway to understanding AI-driven recommendations. 88 For example, a decision tree could outline how viral load and past treatment history influenced the suggested therapy, providing a straightforward map of the model’s logic.
Continuous improvement
Reinforcement learning from human feedback (RLHF) helps fine-tune responses by incorporating real-world patient outcomes and clinician inputs, ensuring the model remains relevant. 89 Continuous monitoring of a model’s performance in real-world settings is essential. Regular audits, including external audits by independent bodies, help maintain compliance with regulatory requirements post-deployment. Periodic reviews of the model’s impact on patient outcomes and healthcare practices ensure it continues to provide benefits without unintended negative consequences.
Training gastroenterologists on LLMs ensures safe and effective integration into clinical workflows. Awareness of limitations is crucial, including potential hallucinations, outdated or biased training data, lack of clinical nuance, and response variability. Best practices include cross-checking AI recommendations with guidelines, assessing confidence scores, using structured prompts, and engaging in feedback loops to refine accuracy. Transparent patient communication about AI’s role reinforces that LLMs support, rather than replace, clinical judgment.
Dissemination of findings
Disseminating AI model findings through peer-reviewed publications, conferences, and preprint servers fosters collaboration and knowledge sharing. Hosting code and data on GitHub ensures transparency, while partnerships with universities and research institutions support clinical validation and refinement. This approach promotes reproducibility, adoption, and continuous improvement in real-world settings.
Discussion
Since AI models are mostly used as CDSS in healthcare settings instead of autonomous interventions, the framework utilized for their deployment plays a key role in impacting patient outcomes.28,90 While certain in silico studies have demonstrated that AI algorithms can match the efficacy of clinicians, there is a lack of convincing evidence suggesting a positive impact on patient outcomes and clinical efficacy.91–93 Bridging this gap requires optimizing the translation process from in silico to real-world settings, with a focus on maintaining feedback loops, continuous learning, and considering human factors and safety.94,95 Regulatory compliance with HIPAA, GDPR, and guidelines like DECIDE-AI, SPIRIT-AI, and CONSORT-AI to ensure responsible deployment and monitoring is essential.94,96–98
LLMs face bias, ethical concerns, susceptibility to cyberattacks, hallucinations, and output variability, often struggling with edge cases, evolving guidelines, and patient variability. 99 Adoption barriers include limited clinician training, workflow integration challenges, and concerns about automation in decision-making. Without continuous updates, LLMs risk stagnation, reducing their long-term utility. Our framework addresses these gaps through fine-tuning with diverse datasets and RAG for real-time adaptability, confidence scoring, calibration, human-in-the-loop oversight, and structured response formats to mitigate errors. Training gastroenterologists in LLM interpretation rather than automation ensures responsible use, while regular audits, RLHF, and iterative updates support long-term accuracy.
In our framework, clinicians are involved from model optimization to implementation, ensuring safe and effective LLM integration in gastroenterology. During model optimization, gastroenterologists contribute by validating fine-tuned outputs, refining prompts for accuracy, and cross-referencing AI-generated recommendations with clinical guidelines. In calibration and hallucination mitigation, they help assess confidence scores, detect errors, and provide real-world feedback to improve model reliability. During implementation, clinicians facilitate EHR integration, ensuring AI recommendations align with workflow needs while participating in pilot testing to evaluate usability and clinical relevance. In continuous improvement, they engage in RLHF refinements, audit LLM performance, and advocate for necessary updates based on evolving Hepatitis C treatment protocols. By actively participating at each stage, gastroenterologists play a crucial role in ensuring LLMs function as reliable, evidence-based decision-support tools in clinical practice. 100
Despite these measures, real-world validation of LLMs remains in its early stages, and regulatory pathways for AI-based clinical decision tools are still evolving. 101 Scaling multi-institutional validation, clinician-AI collaboration models, and AI-assisted clinical trials will be essential in establishing LLMs as reliable tools in gastroenterology. 102
Supplemental Material
sj-docx-1-tag-10.1177_17562848251328577 – Supplemental material for Utilizing large language models for gastroenterology research: a conceptual framework
Supplemental material, sj-docx-1-tag-10.1177_17562848251328577 for Utilizing large language models for gastroenterology research: a conceptual framework by Parul Berry, Rohan Raju Dhanakshirur and Sahil Khanna in Therapeutic Advances in Gastroenterology
Footnotes
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
