Abstract
Background:
Deep learning techniques can accurately detect and grade inflammatory findings on images from capsule endoscopy (CE) in Crohn’s disease (CD). However, the predictive utility of deep learning of CE in CD for disease outcomes has not been examined.
Objectives:
We aimed to develop a deep learning model that can predict the need for biological therapy based on complete CE videos of newly-diagnosed CD patients.
Design:
This was a retrospective cohort study. The study cohort included treatment-naïve CD patients that have performed CE (SB3, Medtronic) within 6 months of diagnosis. Complete small bowel videos were extracted using the RAPID Reader software.
Methods:
CE videos were scored using the Lewis score (LS). Clinical, endoscopic, and laboratory data were extracted from electronic medical records. Machine learning analysis was performed using the TimeSformer computer vision algorithm developed to capture spatiotemporal characteristics for video analysis.
Results:
The patient cohort included 101 patients. The median duration of follow-up was 902 (354–1626) days. Biological therapy was initiated by 37 (36.6%) out of 101 patients. TimeSformer algorithm achieved training and testing accuracy of 82% and 81%, respectively, with an Area under the ROC Curve (AUC) of 0.86 to predict the need for biological therapy. In comparison, the AUC for LS was 0.70 and for fecal calprotectin 0.74.
Conclusion:
Spatiotemporal analysis of complete CE videos of newly-diagnosed CD patients achieved accurate prediction of the need for biological therapy. The accuracy was superior to that of the human reader index or fecal calprotectin. Following future validation studies, this approach will allow for fast and accurate personalization of treatment decisions in CD.
Introduction
Capsule endoscopy (CE) is a prime modality for diagnosis and follow-up in Crohn’s disease (CD). Complete evaluation of the entire small bowel using CE or cross-sectional modality is recommended in all newly-diagnosed CD patients, or patients in need of re-evaluation for non-explained symptoms.1,2 CE is an accurate predictor of pending relapse in CD patients in clinical remission, 3 and can change disease classification in over 50% of CD patients. 4 Location of the lesions is of importance as well, as jejunal CD was demonstrated to be associated with worse long-term prognosis and higher risk of surgery. 5
Artificial intelligence (AI) and deep learning techniques are being rapidly implemented in both research and clinical practice gastroenterology, mostly for real-time polyp detection during colonoscopy.6–8 Additional fields for evaluation of AI in Gastrointestinal tract (GI) assist in early detection of gastric neoplasia,9,10 Barrett’s esophagus, 11 endoscopic ultrasound,12,13 and grading of mucosal inflammation in ulcerative colitis.14–18
An additional field with fast development of AI research is CE, with several publications evaluating deep learning for automated detection of inflammatory lesions,19–26 vascular lesions,27,28 protruding lesions and masses, 29 and scoring of bowel cleanliness. 30 The accuracy for most of these tasks was reported to be at least 90% and frequently much higher.
However, to date the AI research in GI was mostly focused on identification of individual lesions, and this goal seems to be quite successfully achieved in luminal endoscopy. However, the bigger and perhaps more important challenge is the potential utilization of AI for prognostication and personalization of treatment approach. Such evidence is still extremely scarce.
In CD, predictors of disease prognosis and response to treatment are still severely lacking.31–33 The impact of mucosal healing and the lesion size on disease outcomes in CD has been well described.34,35 It is likely that the vast abundance of visual data contained in the complete videos of endoscopic procedures such as CE (containing up to 12,000 still images per film) may contain factors and subtle findings that escape the human eye, however, could be picked by AI algorithms. We hypothesized that AI analysis of CE videos performed at diagnosis could be an accurate predictor of treatment outcomes.
The main aim of our study was to evaluate the utility of AI analysis of complete CE videos of CD patients at inception for prediction of the need for biological therapy.
Methodology
Cohort definition
This was a retrospective cohort study. The reporting of this study conforms to the Strengthening the Reporting of Observational Studies in Epidemiology (STROBE) statement. In this study, we included CD patients followed by the gastroenterology department of Sheba Medical Center between January 2011 and March 2021. Only patients who performed CE within 6 months of diagnosis and before initiation of immunomodulator or biological therapy were included. Patients with less than 6 months of follow-up after CE were excluded. Demographic, clinical, and endoscopic data were extracted from patients’ electronic medical records.
Capsule endoscopy
All CE procedures were performed using PillCam SB2 or SB3 small bowel capsules (Medtronic, Yokneam, Israel) and reviewed using the Reader software version 8 or 9 (Medtronic, Yokneam, Israel). The CE videos were graded using the Lewis score (LS). 36 We extracted complete small bowel videos starting with the first duodenal image and ending with the first cecal image using the RAPID Reader software v9.0 (Medtronic, Yokneam, Israel). The videos were saved in MPEG format.
Study outcome
The study outcome was the initiation of biological therapy within the duration of follow-up.
Machine learning algorithm
Machine learning analysis was performed by Intel Israel. For analyzing videos, we used the TimeSformer, Facebook Research, Menlo Park, California architecture. 37 An overview of the TimeSformer architecture is presented in Figure 1. This is a state-of-the-art algorithm introduced by Facebook. It is based on the Transformer paradigm from the natural language processing domain37,38 (Supplemental Section A).

An overview of the TimeSformer architecture. On the right side: the video self-attention block, applied on the embedded patches. On the left side: the end-to-end architecture. A multilayer perceptron (MLP) is applied both at the end of each block and to the projected and concatenated vectors from all heads.
We used a fivefold cross-validation train-test random split: in each split four folds were used as the train set and the fifth fold was used as the validation set. The raw training videos were augmented (Supplemental Table 1) after the train-test split.
Detailed hyperparameters of the experiments are presented in the supplements. Three TimeSformer models were experimented upon: FT K400, 8 × 32 × 224; FT K600, 8 × 32 × 224; and FT K400, 16 × 16 × 448 (Supplemental Section B).
Model interpretability
To understand the influence of spatial and temporal features, respectively, we used Intel Labs’ InterpreT toolbox. This toolbox allows us to investigate the model behavior and understand its internal structure (Supplemental Section C).
Statistical methods for clinical data
Descriptive statistics were presented as means ± standard deviations/medians with interquartile ratios (IQRs) for continuous variables and percentages for categorical variables. Continuous variables were analyzed by t-test/Mann–Whitney test and categorical variables by chi-square/Fisher’s exact test. A receiver operator curve (ROC) analysis was performed for the models, and LS and fecal calprotectin predictions of the study outcome were analyzed. A two-tailed p-value < 0.05 was considered statistically significant. The analysis was performed using IBM SPSS statistics (version 27.0) (Armonk, NY, USA).
Results
Cohort description
One hundred and one patients were recruited. Clinical and demographic characteristics of the patients appear in Table 1. The median LS was 450 (IQR 225-1012).
Clinical and demographic characteristics of the included patients.

CD, Crohn’s disease; IQR, interquartile ratio.
Thirty-seven patients (36.8%) initiated biological therapy within the duration of follow-up with a median time of 17 months (4.7–38.5) for therapy. The initiated biological treatments included adalimumab (18, 48.6%), infliximab (13, 35.1%), or vedolizumab (6, 16.2%).
On univariate analysis, CD phenotype, LS, and fecal calprotectin were associated with initiation of biological therapy; however, on multivariate analysis, none retained significance (Table 2). For LS, the AUC for predicting biological treatment was 0.70 and for calprotectin 0.74.
Comparison of clinical characteristics between patients that did or did not initiate biological therapy within the duration of follow-up.

CD, Crohn’s disease; IQR, interquartile ratio.
Machine learning results
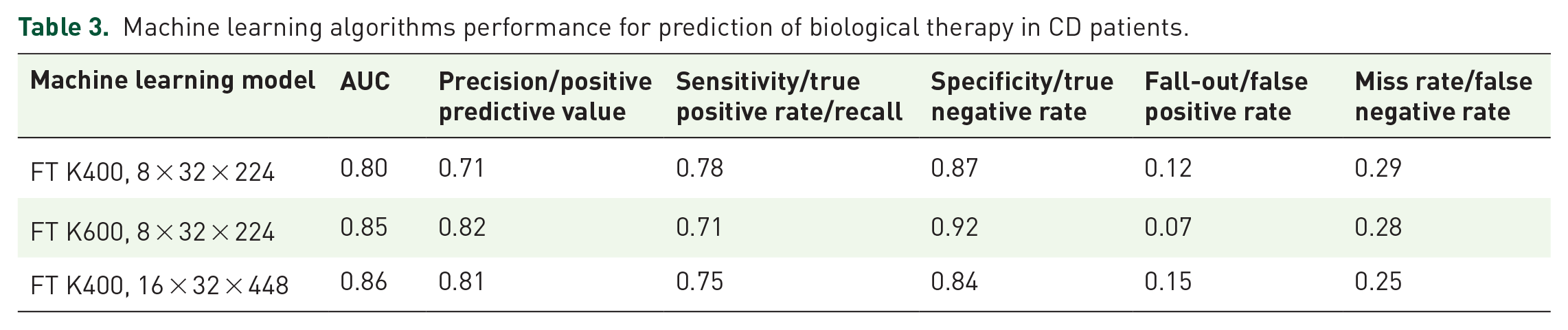
The optimal model achieved AUC 0.86 with accuracy 0.81. Below is a summarization of the TimeSformer experiments with respect to the different model configurations (Table 3, Figure 2).
Machine learning algorithms performance for prediction of biological therapy in CD patients.

The ROC AUC plots for models FT K400 8 × 32 × 224, FT K600 8 × 32 × 224, and FT K400 16 × 32 × 448, respectively.AUC, Area under the ROC Curve.
Model interpretability
Hidden state representations in a two-dimensional space
Supplemental Figure 1 shows the t-distributed stochastic neighbor embedding (t-SNE) representations of the CLS token of each video. This figure shows how positive and negative videos are clustered differently throughout the network’s layers. Especially, two distinct clusters are seen at the last layer.
Then, we focused on the t-SNE representation of the regions of each frame. In Figure 3, pixel id represents the same region in each image, and example ID represents the video level.
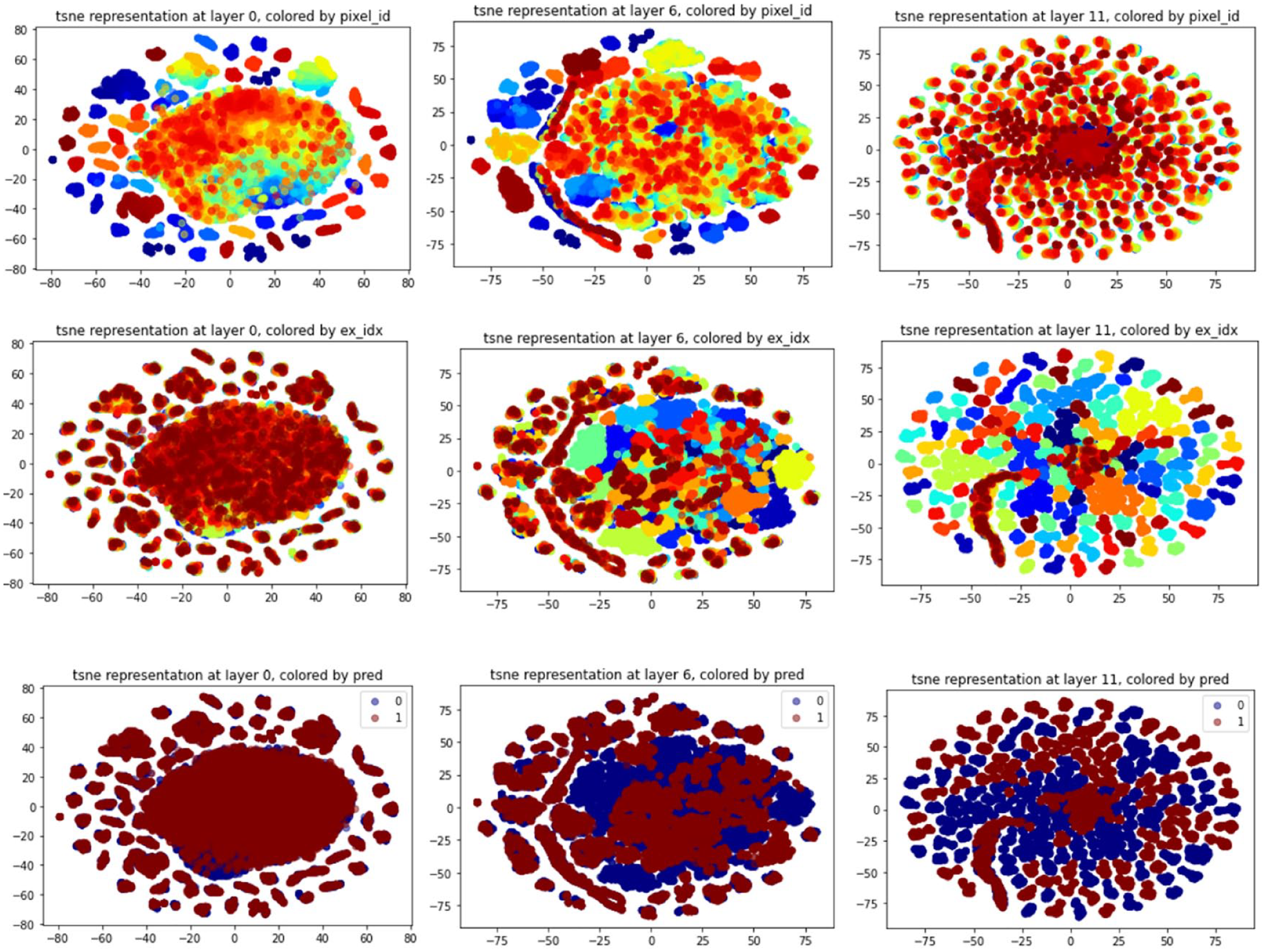
The evolved clustering of the frame regions after t-SNE embedding through the layers. The first row of the figure shows the evolution of the regions’ representation colored by the pixel id while the second row shows this evolution colored by the example ID. These figures show that in the first layers, the clusters are location-specific: pixels of the same part of any different frames will be closer. However, at the end of the network, the clusters are example related. This implies that location embedding is represented in the first layers. In contrast, prediction is at the example level and not at region one, and it is represented in the last layers of the network. The third row shows that there is a separation between positive and negative videos in the last layer, while at the first layers, they were completely mixed and overlapping.
Attention heatmap for ulcer detection
When comparing our results to ResNet101 Convolutional Neural Network classifier trained on a set of colonoscopy images, the TimeSformer attention mechanism was better focused on the ulcers and their localization (Figure 4), as seen in the Grad-CAM heatmaps.

Ulcer detection – comparison of Grad-CAM visualizations to attention head visualizations to the CLS token in the spatial dimension. As can be seen, in the first two examples, the Grad-CAM method succeeds in detecting the lesions but with a diffused heatmap that might not be precise enough. The second two examples show how Grad-CAM also fails to detect ulcers correctly. Focusing on the attentions to the CLS token in the spatial dimension at the head (layer = 3, head = 8), we can see how the model indirectly learned to prioritize the ulcers. Compared to the Grad-CAM heatmaps, we can see how the detection is better focused on the ulcers, and we also see that even when Grad-CAM fails to detect the lesion, our model and this specific attention head succeed in localizing it.
Discussion
Our study demonstrates that machine learning analysis of complete CE videos of CD patients performed at initial diagnosis can accurately predict the need for biological therapy. The performance of TimeSformer algorithm was superior to that of the traditional LS or of fecal calprotectin.
In the recent years, AI algorithms achieved excellent accuracy in identification of findings on still images in multiple fields in gastroenterology, including CE.19,20,22,26,39,40 Moreover, real-time diagnosis of colonic polyps on colonoscopy is currently possible and was demonstrated to be associated with an improved polyp detection rate6–8; AI technology for colonoscopy has already matured into clinically available and FDA-approved tools such as GI Genius (Medtronic, Dublin, Ireland).
However, it is quite plausible that the full-fledged potential of the use of AI in Inflammatory bowel disease (IBD) is yet to be realized. Prediction of disease course and outcomes is among the crucial challenges in CD; despite the massive abundance of research targeting personalization of treatment approach, the practical implication for the daily practice is still scarce.31,41–43 One of the cornerstones of the modern treatment approach in IBD is the achievement of mucosal healing1,2,44; striving for complete mucosal healing in IBD is well justified as patients in mucosal healing have consistently better long- and short-term disease outcomes. 45 CE is a valuable diagnostic and monitoring tool in CD1,2 as it allows for accurate panenteric assessment of bowel inflammation. In established CD patients, proximal disease location is positively associated with a risk of intestinal resection. 5 CE identifies mild to moderate mucosal inflammation in over 80% of CD patients in clinical remission, including the majority of patients in both clinical and biomarker remission 46 ; such smoldering inflammation was an accurate predictor of relapse within 24 months. 3
An average complete CE video may include up to 10,000 images. The existing indices for grading inflammation on CE36,47–52 are based on a small number of semiquantitative parameters such as the number of ulcers (none/few/multiple), disease of ulceration (based on eye-balling of the extent), and presence of structures. Clearly, only a small amount of the available optical information is incorporated and processed in any of these scores. Thus, AI analysis of the entire available visual data may hold additional promise for the complete assessment of mucosal inflammatory burden. However, we assumed that not only the complete extent and morphology of the ulcerations and strictures may be of prognostic importance but the anatomical distribution of the pathologies, which can only be gauged by the temporal location in relation to the first and last images, should also be of importance. Thus, a mere segmentation of videos to individual still images may not have been sufficient for the purpose.
Our approach is unique in that it classifies videos using both space and time features. To this end, we capture the sequential context by learning spatiotemporal features from the video capsule data, giving balanced representations to the local pixel structure as well as relationships across the video frames in time. TimeSformer algorithm was developed in order to capture and identify motion, such as sports activity, so we assumed that could be well suited to address the task of spatiotemporal video analysis.
Our study has several limitations. Primarily, a larger sample size could have resulted in improved accuracy. For classification, we used a relatively ‘soft’ outcome of biological initiation. It is clear that the decision to initiate biological therapy is very heterogenous and could be influenced by a multitude of parameters such as patient and physician preferences, extraluminal disease characteristics (perianal disease, extraintestinal manifestations, etc.). It would have been preferable to select a less-biased outcome such as need for surgery; however, such a cohort would be almost impossible to compile as patients with intestinal strictures are usually excluded from performing CE after evaluation of small bowel patency. In our cohort that included patients followed in a large tertiary center for 9 years, only one patient required surgery. In addition, our cohort included patients with relatively mild disease, as signified by LS and fecal calprotectin levels. In addition, our study focused on prediction of the outcome and did not suggest an algorithm for mucosal burden quantification. Such an algorithm would be of great practical utility and should be investigated in future studies.
Our current effort lays the groundwork for the creation of automated AI-based predictive systems for CE in CD. In fact, completely automated reading of AI can be easily feasible with the currently available tools for detection of pathologies, as they have already matured for excellent accuracy; nonetheless the ultimate goal of any diagnostic tool in early CD would be to predict the treatment outcome and the risk of complications, and for that purpose it appears that a spatiotemporal algorithmic approach that is able to incorporate disease distribution and location as well as the potential impact on GI motility is capable of catering to the task. The current algorithm requires further validation in extensive prospective cohorts, either as a standalone solution or as a component in a multiomic predictive model potentially incorporating multiple biological and visual inputs.
Supplemental Material
sj-docx-1-tag-10.1177_17562848231172556 – Supplemental material for Spatiotemporal analysis of small bowel capsule endoscopy videos for outcomes prediction in Crohn’s disease
Supplemental material, sj-docx-1-tag-10.1177_17562848231172556 for Spatiotemporal analysis of small bowel capsule endoscopy videos for outcomes prediction in Crohn’s disease by Raizy Kellerman, Amit Bleiweiss, Shimrit Samuel, Reuma Margalit-Yehuda, Estelle Aflalo, Oranit Barzilay, Shomron Ben-Horin, Rami Eliakim, Eyal Zimlichman, Shelly Soffer, Eyal Klang and Uri Kopylov in Therapeutic Advances in Gastroenterology
Footnotes
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
