Abstract
Background:
The clinical relevance and biological role of tissular AOC4P in gastric cancer (GC) remains to be clarified.
Methods:
The association between AOC4P expression and clinicopathological characteristics was investigated. In vitro, 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide (MTT), colony formation, wound healing and terminal deoxynucleotidyl transferase dUTP nick end labeling (TUNEL) assays were performed to explore the biological effects of AOC4P on GC cell proliferation, migration, invasion, and apoptosis in MGC-803 and BGC-823 cell lines. In vivo, animal experiments were conducted to confirm the in vitro findings. Quantitative real-time polymerase chain reaction, western blotting, and immunofluorescence were used to investigate the potential mechanisms.
Results:
Expression levels of AOC4P were significantly higher in tumor tissues than in noncancerous tissues, and patients with high levels of AOC4P had poor overall and disease-free survival. AOC4P expression was correlated with lymphovascular invasion. In vitro, knockdown of AOC4P inhibited tumor cell proliferation, migration, and invasion, and promoted apoptosis of MGC-803 and BGC-823 cells. In vivo, BGC-823 cells transfected with AOC4P siRNA formed smaller and lighter tumors than BGC-823 cells transfected with negative control siRNA in severe combined immunodeficiency mice. Additionally, the si-AOC4P group had less proliferating cells and more apoptotic cells in tumor xenografts compared with the negative control. Mechanistically, knockdown of AOC4P decreased the expression of vimentin and MMP9, while increasing the expression of E-cadherin. Immunofluorescence confirmed the relationship between AOC4P expression and E-cadherin, vimentin, and MMP9 levels in clinical GC specimens.
Conclusions:
AOC4P promotes tumorigenesis and progression partly through epithelial–mesenchymal transition in GC. Additionally, AOC4P may serve as a prognostic biomarker for clinical decision making.
Background
Gastric cancer (GC) is a heterogeneous disease with an estimated 5-year overall survival of 27.4% in China. 1 Current approaches for GC management largely depend on multimodal therapeutic strategies including gastrectomy, chemotherapy, and chemoradiotherapy in perioperative settings. However, 25–40% of GC patients have recurrence after treatment.2–4 Hence, more research that focuses on the molecular mechanisms promoting cancer progression is needed, which would aid in the discovery and development of effective diagnostic biomarkers and therapeutic targets for GC and thus provide patients with potentially better outcomes.5,6
Long noncoding RNA (lncRNA) is a class of RNAs of 200 nucleotides in length without a protein-coding ability. Sequencing technologies have shown that only ˂2% of transcripts transcribed from the human genome code for proteins,7,8 leaving much of the noncoding transcripts unexplored. Recent studies have revealed that lncRNAs are associated with GC tumorigenesis and metastasis, and have the potential to serve as diagnostic and prognostic biomarkers.9,10 For example, our previous study established a novel five plasma lncRNA-based panel [terminal differentiation-induced noncoding RNA (TINCR), CCAT2, AOC4P, BRAF-activated noncoding RNA (BANCR) and LINC00857] that discriminates GC from precancerous individuals with relatively high accuracy compared with widely used serum carcinoembryonic antigen, CA19-9, and CA125. 11 However, the molecular mechanism of these lncRNAs in GC initiation and development need to be clarified further.
In the present study, we investigated the clinical relevance and biological role of AOC4P in GC, as the role of TINCR, CCAT2, BANCR and LINC00857 in GC has been previously reported.
Methods
Tissue specimens
GC tissues and adjacent normal tissues were collected from 63 patients who underwent surgery between January 2013 and December 2013 at the Department of General Surgery, Chinese PLA General Hospital. All patients were diagnosed by pathology. None of the patients had received preoperative chemotherapy or radiochemotherapy. Patient characteristics were obtained, including age, sex, T stage, lymph node status, tumor size, tumor differentiation, and TNM (tumor-node-metastasis) stage according to the 7th edition American Joint Committee on Cancer Staging manual. Patients were followed up every 6 months. Patients with suspicion of recurrence were assessed by computed tomography. The last follow-up time was May 2017. Disease-free survival and overall survival times were calculated. All patients provided written informed consent about their tumor specimen for research use. The collection and use of patient’s specimen was approved by the Ethics Committee of the Chinese PLA General Hospital (NO.S2016-057-01).
Cell lines and culture
Human GC cell lines MGC-803 and BGC-823 were purchased from the Chinese Academy of Sciences Committee on Type Culture Collection cell bank (Shanghai, China). The immortalized human gastric epithelial cell line GES-1 was obtained from the Institute of General Surgery at the Chinese PLA General Hospital. The cell lines were cultured as described previously. 11
RNA extraction and quantitative real-time polymerase chain reaction
RNA was extracted from tissues and cultured cells using Trizol reagent (Invitrogen, Carlsbad, CA, USA), according to the manufacturer’s protocol. RNA concentrations and purity were measured by a NanoDrop 2000/2000c spectrophotometer (Thermo Fisher Scientific, Wilmington, USA). cDNA was synthesized from 3 μg extracted RNA using a reverse transcription kit (Invitrogen). Quantitative real-time polymerase chain reaction (qRT-PCR) was performed as described previously. 11 Primer sequences are shown in the supplementary files.
Western blot assay
Western blot assays were performed as described previously. 12 In brief, extracted proteins from tissues and cell lines were separated by sodium dodecyl sulfate polyacrylamide gel electrophoresis and then transferred to polyvinylidene fluoride membranes (Bio-Rad Laboratories, USA). After blocking, the membranes were incubated with a primary antibody overnight at 4°C. Then, the blotted membranes were incubated with a horseradish peroxidase-conjugated secondary antibody (1:2000) for 2 h at room temperature. Labeled proteins were detected using enhanced chemiluminescence following the manufacturer’s protocol. β-Actin (1:1000, Cell Signaling, USA) was used as an internal control. Antibodies against the following proteins were used: E-cadherin (1:1000, Cell Signaling), matrix metalloproteinase-9 (MMP-9; 1:1000, Abcam, USA), vimentin (1:1000, Cell Signaling), cleaved caspase-3 (1:1000, Cell Signaling) and cleaved poly (ADP-ribose) polymerase (PARP; 1:1000, Cell Signaling).
Immunohistochemistry
Immunohistochemistry (IHC) was performed using a standard technique with an avidin-biotinylated peroxidase complex as described previously.12,13 Sections were incubated with an anti-Ki-67 antibody (1:400, Cell Signaling) at 4°C overnight. Diaminobenzidine (DAKO, China) staining was used to detect immunoreactivity. The intensity of immunoreactivity was graded as 0, 1+, 2+, and 3+ for no staining, weak, medium, and strong staining, respectively. Scores of 0 and 1+ were regarded as low expression, while scores of 2+ and 3+ were considered as high expression. The proliferation index of the cancer cells = high expression cells/total cells × 100%.
Immunofluorescence staining
The 5 μm-thick, formalin-fixed, paraffin-embedded tissue sections were incubated with a primary antibody at 4°C overnight. Then, the sections were rinsed three times for 5 min each with phosphate-buffered saline (PBS) followed by incubation with Alexa Fluor-conjugated secondary antibodies at room temperature for 1 h. Fluorescence imaging was performed using a laser scanning confocal microscope (Fluoview FV1000, Olympus, Japan). Fluorescence staining was quantified using Tissue-Quest software (TissueGnostics GmbH). Tumor tissues were classified as high or low expression using a cutoff of the mean expression level of proteins (high expression ⩾ mean; low expression < mean). Antibodies against the following proteins were used: E-cadherin (1:100, Cell Signaling), MMP-9 (1:500, Abcam), and vimentin (1:100, Cell Signaling).
Colony formation assay
A total of 500 cells per well were seeded in a six-well plate in triplicate and maintained in a humidified atmosphere containing 5% CO2 at 37°C. After culture for 10–14 days, cell colonies were washed with PBS, fixed with 4% paraformaldehyde for 30 min, and stained with a 0.1% crystal violet solution for 20 min. Colonies containing more than 50 cells were counted.
Proliferation assay
Proliferation of cells was measured by MTT assays using Cell Proliferation Reagent Kit I (Roche Applied Science, USA), according to the manufacturer’s protocols. A total of 3 × 103 cells/well transfected with the indicated vector were seeded in a 96-well flat-bottomed plate and cultured in normal medium for 24 h. At 0, 24, 48, 72 and 96 h after transfection, MTT solution (5 mg/ml, 20 µl) was added to each well. The relative number of surviving cells was assessed by measuring the optical density of cell lysates at 560 nm. For each treatment group, cells were assessed in triplicate.
Cell migration and invasion assays
Cell migration was measured using a Transwell chamber with an 8 μm pore size membrane according to the manufacturer’s instructions. In brief, 3 × 105 cells in 200 μl serum-free medium were added to the upper chamber. For the invasion assay, 5 × 105 cells in 200 μl serum-free medium were added to the upper chamber coated with 1 mg/ml Matrigel. Subsequently, 500 μl serum-containing medium was added to the lower chamber. Cells were incubated at 37°C for 24 h, and then cells on the upper surface of the membrane were scraped off with cotton swabs. Cells that had migrated and invaded to the lower surface of the membrane were fixed and stained with a 0.1% crystal violet solution. Four random microscopic fields of the membrane were photographed, and cells were counted for statistical analysis.
Wound healing assay
A total of 5 × 105 cells per well were seeded in six-well plates and cultured until 90% confluence. A 200 μl sterile pipette tip was used to make a straight scratch on the culture surface. Detached cells were washed off gently, and images of the scratch were photographed as a baseline. The medium was then replaced, and images of the same location were obtained under a microscope after 48 h. The healing rate was calculated as follows: (Widthbaseline − Width48h)/Widthbaseline.
Flow cytometric analysis
Cells were harvested, washed with PBS and fixed overnight in 4% formaldehyde at −20°C. Then, the cell was stained with propidium iodide using cell cycle kit (BD Biosciences, NJ, USA) according to the manufacturer’s instructions. The cells were analyzed by FACScan (BD Biosicences, Franklin Lakes, NJ, USA).
In vivo tumorigenicity
Animal experiments were conducted in accordance with the recommendations in the Guide for the Care and Use of Laboratory Animals of the National Institutes of Health. Animal experiments were approved by the Animal Care and Use Committee of Chinese PLA Hospital. The 4-week-old severe combined immunodeficiency mice were maintained under specific pathogen-free conditions. BGC-823 cells stably transfected with AOC4P siRNA (si-AOC4P) or the empty vector were harvested and washed. Then, 5 × 106 cells transfected with si-AOC4P or the empty vector were injected subcutaneously into the left and right flanks of each mouse, respectively. Tumor volumes were measured by ultrasound every 7 days, and the mice were euthanized after 4 weeks. Tumor volumes were calculated by the following formula 14 : (width2 × length)/2.
Terminal deoxynucleotidyl transferase dUTP nick end labeling assay
Tumor xenografts were fixed in 4% formalin and embedded in paraffin. An in situ terminal deoxynucleotidyl transferase dUTP nick end labeling (TUNEL) kit (Roche Applied Science) was used to detect cell apoptosis in the implanted tumors according to the manufacturer’s protocol. Apoptotic cells and the total number of cells in five random fields were counted in each group. The apoptotic index of the cancer cells = apoptotic cells/total cells × 100%.
Statistical analysis
Data are expressed as the mean ± standard deviation (SD) and were analyzed with SPSS software version 22.0. Continuous variables were analyzed using an independent Student’s t-test or paired t-test. Discrete variables were compared using the Chi-square test or Fisher’s exact test. Kaplan–Meier plots were used to analyze overall and disease-free survival that was compared by the log-rank test. Univariate and multivariate Cox regression analysis was performed to investigate the prognostic factors. A two-sided p value of less than 0.05 was considered as statistically significant.
Results
AOC4P is upregulated in GC cell lines and tumor tissues
To investigate the expression levels of AOC4P in GC, we performed qRT-PCR in two GC cell lines (MGC-803 and BGC-823) and a human gastric epithelial cell line (GES-1). Consistent with our previous study, as shown in Figure 1(a), AOC4P was highly expressed in GC cell lines compared with the gastric epithelial cell line. Next, we determined the relative expression of AOC4P in 63 paired GC tumor tissues and corresponding adjacent noncancerous tissues. The expression levels of AOC4P were also significantly higher in tumor tissues than in noncancerous tissues, suggesting involvement of AOC4P in the tumorigenesis of GC [Figure 1(b)].

AOC4P expression levels in GC cell lines and specimens, and its prognostic value for GC. (a) Determination of AOC4P expression in GES-1, MGC-803 and BGC-823 cells by qRT-PCR. (b) Determination of AOC4P expression in GC specimens. (c) Relative expression of AOC4P in tumor tissues. (d) Correlation of AOC4P expression with overall survival. (e) Correlation of AOC4P expression with disease-free survival.
High expression of AOC4P correlates with poor prognoses
Using the median expression level of AOC4P in tumor tissues as the cutoff, we divided the patients into two groups: patients with high or low expression of AOC4P [Figure 1(c)]. After a median follow-up time of 41 months (range: 4–49 months), as illustrated in Figure 1(d), patients with a high expression level of AOC4P had poorer overall survival than those with a low expression level of AOC4P. Similarly, patients with high expression of AOC4P had poorer disease-free survival [Figure 1(e)]. Additionally, expression of AOC4P was correlated with lymphovascular invasion (Table 1), a critical step for tumor dissemination. As shown in Table 2, univariate and multivariate Cox regression analysis has revealed that AOC4P expression was strongly associated with disease-free survival and overall survival. These findings suggest that AOC4P is associated with metastasis and can serve as a prognostic biomarker.
Relationship between expression level of AOC4P and characteristics.

lncRNA, long noncoding RNA; TNM, tumor, nodes, metastasis.
Univariate and multivariate Cox regression analysis for DFS and OS.

CI, confidence interval; DFS, disease-free survival; HR, hazard ratio; OS, overall survival; TNM, tumor, nodes, metastasis.
Knockdown of AOC4P inhibits cellular proliferation and colony formation in vitro
To investigate the biological effects of AOC4P on cellular growth and colony formation, loss-of-function experiments were conducted. We first knocked down AOC4P expression by transfection of si-AOC4P into MGC-803 and BGC-823 cells. The results of qRT-PCR analyses revealed that AOC4P expression was knocked down by 89% and 83% in si-AOC4P-transfected MGC-803 and BGC-823 cells, respectively, compared with siRNA negative control (si-NC)-transfected cells (Figure S1). MTT assays showed inhibition of cell growth by knockdown of AOC4P expression in MGC-803 and BGC-823 cells compared with controls [Figure 2(a)]. Additionally, clonogenic abilities were impaired following downregulation of AOC4P in MGC-803 and BGC-823 cells [Figure 2(b)]. In situ TUNEL assays revealed that the proportion of apoptotic cells was higher among si-AOC4P-transfected MGC-803 and BGC-823 cells than si-NC-transfected cells [Figure 2(c)]. Flow cytometric analysis revealed that knockdown of AOC4P significantly increased GC cells in the G0/G1 phase, while reduced cells in the S phase [Figure 3(a)]. Meanwhile, western blot assay showed that the expression of apoptosis-related proteins, including cleaved caspase-3 and cleaved PARP, were significantly increased after knockdown of AOC4P [Figure 3(b)]. Collectively, these data indicate that knockdown of AOC4P inhibits GC cells proliferation and promotes apoptosis.

Knockdown of AOC4P inhibits proliferation and colony formation and promotes apoptosis in vitro. (a) Cell proliferation was measured by MTT assays in MGC-803 and BGC-823 cells. (b) Colony formation assay results. Colonies were photographed and then counted. (c) TUNEL assays to investigate the effect of AOC4P on apoptosis of MGC-803 and BGC-823 cells. Results are expressed as the mean ± SD (n = 3). *p < 0.05, **p < 0.01, ***p < 0.001.
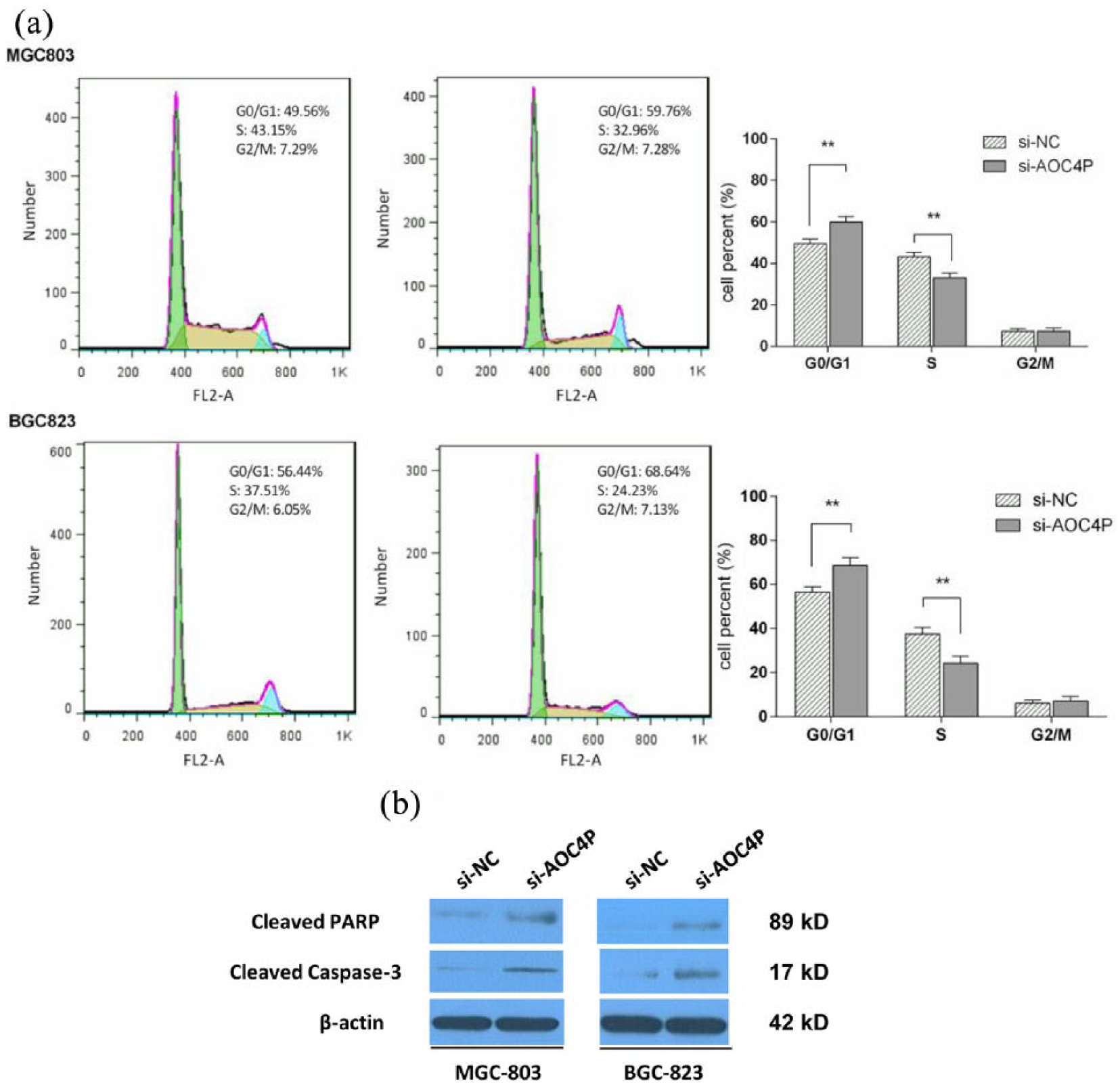
Effect of AOC4P on cell cycle and apoptosis-related proteins. (a) Cell cycle analysis in MGC-803 and BGC-823 cells. Knockdown of AOC4P induced more number of cells in the G0/G1 phase and reduced the number of cells in the S phase. (b) Apoptosis-related proteins detected by western blotting assay. **p < 0.01.
Effect of AOC4P on migration and invasion of GC cells
We used Transwell and wound healing assays to determine the effect of AOC4P on migration and invasion of GC cells. As shown in Figure 4(a) and (b), the migration and invasion abilities of GC cells were significantly decreased when AOC4P expression was reduced in MGC-803 and BGC-823 cells. These results suggested that AOC4P promotes the migration and invasion of GC cells.
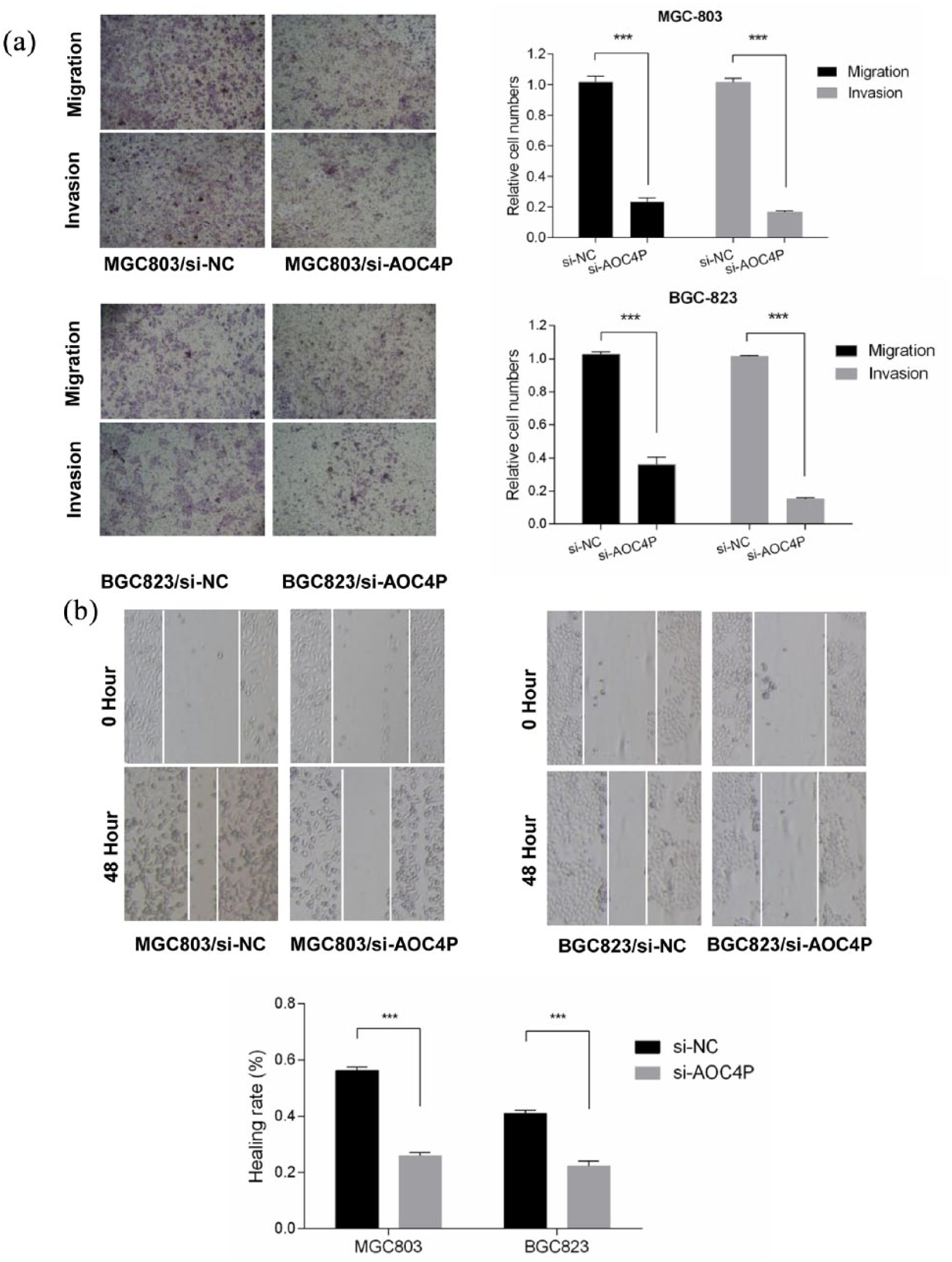
Knockdown of AOC4P inhibits migration and invasion in vitro. (a) Transwell assays were performed to investigate the effect of AOC4P on migration and invasion of MGC-803 and BGC-823 cells. (b) Evaluation of cell motility by wound healing assays in MGC-803 and BGC-823 cells. ***p < 0.001.
Knockdown of AOC4P inhibits GC tumorigenesis in vivo
To explore whether AOC4P affects tumorigenesis in vivo, BGC-823 cells were stably transfected with si-AOC4P or the empty vector and then injected subcutaneously into the left and right flanks of each mouse, respectively. At 14 days after injection, tumors formed in the si-AOC4P group were significantly smaller than those in the si-NC group [Figure 5(a) and (b)]. The tumor weight was also lower in the si-AOC4P group compared with the si-NC group [Figure 5(c)]. qRT-PCR analysis revealed that AOC4P expression was significantly lower in si-AOC4P tumor xenografts [Figure 5(d)]. The results of IHC and in situ TUNEL assays of xenografts showed that si-AOC4P tumor xenografts had reduced proportions of Ki-67-positive cells and increased proportions of TUNEL-positive cells compared with si-NC tumor xenografts [Figure 4(e)]. Consistent with the aforementioned in vitro results, these data indicate that AOC4P affects GC tumorigenesis in vivo.

Knockdown of AOC4P inhibits tumorigenesis in vivo. (a) BGC-823 cells stably transfected with si-AOC4P or the empty vector were injected subcutaneously into the left and right flanks of each mouse, respectively (n = 6). Representative image of an ultrasound is shown. Upper and lower tumor tissues are si-NC and si-AOC4P groups, respectively. (b) Subcutaneous tumor growth curve of the si-AOC4P group compared with the si-NC group. (c) Comparison of tumor weights between si-AOC4P and si-NC groups. (d) Relative expression levels of AOC4P in tumor xenografts. (e) Hematoxylin and eosin staining, Ki-67 expression analysis, and TUNEL assays were performed to evaluate proportions of proliferating and apoptotic cells in tumor xenografts. *p < 0.05, **p < 0.01, ***p < 0.001.
AOC4P modulates GC tumorigenesis via epithelial–mesenchymal transition
A recent study reported that AOC4P suppresses hepatocellular carcinoma via inhibition of epithelial–mesenchymal transition (EMT). 15 Therefore, we investigated whether AOC4P functions in a similar manner in GC. After knockdown of AOC4P in BGC-823 and MGC-803 cells, qRT-PCR analysis revealed an increase in E-cadherin expression, while vimentin and MMP9 expression was reduced [Figure 6(a)]. Western blot assays confirmed these changes at the protein level [Figure 6(b)]. Interestingly, immunofluorescence results of cancer tissues showed that tumors with high expression AOC4P had relatively low levels of E-cadherin and high levels of vimentin and MMP9 (Figure 7). Therefore, these results suggest that AOC4P modulates GC tumorigenesis by regulating EMT.

AOC4P is involved in EMT processes. (a) Determination of E-cadherin, vimentin, and MMP9 mRNA levels by qRT-PCR following knockdown of AOC4P in MGC-803 and BGC-823 cells. (b) Protein expression of E-cadherin, vimentin, and MMP9 analyzed by western blotting after knockdown of AOC4P in MGC-803 and BGC-823 cells.
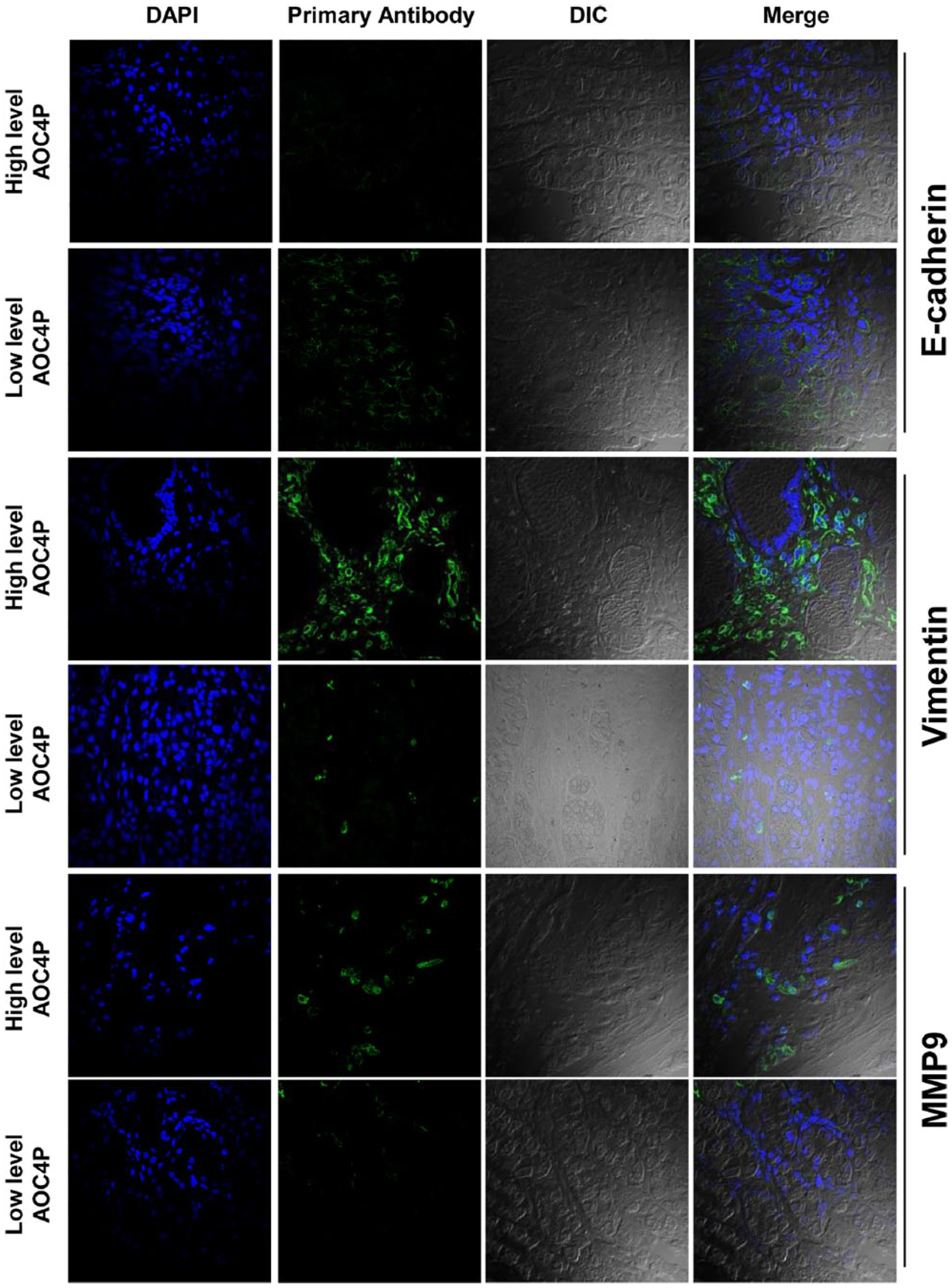
Immunofluorescence of tumor tissues to investigate the association between AOC4P expression and E-cadherin, vimentin, and MMP9 levels.
Discussion
GC is frequently diagnosed as locally advanced in China, leading to the fact that patients in China have a poor estimated 5-year overall survival of 27.4% compared with 73.2% in Korea.1,16 One of the solutions to improve prognoses of patients with GC is early detection and intervention strategies. Therefore, we have focused on investigating novel diagnostic markers and potential therapeutic targets of GC.11,17,18
Recently we identified five differentially expressed lncRNAs between tumor and adjacent normal tissues by lncRNA microarray profiling, including TINCR, CCAT2, AOC4P, BANCR, and LNC00857. 11 Using these circulating lncRNAs, we established a five-lncRNA panel for early detection of GC. In the present study, we investigated the role of tissular AOC4P in GC. We found significant upregulation of AOC4P in GC tissues, and that a high expression level of AOC4P was correlated with poor survival and lymphovascular invasion in patients with GC. Functionally, in vitro and in vivo assays demonstrated that AOC4P promoted tumor growth by inducing proliferation, migration, and invasion, and reducing apoptosis. Mechanistically, the oncogenic effect of AOC4P in GC might be partly attributed to AOC4P-mediated EMT. These data are helpful to explain the significance of AOC4P upregulation in GC and its correlation with clinicopathological characteristics.
Thus far, the functions of AOC4P have only been investigated in colon cancer and hepatocellular carcinoma.15,19 In colon cancer, AOC4P (also termed UPAT) is upregulated in highly tumorigenic colon cancer cells and involved in epigenetic regulation of cancer cells by modulating protein ubiquitination and degradation. 19 Similar to its functions in GC, Taniue and colleagues found that AOC4P plays a critical role in tumorigenicity of colon cancer cells. 19 However, in hepatocellular carcinoma, the expression of AOC4P was downregulated in 68% of tumor tissues. 15 Our results, together with earlier findings, indicated varied expression of AOC4P in different types of carcinomas. A previous study has also reported different expression levels of lncRNA BANCR in GC and colon cancer.20,21 The fact that AOC4P is expressed differentially in various carcinomas might be partly explained by its various functions. In this study, we found that inhibition of AOC4P increased the expression of E-cadherin, while reducing the expression of vimentin and MMP9. Consistent with these findings, immunofluorescence of tumor specimens confirmed the correlation of AOC4P expression with E-cadherin, vimentin, and MMP9 levels. Therefore, AOC4P might exert its oncogenic effect in GC via EMT processes.
As a central driver of tumor malignancy, EMT is involved in cancer cell dissemination, drug resistance, subsequent disease recurrence, and acquisition of immunosuppressive capabilities in a variety of cancers. 22 It has been proposed that essentially all carcinomas develop malignancy-associated characteristics via activation of an EMT process in their constituent neoplastic cells. 22 One of the hallmarks of EMT is replacement of E-cadherin by N-cadherin, which results in the formation of far weaker cell–cell adhesions between adjacent cells. Importantly, a study from the Asian Cancer Research Group has classified GC into four molecular subtypes, among which the EMT type has the worst prognosis, the tendency to occur at an earlier age, and the highest recurrence frequency. 5 Therefore, as a regulator of EMT, AOC4P might be a potential therapeutic target for GC treatment.
Our earlier findings demonstrated a correlation between circulating AOC4P and tissular AOC4P. 11 Circulating lncRNAs in blood are usually incorporated into exosomes, which are small vesicles of endocytic origin that carry a variety of bioactive molecules, including proteins, lipids, RNA, and DNA. 23 Metastasis is a multistep process including invasion of tumor cells into local tissues at a primary tumor site, invasion into blood and lymph vessels, survival in circulation, extravasation from circulation to distant sites, and adaptation and proliferation in the metastatic site. Evidence has shown the involvement of exosomes in all processes of metastasis. 24 Currently, we have only demonstrated that AOC4P promoted tumorigenesis and cancer progression intracellularly. Therefore, our future study will focus on how tissular AOC4P incorporates into exosomes and is released into circulation, and the role of AOC4P-incorporated exosomes in metastatic sites.
Several limitations of present study should be taken into consideration when interpreting the results. First, the prognostic role of AOC4P needed to be validated in another independent cohort. Second, we did not unveil the downstream molecules which AOC4P regulated, therefore the molecular pathway through which AOC4P exerts its function needed a further in-depth investigation.
Conclusion
Taken together, our results revealed a relationship between AOC4P and clinicopathological characteristics and demonstrated the prognostic potential of AOC4P for GC. We also revealed the involvement of AOC4P in proliferation, migration, invasion, and apoptosis of MGC-803 and BGC-823 cells. Our study indicates that AOC4P promotes tumorigenesis and progression partly through EMT.
Supplemental Material
Figure_S1 – Supplemental material for Long noncoding RNA AOC4P regulates tumor cell proliferation and invasion by epithelial–mesenchymal transition in gastric cancer
Supplemental material, Figure_S1 for Long noncoding RNA AOC4P regulates tumor cell proliferation and invasion by epithelial–mesenchymal transition in gastric cancer by Kecheng Zhang, Canrong Lu, Xiaohui Huang, Jianxin Cui, Jiyang Li, Yunhe Gao, Wenquan Liang, Yi Liu, Yang Sun, Hanxuan Liu, Bo Wei and Lin Chen in Therapeutic Advances in Gastroenterology
Footnotes
Acknowledgements
Kecheng Zhang, Canrong Lu, Xiaohui Huang, Jianxin Cui are joint authors. Study design and concept: KCZ, CRL, XHH. Data acquisition: YHG, WQL, YL, YS, HXL. Data analysis and interpretation: KCZ, BW, LC. Collection of clinical data and sample disposal: WQL, YL, YS, HXL. Manuscript preparation: KCZ. Manuscript review: BW, LC. All authors read and approved the final manuscript.
Funding
This work was partly funded by the National Nature Science Foundation of China (No. 81672319, 81773135), National Key Research and Development Plan (No. 2016YFC0905302), and Beijing Municipal Science and Technology Plan projects (Z161100000516237, D141100000414002, Z15110000391547, and Z171100000417023).
Conflict of interest statement
The authors declare that there is no conflict of interest.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
