Abstract
Objective
Our objective was to explore the prognostic role of long noncoding RNA (lncRNA) SH3PXD2A-AS1 in esophageal squamous cell carcinoma (ESCC).
Methods
An SH3PXD2A-AS1 expression dataset was retrieved and analyzed from The Cancer Genome Atlas database, and SH3PXD2A-AS1 expression was determined in our cohort of 134 ESCC patients by using quantitative PCR. The clinical significance of SH3PXD2A-AS1 expression was investigated by the Chi square test and its prognostic value was determined by Kaplan–Meier survival curve analysis and Cox proportional hazards analysis. RNA interference and in vitro functional experiments, including cell viability, migration, and invasion, were used to investigate effects of SH3PXD2A-AS1 on cell malignant phenotype.
Results
SH3PXD2A-AS1 expression was increased in ESCC tissues compared with adjacent normal tissues. A high level of SH3PXD2A-AS1 expression was associated with poor tumor differentiation and advanced T, N, and TNM stages, indicating its oncogenic role in ESCC. Moreover, its high expression predicted poor overall survival in patients with ESCC. Inhibition of SH3PXD2A-AS1 expression significantly suppressed cell viability, migration, and invasion of ESCC cells.
Conclusion
High SH3PXD2A-AS1 expression is a poor prognostic factor for patients with ESCC. SH3PXD2A-AS1 might function as an oncogene that can promote malignant biological characteristics of ESCC cells.
Keywords
Introduction
Esophageal cancer (EC) is a common malignant tumor, with the number of new cases ranking ninth among all malignancies in the world; 1 it is also the sixth most common cause of cancer-related death. 1 Histologically, esophageal cancer consists mainly of squamous cell carcinoma and adenocarcinoma, and approximately 85% of all EC cases are esophageal squamous cell carcinoma (ESCC). 2 Because of the lack of specific methods for early diagnosis, ESCC tends to be diagnosed late. The overall survival of patients with ESCC remains poor, with a 5-year survival rate of 15% to 34% after initial diagnosis.3,4 Therefore, identification of biomarkers to predict survival would be helpful and is urgently needed; these biomarkers can be meaningful for not only clinical monitoring but also therapeutic decision-making.
Recently, long noncoding RNAs (lncRNAs), defined as non-protein-coding RNA molecules longer than 200 nucleotides, have received wide attention. Accumulating evidence indicates that aberrant expression of lncRNAs has a key biological role in various types of cancer.5–8 A number of studies have suggested that aberrant expression of lncRNAs in ESCC is closely associated with clinical pathological characteristics, including histological type, tumor-node-metastasis (TNM) stage, lymph node metastasis, and prognosis.9,10 Some lncRNAs, such as LINC00152, 11 MALAT1, 12 and LEF1-AS1, 13 have been identified as oncogenes that can promote ESCC progression. Other lncRNAs, such as CASC2 14 and MEG3, 15 have been confirmed as tumor suppressors in ESCC. Although the effects of some lncRNAs on ESCC progression have attracted considerable research attention, the functional roles of many lncRNAs in cancer development have not been fully elucidated.
Currently, web-based tools, such as Gene Expression Profiling Interactive Analysis (GEPIA), can perform fast and customizable analyses based on The Cancer Genome Atlas (TCGA) and Genome-Tissue Expression data. 16 In this study, we identified SH3PXD2A-AS1 as an upregulated lncRNA in ESCC tissues of the TCGA database by analysis using GEPIA. However, its functional role in ESCC is still unclear. Thus, we investigated the expression of SH3PXD2A-AS1 in our ESCC cohort and further explored the biological effects on cell phenotype after inhibiting SH3PXD2A-AS1 expression by RNA interference (RNAi). To our knowledge, this is the first report on the expression and role of lncRNA SH3PXD2A-AS1 in ESCC cells.
Materials and methods
Ethical approval
This study was approved by the Ethics Committee of Peking University Cancer Hospital and Institute (no. 2017KT79). Written informed consent was obtained from all participants before collection of specimens.
TCGA dataset and GEPIA
SH3PXD2A-AS1 expression data were retrieved from the TCGA database and analyzed using GEPIA (http://gepia.cancer-pku.cn/) online analysis. The TCGA dataset of esophageal carcinoma (ESCA) and matched normal data included 182 tumor tissues and 13 normal tissues and was selected for this analysis with default parameters.
Tissue specimens
A total of 134 paired ESCC tumor tissues and matched adjacent normal tissues were obtained from the Department of Biobank of Peking University Cancer Hospital and Institute (Beijing, China). Patients with ESCC (mean age 60.86 ± 8.45 years, ranging from 37 to 80 years; 112 male and 22 female) underwent surgery at Peking University Cancer Hospital between July 2012 and September 2015. The following inclusion/exclusion criteria were applied: (1) pathologically confirmed ESCC, (2) no preoperative radiotherapy or chemotherapy, (3) no other history of cancer, and (4) complete and available clinicopathological and follow-up data.
Cell culture
Human ESCC cell lines, including KYSE150, KYSE180, KYSE450, EC109, EC9706, were obtained from the Cell Center of Peking Union Medical University (Beijing, China). All cell lines were cultured in RPMI-1640 medium (Gibco/Thermo Fisher Scientific Inc., Waltham, MA, USA) supplemented with 10% fetal bovine serum (FBS; Gibco/Thermo Fisher Scientific Inc.), 100 U/mL penicillin, and 100 mg/mL streptomycin at 37°C with 5% CO2.
Cell transfection
Short interfering (si)RNAs targeting SH3PXD2A-AS1 were synthesized by Sangon Biotech (Shanghai) Co. Ltd. (Shanghai, China) and transiently transfected into 1 × 105 cells at a concentration of 50 nmol/L using Lipofectamine 2000 (Invitrogen/Thermo Fisher Scientific Inc.) according to the manufacturer’s protocol for 24 hours before subsequent experimentation. Double-stranded RNAi sequences (siRNA1: 5′-CGACGUAACUCCAGACAUATT-3′; siRNA2: 5′-UGAUGGAGCUUCAGAAUUATT-3′; siRNA3: 5′-CCACAUAGGUCUCACUUAATT-3′, and negative control (NC): 5′-UUCUCCGAACGUGUCACGUTT-3′) were designed.
Reverse transcription-quantitative PCR (RT-qPCR)
Total RNA was isolated from tissues and cells using RNeasy Mini kits (Qiagen Inc., Valencia, CA, USA) according to the manufacturer’s protocols. Subsequently, RNA was reverse transcribed with random primers (Takara Biotechnology (Dalian) Co. Ltd., Dalian, China) and Moloney Murine Leukemia Virus Reverse Transcriptase (Invitrogen/Thermo Fisher Scientific Inc.) for 1 hour at 37°C. The following primers were used: SH3PXD2A-AS1 forward, 5′-GCCTCTTTACAGCGTTCCCT-3′ and reverse, 5′-ATCTTTCAGCAAGTCGCGGT-3′; GAPDH forward, 5′-GCCTCTTTACAGCGTTCCCT-3′ and reverse, 5′-ATCTTTCAGCAAGTCGCGGT-3′. The lncRNA expression levels of were evaluated using SYBR Select Master Mix (Thermo Fisher Scientific Inc.) and an ABI 7500 Fast System (Applied Biosystems/Thermo Fisher Scientific Inc.). The cycling conditions were as follows: 10 minutes at 95°C followed by 40 cycles of 95°C for 3 s and 60°C for 30 s. GAPDH was used as the reference gene. The expression level of SH3PXD2A-AS1 was calculated using the 2−ΔCq method, whereby ΔCq = Cq (SH3PXD2A-AS1) − Cq (GAPDH).
Cell viability assay
A total of 5 × 103 cells/well were seeded into 96-well plates and cultured for 12, 24, 36, 48, and 60 hours. Cell growth was evaluated using a Cell Counting Kit-8 (CCK-8; Dojindo Molecular Technologies Inc., Beijing, China), according to the manufacturer’s protocol. Absorbance was measured at a wavelength of 450 nm using a microplate reader (iMark; Bio-Rad Laboratories Inc., Hercules, CA, USA).
Migration and invasion assays
Cells (1 × 104) were seeded into the upper chambers of a Transwell (Corning Inc., Corning, NY, USA) with an 8.0-µm pore size, with or without Matrigel (BD Biosciences, San Jose, CA, USA), in 100 µL of RPMI-1640 medium containing 1% FBS. Then, 500 µL of RPMI-1640 medium containing 10% FBS was placed in each lower chamber as a chemoattractant. After 24 hours, migrated (without Matrigel) or invaded (with Matrigel) cells were fixed with 4% formaldehyde at room temperature for 5 minutes and stained with 1% crystal violet for 5 minutes at room temperature. The number of migrated or invasive cells was counted by light microscopy at 200× magnification in three randomly selected microscopic fields.
Statistical analysis
All data were analyzed using SPSS software (version 22.0; SPSS Inc., IBM Corp., Armonk, NY, USA). Countable data are presented as the mean ± standard deviation of three independent experiments. Statistical significance between two groups was determined by using a two-tailed Student’s t-test, and for multiple comparisons by using one-way ANOVA with Bonferroni post hoc tests. The Mann–Whitney U test was used for two independent sample t-tests on rank data. A two-tailed χ2 test was used to evaluate the association between SH3PXD2A-AS1 expression and clinic-pathological characteristics of patients with ESCC. Overall survival (OS) was analyzed using the Kaplan–Meier method. The cut-off value for SH3PXD2A-AS1 expression was based on the receiver-operator characteristic (ROC) curve corresponding to the maximum Youden index. Univariate Cox analysis was performed, followed by a multivariate Cox regression analysis only including variables that were significant in the univariate Cox analysis.17,18 P < 0.05 was considered to indicate a significant difference.
Results
Expression of SH3PXD2A-AS1 in TCGA dataset
SH3PXD2A-AS1 expression levels were analyzed by using GEPIA tools based on data derived from the TCGA esophageal carcinoma (ESCA) dataset. The results showed that SH3PXD2A-AS1 exhibited high expression in esophageal carcinoma tissues compared with normal tissues (Figure 1a, P < 0.05). To determine whether expression of SH3PXD2A-AS1 was associated with survival time of patients with esophageal carcinoma, we performed survival analysis with different cutoffs (median and quartile). As indicated in Figure 1b and 1c, with either the median or quartile cutoff, patients with a high level of SH3PXD2A-AS1 expression had a shorter disease-free survival time than those with a low level of expression [hazard ratio (HR) = 1.5 cutoff by median; HR = 2.0 cutoff by quartile], but the log rank test did not show statistical significance (P = 0.1 cutoff by median and P = 0.058 cutoff by quartile).

Relative SH3PXD2A-AS1 expression in ESCA tissues from the TCGA database. (a) Higher SH3PXD2A-AS1 expression levels were observed in ESCA tissue samples compared with normal tissues; *P < 0.05. (b, c) K-M survival analysis of disease-free survival time after surgery, with median (b) or quartile (c) value as the cutoff for SH3PXD2A-AS1 expression in ESCA tissues.
Expression of SH3PXD2A-AS1 in patients with ESCC
To further confirm the expression profile of SH3PXD2A-AS1 in ESCC, qPCR was performed to confirm the expression profile of SH3PXD2A-AS1 in 134 pairs of tumor and matched adjacent normal tissues from patients with ESCC. The results indicated significantly increased expression of SH3PXD2A-AS1 in ESCC tumor tissues compared with adjacent normal tissues (paired t-test P = 0.0202; Figure 2a). Then, we evaluated the association between SH3PXD2A-AS1 expression and clinicopathological parameters of patients with ESCC. To facilitate our analysis, patients were divided into two groups according to the maximum Youden index of the ROC curve in the ESCC tissues. As shown in Table 1, the number of patients with high SH3PXD2A-AS1 expression was greater in groups with poor tumor differentiation, advanced T stage, N stage, and TNM stage. The rate of high SH3PXD2A-AS1 expression was higher in tumor tissues of patients with a lower degree of tumor differentiation (χ2 test, P = 0.018), higher T stage (χ2 test, P = 0.011), higher N stage (χ2 test, P = 0.011), and advanced TNM stage (χ2 test, P = 0.000), indicating that high SH3PXD2A-AS1 expression may be associated with tumor progression. However, there was no association between SH3PXD2A-AS1 expression and other clinicopathological features, including sex, age, tumor location, and tumor embolus (Table 1).

Relative SH3PXD2A-AS1 expression in ESCC tissues from a cohort of 134 patients. (a) Increased SH3PXD2A-AS1 expression in ESCC tissue samples compared with adjacent normal tissues. (b) ROC curve predicting the most suitable cutoff value for SH3PXD2A-AS1 by using the maximum Youden index. (c) K-M survival analysis of overall survival time after surgery cutoff by maximum Youden index
Relationship between SH3PXD2A-AS1 expression and clinicopathological characteristics of patients with ESCC.

ESCC, esophageal squamous cell carcinoma; TNM, tumor-node-metastasis.
We then assessed the association the expression level of SH3PXD2A-AS1 and patient OS. The ROC curve was drawn to determine the optimal cut-off value based on the maximum Youden index (Figure 2b). Kaplan–Meier survival analysis revealed that patients with high SH3PXD2A-AS1 expression had a significantly shorter OS than those with low SH3PXD2A-AS1 expression (Figure 2c). Similarly, from the univariate Cox proportional hazard model analysis, location, TNM stage, and SH3PXD2A-AS1 expression were prognostic factors for survival (Table 2; by multivariate analysis). Compared with other patients, patients with upper thoracic esophageal carcinoma, advanced TNM stage, and higher SH3PXD2A-AS1 expression had a significantly higher hazard ratio. However, using multivariate Cox proportional hazard model analysis, only tumor location and TNM stage were identified as independent risk factors for survival (P = 0.017 and P = 0.004, respectively; Table 2). Collectively, these data suggest that high SH3PXD2A-AS1 expression may be a poor prognostic biomarker for patients with ESCC.
Cox univariate and multivariate analyses of overall survival predictors of ESCC.
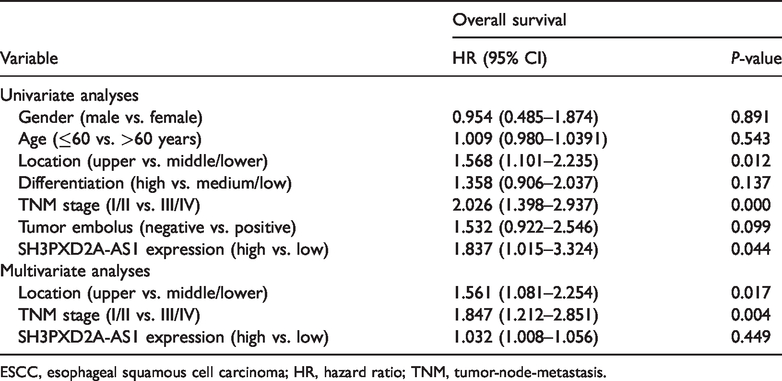
ESCC, esophageal squamous cell carcinoma; HR, hazard ratio; TNM, tumor-node-metastasis.
Expression of SH3PXD2A-AS1 in ESCC cell lines
To determine suitable cell lines to perform RNAi to assess the effects of SH3PXD2A-AS1 inhibition on cell phenotype, SH3PXD2A-AS1 expression was first evaluated in five ESCC cell lines. Two ESCC cell lines, EC109 and KYSE450, had relatively high expression, whereas EC9706, KYSE150, and KYSE180 cells had relatively low expression of SH3PXD2A-AS1 (Figure 3a). Therefore, EC109 and KYSE450 cells were selected for SH3PXD2A-AS1 silencing experiments.
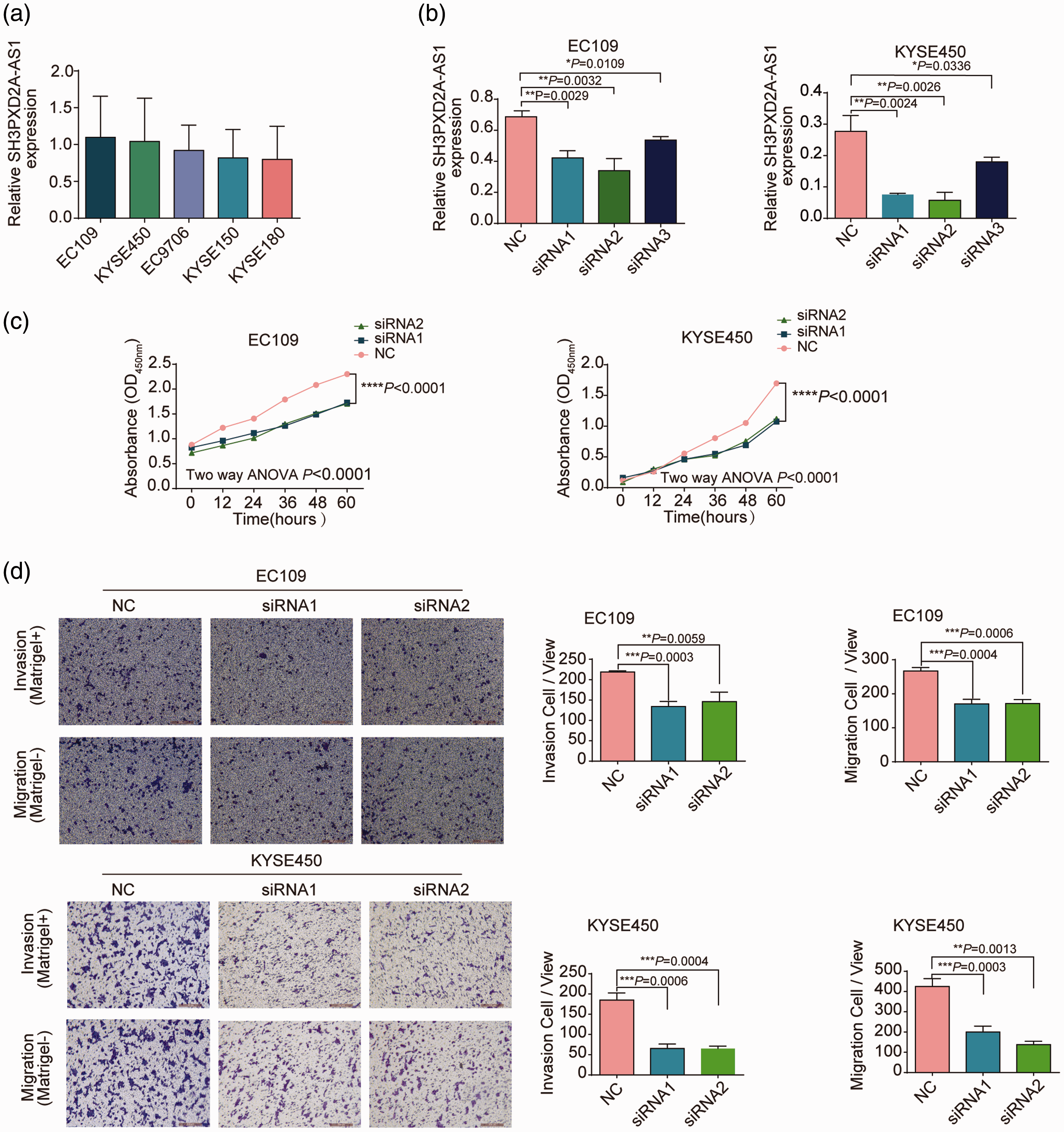
SH3PXD2A-AS1 promotes ESCC cell proliferation, migration, and invasion abilities in vitro. (a) qPCR was used to detect the endogenous expression level of SH3PXD2A-AS1 in five ESCC cell lines. (b) The inhibitory effects of three siRNAs compared with the negative control were evaluated by qPCR. (c) Knockdown of SH3PXD2A-AS1 expression inhibited cell proliferation (measured as optical density at 450 nm) in the EC109 and KYSE450 cell lines. (d) Transwell assays were performed to assess cell migration and invasion capacities of si-SH3PXD2A-AS1-transfected EC109 and KYSE450 cells. Values in a, b, and d are shown as the mean ± SD of three independent experiments with triplicates.
We hypothesized that SH3PXD2A-AS1 would promote progression of ESCC; therefore, we evaluated phenotypic changes following downregulation of SH3PXD2A-AS1 with siRNA in the EC109 and KYSE450 cell lines (that have relatively high endogenous expression; Figure 3a). qPCR analysis demonstrated that SH3PXD2A-AS1 expression was successfully inhibited by three siRNAs (Figure 3b): 39% and 73% inhibition for siRNA1 in EC109 and KYSE450 cells (P < 0.05); 51% and 79% inhibition for siRNA2 in EC109 and KYSE450 cells (P < 0.05); and 22.8% and 32.1% inhibition for siRNA3 in EC109 and KYSE450 cells, (P < 0.05), respectively. siRNA1 and siRNA2 were selected for subsequent experiments because they had greater inhibitory effects than siRNA3. Cell viability was significantly decreased after treatment with the two SH3PXD2A-AS1 siRNAs in both EC109 and KYSE450 cells, as assessed by the CCK-8 assay (Figure 3c). Moreover, compared with transfection with NC, transfection with the two SH3PXD2A-AS1 siRNAs significantly suppressed the migratory and invasive ability of EC109 and KYSE450 cells, as shown by the Transwell assay (Figure 3d). The proportion of invasive cells was decreased by 39% and 65% with siRNA1 (P < 0.05) and by 33% and 65% with siRNA2 (P < 0.05) in EC109 and KYSE450 cells, respectively. The proportion of migrating cells decreased by 36% and 53% with siRNA1 (P < 0.05) and by 36% and 68% with siRNA2 (P < 0.05) in EC109 and KYSE450 cells, respectively. Collectively, these results indicate that silencing SH3PXD2A-AS1 inhibited cell viability and migration and invasion abilities of EC109 and KYSE450 cells in vitro.
Discussion
The lncRNA SH3PXD2A-AS1 is located on chromosome 10; it is 2023 bp long and is an antisense transcript transcribed from SH3PXD2A (SH3 and PX domains 2A), an adapter protein that is involved in invadopodia and podosome formation, extracellular matrix degradation, and invasiveness of some cancer cells. Previously, the SH3PXD2A protein was shown to exert oncogenic roles in multiple cancers.19–21 To our knowledge, there are few reports about the role of lncRNA SH3PXD2A-AS1; we identified only two articles related to SH3PXD2A-AS1 in the PubMed database. In 2019, Li and Zhan 22 reported SH3PXD2A-AS1 as 1 of 16 lncRNAs related to OS of patients with ovarian cancer; in 2018, Ma et al. 23 presented the first evidence that SH3PXD2A-AS1 acts as an oncogene to promote cell progression in colorectal cancer (CRC). In this study, we found that dysregulation of SH3PXD2A-AS1 was correlated with clinicopathological factors in patients with ESCC.
Consistent with previous research in CRC, 23 the results of the present study showed increased expression of SH3PXD2A-AS1 in ESCC tissues compared with adjacent normal tissues, and a high level of SH3PXD2A-AS1 was associated with advanced TNM stage. In CRC, increased SH3PXD2A-AS1 expression was found to be correlated with larger tumor size and positive lymph node metastasis, 23 but in our study, we did not find these relationships to be clinically significant. However, we did demonstrate that increased SH3PXD2A-AS1 expression was correlated with other clinical characteristics in ESCC, including poor tumor differentiation and advanced T and N stage. These inconsistent outcomes may be due to different genetic or biologic tumor characteristics or cohort size; further studies are needed to confirm these findings.
In contrast to reports in ovarian cancer, our results indicated that increased SH3PXD2A-AS1 expression predicted poor OS in patients with ESCC. Li and Zhan 22 retrieved ovarian cancer data from the TCGA database and suggested that SH3PXD2A-AS1 was a good predictor for OS (HR = 0.78) by Kaplan–Meier survival analysis. However, in our study, increased SH3PXD2A-AS1 expression predicted poor survival after surgery by Kaplan–Meier survival analysis, and SH3PXD2A-AS1 was identified as a risk factor (HR = 1.83) for OS of patients with ESCC by univariate Cox proportional hazard model analysis. These contrasting outcomes may be due to the different genetic or biologic backgrounds of diverse tumor types.
In this study, we were unable to obtain normal esophageal epithelium cell lines (due to the COVID-19 pandemic) for use as a control to confirm the upregulation of SH3PXD2A-AS1 in ESCC cell lines. This is a limitation of our study, and further studies are needed with appropriate controls to verify our results. Here, in vitro experiments demonstrated that SH3PXD2A-AS1 silencing had a marked inhibitory effect on ESCC cell proliferation, migration, and invasion abilities, which is consistent with a previous report in CRC. 23 Our results should be validated by in vivo experiments to confirm the role of SH3PXD2A-AS1 in ESCC.
In CRC, knockdown of SH3PXD2A-AS1 promoted G1/G0 phase arrest and increased apoptotic rates in cancer cells. 23 Whether changes in SH3PXD2A-AS1 expression affect the cell cycle and apoptosis in ESCC needs to be confirmed.
Previously, a number of lncRNAs were shown to be aberrantly expressed in ESCC and closely associated with clinical pathological characteristics, including histological type, TNM stage, metastasis, and prognosis.9,10 These results suggest that lncRNAs play important roles in ESCC progression and could serve as prognostic biomarkers. Here, SH3PXD2A-AS1 was found to be a novel prognostic biomarker for ESCC. The combination of SH3PXD2A-AS1 with other lncRNAs in a panel could improve its prognostic prediction value in patients with ESCC. In addition, as a new biomarker, it would be useful to study the underlying mechanism of SH3PXD2A-AS1 as an oncogene in regulating malignant behavior of ESCC cells.
Taken together, our results indicated that upregulated SH3PXD2A-AS1 expression was associated with poor prognosis, poor tumor differentiation, and advanced T, N, and TNM stages in patients with ESCC. Therefore, SH3PXD2A-AS1 may have potential as a prognostic biomarker for ESCC.
Footnotes
Authors' contributions
HH, FX, and JC contributed to the conception and design of the experiments, QL completed most of experiments, SW aggregated the figures, and HH and JC drafted the manuscript.
Availability of data and materials
The datasets used and analyzed during the current study are available from the corresponding author upon reasonable request.
Declaration of conflicting interest
The authors declare that there is no conflict of interest.
Funding
This work was supported by the grants from National Natural Science Foundation of China (No. 81773144, 8172632, 81802943), Science Foundation of Peking University Cancer Hospital 2020-9, Beijing Outstanding Talent training program, the interdisciplinary Medicine Seed Fund of Peking University (No. BMU2018MX019), and Capital Health Research and Development of Special (No. 2018-2-2155).
