Abstract
TNF receptor-associated factor 6 (TRAF6) is a signal transducer, which plays a pivotal role in triggering a variety of signalling cascades. Here, we cloned and identified the TRAF6 gene from the King pigeon. The open reading frame sequence of pigeon TRAF6 (piTRAF6) is 1638 bp long and encodes a 545 aa protein, including a low-complexity domain, RING finger, Zinc finger, coiled coil domain, and meprin and TRAF homology domain. The aa sequence of piTRAF6 shared a strong identity with that of other birds. PiTRAF6 transcripts were broadly expressed in all the tested tissues; piTRAF6 levels were the highest and lowest in the heart and stomach, respectively. Overexpression of piTRAF6 activated NF-κB in a dose-dependent manner and induced IFN-β expression. Upon piTRAF6 knockdown by small interfering RNAs, NF-κB activation was markedly inhibited in HEK293T cells. The expression of piTRAF6, as well as pro-inflammatory cytokines and antiviral molecules, were obviously increased after TLR ligand stimulation and Newcastle disease virus or Salmonella Pullorum inoculation. These results suggest that piTRAF6 may play a key immunoregulatory role in the innate immune response against viral and bacterial infections.
Introduction
TNF receptor-associated factors (TRAF) are a family of cytoplasmic adapters that promote intracellular signal transduction via receptor binding, thereby potentiating the recruitment of signal transduction proteins. 1 Thus far, seven members of the TRAF family have been identified in mammals. Among those, TRAF6 is the only TRAF family member that participates in signal transduction of the TNF receptor superfamily and IL-1 receptor or TLR family. 2 This signal transduction is extremely important in numerous cellular processes, including innate and adaptive immunity, inflammation, bone metabolism and tissue development. 3
The innate immune system provides the first line of host defence against invading pathogens. 4 TLR is considered to be an essential class of PRRs in the innate immune system. TRAF6 acts as an adaptor molecule for various receptors such as IL-1 receptor or TLR, and causes NF-κB activation via the MyD88-dependent signalling pathway. 5 After LPS stimulation, MyD88 is recruited to the TLR4 complex, and IκB kinase is subsequently activated via TRAF6. 6 IL-1 receptor-associated kinase (IRAK)-1 is recruited to the TLR4 receptor complex via MyD88 and then phosphorylated by IRAK4. TRAF6, together with Ubc13/Uev1A (E2 ubiquitin-conjugating enzyme), is recruited onto multi-subunit complexes, where it mediates the formation of free K63-linked polyubiquitin chains and causes the activation of TGF-β-activated kinase 1. 7 Thus, NF-κB and MAPK pathways are activated by phosphorylated TGF-β-activated kinase 1, which in turn induces the expression of pro-inflammatory cytokines and antiviral molecules, including IFN-α, IFN-β, IFN-γ, IL-6, IL-8, IL-12, IL-1β and TNF-α. 8 , 9
Over the last few years, there has been substantial progress in understanding the crucial role of TRAF6 in mammals. 10 , 11 Although PRRs, including pigeon TLR7, are known to play essential roles in antiviral innate immunity, 12 here, we characterised the genomic sequence, phylogenetic relationship and tissue-specific mRNA expression patterns of TRAF6 in pigeons and evaluated the importance of piTRAF6 in NF-κB activation and IFN-β induction. We also analysed the expression of piTRAF6 and pro-inflammatory cytokines in vitro in pigeon PBMC treated with TLR ligands, Newcastle disease virus (NDV) or Salmonella Pullorum.
Materials and methods
Experimental pigeons and tissue collection
Healthy King pigeons (4–5 mo old) were purchased from Jiangyin Wei Tekai Pigeon Co. (Wuxi, PR China), housed in isolators and fed a pathogen-free diet and water. All animal experiments were approved by the Animal Welfare and Ethics Committees of Yangzhou University, and complied with the guide lines of Institutional Administrative Committee and Ethics Committee of Laboratory Animals (IACUC license number: SYXK (Su) 2016–0020). Pigeons were euthanised via inhalation of inert gases (N2). Tissues of 18 pigeons (random sex) from three independent experiments, including the lung, kidney, liver, heart, spleen, large intestine, small intestine, caecum, rib, thigh muscle, brain, bursa of Fabricius and muscular stomach, were snap-frozen in liquid nitrogen and preserved at −70°C before RNA extraction.
RNA extraction and cDNA synthesis
RNA extraction from pigeon tissues was performed using the E.Z.N.A.™ Total RNA Kit I (Omega Bio-Tek, Norcross, GA). Purified RNA was treated with RNase-free DNase I (Takara Biotechnology Co., Dalian, PR China) to remove contaminating DNA, in accordance with the manufacturer’s instructions. The reaction was performed for 50 μl mixture containing 50 μg total RNA, 5 μl 10× DNase I Buffer, 2 μl recombinant DNase I (RNase-free) and 20 IU RNase Inhibitor. The reaction mixture was incubated at 37°C for 30 min. Then, 2.5 μl 0.5 M EDTA was added, followed by incubation at 80°C for 2 min. Next, 10 μl 3 M sodium acetate and 250 μl cold ethanol were added, followed by incubation at −80°C for 20 min. The supernatant was discarded after centrifugation at 13,800 g for 10 min at 4°C. The precipitate was dried and dissolved in DEPC-treated water after cleaning with 70% cold ethanol. Agarose electrophoresis was performed to confirm that genomic DNA had been removed. RNA of several tissues was quantified using a One drop™ spectrophotometer (Wins Technology Co. Ltd, Nanjing, PR China). The PrimeScript RT Reagent Kit (Takara) was used to reverse transcribe total RNA into cDNA, in accordance with the manufacturer’s instructions. The reaction was performed in a total volume of 20 μl, containing 1 μg total RNA, 4 μl 5× PrimeScript Buffer 2, 4 μl RT Primer Mix and 1 μl PrimeScript RT Enzyme Mix I. The reaction mixture was incubated at 37°C for 15 min, followed by incubation at 85°C for 5 s, subsequently diluted to 250 ng/μl and stored at −20°C until further use.
Molecular cloning of pigeon TRAF6
To clone pigeon TRAF6, we used the predicted pigeon TRAF6 coding sequence (GenBank accession number XM_005500241.2) to design the primers piTRAF6-F and piTRAF6-R (Table 1). The fragment of piTRAF6 was amplified using PCR by employing the PrimeSTAR® Max DNA Polymerase (Takara). The reaction was performed in a total volume of 50 μl, containing 25 μl 2× PrimeSTAR Max Premix, 0.2 µM primers piTRAF6-F and piTRAF6-R (Table 1), with 250 ng of cDNA template from spleen tissues as the source of DNA. The PCR programme was as follows: pre-denaturation at 94°C for 5 min, followed by 34 cycles at 95°C for 50 s, 66°C for 30 s and 72°C for 1 min 36 s; the final extension was carried out at 72°C for 10 min. The amplified PCR product was purified using the Takara MiniBEST Agarose Gel DNA Extraction Kit v4.0 (Takara), sub-cloned into the pMD20-T vector with the TA Cloning Kit (Takara). The recombinant plasmid pMD20-T-piTRAF6 was transferred into competent Escherichia coli DH5α cells (Takara), and positive clones were verified by sequencing.
PCR primers used in this study.

Sequence analysis
Conserved N-glycosylation sites (Asn-Xaa-Ser/Thr) in the piTRAF6 protein were predicted by the NetNGlyc server (http://www.cbs.dtu.dk/services/NetNGlyc/), and the conserved phosphorylation sites were predicted by the NetPhos server (http://www.cbs.dtu.dk/services/NetPhos/). The molecular mass and theoretical isoelectric point of piTRAF6 were analysed using a ProtParam tool (http://web.expasy.org/compute_pi/). The simple modular architecture research tool (http://smart.embl-heidelberg.de/) was employed to predict the domain structure of piTRAF6. Homologous TRAF6 sequences were queried using the BLAST programme (http://blast.ncbi.nlm.nih.gov/Blast.cgi) with default settings on the GenBank database. A multiple sequence alignment was constructed using ClustalX2 (http://www.clustal.org/clustal2/), and a phylogenetic tree was constructed using the MEGA 7 programme (http://www.megasoftware.net/mege) that employed the neighbour-joining method.
Plasmid construction
The entire open reading frame of piTRAF6 was amplified using the primers piTRAF6-F and piTRAF6-R, containing restriction enzymes sites KpnI and XhoI (Takara), as shown in Table 1. The pcDNA3.1-piTRAF6 plasmid was constructed by sub-cloning the full-length of piTRAF6 into the same restriction enzyme sites of the pcDNA3.1 (+) expression vector (Invitrogen, Carlsbad, CA). The recombinant eukaryotic expression plasmid pcDNA3.1-piTRAF6 was transferred into DH5α cells and sequenced by the GenScript Company. Recombinant plasmid was extracted using the E.Z.N.A.™ Endo-free Plasmid Mini Kit I (Omega Bio-Tek) according to the manufacturer’s instructions and stored at −70°C until further use.
Cell culture, transfection and luciferase assay
No pigeon cell lines were available. Hence, HEK293T cells were used to analyse the function of piTRAF6. HEK293T cells were cultured at 37°C under a 5% (v/v) CO2 atmosphere in DMEM (Hyclone, Logan, UT) containing 10% (v/v) FBS (Gibco, Grand Island, NY), 100 IU/ml penicillin and 100 μg/ml streptomycin (Gibco). For transfection, cells were seeded onto 24-well plates (2 × 105 cells/well) and cultured for 12 h until they attained a density of 70–80%. PBS was used to wash the cells twice before transfection, and the medium was then replaced with Opti-MEM medium (Invitrogen). Cells were transfected with different concentrations of eukaryotic expression plasmids (12.5, 25, 50, 100 or 200 ng) or an empty vector (50 ng) together with 50 ng luciferase reporter pGL4.32 (luc2P/NF-κB-RE/Hygro) plasmid DNA (Promega, Madison, WI) or luciferase reporter plasmid of human IFN-β promoter (p-125Luc; a gift from Dr. Takashi Fujita at the Laboratory of Molecular Genetics, Institute for Virus Research, Kyoto University, Kyoto, Japan) using the Lipofectamine 3000 Reagent (Invitrogen) in accordance with the manufacturer’s instructions. Cells were harvested at 24 h after transfection. NF-κB or IFN-β-induced luciferase activity was determined using the Dual-Luciferase® Reporter Assay system (Promega) in accordance with the manufacturer’s instructions.
Small interfering RNA interference experiment
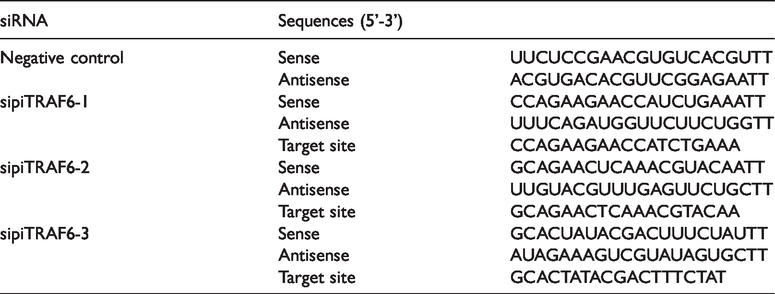
To investigate the function of piTRAF6 during NF-κB activation further, three small interfering RNAs (siRNAs; GenePharma Co., Shanghai, PR China) targeting piTRAF6 mRNA regions were prepared (Table 2). HEK293T cells were transfected with 100 nM siRNA (negative control, sipiTRAF6-1, sipiTRAF6-2 or sipiTRAF6-3) together with 50 ng pcDNA3.1-piTRAF6 plasmid, 50 ng NF-κB promoter luciferase reporter plasmid and Lipofectamine 3000 Reagent (Invitrogen) in accordance with the manufacturer’s instructions. After a 24-h transfection process, cells were harvested for analysis by performing a luciferase assay.
Sequences of siRNAs used in this study.
Western blotting
His tag was added to piTRAF6, and its expression in transfected HEK293T cells was analysed using anti-His Ab. The primers piTRAF6-His-F/R (Table 1) were designed to amplify piTRAF6-His. Cells were lysed in 2× Laemmli buffer (Sigma–Aldrich, St Louis, MO) for Western blotting and subjected to SDS-PAGE. Proteins were subsequently transferred to a nitrocellulose membrane (Pall Corporation, Port Washington, NY). The membranes were blocked with blocking buffer (0.05% Tween 20 and 1% BSA in PBS) for 2 h at room temperature and subsequently incubated for another 2 h with a mAb against the His tag (1:5000 diluted in blocking buffer; Roche, Basel, Switzerland) or anti-β-actin Ab (1:1000 diluted in blocking buffer; Sigma–Aldrich). After washing four times with PBST (0.05% Tween 20 diluted in PBS), the membranes were incubated at 37°C for 1.5 h with goat anti-mouse IgG-HRP secondary Ab (1:8000 diluted in blocking buffer; Sigma–Aldrich). Images of Ab reactions with an ECL chemiluminescence substrate (Thermo Fisher Scientific, Waltham, MA) were acquired using the Amersham Imager 600 Imaging System (GE Healthcare Life Sciences, Pittsburgh, PA).
Preparation and stimulation of pigeon PBMC
Pigeon PBMC (piPBMC) were isolated from the whole blood of pigeons. First, blood samples were collected from healthy King pigeons and mixed with 3.8% sodium citrate solution at a 5:1 (v:v) ratio. This mixture was combined with an equal volume of Dulbecco’s PBS (Gibco). Second, the mixture was layered onto an equal volume of Histopaque-1077 (Sigma–Aldrich) and centrifuged at 650 g for 20 min at 20°C. The white gradient interface was collected, washed with RPMI 1640 media (Invitrogen) and centrifuged at 200 g for 8 min at 4°C. After two additional washes with RPMI 1640, cells were diluted in RPMI 1640 containing 10% (v/v) FBS (Gibco), 100 IU/ml penicillin and 100 μg/ml streptomycin (Gibco). The trypan blue exclusion method was used to detect cell viability and numbers.
After piPBMC were added to 24-well culture plates, the cells were stimulated with the following TLR ligands: Pam3CSK4 (InvivoGen, Toulouse, France) for TLR1/2, 100 ng/ml; Poly (I:C) (InvivoGen) for TLR3, 2.5 μg/ml; LPS (Sigma–Aldrich) for TLR4, 100 ng/ml; flagellin (InvivoGen) for TLR5, 100 ng/ml; FSL-1 (InvivoGen) for TLR2/6, 100 ng/ml; R848 (InvivoGen) for TLR7/8, 2.5 μg/ml; and CpG:ODN (InvivoGen) for TLR9, 2 μM. Cells were collected at 1, 2, 4, 6 and 8 h post stimulation (hps) and stored at −70°C.
Separate preparations of piPBMC were cultured on 24-well culture plates (2 × 105 CFU/well) and exposed to 104 EID50 of LaSota (Wuhan Chopper Biology Co., Wuhan, PR China), an NDV vaccine strain. Cells were collected at 0, 1, 2, 4, 6 and 8 h post exposure (hpe). Salmonella Pullorum S06004 (GenBank accession no. CP006575.1) is a virulent clinical isolate obtained in 2006 from chickens with pullorum disease in the Jiangsu Province of China and stored in our laboratory. 13 Culturing of piPBMC was carried out on 24-well culture plates up to a concentration of 2 × 105 CFU/well and then exposed to S06004 at a concentration of 1 × 106 CFU/well. Cells were collected at 2, 4, 8, 12 and 24 hpe. Control cells were collected at 0 hpe. The cell lysates were processed for subsequently detecting the mRNA of immunologic genes.
Quantitative RT-PCR
The ABI 7500 Real-Time PCR System (Applied Biosystems, Foster City, CA) was used to measure the mRNA expression of piTRAF6, IFN-α, IFN-β, IL-6, IL-8, IL-1β, TNF-α, IFN-γ, IL-4, IL-12 p35, IL-12 p40 and NF-κB. Pigeon β-actin was used as an endogenous control gene. The gene-specific primers used in this analysis are listed in Table 1. The reaction (20 μl total volume) contained 2 μl (500 ng) cDNA, 0.6 μl each of 10 μM forward and reverse primers, 10 μl 2× SYBR Green Master Mix (Takara) and 6.8 μL RNAse Free Water. The comparative threshold cycle (2–ΔΔC(T) method) was used to calculate relative concentrations. All quantitative PCR (qPCR) reactions were performed in triplicate from three independent experiments.
Statistical analysis
All data are presented as the mean ± SE of triplicate samples per experimental condition from three independent experiments using the statistical software Instat v6.01 (GraphPad Software, La Jolla, CA). The differences between the experimental and control groups at each time point were tested using multiple Student’s t-test or one-way ANOVA test followed by Bonferroni’s multiple comparison test, with P values of < 0.05, < 0.01 or < 0.001 being considered statistically significant.
Results
Cloning and characterisation of piTRAF6 gene
The entire coding region of the piTRAF6 gene was successfully amplified from the cDNA of pigeon lung. The open reading frame sequence of piTRAF6 is 1638 bp in length and encodes a putative polypeptide of 545 aa with a molecular mass of 62.03 kDa and a theoretical isoelectric point of 5.74. We deposited the piTRAF6 sequence obtained by us into GenBank under accession number KU243360. There was no difference between the cDNA sequence of piTRAF6 we obtained and the predicted piTRAF6 coding mRNA sequence, according to the sequence alignment. The putative aa sequence of piTRAF6 consisted of an N-terminal low-complexity domain, a RING-type Zinc finger, a TRAF-type Zinc finger, a coiled coil region and a C-terminal meprin and TRAF homology domain. Additionally, piTRAF6 contained 14 potential serine phosphorylation sites, four potential threonine phosphorylation sites, five potential tyrosine phosphorylation sites and two potential N-glycosylation sites.
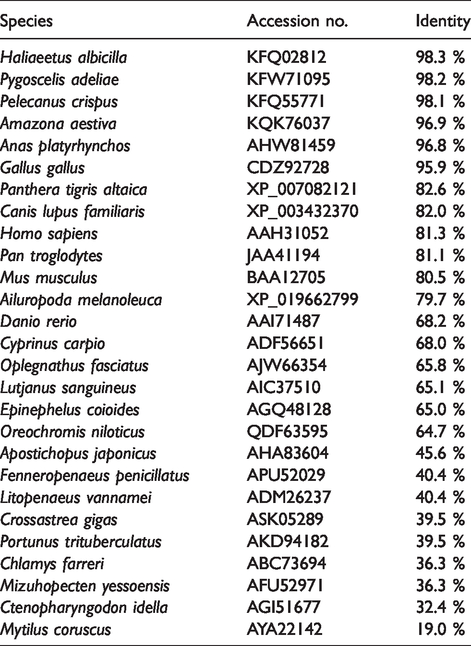
Multiple sequence alignment revealed that the TRAF6 sequence was conserved in those species (Figure 1A). Homology analysis using protein BLAST revealed that piTRAF6 had various degrees of sequence homology with various TRAF6s that were previously submitted to the database (Table 3). As expected, piTRAF6 shared a relatively high level of identity with the corresponding sequences obtained from other birds (98.3–95.9%) but a comparatively low level of identity with corresponding sequences from mammals (82.6–79.7%) and fish (68.2–19.0%). A phylogenetic tree was generated based on TRAF6 protein sequences from different species. The tree revealed three main clusters that corresponded to TRAF6 from birds, mammals and fish, where piTRAF6 was clustered together with TRAF6s from Gallus gallus, Anas platyrhynchos and Anas platyrhynchos, respectively (Figure 1B).
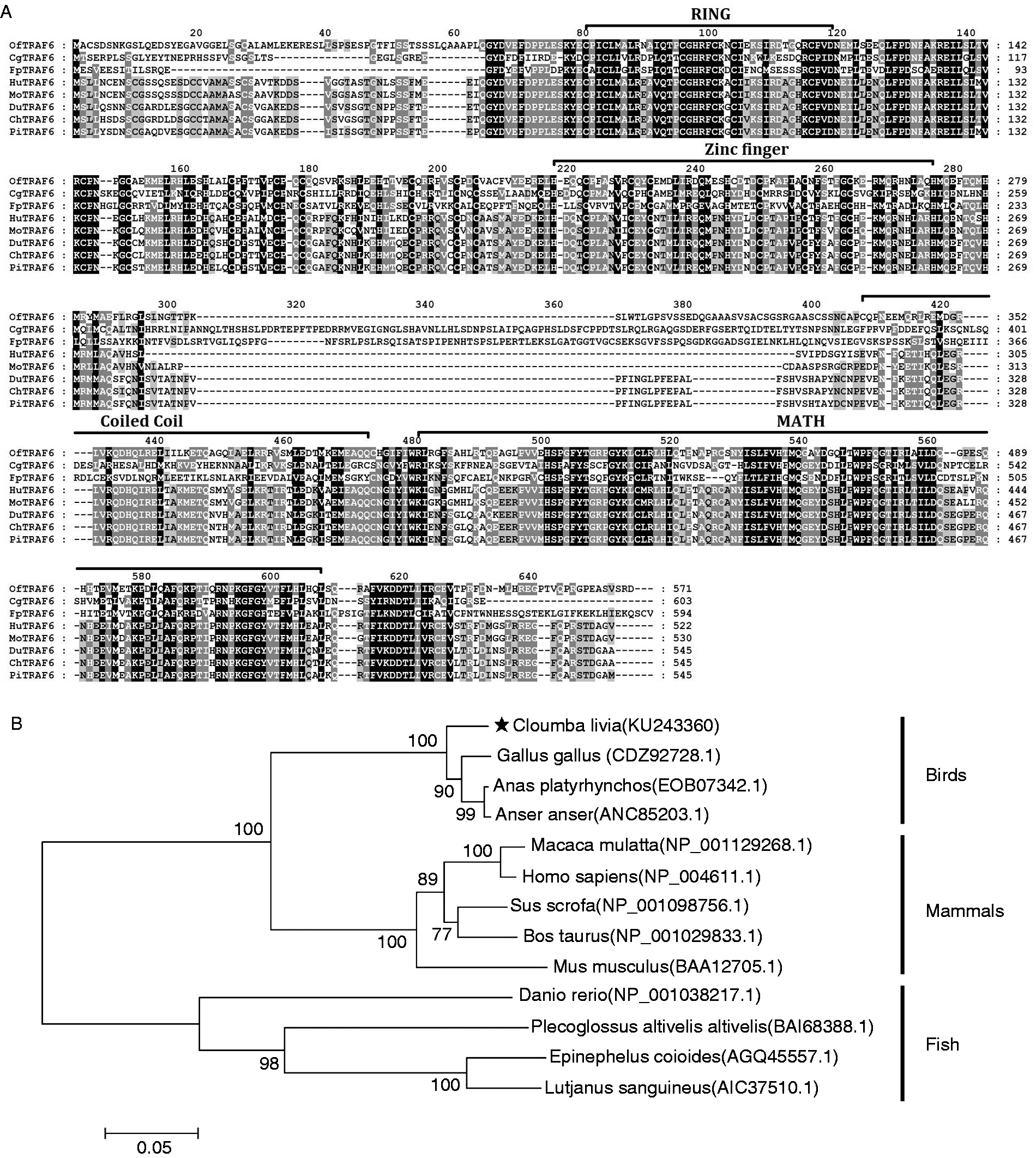
Sequence analysis of piTRAF6. (A) Alignment of Oplegnathus fasciatus (OfTRAF6, accession no. AJW66354), Crossastrea gigas (CgTRAF6, accession no. ASK05289), Fenneropenaeus penicillatus (FpTRAF6, accession no. APU52029), Homo sapiens (HuTRAF6, accession no. AAH31052), Mus musculus (MoTRAF6, accession no. BAA12705), Anas platyrhynchos (DuTRAF6, accession no. AHW81459), Gallus gallus (ChTRAF6, accession no. CDZ92728) and piTRAF6 aa sequences performed using the ClustalX2 programme and edited using the Genedoc program. (B) A phylogenetic tree showing relationships of piTRAF6 and other known TRAF6 proteins. The tree was constructed by the neighbour-joining method using MEGA 7 software. The bar (0.05) indicates the genetic distance.
Protein sequence identity of TRAF6 between pigeon and other species.
Expression of piTRAF6 mRNA in different tissues
To determine the tissue-specific expression of piTRAF6 mRNA, qPCR analysis was performed using pigeon β-actin as the internal control. As shown in Figure 2, piTRAF6 was expressed in all examined tissues, but its level of expression varied, and the highest level of piTRAF6 transcript was approximately 30-fold higher in the heart compared to that in the muscular stomach, the tissue of which exhibit the lowest level of piTAF6 transcript. PiTRAF6 expression in the spleen, lung and liver was approximately 20-fold higher than that in the muscular stomach.
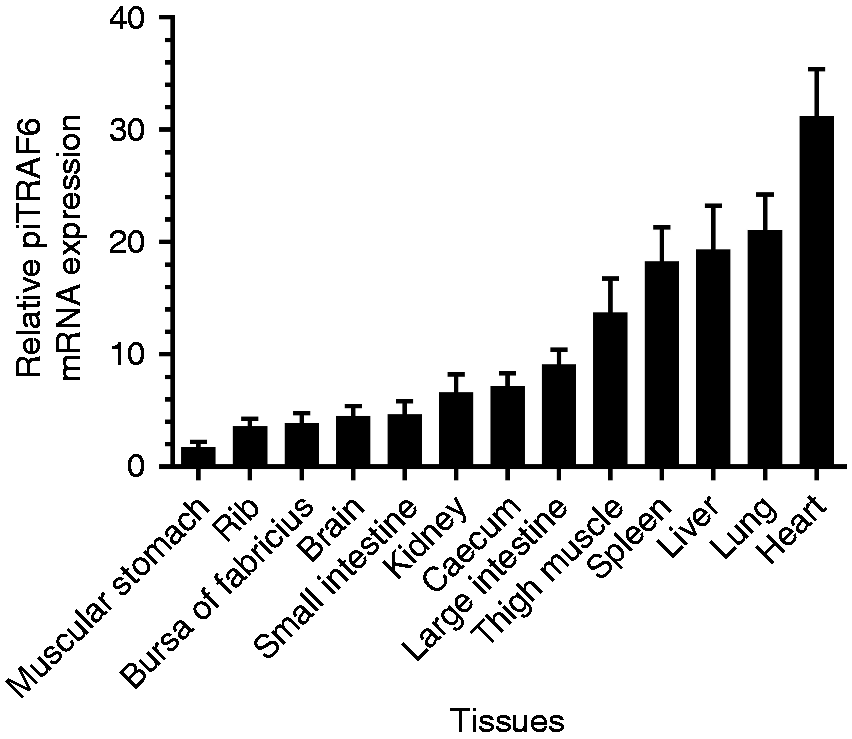
Relative mRNA expression levels of piTRAF6 in various tissues were determined using quantitative PCR (qPCR). The mRNA expression levels were normalised to pigeon β-actin transcript. Tissue of the muscular stomach was chosen as the calibrator (1). Data are presented as the mean ± SE of triplicate samples per experimental condition from three independent experiments.
Functional analysis of piTRAF6
Western blotting using an anti-His mAb confirmed successful expression of piTRAF6 in HEK293T cells transfected with pcDNA3.1-piTRAF6-His plasmid (Figure 3), whereas proteins extracted from control cells and transfected with empty vector cells showed no reaction with the anti-His Ab.
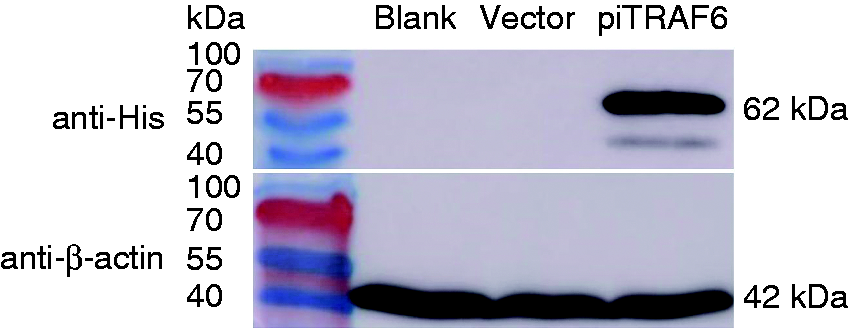
Western blotting of pigeon TRAF6 in transfected HEK293T cells. The control HEK293T cells (lane 1), cells transfected with empty vector pcDNA3.1 (lane 2) and cells transfected with pcDNA3.1-piTRAF6-His plasmid (lane 3) were harvested, and then the proteins were subjected to Western blotting using anti-His or anti-β-actin Ab (as loading control).
As shown in Figure 4A, overexpression of the full-length piTRAF6 activated NF-κB, whereas empty vector did not. Additionally, the overexpression of piTRAF6 in HEK293T cells dramatically activated NF-κB in a dose-dependent manner; high doses (100 and 200 ng) of piTRAF6 further up-regulated the activity of NF-κB by approximately fourfold compared to NF-κB activity associated with low doses (25 and 12.5 ng; Figure 4B). To confirm whether activation of NF-κB was required for the expression of IFN-β in the MyD88- and TRIF-dependent signalling pathways, we tested IFN-β promoter activity in piTRAF6-expressing HEK293T cells using an IFN-β promoter luciferase reporter system assay. We found that piTRAF6 overexpression can significantly activate IFN-β promoter activity in HEK293T cells as opposed to transfection with an empty vector (Figure 4C).
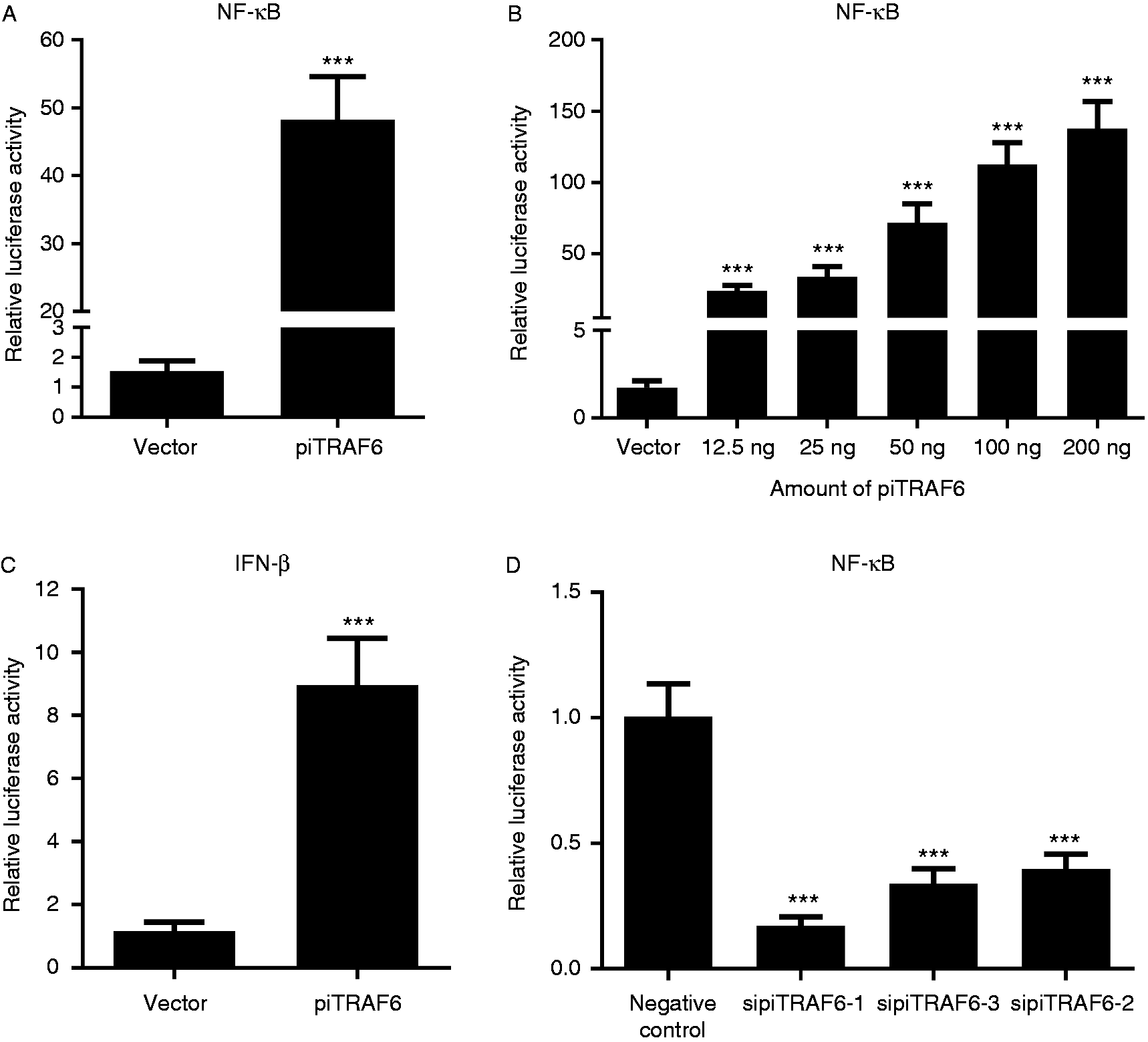
Overexpression of piTRAF6 can activate NF-κB and IFN-β expression in HEK293T cells. (A) NF-κB activity induced by piTRAF6. HEK293T cells were transiently transfected with empty vector or pcDNA3.1-piTRAF6 expression plasmid (200 ng/well), along with NF-κB-Luc reporter plasmid (50 ng/well), using Lipofectamine 3000. (B) HEK293T cells were transfected with empty vector (50 ng/well) or different doses of pcDNA3.1-piTRAF6 expression plasmids (12.5 ng/well, 25 ng/well, 50 ng/well, 100 ng/well or 200 ng/well), together with the NF-κB-Luc reporter plasmid (50 ng/well), using Lipofectamine 3000. (C) HEK293T cells were transiently transfected with empty vector (50 ng/well) or pcDNA3.1-piTRAF6 expression plasmids (200 ng/well), along with IFN-β-Luc reporter plasmid (50 ng/well) and using Lipofectamine 3000. (D) HEK293T cells were transfected with 100 nM sipiTRAF6-1, sipiTRAF6-2, sipiTRAF6-3 or negative control, together with the NF-κB-Luc reporter plasmid (50 ng/well), using Lipofectamine 3000. Multiple Student’s t test (Figure 4A and C) or one-way ANOVA followed by Bonferroni’s multiple comparison test (Figure 4B and D) indicated significant findings in comparison to cells transfected with vector. Statistical significance was defined as P < 0.001 (***). Data are presented as the mean ± SE of triplicate samples per experimental condition from three independent experiments.
Luciferase reporter assays revealed that as opposed to the induction of NF-κB activation by the negative control, transfection with all three siRNAs targeting different piTRAF6 mRNA regions reduced NF-κB activation in HEK293T cells, while sipiTRAF6-1 had the most obvious inhibitory effect, with an activation level approximately fourfold lower than that associated with the negative control (Figure 4D).
Modulation of piTRAF6 and inflammatory cytokine expression following treatment with TLR ligands
To investigate in vitro modulation of piTRAF6 gene expression in response to the stimulation with various TLR ligands, mRNA levels of piTRAF6, IFN-α, IFN-β, IL-6, IL-8, IL-1β and TNF-α were determined by qPCR. First, we explored the optimal time of stimulation for R848. As shown in Figure 5A–C, piTRAF6 as well as IFN-α and IL-8 expression levels increased at 1 hps, peaked at 2 hps and declined to basal level at 8 hps. Therefore, the time point of 2 hps was selected for subsequent TLR ligand stimulation experiments. In Pam3CSK4- and Poly (I:C)-treated piPBMC, a moderate increase (∼4-fold) in TRAF6 expression was observed, and a highly significant induction (∼7.5-fold) was noted in R848-treated piPBMC (Figure 5D). IFN-α and IFN-β mRNA expression was markedly increased after stimulation with Pam3CSK4, Poly (I:C) or flagellin (Figure 5E and F). Compared to control cells, IL-8 and IL-1β expression was significantly up-regulated in piPBMC stimulated with FSL-1 (Figure 5H and I). The mRNA levels of IL-6, IL-8, IL-1β and TNF-α significantly increased in response to Pam3CSK4 or R848 stimulation (Figure 5G–J). Moreover, IL-1β transcription was significantly promoted in flagellin-treated piPBMC as opposed to that in control cells (Figure 5I).
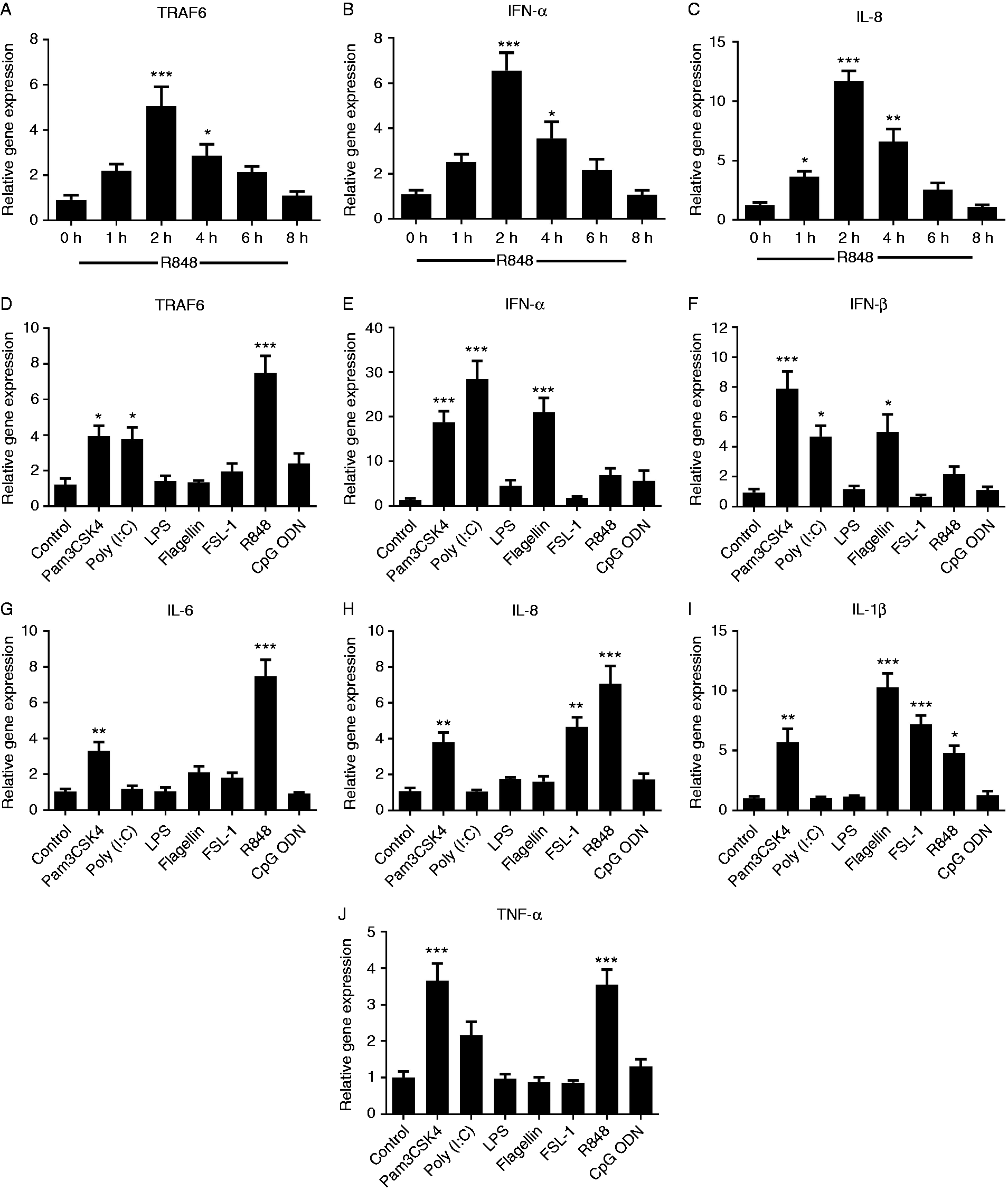
Transcription of piTRAF6 (A), IFN-α (B) and IL-8 (C) in pigeon PBMC after stimulation with TLR7/8 ligand R848 (2.5 μg/ml) for 0, 1, 2, 4, 6 and 8 h. Expression of piTRAF6 (D), IFN-α (E), IFN-β (F), IL-6 (G), IL-8 (H), IL-1β (I) and TNF-α (J) in pigeon PBMC following stimulation with various TLRs ligands for 2 h: Pam3CSK4 (TLR1/2, 100 ng/ml), poly (I:C) (TLR3, 2.5 μg/ml), LPS (TLR4, 100 ng/ml), flagellin (TLR5, 100 ng/ml), FSL-1 (TLR2/6, 100 ng/ml), R848 (TLR7/8, 2.5 μg/ml) and CpG (TLR9, 2 μM). One-way ANOVA followed by Bonferroni’s multiple comparison test indicated significant findings in comparison to the control group. Statistical significance was defined as P < 0.05 (*), P < 0.01 (**) or P < 0.001 (***). Data are presented as the mean ± SE of triplicate samples per experimental condition from three independent experiments.
Expression profiles of piTRAF6 and cytokines in response to NDV and S06004 inoculation
To obtain further insights into the innate immune responses of piTRAF6 against pathophysiological conditions, we subsequently investigated the transcript levels of piTRAF6 and cytokine mRNAs in piPBMC post exposure to the NDV vaccine strain LaSota and S06004, respectively. A significant up-regulation of piTRAF6 was observed in piPBMC at 2 and 4 hpe as opposed to the control cells (Figure 6A). IFN-α and IFN-β expression was enhanced at 1 hpe, peaked at 2 hpe and decreased to the basal level at 6 hpe (Figure 6B and C). The time course analysis of IL-8 and IL-1β in piPBMC showed that the induction of both IL-8 and IL-1β peaked at 1 hpe (∼10- and ∼5-fold, respectively) and thereafter it gradually decreased (Figure 6E and F). In comparison to control cells, IL-6 expression was sustained at a higher concentration in piPBMC after NDV exposure (Figure 6D). TNF-α analysis revealed its significant induction at 2 hpe (∼2.5-fold) and 4 hpe (∼4-fold; Figure 6G) as opposed to that in the control cells.
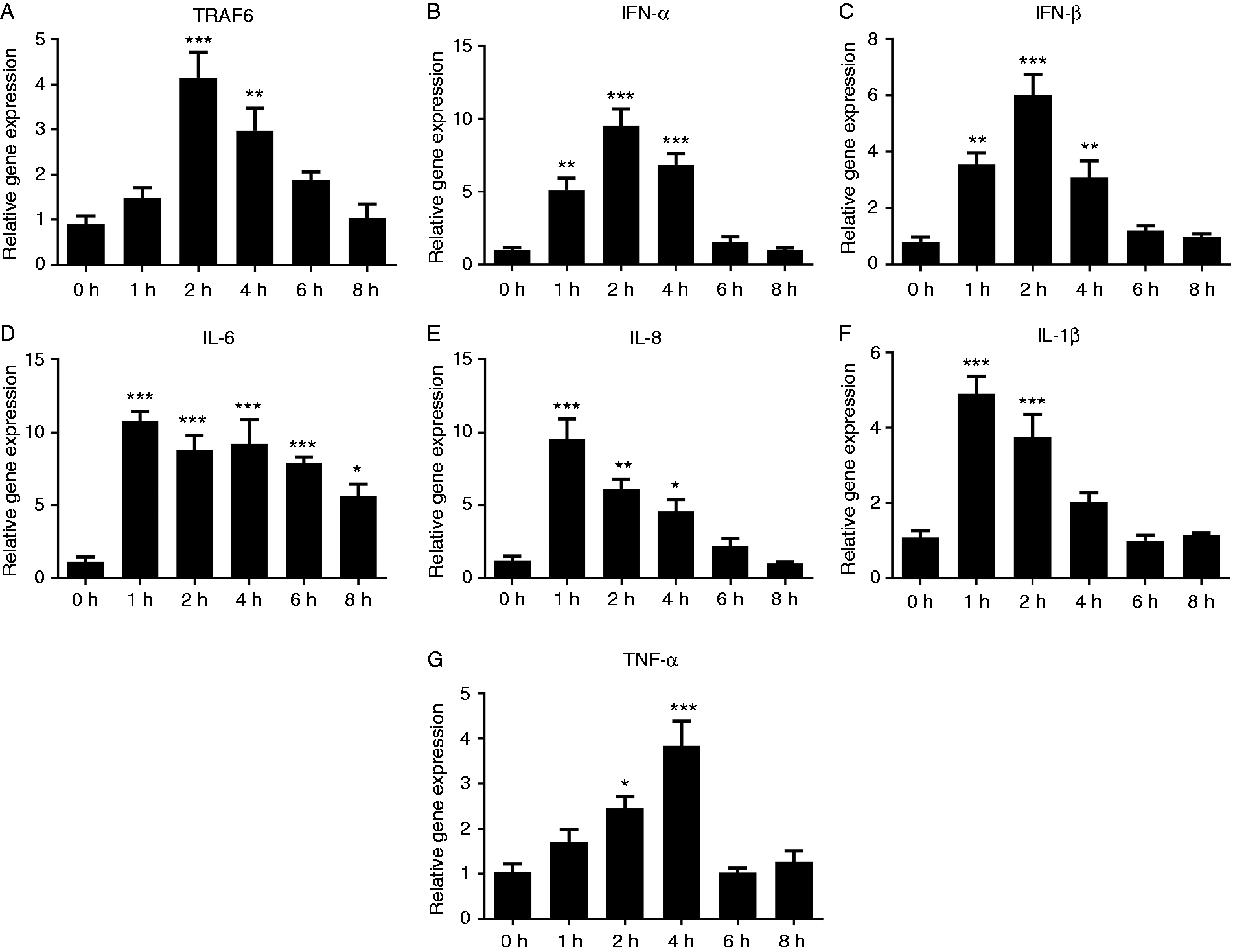
mRNA levels of piTRAF6 (A), IFN-α (B), IFN-β (C), IL-6 (D), IL-8 (E), IL-1β (F) and TNF-α (G) in pigeon PBMC exposed to NDV for 0, 1, 2, 4, 6 and 8 h were measured using qPCR. One-way ANOVA followed by Bonferroni’s multiple comparison test indicate significant findings in comparison to the control group. Statistical significance was defined as P < 0.05 (*), P < 0.01 (**) or P < 0.001 (***). Data are presented as the mean ± SE of triplicate samples per experimental condition from three independent experiments.
As shown in Figure 7A, E and K, the mRNA expression level of piTRAF6 as well as that of IL-4 and TNF-α was significantly up-regulated in piPBMC at 12 h after exposure to Salmonella Pullorum S06004 and then markedly increased at 24 hpe (∼7-, ∼160- and ∼170-fold, respectively). The expression of NF-κB was also significantly promoted simultaneously (Figure 7L). Meanwhile, IFN-α and IL-8 (Figure 7B and G) expression levels were slightly up-regulated at 4 hpe and peaked at 24 hpe (∼90- and ∼12-fold, respectively). Exposure to S06004 also caused significant up-regulation of IFN-β, IFN-γ, IL-12 and IL-1β transcription at 8, 12 and 24 hpe (Figure 7C, D, H, I and J), whereas the expression patterns of these four cytokines were similar. Furthermore, the mRNA level of IL-6 alone was strongly induced at 2 hpe and remained high at 24 hpe compared to that in the control cells (Figure 7F).
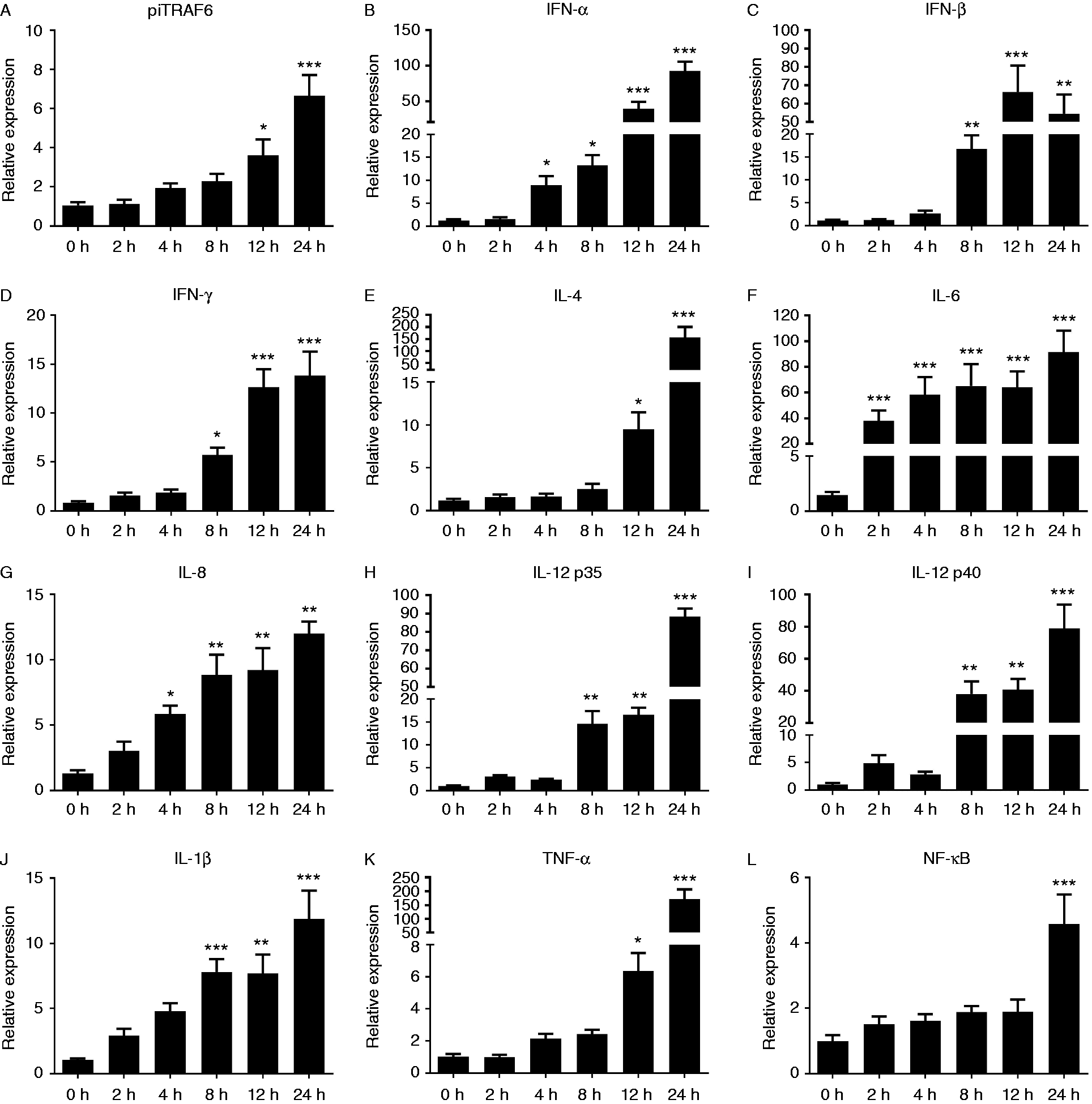
Expression of piTRAF6 (A), IFN-α (B), IFN-β (C), IFN-γ (D), IL-4 (E), IL-6 (F), IL-8 (G), IL-12 p35 (H), IL-12 p40 (I), IL-1β (J), TNF-α (K) and NF-κB (L) in pigeon PBMC exposed to Salmonella Pullorum for 0, 2, 4, 8, 12 and 24 h were measured using qPCR. One-way ANOVA followed by Bonferroni’s multiple comparison test indicate significant findings in comparison with the control group. Statistical significance was defined as P < 0.05 (*), P < 0.01 (**) or P < 0.001 (***). Data are presented as the mean ± SE of triplicate samples per experimental condition from three independent experiments.
Discussion
TRAF6 acts as an important signal transducing adaptor, interacting with cell surface receptors and involved in signalling to regulate the immune and inflammatory processes. 14 The TLR-MyD88-IRAK4/Pelle-TRAF6 signalling pathway has been detected and evolutionarily conserved for antibacterial and antiviral defence mechanisms in organisms ranging from Drosophila to humans. 15 Here, a TRAF6 homolog was identified and characterised from the King pigeon. Like most TRAF proteins, piTRAF6 are composed of a highly conserved carboxy-terminal TRAF domain and a variable amino-terminal domain, containing a RING finger and a Zinc finger. Previous studies revealed that the coiled coil domain was essential for auto-ubiquitination of TRAF6 and downstream signalling events. 16 And the meprin and TRAF homology domain were required for homo- and heterodimerisation of TRAF and interaction with upstream factors, including IRAK-4. 17 Conservation of the domain features and arrangements in the consensus aa sequence of piTRAF6 indicated that piTRAF6 encoded a functional homolog of TRAF6, which thereby suggested similar functions described for other TRAF6s. In addition, the ubiquitination ligase activity of TRAF6 is coordinated via the more variable RING domain, which was required for efficient IκB kinase and NF-κB activation. 18 The different variable regions of TRAF6 from different species may also contribute to host immune defence and may survive in different hostile pathogenic environments.
PiTRAF6 mRNA was detected in all examined tissues using qPCR, suggesting that piTRAF6 is ubiquitously and constitutively expressed. The observed expression profiles suggest diverse biological functions of TRAF6 in each tissue. 19 Extensive tissue expression of TRAF6 has also been reported in Danio rerio, 20 Anas platyrhynchos, 21 Cyprinus carpio, 22 Cirrhinus mrigala 23 and Ctenopharyngodon idella. 24 The most abundant expression was observed in the gills, muscle, liver, kidney and head kidney. PiTRAF6 was highly expressed in the heart tissue, and the pattern of TRAF6 mRNA expression distinctively varied among different species, suggesting that TRAF6 expression pattern is species specific. Abundant expression of piTRAF6 was observed in the immune system–related tissues, namely those of the lung and spleen, as observed in Anas platyrhynchos, 21 indicating that piTRAF6 might be a critical factor in the immune system. These results implied that the expression of piTRAF6 was differentially regulated in various tissues and may contribute to the immune response against bacterial pathogens.
NF-κB is a key regulator of inducible gene expression in the immune system, meaning that both the innate and adaptive immune responses, as well as the development and maintenance of the cells and tissues that comprise the immune system, are under the control of the NF-κB family of transcription factors. 25 Ubiquitination of TRAF6 is an essential event for the activation of the NF-κB pathway. 26 Previous studies in mammals 27 , 28 and fish 20 have demonstrated that TRAF6 is involved in the regulation of the NF-κB signalling pathway. Here, the function of piTRAF6 was indicated by the activation of NF-κB only in piTRAF6-transfected HEK293T cells and not in control cells, as observed in Epinephelus coioides, 29 Crassostrea gigas, 30 Litopenaeus vannamei 15 and Anas platyrhynchos. 21 However, in TRAF6-deficient macrophages, ligands of TLR1, TLR5, TLR7 and TLR9 were all unable to induce NF-κB activation and failed to induce the production of related inflammatory cytokines. 31 As observed in this study, piTRAF6 knockdown blocked the NF-κB pathway. This observation indicated that piTRAF6 might be associated with a diverse array of biological processes, as demonstrated in others species. 32 Furthermore, IFN-β transcription in the MyD88- and TRIF-dependent signalling pathways required NF-κB activation. 33 TRAF6 is believed to play essential roles in the regulation of IFN-β expression in response to viral infection. The potential roles of bird TRAF6 in IFN response have recently been confirmed by the evidence that overexpression of Anas platyrhynchos TRAF6 can induce activation of the IFN-β promoter, 21 consistent with the findings of the present study. Such data indicated that piTRAF6 was essential for NF-κB and IFN-β activation, as well as for the innate immune response.
TLRs play a key role in the activation of innate immune cells, in which their engagement leads to the production of cytokines and co-stimulatory molecules. 34 To induce the activation of NF-κB and MAPK through the same MyD88-dependent signalling pathway, which leads to cytokine production, all TLRs require TRAF6. 35 A recent study revealed that TRAF6 was required for the balanced production of pro-inflammatory cytokines (TNF-α, IL-6, IL-8, IL-1β and type I IFN) by monocyte-derived dendritic cells after stimulation with R848, an agonist of human TLRs 7/8. 34 This is consistent with the findings of the present study. The piTRAF6 and downstream inflammatory cytokines mRNA transcript levels were up-regulated in poly (I:C)- and Pam3CSK4-stimulated piPBMC. Similarly, the release of poly (I:C), Pam3CSK4, LPS, R848 and CpG-mediated IL-6, TNF-α and IFN-β was impaired by blocking the TLR-TRAF6 signalling pathways. 36 Moreover, inhibiting K63-linked polyubiquitination of TRAF6 markedly down-regulated LPS-mediated neutrophil infiltration and TNF-α, IL-6 and IL-1β production. Flagellin could induce the expression of pro-inflammatory cytokines in fish by activating TLR5-TRAF6 signalling pathway. 23 The expression of IL-6, IL-8 and IL-1β was enhanced in FSL-1-stimulated human cells via the MyD88-NF-κB-TRAF6 pathway, 37 consistent with the observation in the present study.
Pigeons are particularly susceptible to the genotype VI genetic variants of NDV, which are also known as pigeon paramyxovirus 1. These viruses are mainly isolated from Rock pigeons (Columba livia). 38 Pigeon paramyxovirus 1 is endemic in domestic and feral pigeons across many regions worldwide, including Europe and the USA, 39 thereby causing significant economic damage to the poultry industry. The morbidity rate of infected pigeons was in the range of 30–70%, and the mortality rate was 10–30%. 40 Infection with NDV can trigger the expression of type I and II IFN, several cytokines and chemokines, IFN effectors and inducible NO synthase genes in the spleen, macrophages, splenic leucocytes and embryo fibroblasts. 41 Previous studies have demonstrated that virus-triggered ubiquitination of TRAF6 was essential for type I IFN induction and cellular antiviral response. 42 Similar results were found in this study, that is, the expression of piTRAF6 and pro-inflammatory cytokines (IFN-α, IFN-β, IL-6, IL-8, IL-1β and TNF-α) was markedly up-regulated in piPBMC after NDV exposure. Additionally, viral RNA and DNA can trigger immediate antiviral responses in human and mouse cells, leading to the rapid production of type I IFN. 43 As observed in this study, piTRAF6 and cytokines were significantly induced in piPBMC after NDV exposure, especially in the early stages. These data suggested that piTRAF6 was essential for the activation of antiviral innate immune responses in pigeons.
Salmonella Pullorum is a highly adapted pathogen that causes pullorum disease in poultry, which in turn causes severe economic losses owing to the reduced production of eggs, low hatchability and high morbidity and mortality rates. 44 Salmonellosis was considered as the most important zoonoses in bacterial diseases that can be transmitted by the domestic pigeons, 45 , 46 making them a public health risk. Salmonella species were frequently isolated from pigeons from several areas worldwide, including Europe, Japan, South America and the USA. 47 , 48 Previous studies proved that Salmonella Pullorum was able to modulate host immunity from a dominant IFN-γ-producing Th17 response towards a Th2 response 49 and that Salmonella Pullorum infection also increased IL-12, IL-4, IL-6, IL-8 and IL-1β expression. These observations were consistent with the findings of the present study. Moreover, IL-8 and TNF-α as well as TRAF6 expression were markedly induced after Edwardsiella tarda infection. 23 In the present investigation, the expression of piTRAF6, NF-κB and related downstream cytokines significantly increased after Salmonella Pullorum infection. This finding was in agreement with the findings of previous studies that TRAF6, IL-6, TNF-α, IL-1β, type I IFN or NF-κB production significantly increased after parasitic or bacterial infections. 24 , 50 , 51
In summary, a new TRAF family member, piTRAF6, was identified from the King pigeon Columba livia. PiTRAF6 was broadly expressed in all tested tissues. Overexpression of piTRAF6 markedly activated NF-κB in a dose-dependent manner and induced IFN-β expression. Furthermore, our results indicated that the expression of piTRAF6, along with inflammatory cytokines and antiviral mediators, was remarkably up-regulated after stimulation with TLR ligands or exposure to NDV or Salmonella Pullorum. These findings suggest that piTRAF6 was a key molecule in the TLR-TRAF6 signalling pathway and that it was involved in innate immune defence against viral and bacterial infections in pigeons.
Footnotes
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship and/or publication of this article: This work was supported by the National Natural Science Foundation of China (Grant Number 31972685), the China Postdoctoral Science Foundation (Grant Number 2018M642333), the Yangzhou University Science and Technology Innovation Team (2016) and the Priority Academic Program Development of Jiangsu Higher Education Institutions (PAPD).
