Abstract
BMAL1 (brain and muscle ARNT-like protein 1, also known as MOP3 or ARNT3) belongs to the family of the basic helix–loop–helix (bHLH)-PAS domain-containing transcription factors, and is a key component of the molecular oscillator that generates circadian rhythms. Here, we report that BMAL1-deficient cells are significantly more susceptible to infection by two major respiratory viruses of the Paramyxoviridae family, namely RSV and PIV3. Embryonic fibroblasts from Bmal1–/– mice produced nearly 10-fold more progeny virus than their wild type controls. These results were supported by animal studies whereby pulmonary infection of RSV produced a more severe disease and morbidity in Bmal1–/–mice. These results show that BMAL1 can regulate cellular innate immunity against specific RNA viruses.
Keywords
Introduction
Essentially, all living organisms display circadian (∼24 h) rhythms in physiology and behavior that are genetically programmed and can be cell intrinsic.1–3 In mammals, circadian system is organized as network of circadian clocks located in every tissue with the central clock located in the suprachiasmatic nucleus of hypothalamus. At a cellular level circadian clocks operate as interconnected transcriptional–translational negative and positive feedback loops. 4 Basic helix–loop–helix Per–Arnt–Sim (bHLH) domain transcription factors BMAL1 and CLOCK (or NPAS2) form a transcriptional complex that positively regulates the activity of multiple E-box-containing promoters,2,3 and expression of other components of the circadian clock such as Periods (Pers), Cryptochomes (Crys), Rev-Erbs and retinoic acid receptor-related orphan receptors (RORs). In turn, PER and CRY act as negative regulators of the BMAL1:CLOCK complex and suppress their own expression, while Rev-Erbs and RORs regulate BMAL1 expression. In addition, circadian clock transcription factors regulate the expression of multiple clock-controlled genes (CCGs), which is believed to produce the circadian clock output in metabolism and physiology. Microarray analyses have revealed that ∼10% of genes in various tissues are directly or indirectly regulated by the circadian clock system.5,6 In fact, it is likely that all physiological functions are under some sort of the circadian control.
BMAL1 is a key component of the circadian clock; 3 mice deficient in BMAL1 exhibit severely impaired circadian behavior and loss of rhythmicity in the expression of BMAL1 target genes. 7 BMAL1-deficient mice also developed many physiological abnormalities such as defects in glucose homeostasis, 8 increased oxidative stress,9,10 increased mTOR signaling, 11 impaired insulin production 12 and accelerated aging.13–15
In view of the pleiotropic phenotype of Bmal1 mutations, we inquired whether BMAL1 may also regulate cellular innate immunity against pathogens. Members of the Paramyxoviridae family of viruses contain non-segmented negative-strand RNA genome and have an intricate relationship with the immune immunity of the host. We have paid special attention to respiratory syncytial virus (RSV) and parainfluenza virus type 3 (PIV3), two pediatric viruses that cause significant morbidity and mortality globally, and for which no reliable vaccine of specific cure currently exists. 16 In the more extensively studied virus, RSV, infection leads to regulation of a plethora of host cell signaling pathways that include apoptosis, NF-κB, miRNA synthesis, reactive oxygen species, cellular stress, cytokines and chemokines, and IFN response, to name only a few.17–26 Very recently, RSV infection has been shown to induce double-stranded DNA breaks and cellular senescence, promoted by reactive oxygen, 27 which is reminiscent of a Bmal1–/– phenotype. It is also well known that RSV is more virulent in older individuals than in young adults. 28 Despite the multimodal nature of BMAL1 function and host–virus interaction, it has remained largely untested whether circadian factors influence cell-based innate immunity against pathogens.
Materials and methods
Cell culture, infection and knockdown studies
Cells were routinely grown in monolayers in DMEM supplemented with l-glutamine, penicillin (100 IU/ml), streptomycin (100 µg/ml) and heat-inactivated FBS (10%). Human RSV (long strain) and vesicular stomatitis virus (VSV; Indiana) were grown in HEp2 cells, and human PIV3, in LLC-MK2 cells. For VSV growth, serum was omitted from the media. For BMAL1 studies, multiple Bmal1 knockout (KO; Bmal1–/–) and wild type (WT; Bmal1+/+) immortalized lung fibroblasts (ILF) clones 11 were used, and representative results of each group were presented. Infectious viral titer was determined by plating serial dilutions on HEp-2 cell monolayers and foci counting.
To knockdown BMAL1 expression, A549 cells were transfected with proprietary duplex Silencer® Select ARNT (Bmal1) siRNA (cat# S1614) or Silencer Negative Control siRNA (cat# AM4611, designed by Ambion) from Invitrogen (ThermoFisher Scientific, Waltham, MA, USA), following the manufacturer's instructions.
Abs
Rabbit polyclonal VSV Ab that detects viral G, N and M proteins was a kind gift from Dr Michael Whitt. RSV Ab was purchased from Chemicon (goat, AB1128) and PIV3 HN Ab from Abcam (mouse monoclonal, M02122321, ab49756). Abs against the following were from Santa Cruz Biotechnology (Santa Cruz, CA, USA): β-actin (mouse monoclonal, sc-81178), GAPDH (G-9) (mouse monoclonal, sc-365062) and BMAL1 (goat polyclonal, N-20, sc-8550). Rat Ab against a synthetic peptide sequence of PIV3 nonstructural protein C was recently made commercially (Biosynthesis, Lewisville, TX, USA). The following HRP-conjugated secondary Abs, used in immunoblot, were all purchased from Santa Cruz Biotechnology: Mouse anti-goat (sc-2354); goat anti-mouse IgG (sc-2031); (sc-2032); goat anti-rabbit (sc-2030).
Animal experiments
Bmal1–/– mice 7 were obtained from Dr C Bradfield (University of Wisconsin), and backcrossed to C57BL/6J inbred strain (The Jackson Laboratory, Bar Harbor, ME, USA) for 12 generations. WT and KO mice were generated by breeding of heterozygous parents. Genotypes were determined using a PCR-based method as previously described. 7 Animals were maintained on a 12:12 light:dark cycle with lights on at 7:00 am, and fed on an 18% protein rodent diet (Harlan, Indianapolis, IN, USA). The experiments were performed in 4–5-mo-old mice, before accelerated aging phenotype developed. Around 12:00 pm on d 0, a total of 12 Bmal1–/– and 10 wild type mice were intranasally infected with 108 pfu of RSV Long as described previously. 29 The mice were weighed and/or sacrificed at indicated days post-infection; the lung were isolated and the infectious RSV titer in the clarified lung homogenates was determined by plaque assay on HEp2 monolayer exactly as described. 29 All animal experiments were IACUC-approved.
Immunoblot
Lysis of cells, SDS-PAGE and immunoblotting were performed as described. 30 HRP, conjugated to the secondary Ab, was developed by chemiluminescence, and the bands were detected either by autoradiography using HyBlot CL film (E3012; Denville Scientific, Holliston, MA, USA) or by the LI-COR Odyssey Fc imaging system.
Densitometry and statistical computation
Densitometric quantification of the immunoblot bands were performed by ImageJ (NIH, Bethesda, MD, USA). 31 All graphs were plotted in Excel (Microsoft, Redmond, WA, USA), each column (bar) representing the mean ± SD of three values. Two-tailed P-value of the fold change from WT to BMAL1-deficient was determined in unpaired t-test, calculated with GraphPad software (GraphPad Inc., San Diego, CA, USA).
Results
Loss of BMAL1 allows better replication of select viruses in cell culture
To determine if Bmal1 has any relation with virus replication, we started with several independent ILF clones isolated from homozygous Bmal1–/– mice, and grew them in cell culture. In parallel, several WT ILF cell lines were also grown. We then tested the ability of multiple viruses and a parasite to replicate in these cells. Owing to our continuing interest in host–virus interaction in RNA viruses, we tested three viruses of the Mononegavirales order (commonly called non-segmented negative-strand RNA virus, or NNR virus), namely RSV, PIV3 and VSV. Whereas both RSV and PIV3 are members of the Paramyxoviridae family, they represent different genera; RSV belongs to the Pneumovirus genus, and PIV3 belongs to the Respirovirus genus. Both viruses are major causative agent of respiratory disease, mainly in children and the elderly. 32 VSV, in contrast, belongs to the Rhabdoviridae family, and is a significant pathogen in cattle and horses.
Regardless of their natural host and tissue tropism, all three viruses are promiscuous in cell culture, being able to infect essentially all types of cells. We infected the ILF cell lines with these viruses and measured virus growth by immunoblotting, as well as by progeny virus assay. We tested four cell lines each of WT and Bmal1–/– mutant, and present representative results of each assay (Figure 1). First, immunoblot of viral proteins revealed that replication of both RSV and PIV3 was higher in the Bmal1–/– cell lines compared with the WT (Figure 1a, b). Remarkably, the results with VSV were the opposite (Figure 1c): while the WT ILF cells robustly supported VSV replication, the Bmal1–/– cells were nearly resistant to this virus. To see if BMAL1 has any role in the replication of pathogens outside the virus kingdom, we tested the growth of Toxoplasma gondii, a unicellular protozoan and an obligatory intracellular parasite of mammalian cells. Clinically, T. gondii is a major human pathogen, capable of replicating in essentially all types of nucleated cells, and is a causative agent of systemic toxoplasmosis, blindness and miscarriage.
33
As shown (Figure 1d), T. gondii replicated equally well in WT and Bmal1–/– cells. We conclude that BMAL1 affects RNA viruses but in a differential manner: it functions against RSV and PIV3, the two members of the Paramyxoviridae family, but in favor of VSV, a prototype member of the Rhabodoviridae family.
Differential effect of BMAL1 on diverse pathogens. Multiple clonal lines of mouse ILF cells isolated from Bmal1–/– (KO) and WT animals were grown in monolayers and infected with either of the following: (a) human RSV (Long strain, serotype A); (b) human PIV3 (JS strain); (c) VSV (Indiana strain); or (d) Toxoplasma gondii (RH strain). In (a), (b) and (d) the cells were harvested at the indicated time points post-infection (0, 12, 18, 24 h). In (c), as VSV growth is much faster, all cells were harvested at 12 h post-infection but infected at different MOIs, as shown. Immunoblotting (Western) of the total cell proteins were performed as described under ‘Materials and methods’. The detected protein bands are indicated on the right. Actin and GAPDH serve as loading controls.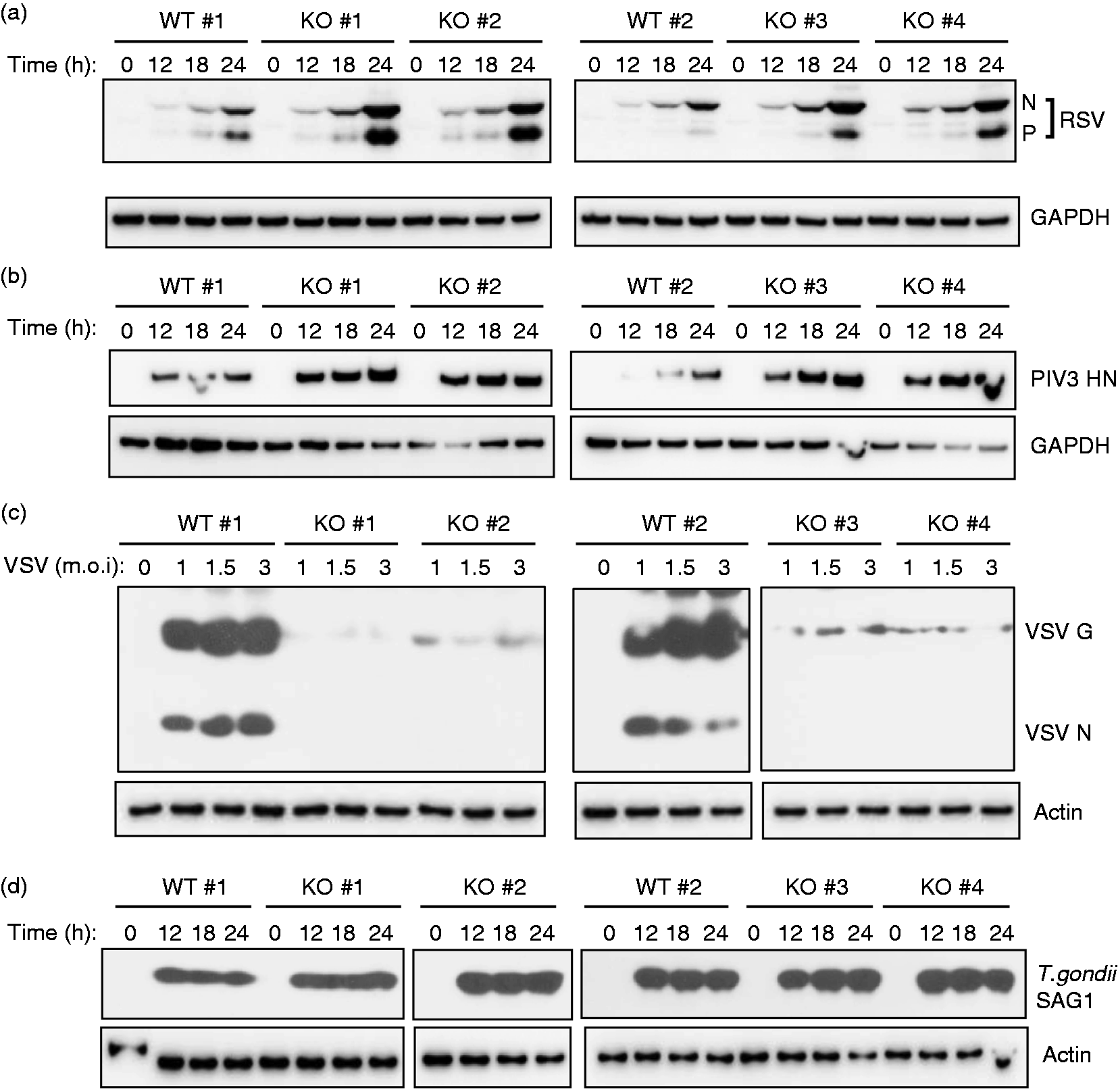
Although the KO cells offered a completely BMAL1-null environment, they were mouse cells that are not natural hosts for human RSV and PIV3 strains. In order to validate the apparently antiviral role of BMAL1 in human cells, we took A549 cells, an alveolar carcinoma cell line of human origin and widely used in human respiratory virus studies. We silenced Bmal1 expression by transfection with specific duplex siRNA (Figure 2a). For optimal silencing without risking non-specific or off-target effects, we then chose a concentration of 100 nM siRNA, which caused 72% knockdown of the BMAL1 protein levels (Figure 2a). We then infected these cells with RSV and PIV3, and tested virus growth by immunoblotting as before. The results (Figure 2b, c) generally mirrored the previous results with KO ILF cells, in that the BMAL1-knockdown A549 cells also supported higher virus growth. However, in visual examination of the blot, the fold stimulation did not appear as high, which we presumed to be owing to the residual amounts of BMAL1 in the knockdown cells (Figure 2a). Nonetheless, we performed a quantitative estimate of the viral band intensities by densitometry, which revealed that RSV and PIV3 growth in the knockdown cells was indeed significantly higher.
Antiviral effect of BMAL1 in human cells. (a) Bmal1 knockdown by siRNA in A549 cells was performed as described in ‘Materials and methods’; 18 h after siRNA transfection, the total cell protein was analyzed by immunoblot with BMAL1 Ab. NS: non-specific siRNA. Residual BMAL1 in siRNA-treated cells were determined by densitometry, normalized against GAPDH band intensities, and expressed as percentage of the siRNA-untreated value. For unknown reasons, the NS siRNA promoted a modest increase of BMAL1 levels. Measurement of (b) RSV and (c) PIV3 growth in A549 cells, treated with 100 nM siRNA (NS or Bmal1), were carried out as described in Figure 1, the only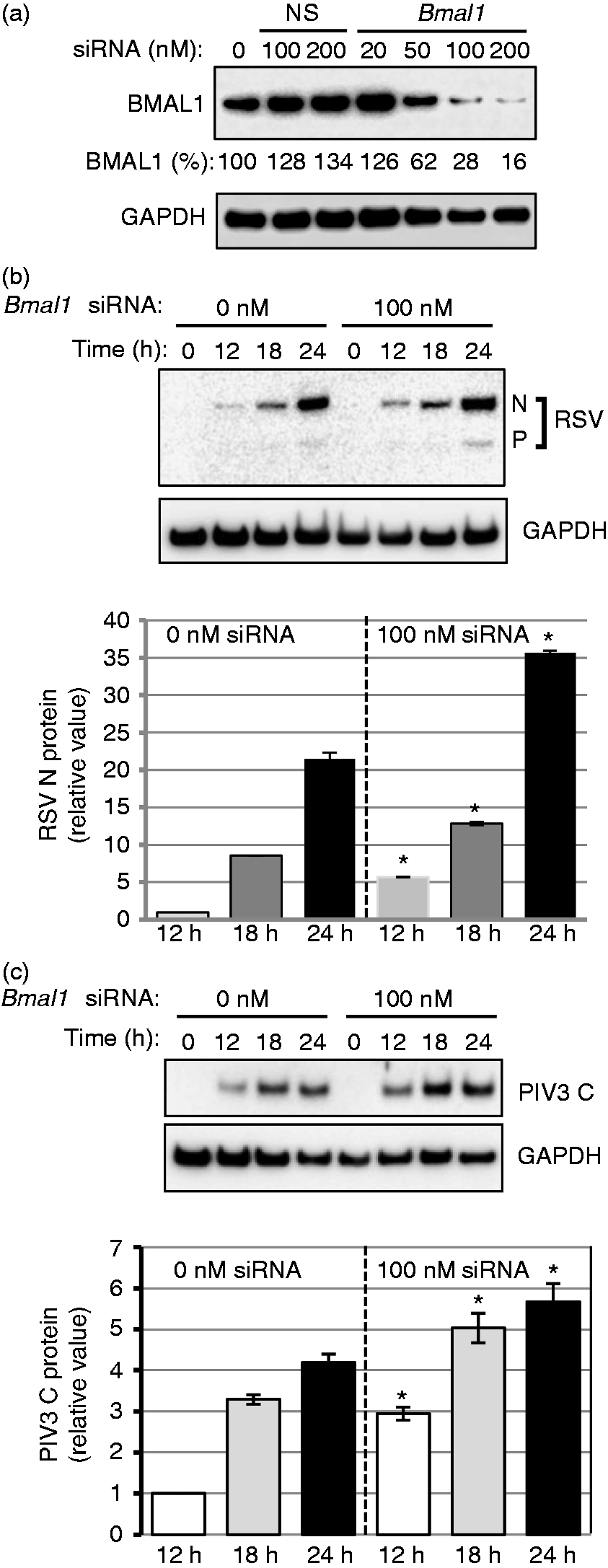
Effect of BMAL1 on liberated progeny virus
The immunoblot results presented above measured intracellular virus growth. To test if the effect of BMAL1 is also reflected in the ultimate release of the progeny viruses, we conducted viral infection for longer periods (48 h) and quantified the infectious liberated RSV in the culture media by plaque assay on HEp2 cells. These results (Figure 3) paralleled the immunoblot results for all three viruses, i.e. the Bmal1–/– ILF cells, as well as the Bmal1-knockdown A549 cells, liberated ∼5–10 times more RSV and PIV3 than the WT, whereas it was the opposite with VSV.
Loss of BMAL1 boosts infectious virus yield. A set of Bmal1–/– (KO) ILF cells (as in Figure 1) and Bmal1 knockdown (using 100 nM siRNA) (as in Figure 2) along with the control cells were infected with the indicated viruses. Media were collected at 30 h post-infection for RSV and PIV3, and at 20 h post-infection for VSV, and infectious viral titer determined as described in ‘Materials and methods’. Viruses from WT (or siRNA-untreated) cells are represented in gray bars, and those from BMAL1-deficient (KO or knockdown) cells, in black bars. *P ≤ 0.001.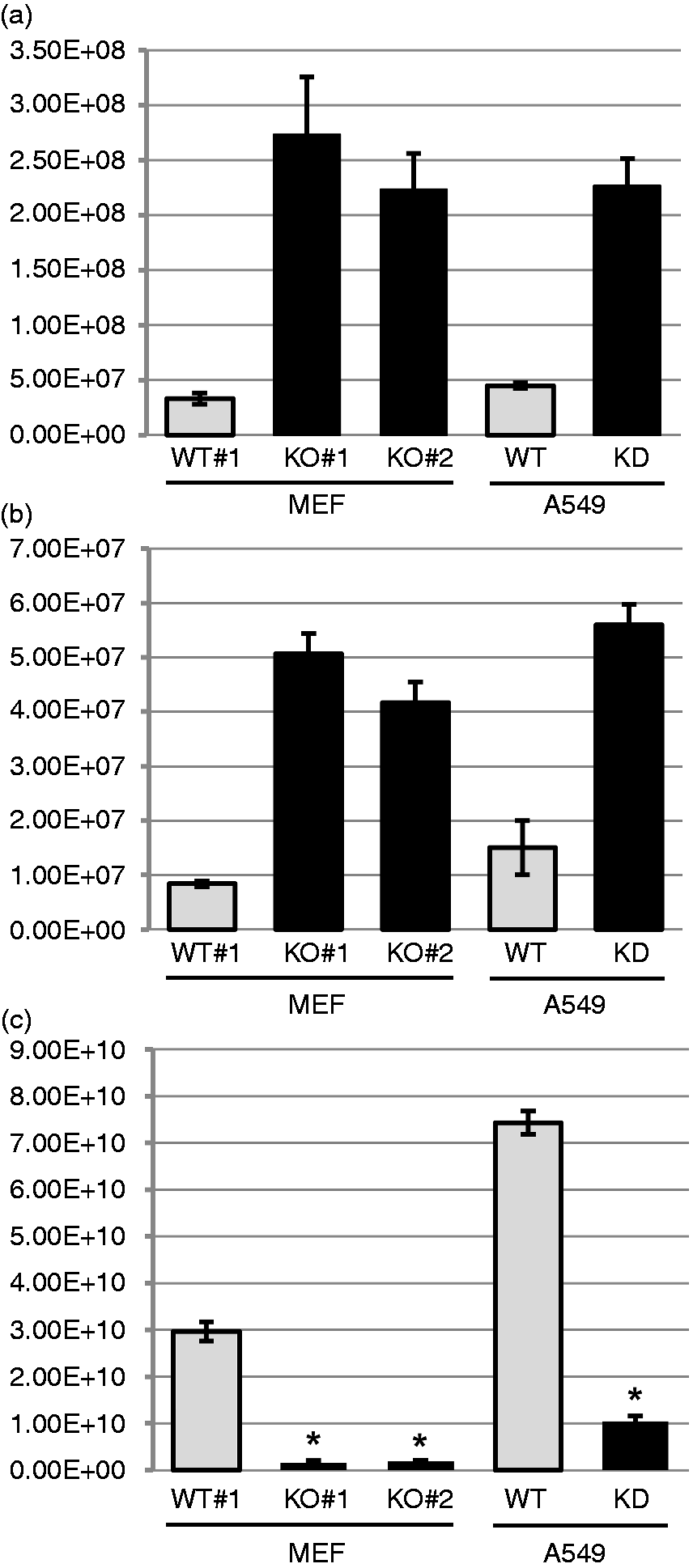
Enhanced severity of respiratory RSV infection in Bmal1 KO mice
Having shown that replication of RSV is higher in Bmal1–/– cells in culture, we asked whether this antiviral phenotype of BMAL1 translates in a live animal infection model. To test this, we conducted intranasal infection of WT and Bmal1–/– mice in the C57BL/6 background,
7
and monitored the animal body masses and RSV levels in the lung tissue, which are commonly used markers of RSV disease progression and severity in mice. As shown by both criteria (Figure 4), the Bmal1–/– mice fared substantially worse than their WT cohorts. Specifically, the Bmal1–/– mice suffered major loss of body mass, from which they never recovered, whereas the WT animal had the standard profile of a small amount of initial mass less, followed by regaining of the mass. The pulmonary RSV titer was higher in the Bmal1–/– mice at all days tested. Together, these results document a more severe RSV infection in the BMAL1-deficient mice.
Severe RSV disease in Bmal1–/– mice. These experiments have been described in ‘Materials and methods’. Two major outcomes of RSV infection were measured as shown: (a) mass loss; (b) viral titer in the lungs.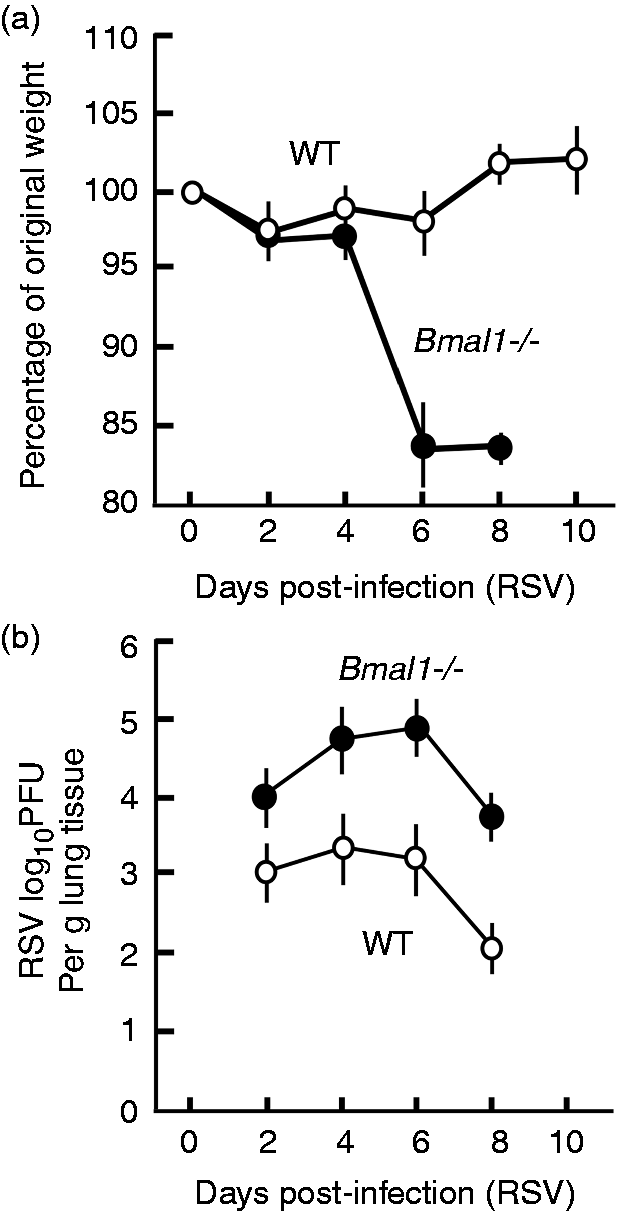
Discussion
Herein, we report that BMAL1, a circadian control protein, can act as a cellular defense factor against certain RNA viruses. Interaction between the circadian clocks and immune system is active field of study. 34 Recently, it was reported that BMAL1 regulates daily rhythms of inflammatory monocytes, 35 connecting circadian system and innate immunity. In present study, as the antiviral effect can be appreciated in cell culture, BMAL1 must have a cell-intrinsic role, likely affecting viral macromolecule synthesis. This can also explain the antiviral phenotype in the animal model, although we cannot rule out roles of additional host factors in the animal context, such as humoral immunity and cytokines that may be regulated by BMAL1. While our manuscript was being reviewed, Edgar et al. published a more detailed study documenting an essentially similar antiviral role of BMAL1. 36 In brief, these authors used murine and human herpesviruses (containing double-stranded DNA genome) to show that the time of day of infection affects the virus replication cycle in both live mice and cells. Interestingly, the timing effect was abolished and viral replication was significantly enhanced in Bmal1-deficient animals and cells, which was also seen with influenza A virus (IAV; an orthomyxovirus with negative-stranded segmented RNA genome). Taken together, our results and those of Edgar et al. 36 establish a relatively broad-spectrum antiviral role of BMAL1 that appears to be cell autonomous.
BMAL1 occupies a central place in the circadian clock regulatory circuit and is the single clock gene whose KO leads to loss of all clock activity. 7 However, as mentioned before, BMAL1 protein forms heterodimeric complexes with several other core clock proteins, notably, CLOCK, REV-ERBα, RORα, DEC1 and DEC2, thus creating complex regulatory loops and also involving various feedback regulatory circuits. 34 Recently, it was reported that circadian clock proteins interact with and regulate the activity of NF-κB pathway,37–39 and hence BMAL1 deficiency may disrupt this regulation. We have not explored the molecular mechanism of BMAL1–virus interaction, but as a transcriptional factor, BMAL1 may induce expression of host antiviral genes (or suppress a proviral gene). A large number of host genes code for antiviral innate immune proteins, such as the family of several hundred genes induced by type I IFN response (IFN-stimulated genes or ISGs).40,41 Although components of the molecular clock have been found in various tissues, mainly liver, skeletal muscle, kidneys and heart, only about 10% of the clock-controlled genes (CCGs) is shared between any two of them, suggesting a high degree of tissue specificity.5,6,42 There is currently no report on the CCGs of the lung tissue, which is the primary site of infection by RSV and PIV3. It will be interesting to find out if a subset of BMAL1-regulated CCGs in the lungs are antiviral, such as the ISGs.
The antiviral effect of BMAL1 on paramyxoviruses (RSV and PIV3; this study) and on herpesviruses and orthomyxoviruses, 36 but not on VSV, is intriguing. As the susceptible viruses span DNA and RNA virus families, BMAL1 is unlikely to directly affect viral polymerases (DNA-dependent RNA polymerase of herpesvirus and RNA-dependent RNA polymerases of RSV, PIV3 and IAV). Although VSV shares the Mononegavirales order with RSV and PIV3, it has important genetic differences with the latter ones. VSV, for instance, is a fast-growing virus with broad host tropism, 43 and its only glycoprotein, G, uses the ubiquitous low-density lipoprotein receptors as the major entry port. 44 PIV3 and RSV, in contrast, are relatively slower growing and encode diverse glycoproteins that interact with cell surface sialoproteins and various co-receptors, such as chemokine receptors and epidermal growth factor receptors.45–47 Furthermore, both RSV and PIV3 encode suppressors of IFN, namely the nonstructural proteins, NS1 and NS2, and the C protein, respectively,48–50 although their mechanism of action is still not fully clear. RSV also activates NF-κB through a novel kinase signaling pathway, which eventually leads to transcriptional activation of pro-inflammatory genes, important in RSV infection.51,52 The status of NF-κB in PIV3- and VSV-infected cells is currently unknown. We speculate that any of these known virus-specific regulatory networks or novel host genes may be modulated by BMAL1 and play opposite roles in RSV/PIV3 and VSV replication. Characterization of such genes and elucidation of how they are regulated by BMAL1 and how they inhibit each virus may lead to novel antiviral regimens, potentially independent of the circadian rhythm. It will also be important to know if the circadian role of BMAL1 is also relevant to this role, and how. As recently conjectured, 36 this may have important bearing on public health and susceptibility to virus infection, relevant in intercontinental travel (‘jet lag’), sleep deprivation, and seasonal or artificial variations in the lengths of light and dark periods.
Footnotes
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was supported, in part, by NIH grants AI109569 (S.B.), 1R01AG039547 (R.V.K.) and CSU start-up funds (S.B.).
