Abstract
Intra-mammary bacterial infections can result in harmful clinical mastitis or subclinical mastitis with persistent infections. Research during the last decades closely examined the pathophysiology of inflamed udders. Initial events after pathogen perception but before the onset of mastitis have not been examined in vivo. The objective of this study was to develop a mastitis model in cows by monitoring initial transcriptional pathogen-specific host response before clinical signs occur. We applied a short-term infection model to analyse transcripts encoding chemokines, cytokines and antimicrobial molecules in the teat cistern (TC) and lobulo-alveolar parenchyma (LP) up to 3 h after challenge with E. and Staphylococcus aureus. Both pathogens elicited an immune reaction by 1 h after challenge. Escherichia coli induced all analysed factors (CCL20, CXCL8, TNF, IL6, IL12B, IL10, LAP, S100A9); however, S. aureus failed to induce IL12B, IL10, LAP and S100A9 expression. The E. coli-induced up-regulation was 25–105 times greater than that after S. aureus challenge. Almost all the responses were restricted to the TC. The short-term mastitis model demonstrates that a divergent pathogen-specific response is generated during the first h. It confirms that the first transcripts are generated in the TC prior to a response in the LP.
Introduction
Intra-mammary bacterial infection is a common yet unresolved problem in dairy cows. It can cause clinical mastitis accompanied by an excessive inflammatory response or subclinical mastitis with persistent infections. 1 These different clinical pictures of disease are often related to the species of the causative pathogen. Intra-mammary infection with Escherichia coli usually leads to clinical mastitis associated with visible changes in milk, swelling and pain of the udder, and possibly a systemic inflammatory response syndrome severely affecting the cow. 2 In contrast, Staphylococcus aureus mostly accounts for subclinical mastitis lacking any visible signs of inflammation in the udder but leading to long-term losses through high milk somatic cell counts (SCC), reduced milk quality and quantity and an increased risk of spreading the pathogen within the herd.3,4
Mastitis still accounts for most of the use of antimicrobial drugs in dairy cows, despite considerable research efforts over the last decades. 5 Recently, better understanding of host–pathogen interaction has revealed that the clinical outcome of mastitis is decided at the stage of pathogen recognition.6–8 This new knowledge about the molecular nature and activation time points of defence mechanisms is crucial for the development of new therapeutic and prophylactic concepts aiming at modulating the cow’s immune response to mastitis pathogens.
Owing to their sheer abundance in the udder, mammary epithelial cells (MEC) and the lobulo-alveolar parenchyma (LP) have predominantly been studied with regard to host response during bovine mastitis. The teat and the gland cistern and its associated tissues have recently been the subject of several studies.9–11 It has been shown that during early stages of infection the transcriptional response is greater in these tissues compared with the LP, demonstrating a sentinel function within these distal compartments of the mammary gland.
The immune response in the udder is dominated by factors of innate immunity. 1 Regardless of the model system applied to analyse the host response, E. coli was always found to trigger a strong inflammatory response in the udder and MEC, while the response to S. aureus was always significantly weaker.6,12,13 Changes in the abundance of mRNA transcripts of immune-related genes are among the first host-associated events after pathogen invasion and recognition. 14 Differentially expressed genes in the early stages of infection mostly encode distinct cytokines, chemokines and antimicrobial peptides.9,15
Previous infection models sequentially challenged several udder quarters of the same animal at different times, allowing the analysis of different stages of intra-mammary infection and host response within one animal.9,10,13 However, at the time of tissue sampling these animals were already affected by clinical and subclinical mastitis. Thus, the results were biased by the systemic response of the host affecting all udder quarters, independent of whether they were pathogen challenged or not. This ‘crosstalk’ between udder quarters was more pronounced in animals affected by clinical E. coli mastitis rather than subclinical mastitis caused by S. aureus. 14
Mastitis research during the last decades has closely examined the pathophysiology of the inflamed mammary gland. However, little is known about the first transcriptional response before the disease becomes apparent by clinical signs (fever, udder swelling) or the increase of SCC in the milk. With regard to transcriptional events during the first 3 h after pathogen contact, knowledge is limited to in vitro experiments with mammary cells or tissue explants.6,8,11 Early stages of intra-mammary infection, in which pathogens have already been recognized but no clinical signs have yet appeared, have, up until now, not been satisfactorily examined in depth in vivo.
The aim of this paper was to develop a sequential mastitis infection model in cows uncovering the very first transcriptional pathogen-specific host responses at different time points of intra-mammary bacterial presence of E. coli and S. aureus before the occurrence of clinical signs. With regard to a location-specific response in the mammary gland, transcripts of genes known to be among the first regulated ones in the MEC and LP after pathogen contact should be analysed in different compartments of the teat and the udder.
Materials and methods
Animals
The animal experiments included 12 healthy, first-lactating Holstein cows, 26–35 mo of age and 59–185 d in milk. The in vivo trials were conducted at the Clinic for Ruminants, LMU Munich (Oberschleißheim, Germany) with the approval of the ethics committee of the regional government of Upper Bavaria, Germany (No. 55.2-1-54-2531-102-09, 21st September 2009). Only animals without previous diagnosis of clinical or subclinical mastitis and a reported SCC <50,000/ml were included in the study. Before the trial, weekly testing of quarter milk samples with a Fossomatic 5000 (FOSS Electric, Hillerod, Denmark) ensured that the milk contained <50,000 somatic cells/ml and was free from pathogens as assessed by plating 15 μl milk on Columbia sheep blood agar, on Violet red bile agar and on Edwards agar (all from Oxoid, Wesel, Germany). During a minimum period of 3 wk before the trial, the animals were transferred to the experimental unit (tie stall on straw) to guarantee an adequate adaption phase to the new environment, including the technical personnel. Three times per wk, the cows were carefully examined by a veterinarian to attest to their general health. They were milked twice daily at 12 h intervals and were fed a total mixed ration based on production level. To achieve comparable hormonal conditions, all the animals were synchronized by insertion of CIDR devices according to the manufacturer’s specifications (Zoetis, Berlin, Germany) so that at the time point of tissue sample collection all the animals were at the stage of oestrus.
E. coli and S. aureus model strains
The bovine mastitis isolates E. coli 1303 and S. aureus 1027 were isolated from cases of clinical and subclinical mastitis, respectively, and have previously been used in different in vivo and in vitro studies.6,13–15 The strain E. coli 1303 belongs to the major E. coli phylogenetic group A, and, based on a multiplex PCR screening for virulence-associated genes, it does not represent an extra-intestinal or intestinal pathogenic E. coli isolate. According to recent structural and genetic analyses, its LPS represents a novel serotype and possesses the K-12 core type. 16
The strain S. aureus 1027 is positive for mannite cleavage, clumping factor and free coagulase. 13 Furthermore, this strain represents the sequence type 133 and the agr-type 1 and expresses leukocidine [lukF‐PV(P83)/lukM] but no superantigens. 17
The second passage of the original isolates was kept cryo-conserved (Mikrobank-System Cryobank; Mast Diagnostika, Reinfeld, Germany) for subsequent preparation of the inoculum dose. The latter was prepared as previously described. 18 Bacteria were plated on Columbia sheep blood agar (Oxoid) and incubated (37℃) for 24 h. A few colonies were transferred to a tube of brain–heart infusion broth (Oxoid) and incubated for 6 h (37℃); then, a 100-μl sample was transferred to a tube containing 9.9 ml trypticase soy broth (Oxoid). Serial dilutions were made after 18 h to prepare the desired inoculum dose of 5 × 106 CFU/2 ml 0.9% sterile, pyrogen-free saline.
Intra-mammary pathogen challenge
A schematic overview of the sequential intra-mammary challenge model is given in Figure 1. The animals were randomly applied to two different treatment groups. In the Ec3h group six cows were experimentally challenged with E. coli, and in the Sa3h group six cows were challenged with S. aureus. At trial start the udder was entirely milked out after i.v. application of 20 IU Oxytocin (Veyx-Pharma, Schwarzenborn, Germany). After cleaning and disinfecting the teat end thoroughly with 70% ethanol, the first challenge dose (5 × 106 CFU/2 ml) was instilled into the hind left quarter intra-cisternally with a sterile syringe and teat cannula via the teat canal. To avoid any unwanted loss of the instilled challenge dose, the teat canal was kept closed by two fingers during removal of the teat cannula. The teat and the udder were not massaged after applying the challenge dose. One h after trial start, the front left udder quarters were challenged; another h later the hind right quarters were challenged as described above. The front right quarters received no challenge and are referred to as control quarters. The trial ended 3 h after the first challenge. At this time point the duration of pathogen presence was 1 h for hind right quarters, 2 h for front left quarters and 3 h for hind left quarters.
Experimental set-up of intra-mammary E. coli and S. aureus challenge. Three of four quarters per animal were sequentially inoculated with either 5 × 106 CFU E. coli (n = 6, group Ec3h) or 5 × 106 CFU S. aureus (n = 6, group Sa3h) for 1, 2 and 3 h. One control quarter (front right quarter) remained untreated and was used to measure baseline expression values. The animals were culled 3 h after inoculation of the first udder quarter and tissue samples from the TC, GC and LP were taken for RT-qPCR analysis.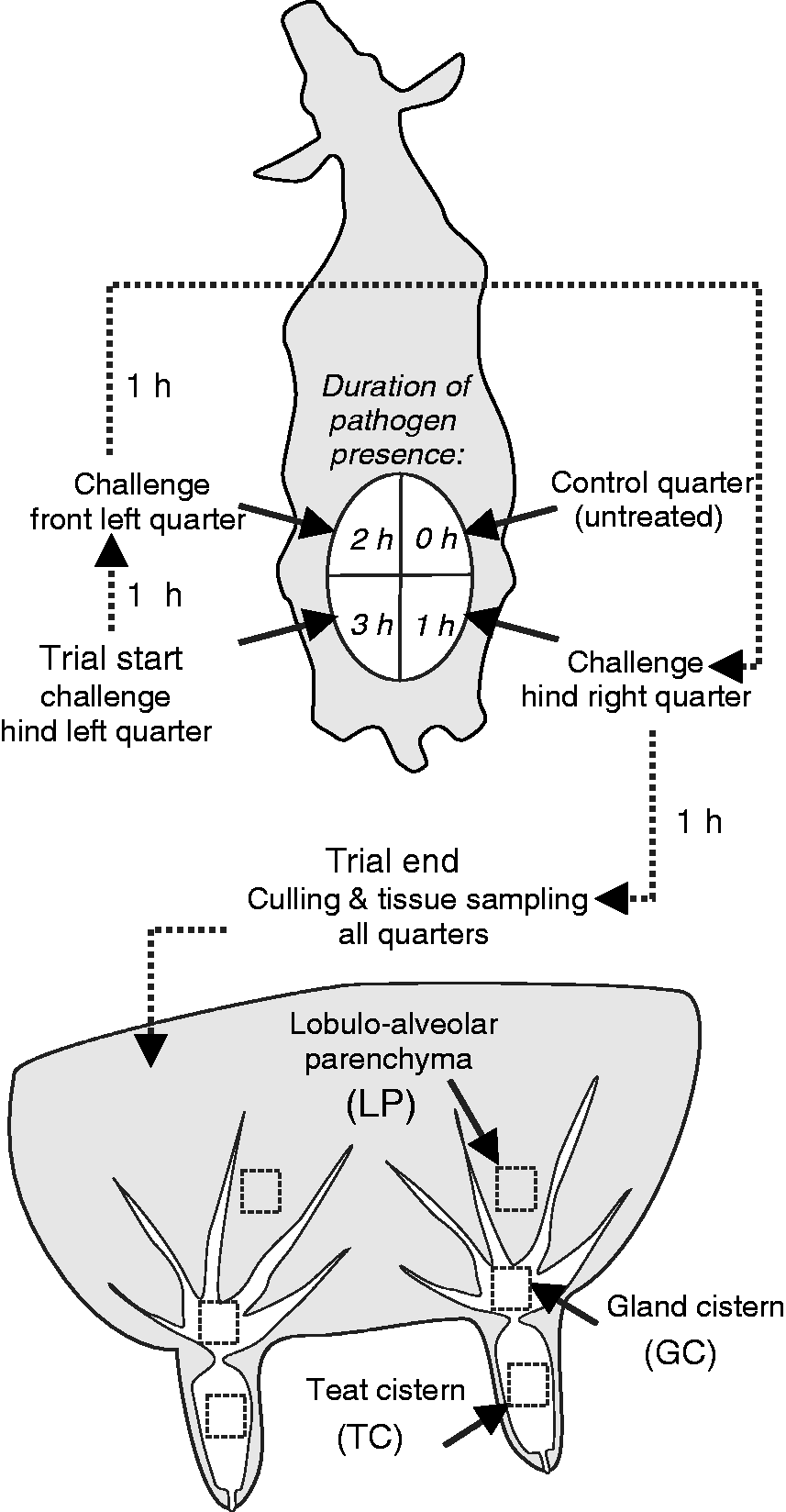
Monitoring local and systemic clinical signs of mastitis
Throughout the trial a complete clinical examination of the animals was carried out in 1-h intervals to guarantee that none of the animals showed either local or systemic signs of mastitis. In addition to monitoring rectal temperature, heart rate, rumen contraction rate, appetite and general attitude every h, the udder was examined for swelling of the parenchyma, subcutaneous oedema, rise in temperature on the udder surface compared with neighbouring quarters and signs of quarter pain. Milk secretion was macroscopically checked for alterations (clots, colour, consistency) at trial start and trial end. Simultaneously, quarter milk samples were aseptically collected in sterile vials for bacteriological examination. Fifteen μl quarter milk samples were plated on Columbia sheep blood agar, on Violet red bile agar and on Edwards agar (Oxoid) to exclude any unwanted bacterial growth during the trial period. The plates were incubated for 48 h at 37℃ and were examined daily for bacterial growth. At trial start and trial end, the animals were milked with a quarter milker (WestfaliaSurge, Bönen, Germany) so that the milk from individual udder quarters was collected in separate containers.
Bacterial recovery from milk
At trial end, 10 ml milk was collected from every separate container of the quarter milker. These samples served to determine the exact amount of CFU of inoculated pathogens within the whole milk fraction of a quarter. A dilution series was made with sterile 0.9% saline and the dilutions 10−1–10−3 were plated on selective media. For the samples from the Ec3h group, the dilutions were plated on Violet red bile agar, and the dilutions from the Sa3h group were plated on Baird-Parker-Agar (Oxoid). After incubating for 24 h at 37℃, the plates were examined for bacterial growth, and the colonies were counted, using an automated colony counter (‘Flash and Grow’) with the software Flash & Go, version 1.0 (both IUL Instruments, Königswinter, Germany). To calculate the amount of CFU/udder quarter milk at trial end, the determined CFU/ml milk was multiplied by the absolute amount of milk gained from the udder quarters at trial end.
Sampling of blood and tissue
Blood samples (10 ml) were taken from the jugular vein aseptically using EDTA-Vacutainers (Becton Dickinson, Heidelberg, Germany) at trial start and trial end. Leukocyte counts in the blood samples were carried out with a Sysmex pocH100i (Sysmex, Norderstedt, Germany). At trial end, the cows were slaughtered with a penetrating captive bolt gun, followed by exsanguination. Tissue samples were collected aseptically from the locations in the teat cistern (TC), gland cistern (GC) LP from the slaughtered cows within 10 min after killing, as shown in Figure 1. Small tissue pieces (0.5 × 0.5 × 0.5 cm) were taken from each location after the right and the left udder side was cut open sagittally. All tissue samples were immediately snap frozen in liquid nitrogen and stored at −80℃ until further processing.
RNA extraction and RT-qPCR
Oligonucleotide primers and fluorescence acquisition temperature for RT-qPCR.

Statistical analysis
The data were entered into Excel (Microsoft Inc., Redmond, WA, USA) and analysed using Excel (Microsoft), Origin (Version 8; OriginLab Inc., Northampton, MA, USA), SAS (Version 9.2; SAS Inc., Cary, NC, USA) and PAST (Version 1.89; Palaeontological Statistics, Oslo, Norway). Description of the data using mean, median, SD, minimum and maximum was performed in Excel. Assessing the normality of the data was performed visually using box-and-whisker-plots, as well as Q-Q-plots. As not all the data were normally distributed, non-parametric tests were used to compare parameters between the two different treatment groups at each individual point in time (Mann–Whitney U-tests).
Within each group the parameters for temperature, leukocytes, bacterial counts at each point in time and the candidate gene expression were compared between the different time points and sampling locations using non-parametric paired tests (Wilcoxon test). A t-test was used to compare the absolute cell counts in the different quarters.
A mixed model (Proc MIXED in SAS) was used to compare the course of the candidate genes, taking into account the repeated measures within the animals, which were considered as a random effect. The significance level was set to P < 0.05.
Results
The absence of clinical signs of mastitis is achieved within the challenge period of 3 h
Rectal body temperature and blood leukocyte counts 0 h and 3 h after intra-mammary challenge with either 5 × 106 CFU E. coli (Ec3h) or 5 × 106 CFU S. aureus (Sa3h).
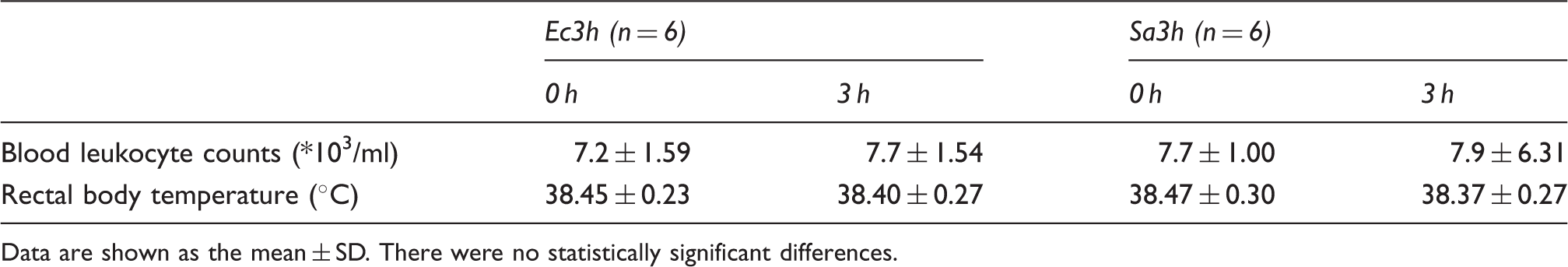
Data are shown as the mean ± SD. There were no statistically significant differences.
E. coli multiplies faster than S. aureus during the first 3 h after pathogen inoculation
During the 3 h period of the challenge trial, the animals of the E. coli challenged group (Ec3h) showed a significant increase in E. coli bacterial counts in milk, dependent on the duration of pathogen challenge, whereas the trend of S. aureus multiplication in the milk of the animals from the S. aureus challenged group (Sa3h) was not significant (Figure 2). E. and S. aureus could be recovered from every challenged quarter and there was no growth of other bacteria at any time point of examination before or after the challenge trial nor in the milk of any of the control quarters (data not shown).
Bacterial counts in milk after intra-mammary E. coli and S. aureus challenge. CFU of E. coli and S. aureus 1, 2 and 3 h after intra-mammary challenge of 6 animals, respectively. The bars display the amount of CFU/udder quarter milk, which were determined by multiplying the CFU/ml milk with the absolute amount of milk gained from the udder quarters at trial end. The dotted line displays the amount of CFU (5 × 106) used for intra-mammary challenge. Different letters denote statistically significant differences between different time points of pathogen presence (P < 0.05).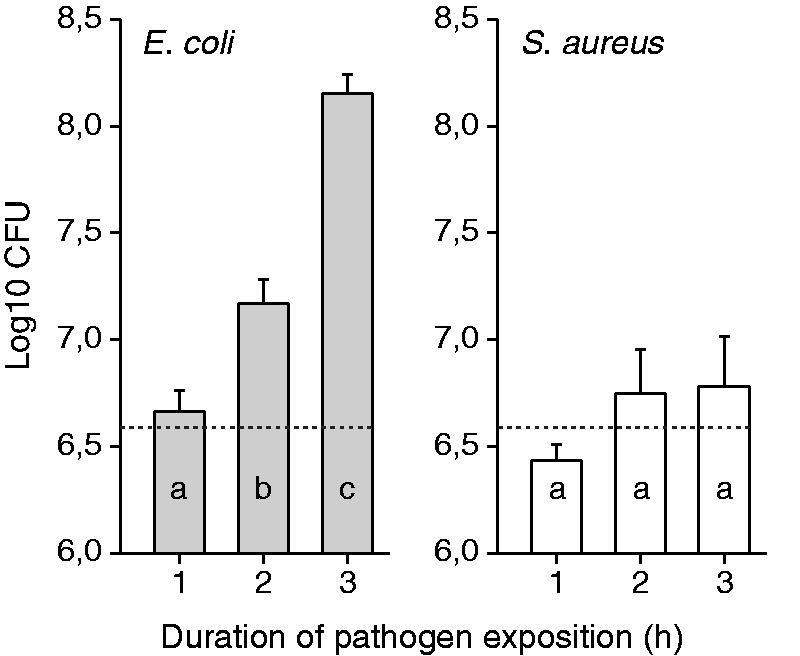
E. coli elicits a more vigorous early immune reaction than S. aureus
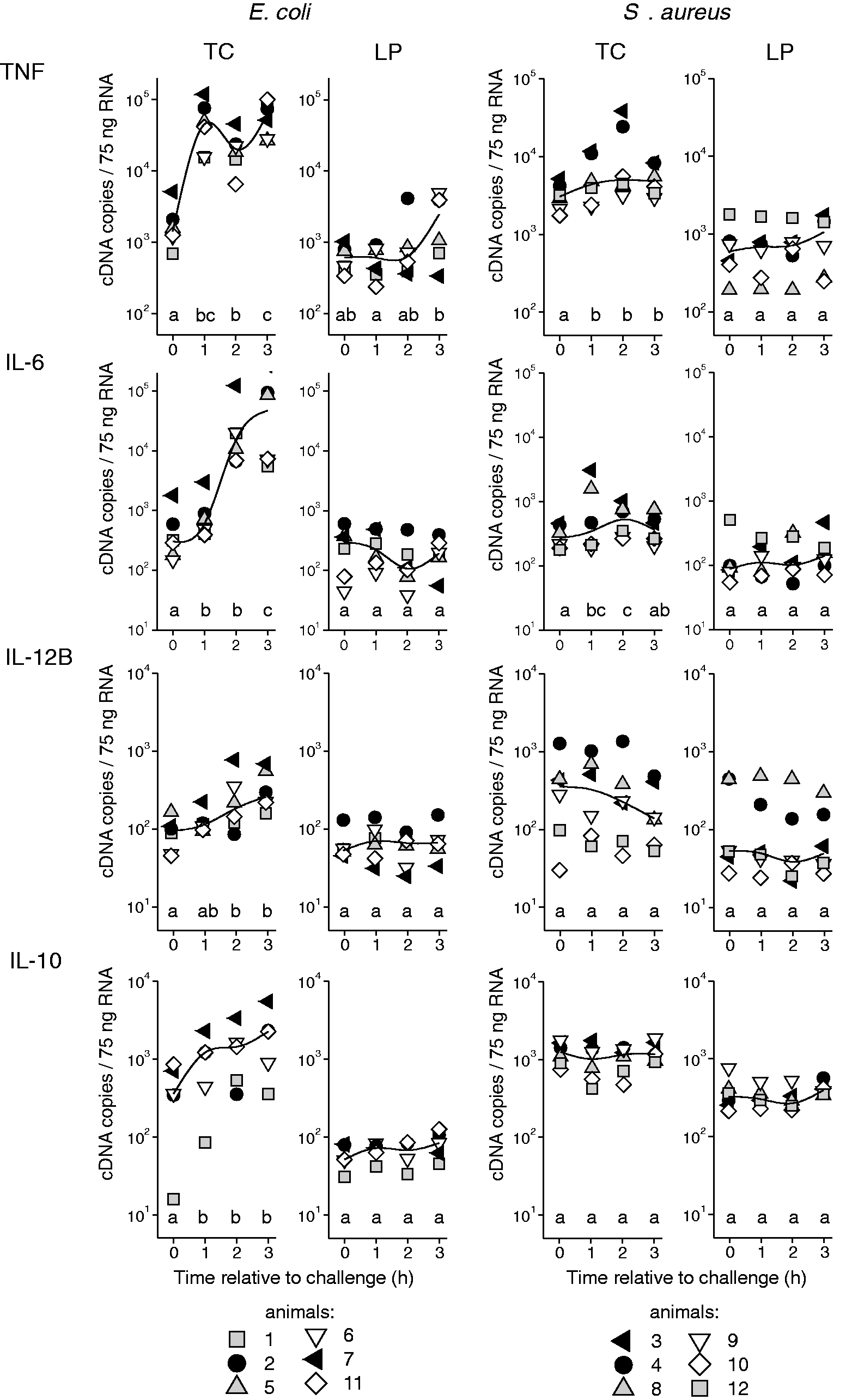
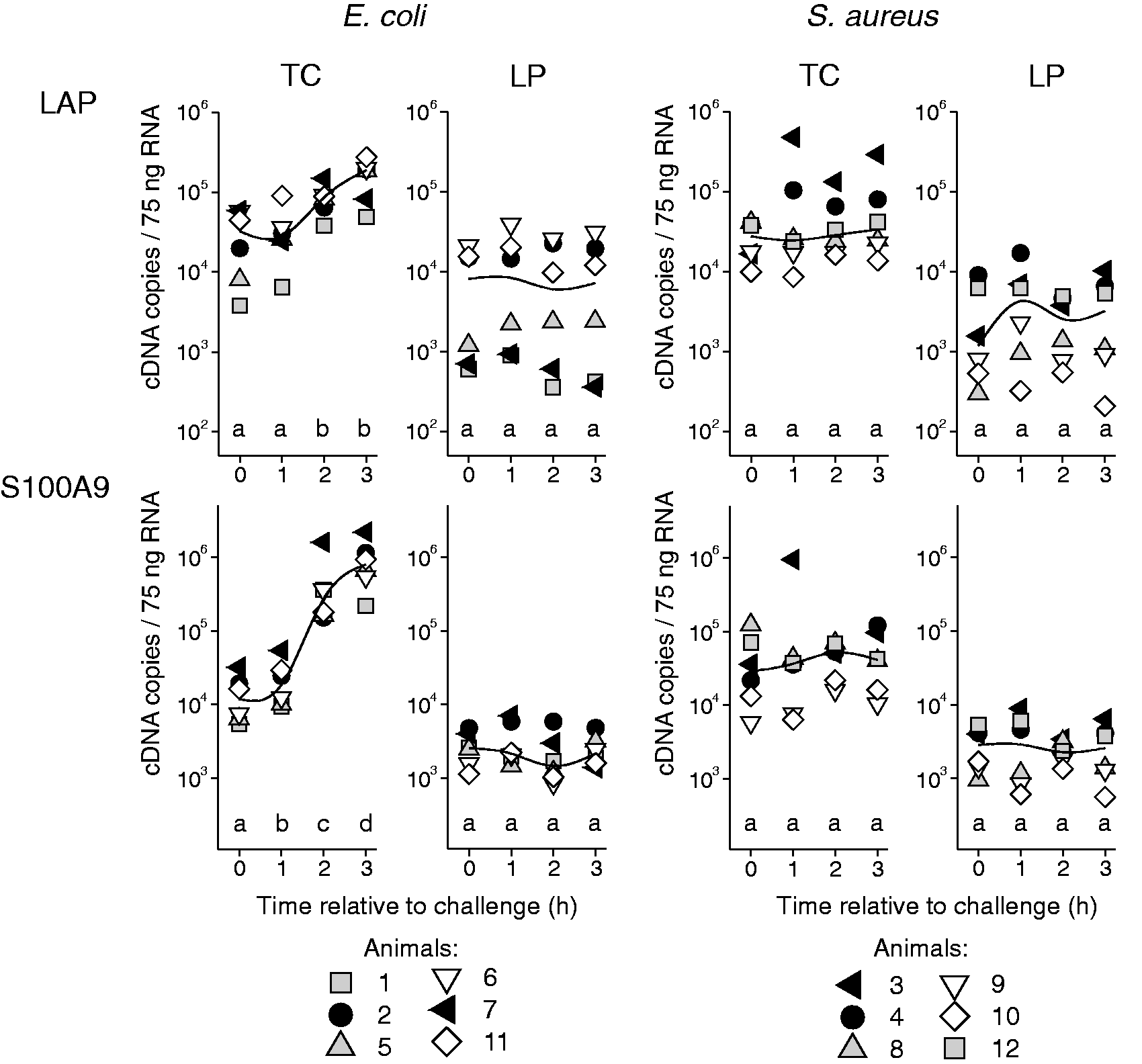
The mRNA copy numbers of the candidate genes measured in the tissue of the teat and the udder are given in Figures 3–5. All the animals displayed a transcriptional response to intra-mammary challenge with either E. coli or S. aureus. This response could already be found in the tissue of the quarters that had been exposed to pathogens for only 1 h. The animals that had received the E. coli challenge responded with up-regulation of all the analysed chemokines (CCL20, CXCL8; Figure 3), cytokines (TNF, IL6, IL12B, IL10; Figure 4) and antimicrobial effector molecules (LAP, S100A9; Figure 5). S. up-regulated the chemokines CCL20 and CXCL8 (Figure 3), and the cytokines TNF and IL6 (Figure 4), but failed to induce IL12B, IL10, LAP and S100A9. Regarding the extent of regulation, we note that E. coli induced a much stronger up-regulation, exceeding that of S. aureus by 25-fold (TNF), 52-fold (CCL20), 76-fold (CXCL8) and 105-fold (IL6).
Gene expression profiles of chemokines after RT-qPCR analysis of tissue from the TC and LP after intra-mammary challenge of six animals with either 5 × 106 CFU E. coli (group Ec3h) or 5 × 106 CFU S. aureus (group Sa3h). Each symbol represents data derived from one individual animal (copy numbers/75 ng mRNA); the curved lines represent median values within the group. Different letters denote statistically significant differences between time relative to challenge (P < 0.05). Gene expression profiles of cytokines after RT-qPCR analysis of tissue from the TC and LP after intra-mammary challenge of six animals with either 5 × 106 CFU E. coli (group Ec3h) or 5 × 106 CFU S. aureus (group Sa3h). Each symbol represents data derived from one individual animal (copy numbers/75 ng mRNA); the curved lines represent median values within the group. Different letters denote statistically significant differences between time relative to challenge (P < 0.05). Gene expression profiles of antibacterial effector molecules after RT-qPCR analysis of tissue from the TC and LP after intra-mammary challenge of six animals with either 5 × 106 CFU E. coli (group Ec3h) or 5 × 106 CFU S. aureus (group Sa3h). Each symbol represents data derived from one individual animal (copy numbers/75 ng mRNA); the curved lines represent median values within the group. Different letters denote statistically significant differences between time relative to challenge (P < 0.05).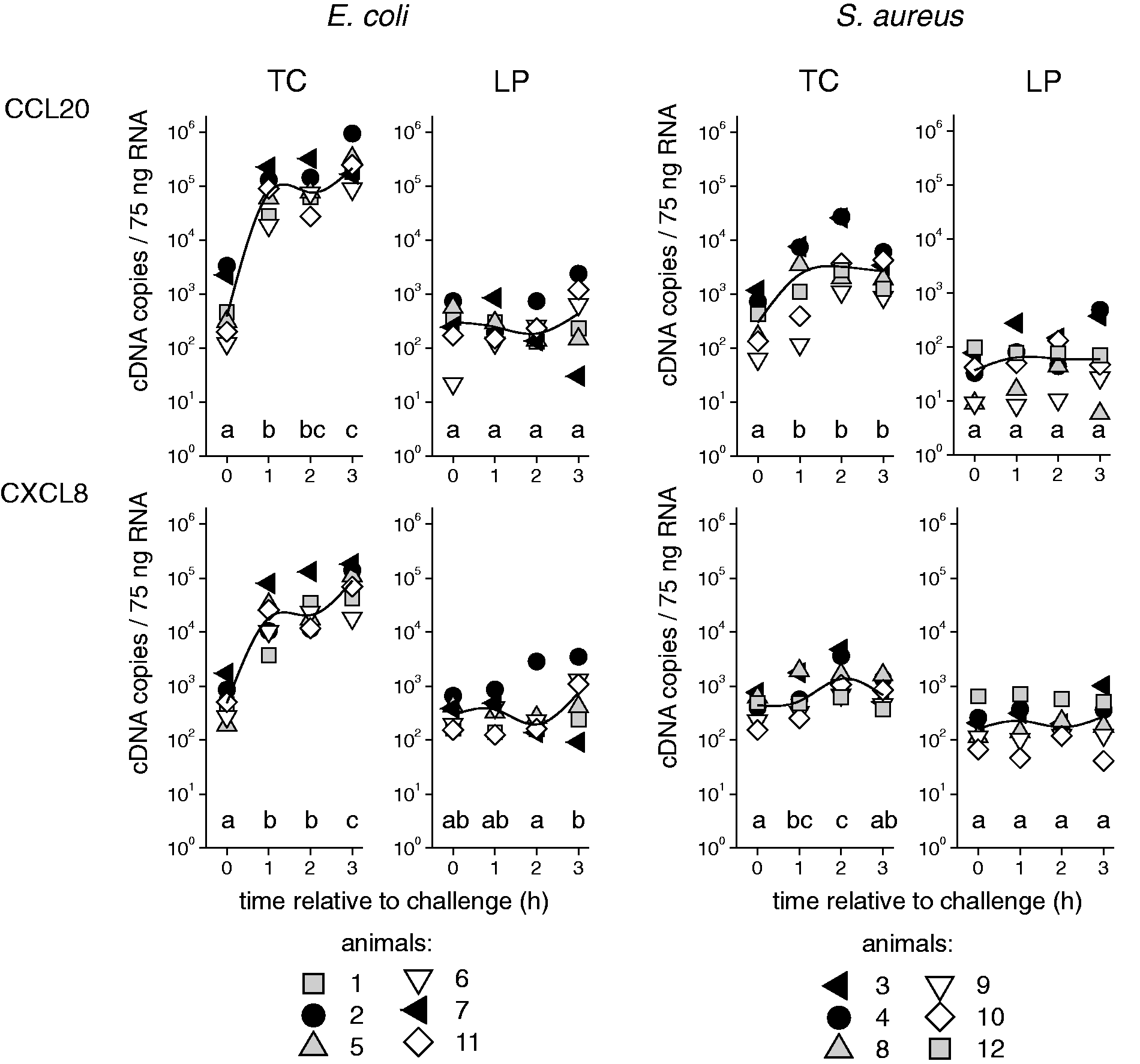
TC and GC respond earlier and with a more pronounced response to pathogens than LP
The TC and GC showed an equal response (data not shown); thus, data are only given for the TC. In the control udder quarters, which had received no pathogen challenge, the constitutive mRNA expressions of S100A9, LAP, TNF and IL10 were significantly higher in the tissue of the TC compared with the LP (P < 0.013; Figures 4 and 5). This was independent of the challenge group (Ec3h or Sa3h) to which the animals had been assigned.
The observed transcriptional response to E. coli and S. aureus challenge was mainly limited to the tissue from the TC. In the animals of the Sa3h group, there was no detectable reaction in the LP. In the Ec3h group, only the factors CXCL8 and TNF were slightly up-regulated in the LP after 3 h of pathogen exposition compared with the unchallenged control quarters (2.5- and 4.0-fold, respectively). However, this up-regulation was not statistically significant (P = 0.22 and 0.09, respectively; Figures 3 and 4).
The sequential animal infection model displays different stages of host response to E. coli and S. aureus
The sequential challenge model reflected the induction kinetics of the selected immune-related genes in the early phase of mastitis in the TC (Figure 1). In the E. coli-challenged animals, TNF, IL10, CCL20 and CXCL8 were among the immediate early up-regulated genes and had already reached a more or less distinctive expression plateau 1 h after challenge (Figures 3 and 4). The expression kinetics of IL6, IL12B, LAP and S100A9 did not reach a plateau level during the challenge period (Figures 4 and 5). A gene-specific induction kinetic could not be observed after S. aureus challenge. The significant induction in the TC of TNF, IL6, CCL20 and CXCL8 reached a plateau level in the 3-h challenge period, but the level of induction was much less marked than after E. coli challenge (Figures 3 and 4).
Discussion
The pathogens E. coli and S. aureus play a major role in clinical and subclinical mastitis, respectively. We applied a novel short-term infection model to analyse initial transcripts in the udder, dependent on time, location and the causative pathogen. These analysed factors are among the first genes regulated in the LP and MEC after challenge with E. coli or S. aureus or their associated molecules and are held responsible for shaping the immune response towards these pathogens.6,8
By 1 h after pathogen challenge, E. coli had already elicited a stronger transcriptional response of all the examined genes than S. aureus in the TC. This observation is in line with all previous reports from in vivo and in vitro experiments comparing the response towards these two pathogens.6,12,14 However, up until now, animal experiments comparing the response to E. coli and S. aureus were limited to trials, where signs of an inflammatory response were already present with regards to either a systemic response (fever, leukopenia) or a local response (udder swelling, altered milk appearance, increased SCC). We validated that our new in vivo model, as presented here, enabled us to analyse the initial response before any signs of clinical and subclinical mastitis emerged. A previous study had shown that by increasing the dose of intra-mammary E. coli challenge, the period to onset of clinical signs could be shortened. 20 Based on these observations, we applied a relatively high challenge dose (5 × 106 CFU) of either E. coli or S. aureus into the udder to guarantee that the bacterial load was sufficiently abundant to be recognized. Furthermore, we kept the duration of the experiment short enough to avoid the emergence of a clinical response.
The mastitis pathogens E. coli and S. aureus differ in their ability to multiply within the mammary gland during infection. 1 E. coli is known to multiply very fast in milk and may display a generation time of up to 20 min. 2 This may be partially causative for the observed differences in the pathogen-specific host response in numerous studies. Within this study, we could clearly demonstrate that the inoculated E. coli strain significantly multiplied, while there was no significant increase of S. aureus in the milk of the challenged cows during the first 3 h after challenge (Figure 2). This may have accounted for the more vigorous response of the E. coli challenged cows; however, in udder quarters challenged for only 1 h, a comparable bacterial load of E. coli and S. aureus elicited a much stronger response in the tissues of the E. coli challenged animals (Figures 2–5).
Our study shows that both applied pathogens initially elicit a transcriptional response in the TC but not in the LP (Figures 3–5). This points to a time- and compartment-dependent response induction after intra-mammary challenge. Because the TC is confronted earlier than the LP with invading pathogens, a more rapid response was expected. The present but comparably weak response to S. aureus challenge was limited to the tissue of the TC, and there was no significant regulation of the analysed transcripts in the LP. This observation demonstrates the necessity for applying multiple sampling locations of tissue in mastitis infection trials, as previously demonstrated. It was shown that the host response in the udder towards E. coli 12 h after challenge was mostly found in the tissues of the teat and gland cistern; although, in contrast to our study, these animals were already affected by clinical mastitis. 9 We could confirm this observation for the E. coli and also for the S. aureus-challenged animals; however, within the applied model system here we were even able to show this location-specific response before the onset of clinical signs. In a previous S. aureus challenge model, it was shown that the factors of the innate immune response were up-regulated in a more pronounced manner in the LP. 10 However, sample collection was carried out 21 d after the first S. aureus challenge, which possibly explains the differences in the data of the present study.
All eight selected candidate genes were significantly up-regulated in the TC after E. coli challenge. The sequential challenge model allowed us to describe a gene-specific kinetic of the immune gene induction (Figure 1). Roughly two groups of genes could be distinguished: the first group (TNF, IL10, CCL20, CXCL8) was characterized by an immediate early induction, which rapidly reached a plateau level that was sustained during the 3-h challenge period. This is in line with previous reports where TNF, CCL20 and CXCL8 were among the immediate early regulated genes in bovine MEC. 6 The subsequently induced group of genes (IL6, IL12B, LAP and S100A9) did not show a plateau level of mRNA copies during the 3-h E. coli challenge (Figures 4 and 5). The antimicrobial effectors LAP and S100A9 are known to be late-induced genes in E. coli-stimulated MEC. 6 The cytokines IL6 und IL12B are well-known secondary response genes, 21 which require new protein synthesis for their induction. In E. coli-stimulated MEC, IL6 is among the delayed early induced inflammatory response genes. 6
Except for IL10, all the other immediate early induced genes (TNF, CCL20 and CXCL8) were also significantly up-regulated in the TC after S. aureus challenge (Figures 3 and 4). Although the induction level in the TC was only 1–7% compared with the E. coli-challenged animals, both pathogens elicited a response within the first h after challenge. The strong difference in the level of induction between E. coli and S. aureus is in line with previous reports, where the S. aureus-mediated induction in MEC was <5% compared to the E. coli-mediated induction.6,7
In addition to CXCL8, TNF and CCL20, the cytokine IL6 was significantly induced in the TC after S. aureus challenge (Figure 4). The importance of this delayed early cytokine for the inflammatory response of the MEC after S. aureus stimulation has recently been demonstrated. 6 Although we were not able to show within this study which cell types were responsible for the induction of the gene transcripts, the induction pattern of those four inflammatory factors in the TC after S. aureus infection, including an absent induction of IL-10, resembles the previously described response of S. aureus-stimulated MEC and could now be demonstrated in vivo. 6 The observation suggests a major role of the epithelial cells in the response against S. aureus in the TC. However, after E. coli challenge, IL10 and IL12B were also up-regulated in the TC. These cytokines are reported to be primarily expressed by myeloid cells (e.g. macrophages, dendritic cells) and T cells but not by MEC,7,22 suggesting their contribution in the early phase of the immune response in the TC against E. coli.
In addition to the strong induction of inflammation related genes in the TC after E. coli challenge, we observed the beginning (not significant) of the induction of the inflammatory cytokines TNF-α and CXCL8 in the LP 3 h after E. coli challenge (Figures 3 and 4). This suggests that within the 3-h timeframe after E. coli challenge, the observed immediate early regulation of these genes subsequently starts in tissue compartments more distant to the site of pathogen entry and may be caused by a later contact with the pathogens. Whether the observed higher constitutive expression of S100A9, LAP, TNF and IL-10 in the TC compared with the LP in the untreated udder quarters were caused by a different cellular composition was not analysed within this study (Figures 4 and 5). However, it underlines the necessity for multiple sampling locations in mastitis in vivo trials. The higher expression of antimicrobial effector molecules (S100A9, LAP) in the TC compared with the LP may act as a natural defence mechanism against pathogens invading via the teat canal. This is underpinned by a previous report showing even higher constitutive expression levels of antimicrobial S100-proteins in the Furstenberg’s rosette compared with the TC. 11
Experimental mastitis models with cows usually involve very limited numbers of animals owing to comparably high expenses. Thus, individual differences in the host response have a major impact on the achieved results. Although the selection of comparable experimental animals within this study was based on well-proven and strict criteria, 15 the transcriptional response to intra-mammary challenge displayed a certain variance in the expression levels of the analysed genes. A continuous pattern became obvious when individual animals were classified with regard to expression levels in the teat cistern. Animals with comparable high levels of all the analysed genes in the Ec3h group (animals 2 and 7, black symbols in Figures 3–5) consistently showed the strongest transcriptional response to E. coli challenge. Within the Sa3h group this could also be observed in two animals for CCL20 and TNF (animals 3 and 4, black symbols in Figures 3 and 4). These animals also displayed the highest levels of LAP after intra-mammary S. aureus challenge (Figure 5, animals 3 and 4). Whether this stronger transcriptional response is related to a better defence against pathogens in the udder could not be analysed within this study. However, we previously showed that Holstein heifers could be divided into ‘fast’ and ‘slow’ responding animals based on their SCC increase after intra-mammary S. aureus challenge, and the ‘fast’ responding animals cleared infection more effectively than the ‘slow’ responders. 13
The interdependence of a transcriptional response between udder quarters by systemic effects within one animal have been shown to a high extent in E. coli-challenged cows, 15 and to a lesser extent during experimental S. aureus mastitis.10,14,15 Thus, the data of the intra-animal controls from uninfected udder quarters must be interpreted with caution. Within the herein presented sequential challenge model, crosstalk may have affected the transcriptional response of the udder quarters that received pathogen challenge for 1 or 2, or no challenge. However, there was no difference in the expression levels of the analysed genes in the unchallenged quarters regardless of the challenge group (Ec3h or Sa3h) to which the animals belonged, suggesting that systemic effects were absent. Whether the onset of systemic clinical signs (fever, leukopenia) would have been coincidental with the transcriptional systemic effects between udder quarters, while likely could not have been shown within this study.
Altogether, the applied model is a suitable tool to examine very early events during pathogen recognition in the udder and may serve in the future for holistic approaches to analyse the pathways of immune activation during the onset of clinical and subclinical mastitis.
In conclusion, this short-term model clearly demonstrates that a pathogen specific response to intra-mammary challenge is already generated at transcriptional level during the first hour. It confirms that the first transcripts in response to pathogenic threats are generated in the tissues of the teat while a response in the udder parenchyma is still absent. The typical divergent host response patterns caused by E. coli and S. aureus with regard to clinical and subclinical mastitis are already reflected in quantity and quality at this early stage in the teat.
Footnotes
Acknowledgements
We thank Dr Katharina Zimprich, Dr Monique Lind-Böhm, Sandra Kirsch, Angelika Deike and Bärbel Pletz for their technical assistance.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was funded by the German Research Foundation (DFG FOR 585).
