In May 2007, a combined milk sample from all 4 quarters from a cow in the second month of
pregnancy in her second lactation was submitted to the laboratory due to a high somatic
cell count (SCC), enumerated on a commercial counter,
a
of 1,008,000. Gel formation at the California mastitis test (http://www.infovets.com/demo/demo/dairy/d100.htm) was
observed only in the sample from the left forequarter, indicating that this quarter was the
source of the high SCC in the combined sample.
The significance of SCC as an indicator of udder infection and its deleterious impact on
dairy products has been reviewed.
8
Briefly, SCC higher than 200,000/ml may indicate an intramammary infection. A SCC
higher than 600,000 is often caused by major mammary pathogens, such as
Staphylococcus aureus and Streptococcus spp. A high
SCC causes significant economic damage to the dairy industry by decreased product quantity
and quality, due primarily to proteolytic activities. Moreover, the shelf life of milk and
dairy products is shortened. The bulk tank SCC is an indication of the prevalence of
subclinical mastitis in the herd, because milk from cows suffering from clinical mastitis
is discarded.
The cow in the current study was located on a family-managed dairy farm consisting of 48
Israeli Holstein dairy cows, 18 of which were primiparous cows. The management on the farm
and the hygienic conditions were very good. Milking hygiene included predipping, wiping,
and postmilking teat dip. The farm was under continuous monitoring by the National Service
for Udder Health and Milk Quality. A microorganism, later identified as Mannheimia
granulomatis, was isolated in pure culture from the left forequarter of the cow
in question. According to the policy adopted in Israel to abstain from antibacterial
therapy in cases of subclinical mastitis, except for infections with specific bacteria in
which a positive outcome is expected, the cow was not treated. It kept shedding the
microorganism, exclusively from the same quarter, at least until March 2008. Elevated SCC
was found throughout this period, even in a sample from which M.
granulomatis was not isolated. Mannheimia granulomatis was not
isolated from other cows on the farm in spite of the affected cow's not being isolated. In
fact, between April and June 2007, 40 milk samples from other cows, all from subclinical
mastitis cases, were submitted to the laboratory; 26 yielded no growth, and 1 was
contaminated. The following bacteria were isolated from the other samples:
coagulase-negative Staphylococcus spp. (9), Streptococcus
dysgalactiae (1), Enterococcus faecalis (1),
Escherichia coli (1), and Corynebacterium bovis (1). A
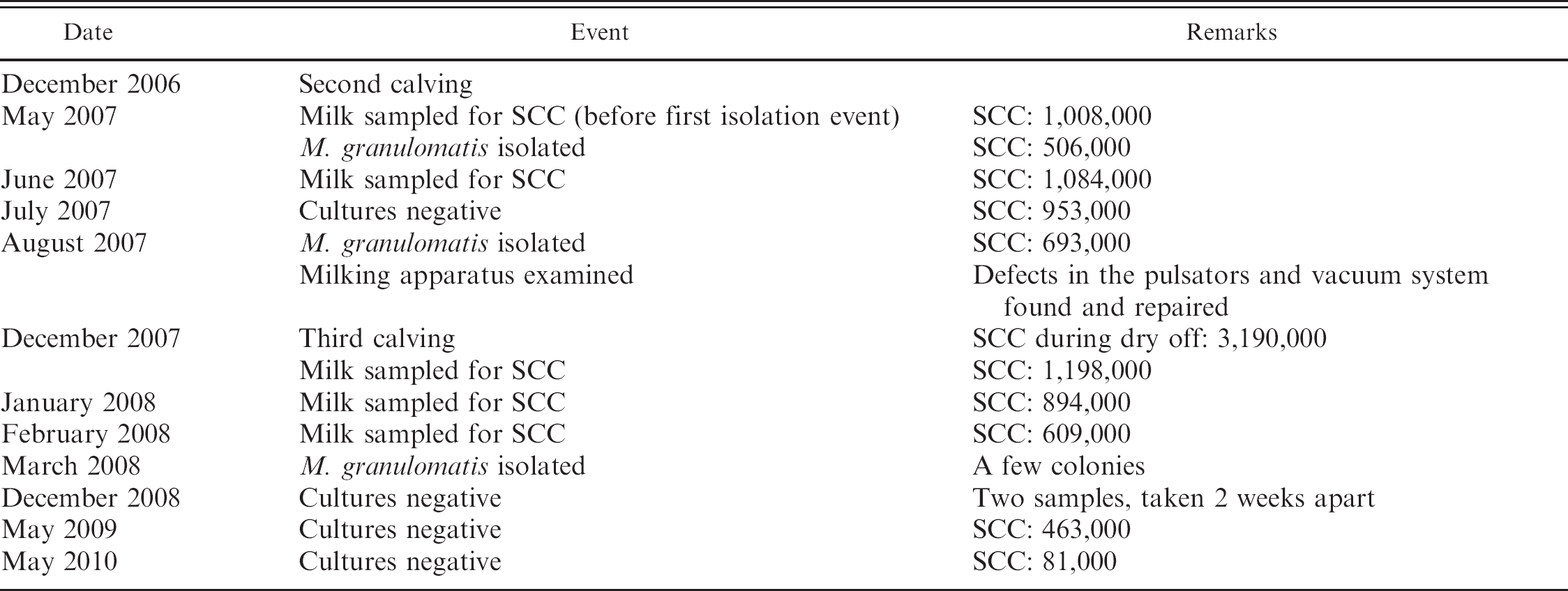
detailed timeline of the event and follow up are presented in Table 1.
Bacteriologic examination of the milk samples and presumptive identification of bacteria
were performed as previously described.
10
The isolate stained Gram negative, was oxidase and catalase positive, did not grow
on MacConkey agar, was hemolytic on 5% sheep blood agar, and was inert and nonmotile in
semisolid glucose–cystine–trypticase agar. The following phenotypic characteristics were
found.
Gram-negative rods.
b
Positive results: potassium nitrate reduction, esculin,
4-nitrophenyl-βD-galactopyranoside. Negative results: indole production, D-glucose
(fermentation), L-arginine, urea, gelatin, assimilation of D-glucose, L-arabinose,
D-mannose, D-mannitol, N-acetylglucosamine, D-maltose, potassium gluconate, capric acid,
adipic acid, malic acid, trisodium citrate, phenylacetic acid. Identification:
Mannheimia haemolytica (85.9%, T = 0.56) at low discrimination with
Brevundimonas vesicularis (11.7%, T = 0.53).
Corynebacteria.
c
Positive results: potassium nitrate reduction, 2-naphtyl-phosphate,
2-naphthyl-βD-galactopyranoside, esculin, fermentation of D-glucose, D-ribose, D-xylose,
D-mannitol, D-maltose, and D-saccharose. Negative results: pyrazine carboxamide,
pyroglutamic acid-β-naphthylamide, naphthol ASBI-glucuronic acid,
2-naphthyl-αD-glucopyranoside, 1-naphthyl-N-acetyl-βD-glucosamine, urea, gelatin,
fermentation of D-lactose and glycogen.
Anaerobes.
d
Positive results: 4-nitrophenyl-βD-galactopyranoside,
4-nitrophenyl-βD-glucopyranoside, potassium nitrate reduction, 2-naphtyl-phosphate,
L-arginine-β-naphthylamide, L-alanyl-L-alanine-β-naphthylamide, L-glycine-β-naphthylamide.
Negative results: urea, L-arginine, 4-nitrophenyl-αD-galactopyranoside,
4-nitrophenyl-βD-galactopyranoside-6-phosphate-2CHA, 4-nitrophenyl-αD-glucopyranoside,
4-nitrophenyl-αL-arabinofuropyranoside, 4-nitrophenyl-βD-glucuronide,
4-nitrophenyl-N-acetyl-βD-glucosamine, glutamic acid decarboxylase,
4-nitrophenyl-αL-fucopyranoside, indole production, L-proline-β-naphthylamide,
L-leucyl-L-glycine-β-naphthylamide, L-phenylalanine-β-naphthylamide,
L-leucine-β-naphthylamide, pyroglutamic acid-β-naphthylamide, L-tyrosine-β-naphthylamide,
L-histidine-β-naphthylamide, L-glutamyl-L-glutamic acid β-naphthylamide,
L-serine-β-naphthylamide, fermentation of D-mannose and D-raffinose.
Enterobacteriaceae.
e
Positive results: esculin, para-nitro-phenyl-phosphate,
para-nitro-phenyl-α-β-glucoside, para-nitro-phenyl-β-galactoside, para-nitro-phenyl
bis-phosphate, para-nitro-phenyl-arabinoside, fermentation of mannitol, saccharose, and
sorbitol. Negative results: L-arginine, urea, para-nitro-phenyl-xyloside,
para-nitro-phenyl-phosphorylcholine, para-nitro-phenyl-glucuronide,
pa-ra-nitro-phenyl-N-acetyl glucosamide, proline nitroanilide, γ-L-glutamyl p-nitroanilide,
p-nitro-DL-phenylalanine, glycine, citrate, malonic acid, triphenyl tetrazolium chloride,
lysine, arabinose, mannose, melibiose, rhamnose, adonitol, galactose, inositol.
Identification: Mannheimia haemolytica, 0.9899.
When the same analyte was tested with different kits, results were consistent, except for
D-glucose fermentation. This reaction was negative with the Gram-negative rod kit
b
and positive with the corynebacteria kit,
c
which was confirmed by performing the tests in triplicate.
Results lead to a tentative identification of the isolate as M.
granulomatis. To confirm this identification, 2 isolates from 2007 were
submitted to molecular identification by 16S ribosomal DNA (rDNA) sequencing. Briefly,
total bacterial DNA was extracted from colonies using a commercial kit,
f
according to the manufacturer's instructions. Polymerase chain reaction
amplification of the first 800 base pairs of the 16S rDNA was performed with the following
universal primers
6
: forward sequences (4F: TTGGAGAGTTTGATCCTGGCTC), reverse sequences (801R:
GGCGTGGACTTCCAGGGTATCT). The polymerase chain reaction was started with an activation step
at 90°C for 15 min, followed by 35 cycles of denaturation at 94°C for 30 sec, annealing at
50°C to 60°C for 90 sec (60°C for the first 5 cycles, 55°C for next 5 cycles, and 50°C for
the last 25 cycles), and extension at 72°C for 4 min, ending with a final extension step at
72°C for 10 min.
The amplicon product was verified with agarose gel electrophoresis, purified according to
the manufacturer's instructions,
g
and sequenced using primer 4F on an automated DNA sequencer.
h
Obtained sequences were aligned and compared with archived NCBI sequences and with
the Bioinformatics Bacterial Identification database (http://pbil.univ-lyon1.fr/bibi/). In addition, a partial sequence of the
rpoB gene of the original isolate was obtained as previously
described.
5
The sequence obtained from the molecular analysis of the 16S ribosomal RNA (rRNA) gene was
deposited in GenBank (accession no. GQ844959). A match of 99% was found between this
sequence and the 16S rRNA sequence from M. granulomatis strain W4672/1 16S
and M. granulomatis strain ATCC 49244 16S. The sequence obtained from
partial amplification of rpoB was deposited in GenBank (accession no.
GQ856691). A match of 97% was found between this sequence and the rpoB
sequence of M. granulomatis strain ATCC 49244.
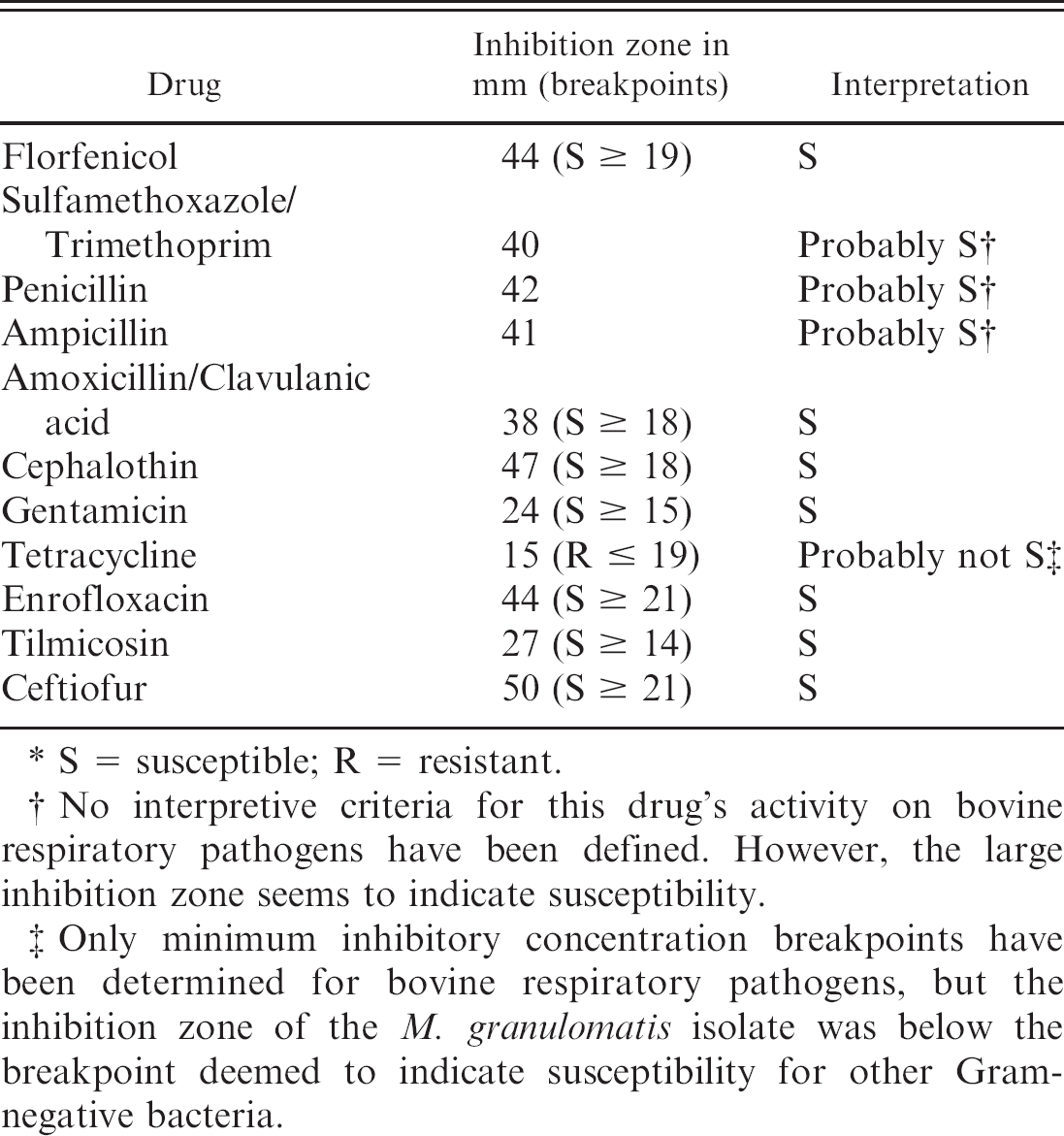
In vitro antibacterial susceptibility testing was performed by the disk diffusion method,
according to Clinical and Laboratory Standards Institute standards.
7
Inhibition zones were determined with an imaging instrument.
i
Interpretive criteria applied were those defined for M.
haemolytica. The isolate was susceptible to all the antibacterial drugs tested
except tetracyclines (Table
2).
The Pasteurellaceae family has undergone substantial changes in the past
decade due to the application of molecular taxonomical tools. The genus
Mannheimia comprises M. haemolytica, M. glucosida, M.
ruminalis, M. varigena, and M. granulomatis.
2
The latter species has been associated with a potentially fatal infection defined
as bovine focal proliferative fibrogranulomatous panniculitis, or “lechiguana” in south
Brazil.
9
Isolates from this syndrome were shown to produce leucotoxin.
11
In addition, the microorganism was isolated in Australia from cattle with
suppurative bronchopneumonia, jaw abscess, and actinobacillosis-like lesion of the
tongue
3
; in Denmark from a hare with pneumonia
1
; and in Sweden from various infections, especially in the respiratory tract and the
mouth, in Western roe deer (Capreolus capreolus).
4
To the authors' knowledge, this is the first report of M. granulomatis
causing subclinical mastitis in a cow. Although direct causality between the
microorganism's presence and the high SCC counts was not established, the fact that they
were both found in the same single quarter suggests the possibility that they are related.
The fluctuations in SCC observed during the follow-up period are an accepted
phenomenon.
8
The source of the microorganism's introduction to the farm and possibly to Israel
is unknown. The intensive routine monitoring of Israeli dairy cattle by the National
Service for Udder Health and Milk Quality makes its previous presence in the country very
unlikely. Because this is thus far the only case in Israel, its importance relative to
other causes of environmental or infectious mastitis cannot be established.
The fact that this microorganism was previously not isolated from similar cases may be due
to its similarity to other Mannheimia spp. and
Actinobacillus spp. Thus, the possibility of misidentification cannot
be excluded, especially because some commercial kits that do not include M.
granulomatis in their databases would identify the isolates as M.
haemolytica. In fact, strains identified in Australia as M.
granulomatis were previously thought to belong to other
Mannheimia spp. or Actinobacillus spp.
3
Consequently, M. granulomatis should be considered as the possible
etiology of typical actinobacillary lesions of the jaws and tongue and of infections caused
by M. haemolytica, such as pneumonia and udder infections.