Abstract
Chronic periodontitis (CP) is a prevalent pathogen-associated inflammatory disorder characterized by the destruction of tooth-supporting tissues, and linked to several systemic diseases. Both the periodontopathogen Porphyromonas gingivalis (Pg), and the genetically determined host immune response, are hypothesized to play a crucial role in this association. To identify new target genes for CP and its associated systemic diseases, we investigated the transcriptome induced by Pg in human monocytes using a genome-wide approach. Monocytes were isolated from healthy male volunteers of European origin and challenged with the Pg virulence factor LPS. Array-based gene expression analysis comprising >47,000 transcripts was performed followed by pathway analyses. Transcriptional data were validated by protein and cell surface markers. LPS Pg challenge led to the significant induction of 902 transcripts. Besides known periodontitis-associated targets, several new candidates were identified (CCL23↑, INDO↑, GBP 1/4↑, CFB↑, ISG20↑, MIR155HG↑, DHRS9↓). Moreover, various transcripts correspond to the host immune response, and have been linked to cancer, atherosclerosis and arthritis, thus highlighting the systemic impact of CP. Protein data of immunological markers validated our results. The present findings expand understanding of Pg elicited immune responses, and indicate new target genes and pathways of relevance to diagnostic and therapeutic strategies.
Introduction
Chronic periodontitis (CP) is a prevalent inflammatory disease characterized by the irreversible destruction of tooth-supporting tissues. 1 CP has become the focus of increasing interest, owing to its rising prevalence and its association with life-threatening systemic disorders such as atherosclerosis, autoimmune disease and cancer. 2 DNA-based phylotyping studies have indicated that bacterial diversity has been influenced by the introduction of Neolithic and industrialization diets, resulting in an increase in periodontal pathogens, which are key players in CP. 3
Porphyromonas gingivalis (Pg), a Gram-negative anaerobic microbe, is highly related to the severity of CP. 4 However, there is evidence that a certain genetic predisposition also plays a role in CP, reinforcing the notion of host-derived pathogen-associated periodontal destruction. 5 In the last decade, research has investigated associations between periodontitis and various single nucleotide polymorphisms (SNPs). Based on the hypothesis that a pro-inflammatory environment and immune cell recruitment are implicated, the primary focus of this research was the investigation of SNPs located in immunological, pro- and anti-inflammatory genes like IL-1 and receptor antagonist (IL1RN), TNF-α or IL-10. 6 Specific innate immune receptors such as TLRs may also play a role in CP. TLRs are highly conserved pathogen recognition receptors, and are responsible for the detection of PAMPs such as LPS. Toxic LPS, a cell envelope component of Gram-negative bacteria, activates TLR4–MD2 complex. This activation is followed by the nuclear translocation of transcription factors, like NF-κB, and the secretion of pro-inflammatory cytokines and chemokines. The LPS that is derived from Pg is structurally and functionally distinct from the LPS of Escherichia coli, an enteric bacterium, and research suggests that LPS Pg interacts with TLR2, TLR4 and TLR7. 7
Monocytes play a crucial role in systemic pathogen signaling as they migrate to the sites of infection via PAMP recognition, and differentiate into macrophages or dendritic cells (DCs). Recent investigations have shown that periodontal pathogens reach the bloodstream (bacteremia), and that the level of bacteremia is related to the periodontal pathogen density and the severity of periodontal destruction. 8 Hence, the interaction between periodontal pathogen bacteria and the host immune response is crucial. Hypo- and hyper-responsiveness of the host immune response may predispose to, and aggravate, tissue destruction, thus influencing both periodontal and systemic disease.
The aim of the present study was to identify new target genes and pathways of relevance to CP disease and the associated systemic immune response. The investigation focused on the transcriptome induced by LPS Pg in human monocytes. Therefore, >47,000 transcripts were analyzed to provide genome-wide transcriptional coverage of well-characterized genes derived from the National Center for Biotechnology Information Reference Sequence. The present findings expand understanding of the Pg-elicited immune response, and indicate new target genes and pathways of relevance to diagnostic and therapeutic strategies, and explain the development of the associated systemic disorders.
Materials and methods
Monocyte isolation
The study was approved by the ethics committee of the University of Bonn (Nr. 352/13), and all participants provided written informed consent prior to inclusion. Ten healthy male volunteers of European origin (18–35 yr) were recruited, and peripheral blood samples were obtained. Females were excluded, as hormonal factors may influence transcriptional expression patterns. The following inclusion criteria were applied: non-smoker; no acute or chronic infection; no vaccination within the 4 wk prior to blood sampling; C-reactive protein levels < 2.5 mg/dl; monocyte purity > 95%; and overnight survival > 85%. On the basis of these criteria, three individuals were excluded. The cells of the remaining seven individuals underwent further processing (Figure 1A–C).
Experimental setting and methods. (a) Blood of healthy volunteers was collected for CD14+ monocyte isolation. Purity (>95%) was determined using CD14-microbeads (Miltenyi Biotec GmBH) and flow cytometry. (b) Gating strategy to identify CD14+ expressing human primary monocytes. (c) Detection of monocyte survival and activation: Prior to LPS stimulation (24 h after cell isolation), human monocyte survival was determined using the Annexin V-FITC Apoptosis Detection Kit (eBioscience) and flow cytometry. Monocyte activation was evaluated using specific surface markers [CD86 (eBioscience) and HLA-DR (BD Bioscience)]. SSC: side scatter; FSC: forward scatter.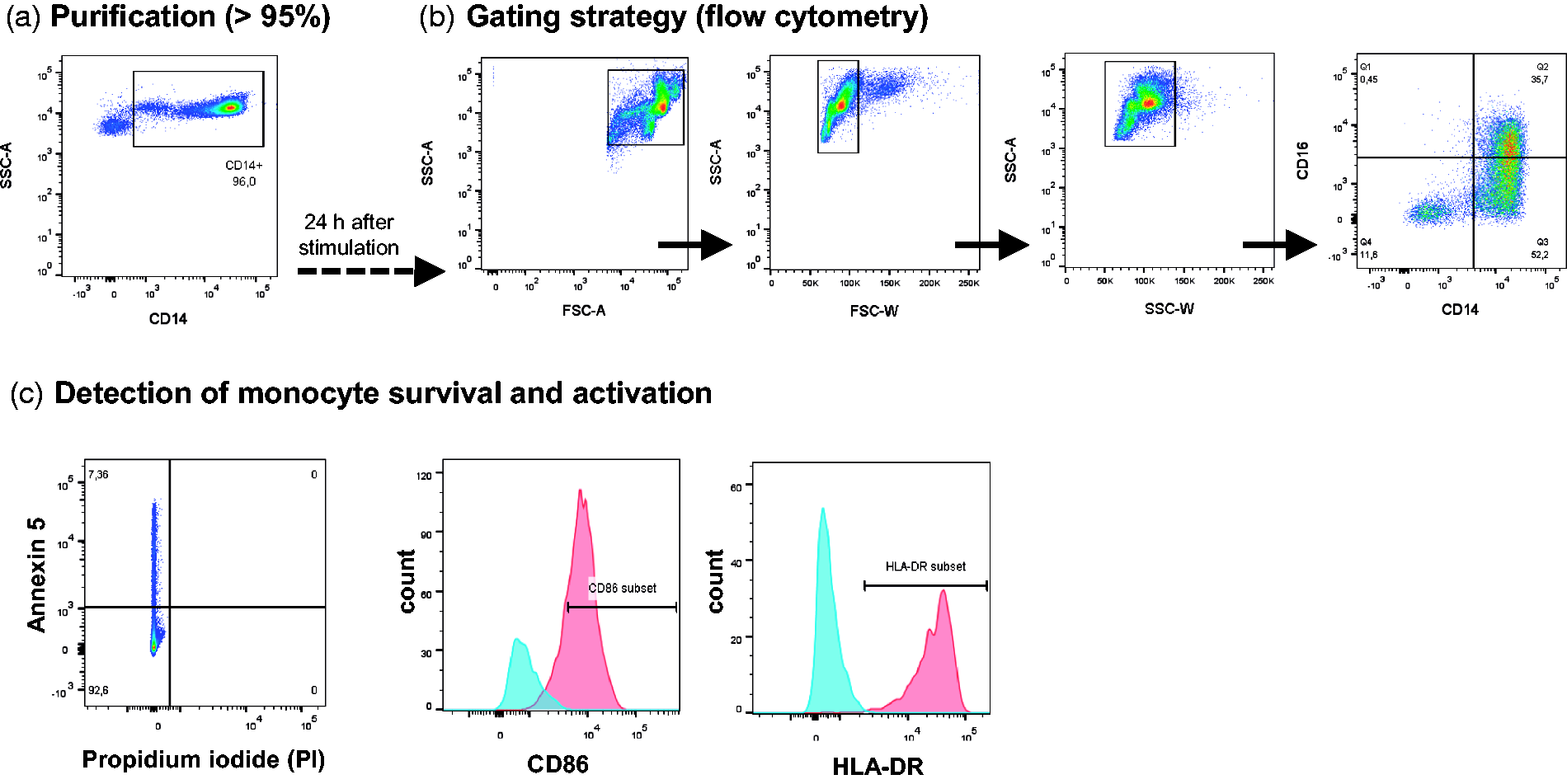
Peripheral blood mononuclear cells from 100 ml whole blood were purified using a Ficoll-Plaque density gradient. CD14+ monocytes were then separated using magnetic-activated cell sorting and CD14-microbeads (Miltenyi Biotec GmbH, Bergisch Gladbach, Germany), in accordance with the manufacturer’s instructions. Cell purity was determined by flow cytometry followed by cell resuspension in RPMI 1640 GlutaMAX. The latter was supplemented with 10% heat-inactivated FBS (all from Gibco, Life Technologies, Waltham, MA, USA); 100 U/ml penicillin; and 100 µg/ml streptomycin (PAA Laboratories/GE Healthcare, Pasching, Austria).
Monocyte stimulation
Monocytes were cultured overnight in 96-well round-bottom wells at a density of 500,000 cells/well in 100 µl. The next morning (approximately 18–20 h after cell isolation) cell survival was analyzed optically using trypan blue staining, followed by Annexin V (Apoptosis Detection Kit) and propidium iodide detection (eBioscience, San Diego, CA, USA) via flow cytometry. The cells were then stimulated with ultrapure LPS from Pg (InvivoGen, Darmstadt, Germany) for 90 min or 6 h. On the basis of the proliferation assay results (XTT assays; PromoKine, Heidelberg, Germany), a 200-ng/ml LPS challenge was selected for cell stimulation. Untreated monocytes served as controls.
After stimulation, the supernatants were collected, cells were lysed in RLT Plus buffer (Qiagen, Venlo, the Netherlands), and stored at −80℃ until analysis.
RNA extraction
RNA was extracted from lysed monocytes using the Qiagen AllPrep DNA/RNA Mini Kit in accordance with the manufacturer’s instructions. For quality control purposes, RNA concentrations were measured using NanoDrop (PeqLab, Erlangen, Germany), and RNA degradation was measured using Bioanalyzer (Agilent Technologies, Santa Clara, CA, USA).
Transcriptome analysis
RNA was amplified and biotinylated using the Illumina TotalPrep RNA Amplification Kit (Life Technologies). Subsequent array-based gene expression analysis was performed using the Human HT-12 v4 Expression BeadChip (Illumina, San Diego, CA, USA) and the automated protocol, in accordance with the manufacturer’s manual. The HT-12 v4 Expression BeadChip comprises 47,231 probes. The arrays were scanned with an iScan microarray scanner (Illumina). Statistical analysis was performed using functions implemented in the statistical software packages R and Bioconductor. All data were subjected to quantile normalization using the limma package.9,10 Selection of differentially expressed genes was performed using the following statistical filter criteria: fold change (FC) ≥ 1.7 and P ≤ 0.05. The FC is the ratio of the group mean values. Transcripts with at least a 1.7-fold change were selected. All adjusted P values were calculated using a Student’s t-test, and adjusted for multiple testing using Benjamini–Hochberg correction.
Protein analysis
To determine the impact of specific transcripts, ELISAs or Multiplex Immunoassays were performed in accordance with the manufacturer’s instructions (eBioscience or R&D (Minneapolis, MN, USA). The supernatants of monocytes challenged with LPS Pg for 6 h were analyzed for IL-10, TNF-α, TIMP-2, CXCL8 (IL-8), CCL8 (MCP-2), CCL23 (variant Ckβ8-1, MIP-3) and CCL4 (MIP-1) concentrations.
Cell surface markers
To assess monocyte activation, surface markers were determined using flow cytometry. First, monocytes were directly blocked with PBS/1% BSA containing 0.1% Fc block (Sigma-Aldrich, St. Louis, MO, USA). Second, monocytes were stained with the following Abs: CD14 FITC (Clone: 62D3); CD16 APC (Clone: eBioCD16); human leukocytes Ag (HLA)-DR PerCP-Cy5.5 (Clone: LN3) (eBioscience); and CD86 (known as B70/B7-2) PE [Clone: 2331(FUN-1)] (BD Bioscience, San Diego, CA, USA). Flow cytometry was performed using BD FACS Canto followed by analysis with FACS Diva software (BD Biosciences) and FlowJo X software (FLOWJO, Ashland, OR, USA).
IPA analysis
Transcriptional data were evaluated using Ingenuity Pathway Analysis (IPA, Systems Inc., Redwood City, CA, USA). 11 Here, each gene is represented in a global molecular network that is designed by information from the Ingenuity Pathways Knowledge Base. ‘Networks’ were generated algorithmically, based on their connectivity in terms of activation, expression and transcription. Molecular relationships between genes are represented by connecting lines between nodes, as supported by published data stored in the Ingenuity Pathways Knowledge Base and/or PubMed.
Data analysis
Protein and surface marker expression profiles were analyzed using Student’s t-test and GraphPad Prism Version 5.0 (GraphPad Software, San Diego, CA, USA). Data are shown as the mean with SEM. P ≤ 5% was considered statistically significant.
Results
Transcriptome induced by LPS Pg in CD14+ monocytes
Initial experiments demonstrated that 90 min of LPS Pg challenge induced 423 differentially regulated transcripts (260 overexpressed and 167 repressed genes; see Supplementary Table S1). The ‘immediate’ genes included candidates of inflammasome activation and immunity (e.g. TNF-α↑, IL-6↑, NLRP3↑, CCL14↑, CCL20↑, ICAM1↑). Comparison of these results with those obtained after 6 h of LPS Pg stimulation revealed distinct differences in expression profiles (Figure 2A). In view of the higher outcomes of prolonged stimulation, LPS challenge for 6 h was further investigated.
Genome-wide transcriptome induced by LPS P. gingivalis challenge in human monocytes. (a) Venn diagram of differentially expressed transcripts (FC ≥ 1.7 and P < 0.05) after LPS Pg challenge, demonstrating differences and similarities between 90 min and 6 h exposure. (b) Volcano plot analysis of LPS Pg-induced genes after 6 h. Red indicates overexpressed and blue indicates repressed genes (n = 902). For the complete list of differentially regulated genes and detailed information on the individual probes, see Supplementary Table S2. (c) Hierarchical cluster analysis of differentially expressed genes (n = 902) after 6 h of LPS exposure. Data are based on seven individuals and independent experiments. Columns represent subjects and rows represent transcripts. Red indicates high relative expression levels and blue indicates low relative expression levels. A hierarchical cluster analysis was performed using Pearson’s correlation as distance measurement and average linkage. (d) Overview of diseases associated with LPS Pg-induced genes visualized by bar plot constructed using IPA (www.ingenuity.com).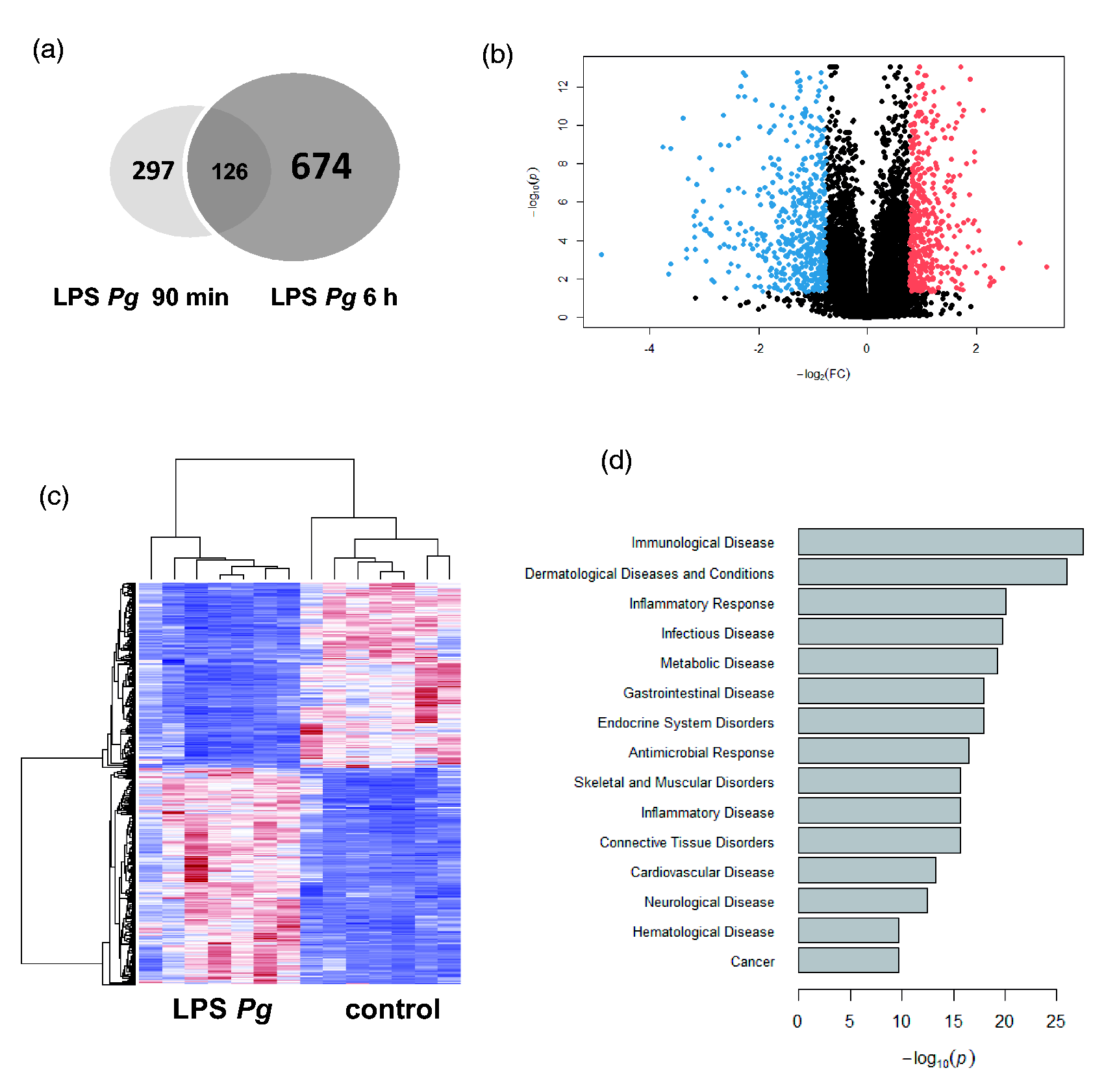
Top transcripts (FC > 5) significantly induced in human monocytes (n = 7 different donors) after 6 h of LPS Pg challenge.

A total of 902 genes showed significant differential expression at a false discovery rate (Benjamini–Hochberg adjusted P-value). The complete list of differentially regulated genes and detailed information on the individual probes are provided in Supplementary Table S2.
Pathways and associated diseases
According to IPA, a total of 231 canonical pathways/signaling were activated during 6 h LPS Pg challenge (Table 2; Supplementary Table S3). These included TREM1, NF-κB, TLR, IL-10, IL-8, HMGB1, VEGF, erythropoietin, CD40, TNFR, iNOS, Notch and leptin signaling. For example, Figure 3A shows the triggering receptor expressed on myeloid cells 1 (TREM1) signaling, which was highly induced by LPS Pg. Periodontitis and rheumatoid arthritis share common host responses,
12
and this was reflected in several LPS Pg-induced pathways (Table 2).
LPS Pg-related gene networks designed using information obtained from the Ingenuity Pathways Knowledge Base. ‘Networks’ were generated algorithmically based on their connectivity in terms of activation, expression and transcription. Direct molecular relationships between genes are represented by connecting lines between nodes/genes, as supported by published data stored in the Ingenuity Pathways Knowledge Base and/or PubMed. Red symbols indicate upregulated gene expression, whereas green symbols indicate downregulated gene expression. (a) TREM1 signaling pathway significantly activated by LPS Pg in human monocytes according to IPA (www.ingenuity.com). (b) Immunologically relevant genes induced by LPS Pg in human monocytes were merged into one network showing the relationship between LPS Pg induced transcripts. (c) Depiction of chemokine networking to display inter-relationships and the interplay between CCL23, MMP2 and TIMP2. Highly induced canonical pathways in human monocytes after 6 h of Pg challenge, as determined using the Ingenuity Pathways Knowledge Base (IPA). The complete list and detailed information are provided in Supplementary Table S3.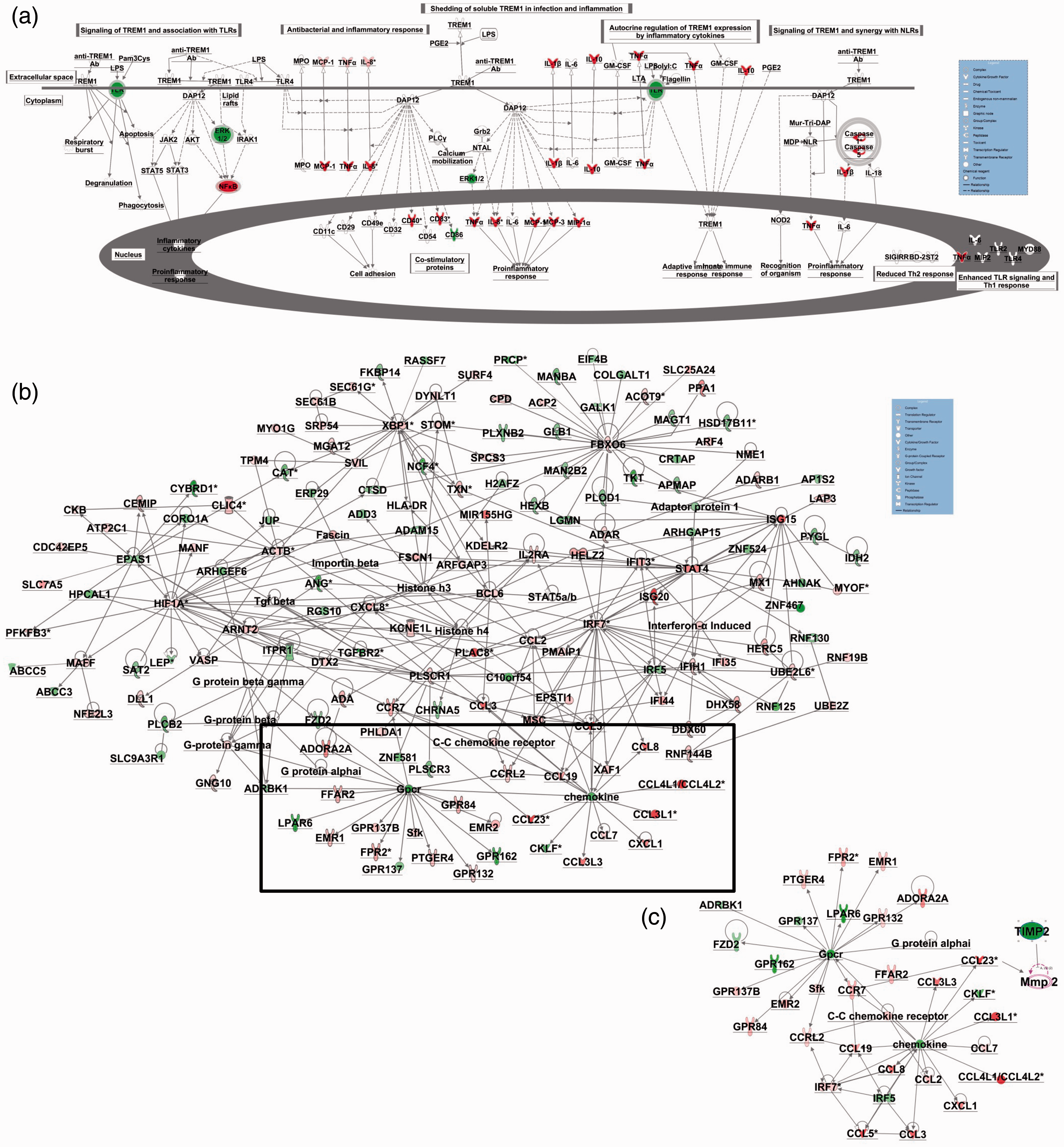
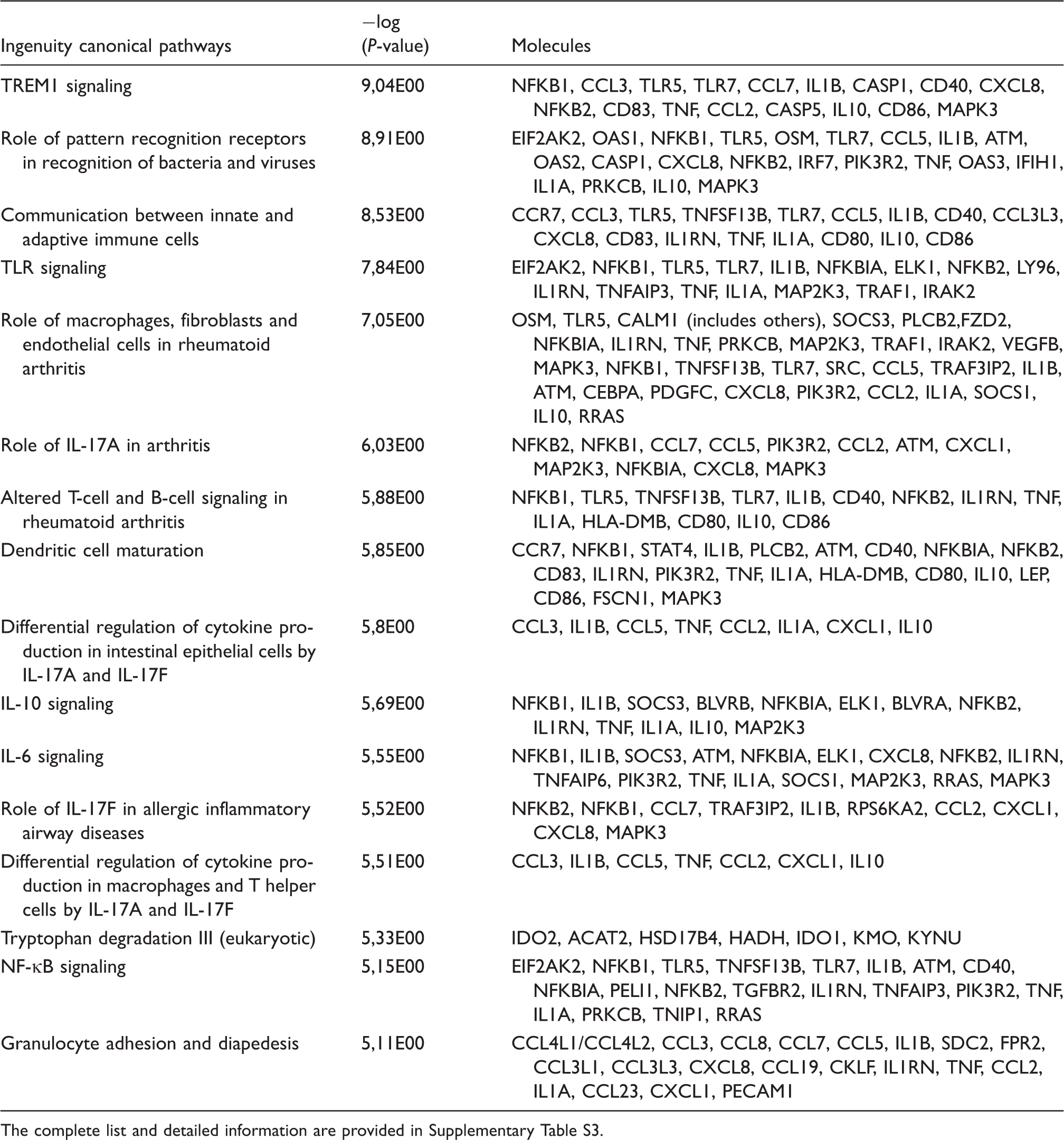
Figure 2D shows diseases with the most pronounced association with LPS Pg-induced genes. These comprise immunological, dermatological, inflammatory, infectious and cardiovascular diseases, as well as cancer. Supplementary Table S4 provides a list of 500 pathologies and functional annotations regulated by LPS Pg. These include systemic autoimmune diseases; the quantity and function of several immune cells (neutrophils, lymphocytes, DCs, macrophages); diabetes mellitus; rheumatic diseases; atherosclerosis; obesity; the synthesis and metabolism of reactive oxygen species; lipid and fatty acid concentrations; Ag presentation; cancer; osteoclast differentiation; and bone resorption.
Upstream analysis
A total of 2149 upstream regulators were determined by IPA (R)-analysis (see Supplementary Table S5). These included various cytokines (IL-1B, TNF-α, IFNs, IL-10); transcription regulators (IRFs, STATs, NF-κB1); transmembrane receptors (TLRs, FAS); and others (NOD2, LDL), which provide additional insights into Pg infection.
Networks
IPA networks were used to depict inter-relationships between diseases, functional annotations and significantly regulated transcripts (Figure 3B, C; Supplementary Figure S1A–E). Immunologically relevant genes with known chemoattractant capacities were merged into one network, demonstrating a direct relationship between these genes (Figure 3B, C). As monocytes differentiate into phagocytes, which are primarily responsible for tissue destruction; genes involved in phagocyte metabolism are networked in Supplementary Figure S1A. Systemic inflammatory and rheumatic diseases associated with LPS Pg infection are networked in Supplementary Figures S1B and C, respectively. Candidates for lipid metabolism, cardiovascular disease, and cancer were also networked, and this identified distinct affected targets such as MMP8 and MIR-124 (see Supplementary Figure S1D). Consideration of genes implicated in cell development, movement and homoeostasis revealed additional targets, such as heat shock proteins (HSPs; see Supplementary Figure S1E).
Protein data
To validate the transcriptome results, protein levels of pro- (TNF-α), anti-inflammatory cytokines (IL-10) and chemoattractants were determined in supernatants of monocytes challenged with LPS Pg for 6 h (Figure 4A–E). In accordance with transcriptional data, TNF-α (1000-fold) and IL-10 (158-fold) proteins were upregulated by LPS Pg (Figure 4A, B). Moreover, LPS Pg induced a significant increase in CXCL8/IL8 protein levels (2660-fold) (P < 0.001, Figure 4C). In the case of CCL8, a marked protein elevation was observed (Figure 4D). The CCL4 data were consistent with our mRNA findings (15-fold) (Figure 4E). CCL23 and TIMP2 levels were below the detection limit after 6 h of LPS Pg challenge (data not shown).
Protein expression of cytokines, chemokines, and markers of differentiation in human monocytes. (a) TNF-α, (b) IL-10, (c) CXCL8/IL8, (d) CCL8 and (e) CCL4 protein levels in the supernatants of human monocytes stimulated with LPS Pg for 6 h. Frequency and mean fluorescence intensity (MFI) of CD14+ monocytes expressing (f) CD86 and (g) HLA-DR/MHCII induced by LPS Pg challenge for 6 h. Data are shown as the mean with SEM and were analyzed using Student’s t-test (n = 4–7; *P < 0.05, **P < 0.01, ***P < 0.001).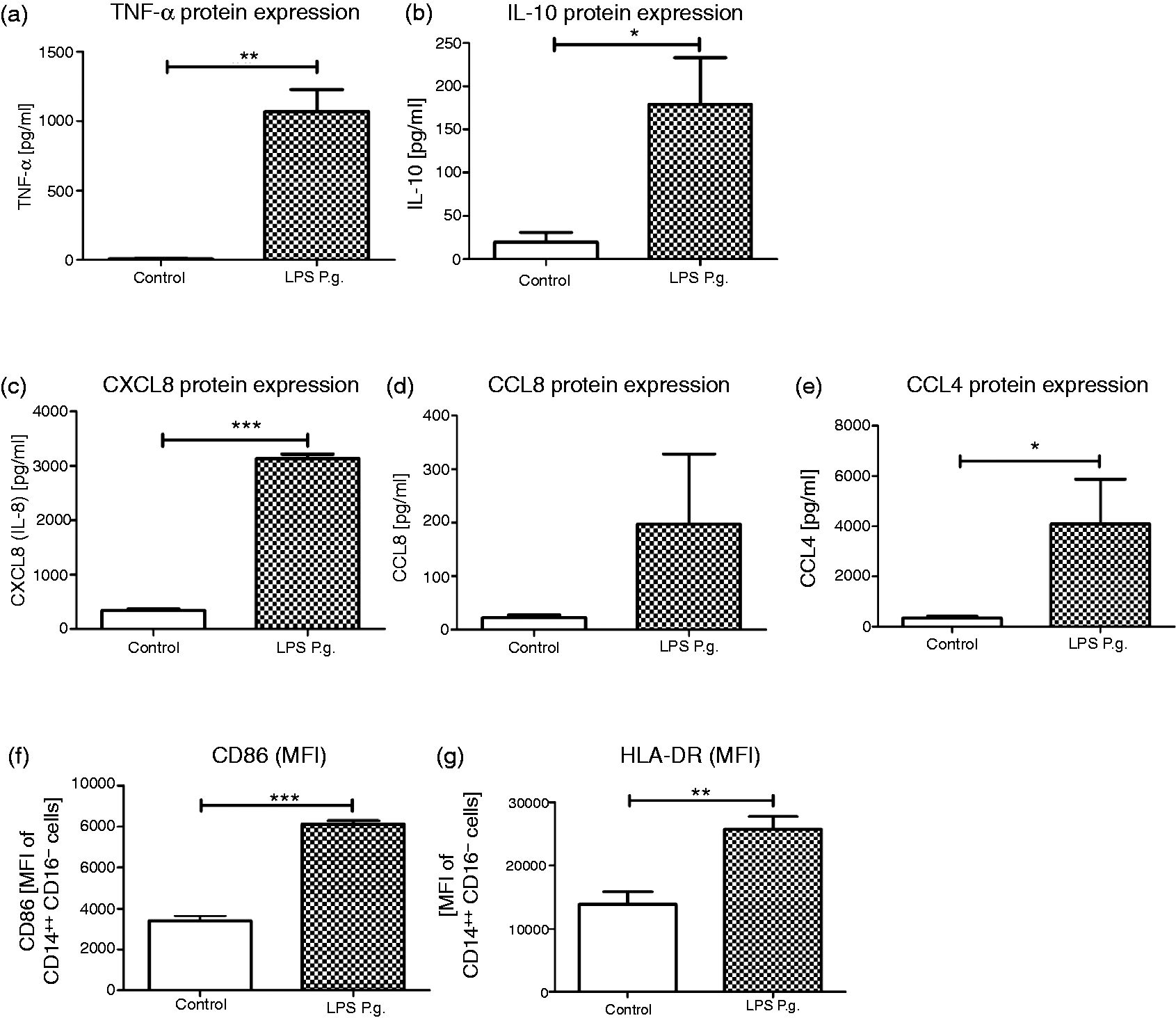
CD14+ monocyte activation and differentiation
As monocytes are key players in periodontal and systemic diseases, their activation and differentiation were investigated via specific surface markers (Figure 4F, G). LPS Pg stimulation induced an upregulation of CD14 and repressed CD16 expression as a consequence of monocyte activation (data not shown).
Moreover, LPS Pg stimulation caused a significant increase in CD86 expression on classical CD14++CD16− monocytes (P < 0.001; Figure 4F). This was accompanied by an overexpression of HLA-DR/MHCII (P < 0.001; Figure 4G). Intermediate/non-classical CD14++CD16+ monocytes displayed similar expression profiles (data not shown).
Discussion
CP is a highly prevalent inflammatory disease, and Pg is a keystone pathogen in its manifestation and severity. Extensive research evidence demonstrates that Pg-induced CP is associated with systemic diseases. 2 As monocytes constitute one of the main immune cells in the blood and play a pivotal role in systemic and periodontal inflammation, 13 we applied a genome-wide transcriptional approach in human monocytes challenged with LPS derived from the periodontal pathogen Pg. In this analysis, 902 genes were significantly induced after 6 h of LPS Pg stimulation. Some of these genes have been associated, in previous reports, with periodontitis (IL-8, NAMPT, IL-1RN, IL-10, TNFA or CD38),6,14–16 thus validating the present findings. Interestingly, several new candidates, such as CCL23, ISG20, IRG1, GBP1, MIR155HG and DHRS9, were detected, accomplished by underlying pathways, upstream regulators and networks corresponding to host immune response. These findings thus support the notion that host immune response is implicated in Pg-induced CP and its association with systemic disease.
The top transcripts included several chemoattractants (CCL3, CCL4, CCL5, CCL8) previously associated with periodontal disease. 17 These chemoattractants are responsible for immune cell recruitment and activation. CCL23 (also known as Ckβ8-1, MPIF1 and MIP3) plays a key role in immunoregulation. To our knowledge, the present study is the first one to implicate CCL23 in Pg-induced immune response (9.91-fold increase). Interestingly, CCL23 is positively correlated with RA, a disorder that has also been linked to periodontal disease.12,18 Furthermore, investigation of a large population-based sample revealed that circulating CCL23 was associated with coronary atherosclerosis. 19 Some of these effects may be attributable to increased matrix metalloproteinases (MMP)2 levels. 20 In addition, CCL23 has chemoattractant effects on immune cells, such as monocytes, DCs and neutrophils, as well as on osteoclast precursor cells. These chemoattractant effects promote diapedesis and bone resorption, processes that are highly implicated in periodontal destruction.21,22 Hence, CCL23 may constitute an interesting new target for CP and its associated disorders, even though protein levels in the present study remained unaltered after 6 h, probably because of kinetics. This hypothesis is supported by Novak et al., 23 who found the highest CCL23 mRNA concentrations in human primary monocytes after 24 h and the highest protein levels after 4 d.
Notably, CXCL8/IL8 transcriptional and protein levels were significantly induced by LPS Pg. CXCL8 acts as chemoattractant for neutrophils, which are key players in periodontal health. 24 Interestingly, a previous report showed that subpopulations of neutrophils in patients with CP displayed enhanced responsiveness to CXCL8, thus supporting the notion of host-derived pathogen-associated periodonal destruction. 25
Additionally, we observed an up-regulation of pro- and anti-inflammatory transcripts accomplished by increased TNF-α and IL-10 protein. In contrast, the expression of TGF-β-related genes (TGFBR2 and TIAF1) was significantly reduced secondary to LPS Pg challenge. These mediators determine inflammatory processes, which favor the recruitment and activation of immune cells. Therefore, reduced or enhanced levels of these mediators predispose to an imbalanced immune response, which may aggravate CP severity. In accordance with this, SNPs in these genes have been associated with CP. 6 Furthermore, initial experiments with 90 min of LPS Pg challenge revealed an increase in NLRP3 inflammasome transcription (eightfold). Thereby, NLRP3 activation triggers IL1β production and release via procaspase-1. This critical defense mechanism has been linked to various pathologies, including periodontitis.26,27
As monocytes are confronted with periodontal pathogens and are key players in systemic disease, we focused on their surface markers. After LPS Pg challenge, transcriptional changes of CD38, CD40, CD86 and HLAs were found, accompanied by elevated CD86 and HLA-DR surface expression. CD86 and HLA-DR are typical mediators APC that stimulate the activation and priming of T and NK cells. 28 The increase in these markers indicates that monocytes were highly activated promoting T-cell regulation and monocyte differentiation to macrophages or DCs related to periodontal and systemic disorders. 2
The top genes included several of relevance to immunity and bacterial infection. Immunoresponsive gene 1 (IRG1) plays a role in innate immunity, inflammatory processes, regulation of NF-κB signaling, oxidative stress and LPS-induced tolerance, and thus influences disorders such as sepsis and carcinoma. 29 IFN-stimulated exonuclease gene 20 kDa (ISG20) has antiviral and immunological capacities and is implicated in diseases of the skin. 30 In accordance with this, the significantly induced transcripts included several IFN-related genes, such as guanylate-binding protein (GBP)1. GBPs are a family of IFN-inducible antiviral and antimicrobial GTPases primarily responsible for bacterial growth restriction. 31 Notably, GBP1 has been shown to inhibit the expression of MMP1, which is implicated in periodontitis.32,33 Dysregulated IFN-related genes might therefore promote bacterial proliferation and periodontal destruction.
MMPs and their inhibitors (TIMPs) are responsible for cell invasion, tissue remodeling and degradation. Excess MMPs activity has been associated with the progression of periodontal disease, cancer metastasis and atherosclerosis. 34 The present analyses identified reduced transcriptional levels of TIMP2 (3.8-fold), whereas altered TIMP2 protein levels were not detectable after 6 h of LPS Pg challenge. TIMP2 is an inhibitor of MMP2, which has recently been identified as candidate for periodontitis. 35 Notably, CCL23 induction has been linked to increased MMP2 expression, thus reinforcing the impact of TIMP2, MMP2 and CCL23 in Pg-induced periodontal destruction.
Furthermore, several genes implicated in oxidative stress (HIF1A, SOD2, CAT and HYOU1, also known as HSP12A) were significantly regulated by LPS Pg. Recent investigations have highlighted the importance of oxidative stress in periodontal disease.33,36,37 Therefore, hypoxic inducible factor (HIF)1α, HSPs and redox systems such as superoxide dismutase (SOD) or catalase (CAT) appear to play a central role in these events, and the present findings support these studies.
Interestingly, overexpressed transcripts included genes involved in tryptophan catabolism, i.e. IDO1 and INDO. Both genes are implicated in NAD biosynthesis and tryptophan degradation. They regulate immunological, inflammatory and apoptotic processes, as well as T-cell proliferation and tolerance induction, all of which are associated with systemic disorders. 38 Current results indicate that these two genes may also be new candidates for Pg-induced CP.
Concerning microRNAs like MIR155HG (6.91-fold increase), current knowledge indicates that they have emerged as key negative post-transcriptional regulators referred to as ‘fine-tuners’ of gene expression. Thereby, each miRNA influences several target mRNAs, 39 and research suggests that miRNAs are also involved in Pg-induced changes in innate immunity.40,41 MIR-155 is encoded within a region known as the B-cell integration cluster gene. Altered expression patterns have been associated with autoimmune diseases and several malignancies of B and myeloid cell origin. 42 Recent studies have generated evidence for the impact of B cells and miRNA-155 on periodontitis.16,43 Moreover, our findings are validated by Tili et al., 44 who found that putative inflammation-related targets of miR-155 include the suppressor of cytokine signaling 1 (SOCS-1), which has been linked to periodontitis. 45 We hypothesize that the application of specific human expression chips for miRNA analysis would clarify the impact of miRNAs in systemic Pg infection.
As for the most repressed genes, DHRS9 (6.58-fold) and MS4A6A (8.53-fold), their roles remain unclear. MS4A6A encodes specific membrane proteins with an involvement in cancer. 46 Dehydrogenase/reductase member 9 (DHRS9) is an enzyme responsible for androgen and progesterone metabolism, which might explain reports of impaired reproduction in patients with periodontal disease. 47 As steroids have anti-inflammatory properties and inhibit the immune response, altered DHRS9 expressions may exacerbate inflammatory disorders such as periodontitis.
Our data complement recent investigations, which applied microarray technology and different cell types, stimuli and exposure durations.48–50 For instance, Yu et al. investigated human macrophages isolated from buffy coats that were challenged with 10 µg/ml LPS or live Pg for 2 h. 48 The authors detected 575 differentially regulated genes in the LPS group based on three independent macrophage experiments, and identified 13 pathways, most of which correspond to host immune defense. 48
The present study identified >900 genes, >2149 upstream regulators and 200 canonical pathways implicated in the Pg-induced immune response. Differentially regulated genes were associated with immunological, inflammatory and infectious disorders; insulin and leptin signaling; diabetes; arthritis; carcinogenesis; and cardiovascular diseases. This confirms that Pg-associated CP may contribute to systemic disorders, and that this is likely to occur via dysregulation of the host immune response.1,12 As the present data are based on human monocytes of relatively young males of European origin challenged with a virulence factor of Pg, our results might differ with that obtained from females, older individuals, people of other ethnicities, different cells or the whole bacterium. Moreover, while gene expression patterns are essential in terms of determining molecular mechanisms, they provide no insights into functional variability, as post-transcriptional and post-translational modifications can compensate changes in RNA expression. However, our data suggest that altered gene expression, or SNPs in the identified genes, may predispose to CP and systemic disorders. In addition, we validated our transcriptional findings by evaluating the protein levels of several mediators and cell surface markers.
In conclusion, the present systematic genome-wide transcriptional approach in human monocytes explored LPS Pg-elicited immune responses and identified new target genes and pathways of relevance to diagnostic and therapeutic strategies. SNPs in these genes may predispose to, or aggravate, periodontal disease due to immune hypo- or hyper-responsiveness. The present findings also support the hypothesized interplay between periodontal infection and systemic diseases, and further research is therefore warranted.
Footnotes
Acknowledgements
We thank all study participants for their cooperation. We acknowledge the role of our laboratory technicians in database management, and thank Jessica Becker, Bonn, Germany, for her helpful advice.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: LG was awarded a Gerok Fellowship in the Clinical Research Unit 208 (‘Aetiology and Sequelae of Periodontal Diseases – Genetic, Cell Biological, and Biochemical Aspects’), which was supported by the German Research Society (DFG, KFO 208), as well as intramural funding from the University Hospital of Bonn (BONFOR 2014-1-34). BB was supported by the Jürgen Manchot Stiftung, Düsseldorf. AH and MN are members of the Excellence Cluster Immunoscience (DFG, EXC 1023), which supported parts of this study. The funders had no role in study design, data collection and interpretation, or the decision to submit the work for publication.
