Abstract
Lipopolysaccharides are structural components within the cell walls of Gram-negative bacteria. The LPSs as microbe-associated molecular pattern (MAMP) molecules can trigger defense-related responses involved in MAMP-triggered immunity and basal resistance in plants, presumably from an initial perception event. LPS from Burkholderia cepacia as well as two fragments, the glycolipid, lipid A and the polysaccharide (OPS-core) chain, were used to treat Arabidopsis thaliana seedlings to evaluate the eliciting activities of the individual LPS sub-domains by means of Annealing Control Primer-based Differential Display transcript profiling. Genes found to be up-regulated encode for proteins involved in signal perception and transduction, transcriptional regulation and defense – and stress responses. Furthermore, genes encoding proteins involved in chaperoning, secretion, protein–protein interactions and protein degradation were differentially expressed. It is concluded that intact LPS, as well as the two sub-components, induced the expression of a broad range of genes associated with perception and defense as well as metabolic reprogramming of cellular activities in support of immunity and basal resistance. Whilst the lipid A and OPS moieties were able to up-regulate sub-sets of defense-associated genes over the same spectrum of categories as intact LPS, the up-regulation observed with intact LPS was the more comprehensive, suggesting that the lipid A and glycan molecular patterns of the molecule act as partial agonists, but that the intact LPS structure is required for full agonist activity.
Introduction
In addition to preformed defenses, plants can specifically recognize pathogenic microbes and then respond with appropriate defense mechanisms by activating a multicomponent defense response.1,2 Complex signaling pathway(s), involving cytosolic Ca2+ and H+ ions, reactive oxygen intermediates (ROI), jasmonic acid (JA), salicylic acid (SA) and ethylene, trigger inducible defense responses 3 characterised by defense gene activation, activation of programmed hypersensitive response (HR) cell death to restrict the invading pathogen and production of phytoalexins and pathogenesis-related (PR) proteins. 4 – 6
Activation of the signal transduction cascades depends on the initial recognition of the plant pathogen, perceived by receptor proteins encoded by receptor-like protein/kinase (RLP/K) genes to trigger pathogen-triggered immunity (PTI). Microbe-associated molecular pattern (MAMP) molecules have several common characteristics including highly conserved structures, indispensability to the micro-organism and their absence in host organisms.7,8 To date, several MAMPs from different micro-organisms (bacteria, fungi, and oomycetes) have been reported with the potential to trigger plant defense responses, and include flagellin, chitin, cell wall derived oligogalacturonides, elongation factor Tu, peptidoglycan and LPSs.2,9 The corresponding receptors for some of these MAMPs in plants have been identified and characterized as pattern recognition receptor-like proteins/kinases (PRR-RLP/Ks). Such include, amongst others, the flagellin sensing 2 (FLS2) receptor for flagellin and the EFR for the elongation factor Tu; their functions towards enhanced plant pathogen responses are well documented. 10 – 12 Although no receptor for LPSs has been identified thus far, it can be recognized by plants to trigger defense-related responses. 13 – 17 These biochemical changes may be associated with the HR-preventative and priming activities reported for LPSs. 16
On-going research on the LPS from a plant-associated endophyte (Burkholderia cepacia, strain ASP B 2D) has shown that it exhibits inducing capacity as a modulator of defense-related responses. 18 We have previously shown that LPSB.cep. can elicit defense responses on a cellular level. These responses include influx of calcium, an oxidative burst, an NO burst, extracellular alkalinization (K+/H+ exchange), and changes in reversible protein phosphorylation, including MAP-kinase activation.14,19– 23 In addition, the synthesis of inducible PR proteins was observed in tobacco. 18 The identification of RLK and resistance (R) genes up-regulated by LPSB.cep. in tobacco cells was indicative of the operation of an innate immune recognition system and expression of basal resistance. 18 The altered gene expression in response to LPS may be responsible for activation of surveillance mechanisms and enhancement of the non-self recognition capacity 24 supporting the induction of an enhanced defensive capacity 18 or systemic acquired resistance (SAR) by LPSs. 25
Up to now, the eliciting or MAMP-active part(s) of LPSs have not been fully identified. As lipoglycan macromolecules, LPSs can potentially contain MAMP structures within the lipid A, oligosaccharide core and O-polysaccharide (OPS). Lipopolysaccharides from bacteria have developed a wide spectrum of compositionally different LPSs 26 and not only the OPS, but also the attached core and the lipid A region can vary in structure. The OPS extending outward from the bacterial cell surface are exposed to the environment and the defense system of a potential host, and have been found to play a role in triggering plant defenses. The structural variation of the OPS of bacterial pathogen and symbiont LPSs and their roles in plant-microbe interactions was previously reviewed. 27 The composition and/or size of the OPS might be reliable indicators of virulence potential and appears to be part of a molecular communication between the bacterium and the host plant.27,28
It is increasingly believed that the lipid A of LPSs from different species lies at the center of the biological activity of LPSs;16,23 for example, lipid A has been implicated to be the inducing part of LPS from Xanthomonas campestris pv. campestris.16,29,30 This is believed to be due to the fact that lipid A is relatively conserved in many Gram-negative bacteria, while the OPS shows variations from one organism to another.28,31 Rough-LPS (LPS lacking an OPS), i.e. lipooligosaccharides (LOSs), were found to exhibit biological activity similar to those of intact LPSX.cam. These activities include induction of plant defenses, monitored by the expression of the PR1 and PR2 genes. 32
It has been observed that certain bacteria can manipulate the composition of their lipid A in response to environmental cues and thereby modulate or even antagonize the triggering of innate host responses.16,30,33,34 Variations in lipid A structures are exhibited by different fatty acid and phosphate numbers and locations and can lead to a heterogeneity even in the same bacterium. 35 The chemical composition, structure and conformation of lipid A may thus be important determinants during a plant’s interaction with pathogenic or beneficial bacteria.
Support for multiple MAMPs within the LPS structure comes from Silipo et al., 32 that showed that the LOS from X. campestris was able to induce defense-related gene transcription in two temporal phases. This LOSX.cam. did not contain an OPS chain, but only a very peculiar core oligosaccharide ending up with a terminal mannose disaccharide. The core oligosaccharide induced the earlier phase only and, in contrast, the lipid A induced the latter phase only. The data suggested that not only can plant cells recognise lipid A and core oligosaccharide structures within LOS in order to trigger defensive cellular responses, but that this recognition may occur via two distinct recognition events.28,32
Using a different approach with LPS mutants of X. campestris, Braun et al.
29
reported that the LPSX.cam. substructure recognized by tobacco cells is localized in the inner core region of the LPS, consisting of the glucose, galacturonic acid and 3-deoxy-
In the current study, the MAMP-active eliciting determinant(s) of LPSB.cep. were investigated by comparing the genes of which the expression is increased at a transcriptional level by lipid A and the OPS to those induced by the intact LPS in A. thaliana. Here, a very sensitive variant of differential display mRNA profiling, the Annealing Control Primer (ACP)-based Differential Display Reverse Transcription Polymerase Chain Reaction (DDRT-PCR)36,37 was used to obtain information about the respective transcriptomes.
Materials and methods
Plant material
Arabidopsis thaliana (Colombia ecotype) seeds were surface sterilized and grown in 0.5× Murashige and Skoog (MS) medium containing 2% glucose at pH 6. 38 Seedlings were grown in horizontal Falcon tubes on an orbital shaker at approximately 25 rpm at 24°C in a 9-h light/15-h dark photoperiod.
Lipopolysaccharide, lipid A and OPS preparation
Burkholderia cepacia strain ASP B 2D was grown in nutrient broth liquid medium and LPS was extracted, purified and characterized as previously described.
18
The OPS was prepared and purified as previously described.
31
Briefly, LPS was hydrolyzed with aqueous 1% acetic acid for 2 h at 100°C and centrifuged (6000 g, 4°C, 1 h). The supernatant (OPS-core fraction, 55 mg, 90% LPS), was further purified by gel permeation chromatography on a Sephacryl S100-HR column (90 × 1.5 cm) using 0.05
Plant elicitation
Ten-day-old seedlings were harvested and placed in fresh 0.5× MS medium. Stock solutions of LPS, lipid A and OPS in MS medium were added to final concentrations of 100 µg/ml, 20 µg/ml and 50 µg/ml respectively, in separate tubes. Plant elicitation was allowed to take place for 8 h while the tubes were continuously rotated at 25 rpm as described above.
Isolation of RNA
After 8 h of elicitation, seedlings were harvested and collected using vacuum filtration. Total RNA was isolated using the Trizol reagent method (Sigma-Aldrich, Germany). RNA integrity and concentration were assessed by denaturing RNA gel electrophoresis and NanoDrop Spectrophotometry (NanoDrop Technologies Inc., Wilmington, USA). The most important aspect for the success of ACP-DDRT-PCR is the quality of RNA. The RNA used in the study was of a high standard, and DNA-free, following treatment with RNase-free DNase (Promega) according to the manufacturer’s instructions.
Differential display
Annealing Control Primer-based DDRT-PCR was performed on the total RNA using the GeneFishing DEG premix kit (Seegene, Seoul, South Korea) according to the manufacturer’s instructions, using 40 different arbitrary primers in combination with a oligo-dT anchor primer. Annealing Control Primer-based DDRT-PCR amplicons were evaluated on 1.5% agarose gels stained with Gel Green stain (Biotum Inc., Hayward, USA). In order to identify differentially expressed bands, LPS, lipid A, and OPS-responsive amplicons were analyzed parallel to the control amplicons. Display patterns obtained with biological replicates were reproducible. Bands which represented up-regulated genes were excised from the gel with a sterile scalpel blade and kept at –80°C until further use. The DNA was extracted from gel slices using the Zymoclean Gel DNA Recovery kit (Zymo Research, USA). The extracted cDNAs were re-amplified using two universal primers with sequence complementary to the 5’ end of the ACP-primers used in the ACP-DDRT-PCR. The sequences of the primers were: upstream primer, 5’-GTCTACCAGGCATTCGCTTCAT-3’ and downstream primer, 5’-CTGTGAATGCTGCGACTCGAT-3’.
Sequencing of DNA and bioinformatic analysis
Approximately 12 µg of DNA derived from either LPS, lipid A or OPS treatments were labelled with different tags in order to distinguish them from one another. Sequencing of DNA was performed with a GS-FLX sequencer (Inqaba biotec, RSA), with the aid of 454 high-throughput pyrosequencing technology using the Genome Sequencer 20 system (Roche diagnostics, Germany). Using the different sequencing tags, the source of the amplicon after sequencing could be identified, i.e. whether it was derived from the LPS, lipid A or OPS pool. Gene annotation or identification was done using the basic local alignment search tool (BLAST) programme from NCBI (National Center for Biotechnology Information), the Arabidopsis Information Resource (TAIR; <http://arabidopsis.com>), Genevestigator Response Viewer (<http://www.genevestigator.com>, 40 and the InterPro database (<http://www.ebi.ac.uk/interpro/>).
Reverse transcriptase-PCR analysis
In order to confirm the up-regulation observed with ACP-DDRT-PCR, four genes were selected and investigated with RT-PCR in comparison to 18S rRNA as a reference gene. Total RNA was isolated following treatment with 20 µg/ml lipid A or 100 µg/ml LPS for 4 h and 8 h, using the Trizol reagent method according to the manufacturer’s instructions (Sigma-Aldrich, USA). Following DNase treatment (Promega, USA), semi-quantitative RT-PCR was performed with the Transcriptor-one-step RT-PCR kit (Roche, Germany). Transcription profiles of the lipid transfer protein (LTP, At4g12470) and oxysterol-binding protein (OBP, At4g08180), serine hydroxymethyl transferase 1 (At4g37930) and the ACT domain-containing protein (At2g39570) were obtained. Primer pairs were designed using the Primer3 programme and their sequences were as follows:
LTP-Fw TAATCCTAGGCCGGTCACAC LTP-Rv GACTCCCGATTGCAAGAACC OBP-Fw AAGGCACTGCCACTGAGTCT OBP-Rv TCATCGGACGAAAATGATGA ACT-Fw GGGTTGTGATTGCTGTTGTG ACT-Rv GGGTCAACTCTGTCGCTCTC SHT-Fw CCAGGAACCAAGTTGAAGGA SHT-Rv CCAATTGTTGGGAACTGCTT 18S-Fw CGGGTACCACATCCAAGGAA 18S-Rv GCTGGAATTACCGCGGCTCT
The thermocycler RT-PCR programme was as follows: reverse transcription at 50°C for 30 min, initial denaturation at 94°C for 2 min, denaturation at 94°C for 30 s, annealing at 58°C for 30 s, elongation at 68°C for 30 s and final elongation at 68°C for 7 min. The samples were then analyzed on a 1.5% agarose gel and stained with ethidium bromide.
Results and discussion
Molecular patterns within LPS
The LPS from this plant-associated B. cepacia
18
is a macromolecular (Mr > 12 kDa), hydrophilic heteropolysaccharide, that is covalently linked through a core oligosaccharide (Mr approximately 1.8 kDa) to the glycolipid moiety, lipid A (Mr approximately 1.6 kDa). An LPS-specific SDS-PAGE analysis of LPSB.cep. is shown in Figure 1. The lanes on the gel loaded with purified LPS displays bands representing LPSB.cep. of various molecular sizes. The characteristic ladder represents LPS biosynthesis intermediates, consisting of the lipid A and core oligosaccharide, but containing a different number of oligosaccharide repeating units that forms the extended OPS.
Electrophoretic analysis of purified LPS from B. cepacia by 12.5% denaturing SDS-PAGE gel containing 4 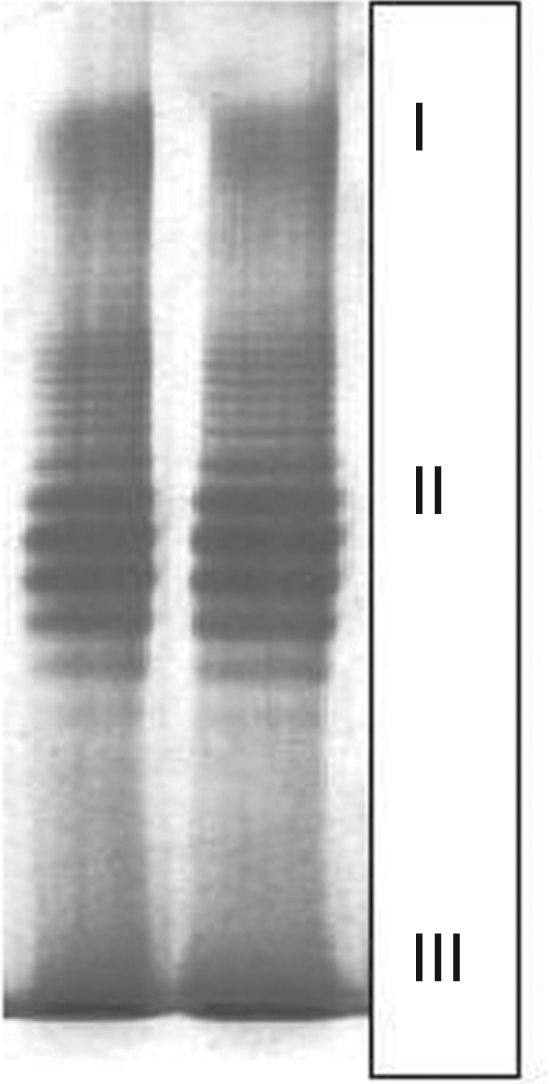
The repeat unit of the OPS of the LPSB.cep. lipoglycan was recently characterized in our laboratories as [→2)-β-
Transcriptional reprogramming in response to LPS, OPS and lipid A
Messenger RNA differential display has been successfully used to investigate alterations in transcriptome profiles and to identify and isolate genes involved in plant stress responses, including plant–microbe interactions.15,42,43 Annealing Control Primer technology is an improved method for identification of differentially expressed genes (DEGs) using a novel, patented annealing primer system. Because of high annealing specificity during PCR using the ACP system, the application for DEG discovery generates highly reproducible, authentic, and long PCR products. Furthermore, it generates no false positives.36,37 A representative electrophoretogram which shows the outcome of the DDRT-PCR and re-amplification PCR of selected amplicons is shown in Figure 2. It was also found that the sequences aligned perfectly to the subject sequences during BLAST (basic local alignment search tool; National Centre for Biotechnology) searches, with ‘E-values’ for alignments below 10–50.
(A) Representative electrophoretogram of DDRT-PCR amplicons derived from treated versus control samples using arbitrary primer pairs 35–40. M, C, T represent DNA molecular size ladders (1000–50 bp), control and treated, respectively. Only bands that exhibited strong differential expression were selected for re-amplification and further analysis. (B) Electrophoretogram of selected re-amplified cDNA amplicons.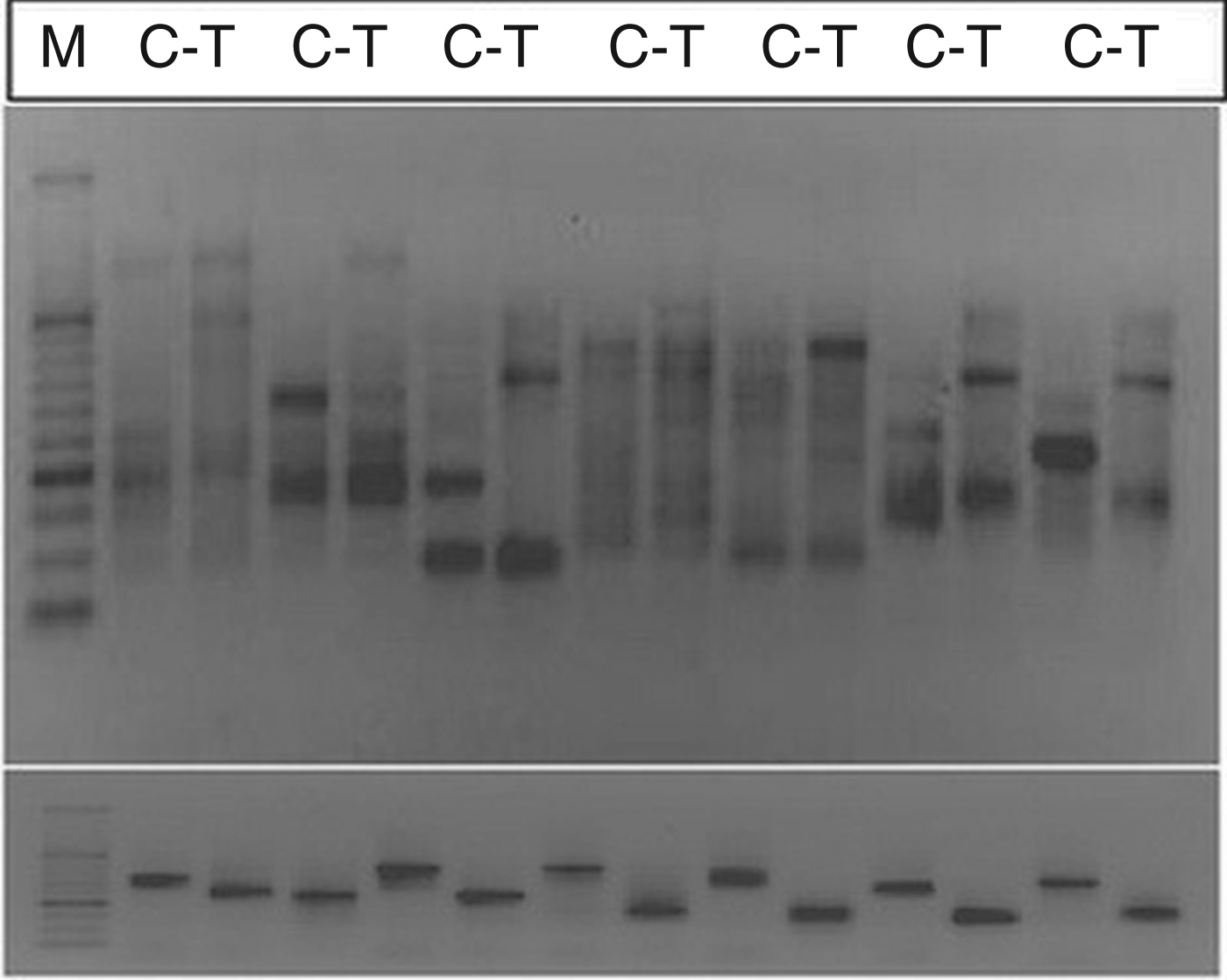
List of A. thaliana genes found to be up-regulated in response to elicitation by LPS, OPS and lipid A, 8 h after treatment of 10-day-old seedlings grown in 0.5× MS medium
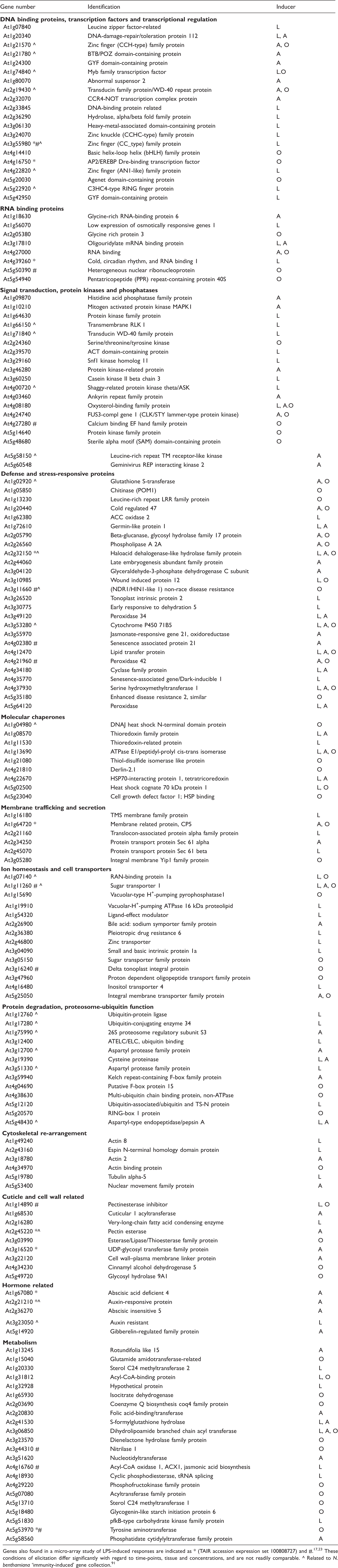
Genes also found in a micro-array study of LPS-induced responses are indicated as * (TAIR accession expression set 100808727) and #.17,23 These conditions of elicitation differ significantly with regard to time-points, tissue and concentrations, and are not readily comparable. ^ Related to N. benthamiana ‘immunity-induced’ gene collection. 91
In order to show the overall number of genes which were found to overlap between all three elicitors, a Venn diagram was constructed (Figure 3). This shows the number of genes which were found to be uniquely expressed in response to individual elicitors. Out of 97 genes induced by LPS, 67% were not overlapping (thus were uniquely expressed). Out of 82 genes induced by lipid A, 51% were not overlapping, and out of 85 genes induced by the OPS, 63% were not overlapping. Figure 3 also shows the relatively small number of genes which were found to be co-regulated by all three elicitors, thus indicating that the responses might be due to distinct perception events. These observations are in support of that made by Silipo et al.
32
with LOS and lipid A from X. campestris.
Venn diagram of genes induced in response to elicitation by LPS, lipid A, and the OPS. The diagram shows both overlapping (common) and non-overlapping (distinct) genes, as indicated by the numbers in the interceptions and within the circles, respectively.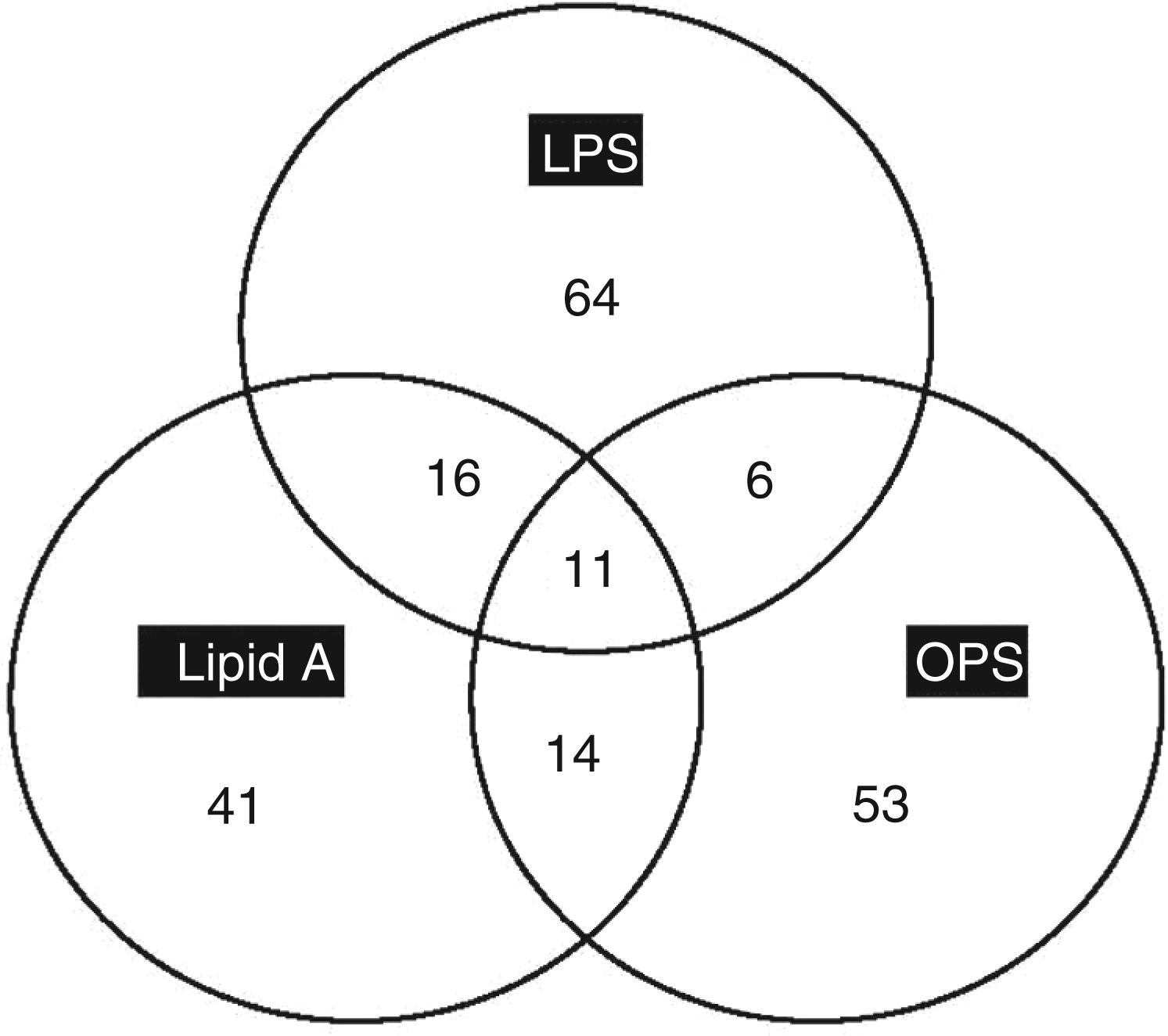
Transcriptional reprogramming of plant cells upon pathogen attack can be extensive, potentially affecting between 3–12% of the 24,000 tested A. thaliana genes upon fungal or bacterial challenge, respectively. 45 In the case of tomato responding to Pseudomonas syringae pv. tomato, more than 400 genes were found to be up-regulated as part of PTI and effector-triggered immunity (ETI) responses. 46 However, using intact LPS and its sub-components as single, defined entities, these numbers are expected to be much lower. The response of the plant to potential MAMPs will depend on the structural features/molecular patterns of the elicitors, the corresponding perception capabilities of the plant cells and the time required for signal transduction processes to exceed a threshold value.
After each amplicon was identified and annotated, genes which code for proteins with similar functions were grouped into functional categories based on described gene ontologies with regard to cellular component, molecular function and biological process. Members from these functional categories are discussed below with reference to their reported involvement in plant defense responses, as supported by previously published data, with the gene number followed by the inducing agent (L, A or O) in brackets. This enables a comparison of lipid A (A) and the OPS (O), based on their eliciting potential, judged from the genes that they have induced in comparison to those induced by intact LPS (L).
Resistance and defense-related proteins
The PTI-induced transcriptional responses include a large number of genes, including many that encode RLKs, some of which may be involved in MAMP perception.47,48 At1g66150 (L) and At5g58150 encode a ‘receptor-like transmembrane kinase 1’ and a ‘leucine-rich repeat (LRR) transmembrane receptor-like kinase’. The RLKs are known to act as surveillance sentinels, alerting the plant to pathogen attack.2,9 Related to this function, At1g13230 (O) encodes a ‘LRR family protein’ of which the role in plant defense responses is well reviewed. 49 At3g11660 (O) encodes the ‘NDR1/HIN1-like protein’, i.e. the tobacco harpin-induced protein and A. thaliana non-race-specific disease resistance gene and At5g35180 (O) encodes an ‘enhanced disease resistance 2’ protein.
Ion homeostasis
Changes in pH of cellular compartments and alkalination of extracellular compartments are due to influx of H+ and efflux of K+ across the cellular membrane. This process is one of the earliest cellular responses involved in signal perception and transduction associated with defense 50 and is also a response induced by LPSB.cep. in tobacco. 14 Two of the identified genes, At1g19910 (L) encoding a ‘vacuolar H+-pumping ATPase proteolipid’ and At1g15690 (O), encoding a ‘vacuolar-type H+-pumping pyrophosphatase 1’ support changes in intracellular pH. In addition, an influx of calcium is another early response induced by LPSB.cep. 14 and At4g27280 (A, O), encoding a ‘calcium-binding EF hand family protein,’ might be involved in the calcium second messenger system activated in response to LPS.
Signal transduction and protein kinases
Cytoplasmic kinases do have the potential to integrate immune signaling from multiple immune receptors and genes encoding intracellular protein kinases acting downstream from RLKs, were found to be induced by all three elicitors. The genes responsive towards intact LPS include: At1g64630 (L), a ‘protein kinase family protein’, At3g60250 (L), a ‘casein kinase II beta chain 3’, At3g29160 (L), the ‘SNF1 kinase homolog 11’. In non-plant systems, it represents a central regulator of metabolic and stress responses. 51 At4g00720 (L) encodes a ‘shaggy-related protein kinase theta/ASK-theta’; the product of this gene is believed to play a vital role in signal transduction and was up-regulated in A. thaliana plants challenged with Bradyrhizobium spp. strain ORS278 and P. syringae. 52 The OPS induced At5g14640 (O), encoding a ‘protein kinase family protein’ and At2g24360 (O), a ‘serine/threonine/tyrosine kinase protein’.
The MAPKs are the most classical examples of those whose activities in signal transduction are well documented and At1g10210 (A), a ‘mitogen activated protein kinase 1’ was up-regulated in response to lipid A. Recently, AtMAPK1 was found to be induced by different stresses such as wounding, ABA, JA and H2O2. 53 lipid A induced other protein kinase genes as well, such as At3g46280, At1g10210 and At5g60548.
The up-regulation of the oxysterol-binding protein gene, At4g08180 (L, A, O), is significant (Figure 4). These proteins also bind phosphoinositides and play roles as lipid sensors/transporters in signal transduction by controlling the extracellular signal-regulated (ERK/MAPK) pathways in mammalian cells.
54
In A. thaliana, the gene was found to be up-regulated in response to pathogens (Botrytis cinerea and P. syringae – Genevestigator) and in potato in response to Phytophthora infestans and oligogalacturonides.
55
Semi-quantitative RT-PCR of the expression of the genes encoding the lipid-transfer protein (LTP – At4g12470) and the oxysterol-binding protein (OBP – At4g08180) in response to lipid A elicitation and serine hydroxymethyltransferase (SHT – At4g37930) and ACT domain-containing protein (ACT – At2g39570) in response to LPS elicitation. 18S rRNA was used as reference gene.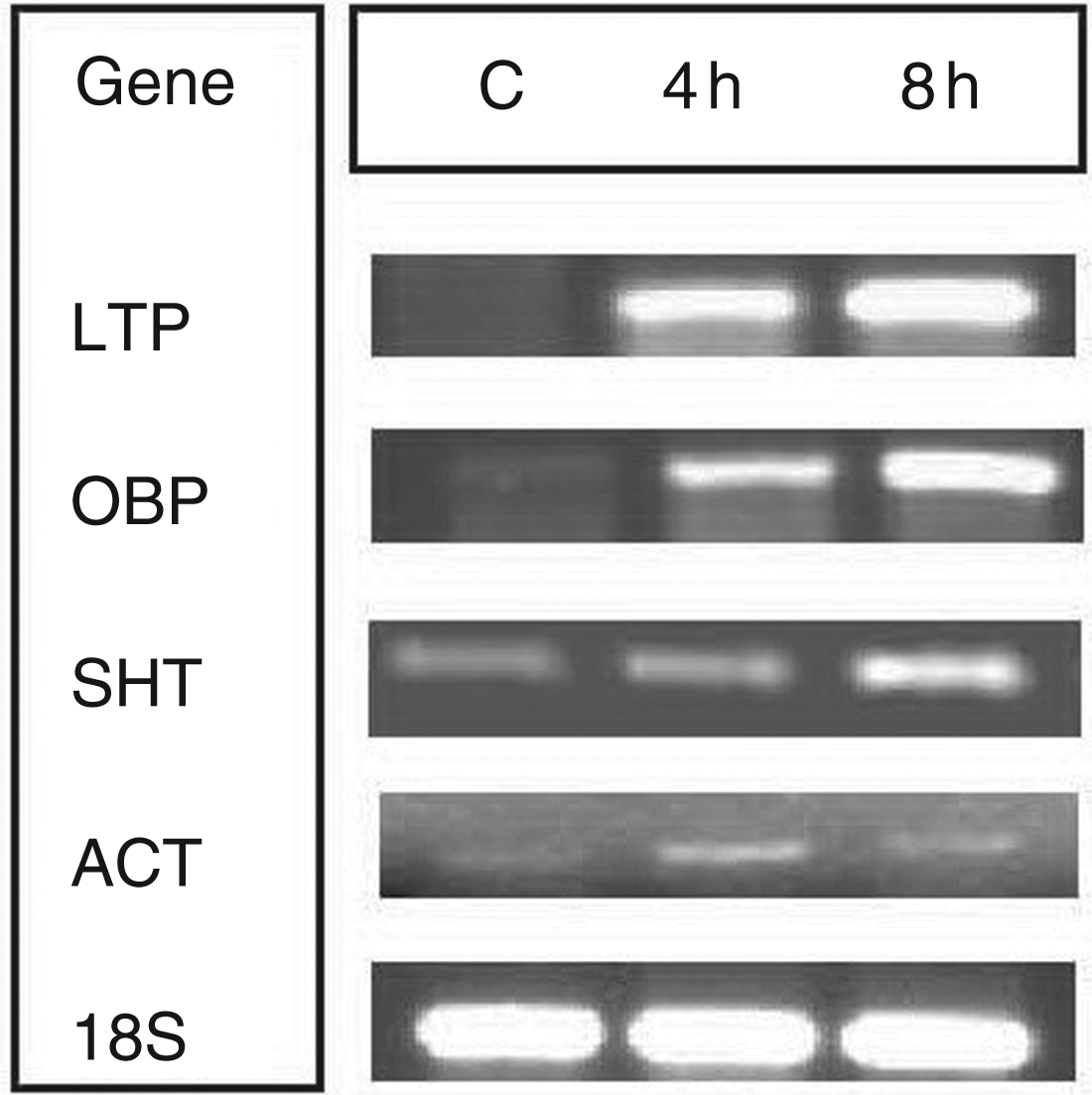
Transcription factors
Microbe-associated molecular pattern-responsive transcription factors have been proposed to be involved in amplifying the PTI response 45 and several transcription factor genes were responsive to all three elicitors (Table 1). These include: At1g21570 (A, O), a ‘zinc finger (CCH-type) family protein’; At3g55980 (L), a ‘zinc finger (CC-type) family protein’; At5g22920 (L), a ‘C3HC4-type RING finger’; At1g07840 (L) a ‘leucine zipper factor’; At4g22820 (L), a ‘zinc finger (AN1-like) family protein’; At3g24070 (L), a ‘zinc knuckle (CCHC-type) family protein’ and At1g74840 (O), a ‘Myb transcription factor’. Furthermore, At4g16750 (O), the ‘ethylene responsive transcription factor (ERF)/dehydration-responsive element (DRE)-binding transcription factor’ and At4g14410 (O), a member of the ‘basic helix-loop-helix (bHLH) family, were up-regulated by the OPS. From this list, it is noticeable that more were induced by intact LPS as compared to the other two elicitors. The above-mentioned genes were also amongst the ones identified by Truman et al. to modify basal defence responses during pathogenesis and resistance. 47
Protein synthesis, folding and secretion
Ten genes with functions related to ribosomes were identified (Supplementary Table S2), presumably in support of enhanced protein biosynthesis required for defense. Chaperones play a vital role during plant–pathogen interactions, being capable of binding and stabilizing proteins such as PRRs at intermediate stages of folding, assembly and translocation from endoplasmic reticulum to plasma membranes.56,57 The LPS and its sub-components were found to induce genes encoding proteins with chaperonin activity. These proteins include At5g02500 (L), a ‘heat shock cognate 70 family protein’ and At4g22670 (L, A), a ‘HSP 70 interacting protein 1’/tetratricoredoxin. Members of the HSP70 family proteins are known to be involved in protection of cellular homeostasis following stressful stimuli, including biotic stressors 58 and, together with HSP90, to be involved in the HR and non-host resistance. 59
Chaperonin-related genes include At5g53400 (A) that encodes a small heat shock protein; At1g13690 (A, O), that encodes ‘(ATPase1) ATE1’, known to stimulate the ATPase activity of the heat shock proteins DNAK/DNAJ, and At1g21080 (O), a ‘DNAJ heat shock N-terminal domain containing protein’ gene. The up-regulation of a related ‘DNAJ heat shock N-terminal domain containing protein’ At1g56300, was found in A. thaliana after treatments with bacteria (P. syringae), PAMPs and oomycetes. 60
In addition to their involvement in protein folding, thioredoxins have also been implicated in plant defense responses. The CITRX thioredoxin may directly interact with the tomato resistance protein, Cf-9, to regulate the defense, enhancing resistance. 61 Two genes which encode thioredoxin-related proteins were found to be induced: At1g08570 (L), a ‘thioredoxin family protein’ and At1g04980 (O), a ‘thiol-disulphide isomerase-like protein’.
Protein translocation and vesicle trafficking are important aspects of early signaling and defense in response to general elicitors. 62 Lipid A induced genes encoding transport proteins with distinctive functions in plant stress responses. At2g34250 (A) and At2g45070 (L) encode ‘transport protein sec61α and β’ respectively, secretion-related genes of the Sec61 translocon complex that provides a channel for proteins to cross the endoplasmic reticulum membrane. These genes and the induction of a protein secretory pathway was reported to be required for SAR. 63
Stress response proteins
Several genes encoding stress-responsive proteins were found to be induced by all three elicitors. At1g02920 (A, O), encoding a ‘glutathione S-transferase enzyme 11’ was reported to be induced in A. thaliana cell suspensions treated with fungal elicitor and in plants challenged with Peronospora parasitica or treated with SA.64,65
Senescence-associated genes (SAGs) are induced by biotic and abiotic stressors.66,67 Here, SAGs playing roles in oxidative stress tolerance, wounding and necrosis were found to be induced by LPS, lipid A and OPS: At4g35770 (L), At4g02380 (A) and At3g10985 (O), respectively.
At4g37930 (L, A, O), a ‘serine hydroxymethyltransferase 1’ (Fig. 4), is involved in controlling cell damage caused by abiotic stressors and the HR in plants, and was reported to influence resistance in A. thaliana towards biotic and abiotic stresses. 68 Another stress-responsive gene At1g72610 (L, O), a ‘germin-like protein 1’, plays a significant role during various stresses, including pathogen elicitation. 69 At3g53280 (L, A, O) encodes a cytochrome P450 71B5 monooxygenase, involved in stress metabolism and also responsive towards flg22, elf26, hrpZ, and chitin elicitation (Genevestigator).
Peroxidases and PR-proteins
Three peroxidases were found to be induced: At3g49120 (L, A), At5g64120 (L) and At4g21960 (A, O). Their involvement during stress responses is well documented, and some are regarded as pathogenesis-related (PR) proteins. 70 An endochitinase, At1g05850 (O), was responsive to the OPS and At2g05790 (A, O), a β-glucanase, to both lipid A and OPS. A third PR protein gene, At4g12470 (L, A, O), a lipid transfer protein (LTP), was found to be induced by all three agents (Figure 4), as well as P. syringae, flg22, hrpZ, ethylene and wounding (Genevestigator). This gene, azi-1, was reported to be involved in priming of SA induction and systemic immunity triggered by pathogens or azelaic acid 71 and to confer disease resistance when overexpressed. 72 A related LTP (At5g48485, dir1-1, defective in induced resistance 1-1), was reported to be involved in systemic resistance through interaction with lipid-derived molecules to promote long-distance signaling. 73
Cellular transporters
A transport-related gene, At2g26900 (A), encodes a ‘bile acid: sodium symporter family protein’, associated with several stress responses. 74 Bile acids were previously reported to elicit both HR cell death, PR proteins, and phytoalexin accumulation in rice. 75 At1g11260 (L, A, O), a gene encoding a H+/hexose transporter, has been identified as a target for early elicitor signaling in A. thaliana 76 and is also responsive to flg22, hrpZ, and induced programmed cell death (PCD) (Genevestigator).
Ubiquitin and the proteasome
The proteasome facilitates the degradation of ubiquitinated proteins, the functioning of which is connected to different processes allowing plant cells to respond to environmental cues, including disease resistance and the regulation of plant defense. 77 – 82 The inhibition of ubiquitin-26S proteasome system suppresses the elicitation of defense 81 and ubiquitin-ligase genes were induced by a plant defense elicitor, chitin. 83 The ubiquitination and proteasomal degradation of repressor proteins controlling the expression of the plant defense response as well as increased turnover of R proteins was observed.83,84
The LPS elicited At1g17280 (L) and At1g12760 (L) that encode for a ‘ubiquitin-conjugating enzyme 34’ and ‘ubiquitin-protein ligase’ respectively; while At3g12400 (L) and At5g12120 (L) encodes ‘ATELC/ELC, ubiquitin binding’ and ‘ubiquitin-associated/ubiquitin and TS-N domain-containing protein’ respectively. The OPS elicited At4g04690 (O), At5g20570 (O) and At4g38630 (O), that encode a ‘F-box protein 15’, a ‘RING-BOX 1’ and ‘regulatory particle non-ATPase (multiubiquitin chain binding protein), respectively. In the case of lipid A, it induced At1g21780 (A), encoding a BTB domain-containing protein reported to associate with cullin proteins to form ubiquitin protein ligases: 85 At1g75990 (A) encodes a ‘26S proteasome regulatory subunit S3’ (RPN3) and At3g59940 (A) a ‘Kelch repeat-containing F-box family protein’. The F-box containing proteins are also involved in pathogen defense responses.78,86
The proteasome complex proteins were also found to be responsive towards LPSB.cep. treatment on a proteomic level. 21
Cell wall modification
Cell wall remodelling and re-inforcement are critical plant defenses and all three elicitors had an effect on this response. These genes include: At1g14890 (L, O), encoding a ‘pectinesterase inhibitor’; At2g45220 (A) and At3g49220 (O) encoding ‘pectinesterases’; and At5g49720 (O) encoding ‘glycosyl hydrolase 9A1’. Genes related to this category whose expression was increased by lipid A also include At3g16520 (A), an ‘UDP-glycosyl transferase family protein’ and At4g34230 (O), a ‘cinnamyl alcohol dehydrogenase 5’ protein involved in synthesis of lignin precursors.
Cytoskeletal re-arrangements
The expression of non-host resistance is associated with the dynamic re-organization of microfilaments and microtubules, 87 and genes encoding proteins involved in cytoskeletal re-arrangements were induced by all three agents. At5g19780 (L), encodes ‘tubulin alpha-5’ and At1g49240 (L) an ‘actin 8’. At4g34970 (O) codes for an ‘actin binding protein’ and At3g18780 (A) an ‘actin 2’ protein. Actin and tubulin were previously reported as targets for LPS-responsive post-translational modification in tobacco cells by us. 21
The cytoskeleton and cell wall–plasma membrane connectivity have been identified as important responsive elements of non-host resistance. 88 In this context, it is of interest that At3g22120 (A) encoding a ‘cell wall–plasma membrane linker protein’ was up-regulated by lipid A.
Metabolism and energy production
Several genes which have a role in metabolism have been found to be up-regulated by all three elicitors. During plant–pathogen interactions, the normal metabolic flow in the host plant is altered in support of defense responses but also, potentially, as means whereby pathogens channel the nutrients for their own benefit in order to survive. 89 Phospholipase A 2A, At2g26560 (A, O), that accumulates upon infection by fungal and bacterial pathogens is an example of this category as well as At3g04120 (L), glyceraldehyde-3-phosphate dehydrogenase C, that is responsive towards a signaling cascade induced by ROS. The sterol C24 methyltransferases At1g20330 (L) and At5g13710 (O) encode enzymes that control carbon flux into sterol synthesis, potentially influencing membrane fluidity, permeability and microdomains. Interestingly, the application of β-sitosterol to A. thaliana leaves results in enhanced resistance to P. syringae. 90 The importance of members of this category should not be overlooked, as several of the 14 Nicotiana benthamiana genes identified that, when silenced, compromised PTI, were grouped in this category. 91
Lastly, genes grouped to the categories ‘mitochondria and chloroplast function’, are listed in Supplementary Table S2.
Concluding discussion
Previous micro-array data with LPSB.cep. (TAIR accession expression set 100808727), although obtained under different experimental conditions and plant development stages, are consistent with the data obtained in this study. 23 The results indicate that perception of the LPS-derived MAMPs activate signal transduction cascades that result in reprogramming of the transcriptome. This lead to derepression or induction of defense responses, metabolic reprogramming, redeployment of cellular energy and changes in cellular activities supporting protein synthesis, folding, vesicle trafficking and secretion, accompanied by changes to the cytoskeleton. Many of the corresponding proteins are known to be interconnected at various levels through an intricate web of activation/de-activation, protein–protein interactions, complex formation and chaperoning reactions. Several lines of evidence suggest that MAMPs stimulate defense pathways that are independent of the signal molecules, SA, JA and ethylene. 44 The results obtained here seem to support this observation as few SA- or JA-responsive genes were identified. The broad classes of genes identified were strikingly similar to those reported to be increased in tomato responding to P. syringae pv. tomato.46,91
Different MAMPs induce overlapping responses to bacteria within leaf tissue 45 and current models of MAMP perception indicate that different MAMPs can trigger specific sets of both individual as well as generic cellular responses with possible cross-talk of the signaling pathways. Moreover, different MAMP perception systems might rely on different classes of receptor molecules,2,24 each possibly initiating a specific subset of downstream responses. 92
As MAMPs, LPSs are structurally complex molecules, comprising different moieties with distinct chemical patterns and both the lipid A and the OPS from LPSB.cep. were found to induce genes with known functions in plant defense responses. The results can be interpreted as evidence to support the concept that lipid A and the OPS are MAMP-active structures in their own right. Similar genes were found to be induced by all three elicitors and this may be due to the fact that the responses triggered by these elicitors converge later at the level of signal molecules or transcription factors to form a common response which is aimed at activating defense. However, lipid A and the OPS also induced genes not found/different to those induced by the intact LPS molecules, suggesting that different perception and signal transduction mechanisms might be operative.
Whilst the lipid A and OPS moieties were able to up-regulate genes over the same spectrum of categories as intact LPS, the up-regulation of defense-associated genes observed with intact LPS was the more comprehensive; suggesting that the lipid A and glycan moieties of the molecule act as partial agonists, but that the intact LPS structure is required for full agonist activity. Conjugation to the long hydrophilic OPS glycan may assist in maintaining hydrophobic lipid A in a form in which it can be presented to the plant membrane. Lipid A has the ability to intercalate into phospholipid cell membranes, 93 thereby allowing the free carbohydrate-based OPS to bind to putative lectin-like or carbohydrate-recognising signaling receptors, 24 and contributing to the defence response. However, it is also possible that the carbohydrate and lipid molecular signatures are perceived by separate, but related or interconnected mechanisms. Indeed, results from animal systems indicate that delipidated LPS appears to have its own binding site in the LPS recognition pathway 94 and that the lipid A fraction could induce discrete MAPK activation different from that due to LPS. 95
The presented data offer novel insights and further evidence for the biochemical action of macromolecular LPSs as lipoglycan elicitors of PTI. It can be concluded that the LPS-derived lipid A and OPS from B. cepacia were perceived by the seedlings, leading to the induction of a range of genes; some known to be associated with defense responses as well as biotic and abiotic stresses, whilst others are associated with metabolic reprogramming of cellular activities in support of immunity and defense.
Supplementary data
Table S1.1–1.4. Induced genes of A. thaliana found to be co-regulated in response to LPS, Lipid A and the OPS, 8 h after treatment of 10 d old seedlings.
Table S2. List of ribosomal, mitochondrial and chloroplast genes from A. thaliana found to be up-regulated in response to elicitation by LPS, OPS and Lipid A, 8 h after treatment of 10 d old seedlings.
Footnotes
Acknowledgements
The South African National Research Foundation (NRF) is acknowledged for research funding (Grant number 206492) and the authors thank Dr Lizelle Piater for critical reading of the manuscript.
