Abstract
Colonization of the gastric mucosa by Helicobacter pylori can lead to peptic ulcer and gastric adenocarcinoma. TLRs are signaling receptors involved in the recognition of microorganisms, and polymorphisms in their genes may influence the innate and adaptive immune response to H. pylori, affecting the clinical outcomes of the infection. We assessed the association between single nucleotide polymorphisms in TLR9 and TLR5 and gastroduodenal diseases. All patients were genotyped by allelic discrimination in regions 1174C > T and 1775A > G of TLR5 and −1237T > C and 2848G > A of TLR9. The 2848A allele of TLR9 was more frequent in duodenal ulcer and showed an association of risk with this pathology. Polymorphisms in TLR5 were not found to be associated with disease. Patients with polymorphisms in TLR9 and TLR5 expressed significantly lower levels of IL-1β and TNF-α, whereas polymorphisms in TLR5 also decreased the expression of IL-6 and IL-10. Our findings suggest that 2848G > A polymorphism in TLR9 increases the risk for the development of duodenal ulcer probably by modifying the inflammatory response to H. pylori infection. This is the first study to show an association of 2848A allele of TLR9 with duodenal ulcer and with altered expression of inflammatory cytokines in the gastric mucosa.
Introduction
Helicobacter pylori is a Gram-negative bacterium that infects more than half of the world’s population and is the major cause of gastric and duodenal ulcers, gastric carcinoma and mucosal-associated lymphoid tissue lymphoma.1,2
Microbial, host and environmental factors have been associated with the diverse clinical outcomes associated with H. pylori infection. 3 It has been shown that polymorphisms in genes of the inflammatory response, such as the pro-inflammatory cytokines IL-1β and TNF-α, increase the risk for gastric cancer in some populations but not in others. These results show the importance of the immune response in the development of the pathologies associated with this infection and the need to further study the influence of the immune response and its genetic factors in association with H. pylori infection. The number of human genes involved in the immune response is very large and keeps increasing as we learn more about the process.
The recognition of microorganisms colonizing human epithelial mucosa, such a H. pylori, is mediated by a set of receptors referred to as pattern recognition receptors (PRRs), 4 critical in the innate immunity, which recognize conserved molecular patterns (PAMPs) shared by most microorganisms. 4 An important family of PRRs is the TLRs, trans-membrane receptors that recognize a wide range of ligands present in microorganisms. TLRs play an essential role in innate immunity, being involved in the regulation of inflammatory responses and the activation of the adaptive immune system.4,5
There is strong evidence that gastric epithelial cells express TLR5 and TLR9, both in patients with non-inflamed gastric mucosa and in patients with H. pylori infection.6,7 In fact, in H. pylori gastritis, TLR5 and TLR9 expression on the gastric epithelium changes to a basolateral localization without detectable expression at the apical pole, and TLR5 and TLR9 polarization seems to be a process dynamically influenced by H. pylori infection. 7 In addition, flagellin from H. pylori-activated NF-κB in cells transfected with TLR5, and expression of IL-8, macrophage inflammatory protein-3α and growth regulated oncogene-α, was induced after H. pylori was recognized by TLR5, 8 probably through p38 activation and phosphorylation of the transcription factor activating transcription factor 2. 9 However, TLR9 is responsible for initiating an inflammatory response to bacterial CpG DNA, and H. pylori has been shown to induce the expression of TLR9 on gastric epithelial cells and to activate MAPKs, resulting in cyclooxygenase-2-dependent release of prostaglandin E2, which, in turn, contributes to cell invasion and angiogenesis.6,10,11 Thus, it is well documented that TLR5 and TLR9 play an important role establishing an inflammatory response to H. pylori infection.
Polymorphisms in TLR genes have been associated with increased risk for infections and inflammatory diseases, and, in conjunction with exposure to environmental risk factors, can promote the development of malignant disease, 12 although the functional consequences of these genetic variations are still to be uncovered. 13 The effect of polymorphisms in TLRs on disease susceptibility during H. pylori infection has been poorly studied. In TLR4, the single nucleotide polymorphisms (SNPs) D299G and T399I have been associated with severe H. pylori-associated disease,13,14 and with a modified pattern of inflammatory cytokines and chemokines in the gastric mucosa. 14 Recently, Zeng et al. reported that TLR5 rs5744174 (2487C allele) carriers had a higher risk of developing gastric cancer. 15 The TLR9 −1237C allele is significantly associated with the development of H. pylori-induced premalignant gastric changes. 16 These data show that genetic modifications in TLRs may lead to an altered inflammatory response to H. pylori and a modified mucosal cytokine secretion, thus modifying the risk for the development of associated diseases. However, to date, there are still few reports on polymorphisms in TLRs and H. pylori-associated disease.
Accordingly, the aim of this study was to determine whether polymorphisms in TLR5 and TLR9 are associated with H. pylori-induced duodenal ulcer or gastric cancer, and to analyze the effect of these polymorphisms on the expression of inflammatory cytokines in the gastric mucosa of patients.
Materials and methods
Study design and recruitment of patients
This case–control study included 561 Mexican patients from the Hospital General de Mexico, Secretaría de Salud (SS), and the Hospital de Oncología, Instituto Mexicano del Seguro Social (IMSS), Mexico City. Patients attended the gastroenterology service for diverse gastroduodenal symptoms. Patients aged <18 yr, with any autoimmune disease, diabetes or cancer, other than gastric cancer, were excluded from the study. Informed consent was obtained from all patients prior to enrollment in the study. The ethics committees of the two participating institutions, IMSS and SS, approved the study.
Diagnosis
All patients were endoscoped as part of the diagnostic protocol, and gastric biopsies were taken for pathology and H. pylori culture. Diagnoses of gastritis, peptic ulcer, precancerous lesions or gastric cancer were based on endoscopy and histopathology findings. Diagnosis of H. pylori infection was done by serology (total H. pylori extract and CagA protein) with an ELISA, as previously described. 17
Specimen collection
Biopsies were obtained from both lesser and greater curvatures of the antrum and corpus. In order to increase sensitivity in the detection of precancerous lesions and H. pylori infection, seven biopsies, four from the antrum and three from the corpus, were processed for histology. One biopsy from the antrum and another from the corpus were immediately placed in liquid nitrogen and stored at –80℃ until processed for determination of cytokines and chemokines.
Preparation of genomic DNA
A 10-ml sample of blood was drawn from each patient and mononuclear cells were purified by centrifugation in a Ficoll-Hypaque density gradient. DNA was isolated from these cells using the salting out microtechnique and stored at −20℃ until tested. 18
Detection of SNPs in TLR genes
Primers and probes used to analyze SNPs in TLR5 and TLR9 in patients with gastroduodenal diseases.

MGB: minor groove binder at 3′end.
Quantitative RT-PCR assays for cytokines and chemokines in gastric tissue
For the study of the levels of cytokines and chemokines in gastric lesions, we selected a sample of 24 biopsies from patients with the −1237 SNP and 24 from patients with the 2848 SNP in TLR9, six of each pathology: non-atrophic gastritis, intestinal metaplasia, gastric cancer and duodenal ulcer. In addition, biopsies from all patients with the 1174T allele in TLR5 (15 patients) were also studied. As controls, we selected the same number of biopsies from patients with the ancestral allele, matched by sex, age and pathology. Total RNA was extracted from gastric biopsies using a RNA extraction kit (RNAqueous; Ambion, Austin, TX, USA). After DNase treatment, 1 µg total RNA was subjected to reverse transcription with 200 U of Super Script II reverse transcriptase (Invitrogen, Carlsbad, CA, USA). For quantitative PCR, we normalized the cytokine mRNA expression levels by using cDNA of the housekeeping gene, β-actin. The quantification of cytokine and β-actin mRNA levels was performed using a Light Cycler 480 real time PCR system (Roche, Indianapolis, IN, USA). Specific primers and fluorescent hybridization probes [Universal Probe Library (UPL); Roche, Basel, Switzerland] were used for IL-1β, IL-6, IL-10 and TNF-α expression. Reaction mixtures for PCR (10 µl) were prepared by mixing 1 µg synthesized cDNA solution with 1 × Light Cycler TaqMan Master (Roche, Indianapolis, IN, USA), 1 nM of each primer and 0.5 nM UPL probes. PCR was carried out at 50℃ for 2 min and 95℃ for 10 min, followed by 50 cycles of 95℃ for 15s and 60℃ for 60s, according to the manufacturer’s instructions. The expression levels of cytokine mRNA were expressed as the ratio of cytokine mRNA to β-actin mRNA. Each assay was performed in duplicate, and each cytokine assay was performed testing all RNA samples in the same experiment.
Statistical analysis
Data were analyzed using the Hardy–Weinberg distribution for all alleles. Frequency of polymorphism among the different groups was analyzed using the Fisher test. The association between the polymorphisms and disease was analyzed with the odds ratio (OR) test; results are presented as ORs with 95% confidence intervals (CIs). Cytokine expression is presented as mean, and differences between polymorphic and no polymorphic groups were analyzed using the Mann–Whitney test; a P-value of 0.05 was considered statistically significant. Data were analyzed using the statistical software Stata 9.0 (Stata Corp., College Station, TX, USA).
Results
Diagnosis and characteristics of the patients with gastroduodenal diseases studied for SNPs in TLR5 and TLR9.

Infection determined by serology.
Frequency of SNPs in TLR5 and TLR9 and their association with disease
Owing to the reported role of TLR5 and TLR9 in the initiation of the innate immune response to H. pylori infection, we studied polymorphisms 1174C > T (R392*) and 1775A > G (N592S) in TLR5, and −1237T > C and 2848G > A in TLR9. The allelic frequencies for the different polymorphisms in all studied samples were as follows: for TLR5, 6.1% for the 1174T allele and 0.76% for the 1775G allele; for TLR9, 16.1% for the −1237C allele and 49.7% for the 2848A allele (all were in Hardy–Weinberg equilibrium).
For TLR5, carriage of the 1174T allele tended to be more frequent among patients with cancer (7.89%) than in patients with gastritis (6.61%), with metaplasia (6.67%) or patients with duodenal ulcer (2.38%), although differences were not statistically significant (Supplementary Table 1). The SNP in 1775A > G was present in only two of the patients studied.
The −1237C allele of TLR9 tended to be more frequent among patients with metaplasia (19.35%) than in patients with gastritis (15.63%), with cancer (15.93%) or patients with duodenal ulcer (12.82%), although differences were not statistically significant (Supplementary Table 2).
Frequency of TLR9 2848G > A genotype in patients with gastroduodenal diseases and H. pylori infection.
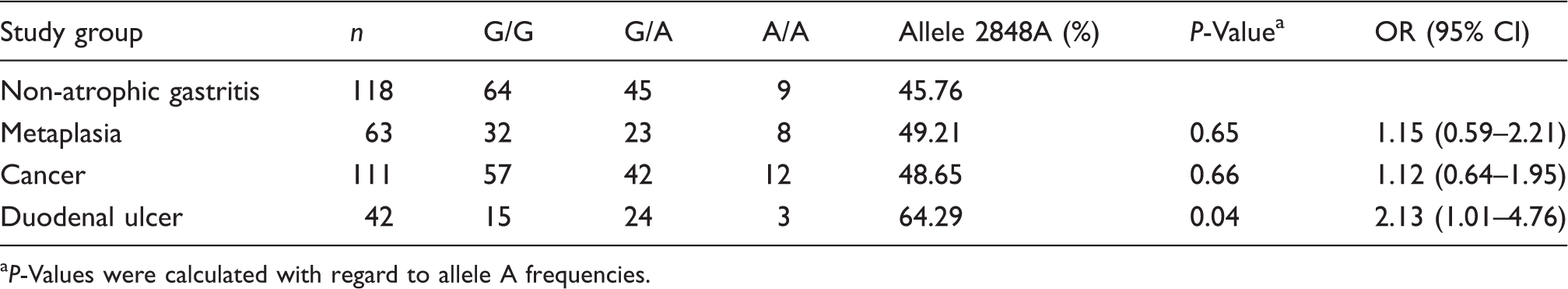
P-Values were calculated with regard to allele A frequencies.
The association of polymorphisms in TLR5 and TLR9 with the pattern of cytokines expression in the gastric mucosa
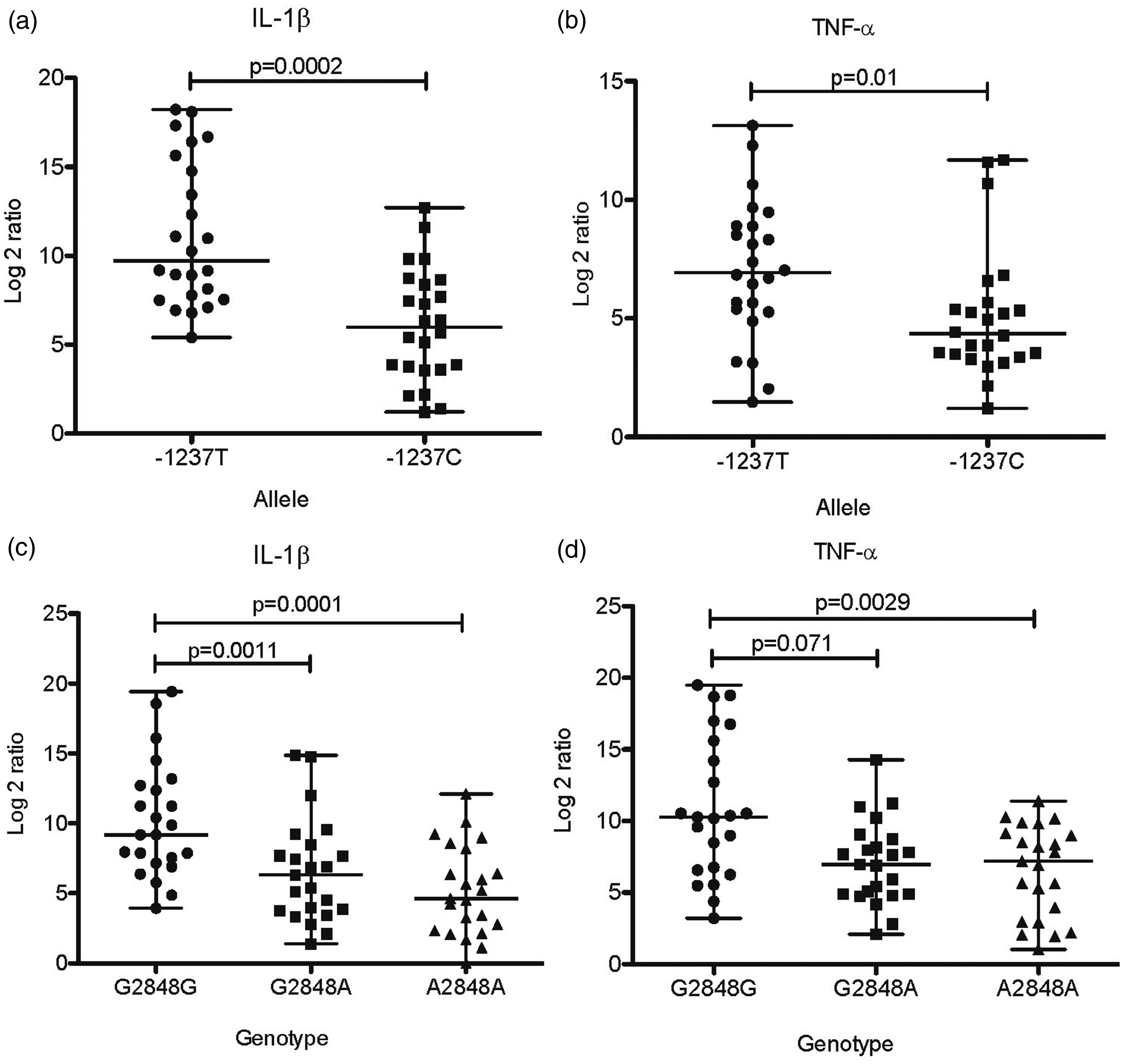
As polymorphisms in the TLR genes may affect their function, we hypothesized they might alter the expression of cytokines in the gastric mucosa. Thus, we studied whether the presence of variants of TLR5 and TLR9 alleles modified the expression of cytokines in the H. pylori-infected gastric mucosa. As shown in Figure 1, the expression of TNF-α (P = 0.001), IL-1β (P = 0.01), IL-6 (P = 0.005) and IL-10 (P = 0.04) was significantly lower in patients with the 1174T variant allele of TLR5. For variants in TLR9, we found that expression of IL-1β (P = 0.002; Figure 2A) and TNF-α (P = 0.01; Figure 2B) was significantly lower in patients with the −1237C variant allele than the group with the −1237T ancestral allele. In the case of the 2848G > A SNP, we observed a significant gene–dose effect with a reduction in the expression of both IL-1β (P = 0.0011; Figure 2C), and TNF-α (P = 0.071; Figure 2D) from 2848G/G to 2848G/A, to 2848A/A (P < 0.0001 and P = 0.0014, respectively). Finally, in contrast to what we observed with TLR5 SNPs, none of the SNPs in TLR9 were associated with a modified expression of IL-6 or IL-10 (Supplemental Figure 1).
Expression of cytokines in biopsies of H. pylori infected patients with C1174T SNPs in TLR5 gene. As controls, we selected the same number of biopsies from patients with the ancestral allele, matched by gender, age, and pathology. Reverse transcription and quantitative PCR were performed to estimate the relative expression levels of each cytokine. Expression of TNF-α (a), IL-1β (b), IL-6 (c) or IL-10 (d) was normalized with expression values of human β-actin. Difference in mRNA expression between patients with SNPs in TLR5 and patients with the ancestral allele were significant in all cases. A p value <0.05 was considered statistically significant using the U of Mann–Whitney test. Expression of cytokines in biopsies of H. pylori infected patients with T-1237C and G2848A SNPs in TLR9 gene. As controls, we selected the same number of biopsies from patients with the ancestral allele, matched by gender, age, and pathology. Reverse transcription and quantitative PCR were performed to estimate the relative expression levels of each cytokine. Expression of IL-1β (a) (c), TNF-α (b) (d) was normalized with expression values of human β-actin. Difference in mRNA expression between patients with SNPs in TLR9 and patients with the ancestral allele were significant in all cases. A p value <0.05 was considered statistically significant using the U of Mann–Whitney test. In 2848G>A SNP, there was significant gene–dose effect with a reduction in the expression of both IL-1β (c), and TNF-α (d) from 2848G/G to 2848G/A (p = 0.0011 and p = 0.071 respectively), to 2848A/A (p<0.0001 and p = 0.0014 respectively).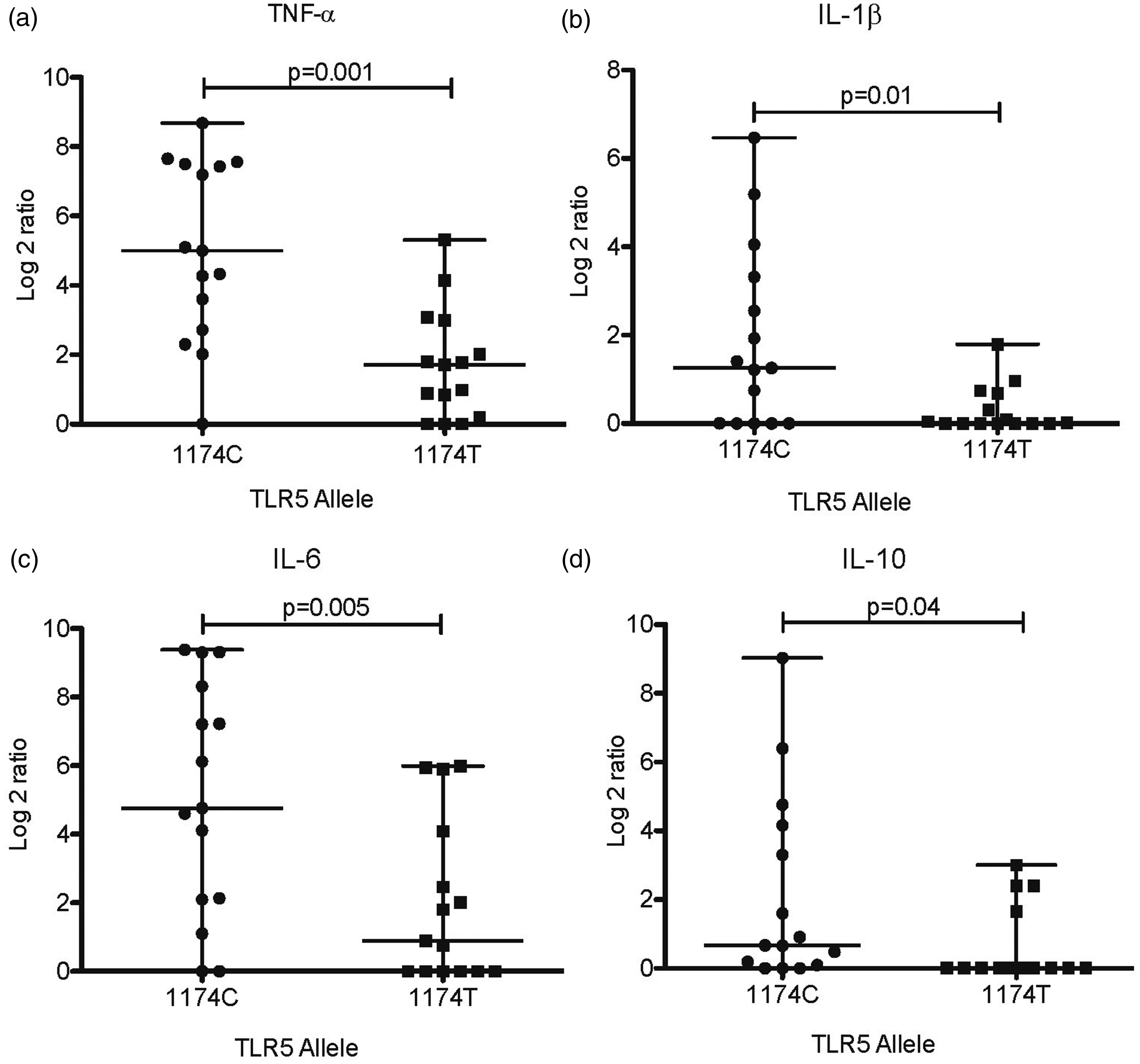
Discussion
It has been suggested that functional polymorphisms in genes linked to the immune response are important factors for the development of gastroduodenal diseases associated with H. pylori and are thought to contribute to differences in the long-term outcomes of H. pylori infection. TLRs are involved in the recognition of H. pylori and might play an essential role in the regulation of inflammatory reactions and in the activation of the adaptive immune response; thus, they may play a role in the pathogenesis of gastroduodenal diseases. However, there are only few and controversial studies that address the possible role of polymorphisms in TLR genes as risk factors for H. pylori-associated diseases, and they did not assess the effect of these polymorphisms in the expression of inflammatory mediators in the gastric mucosa of patients. Accordingly, we chose to study the possible association of polymorphisms in TLR5 and TLR9 with disease and with the pattern of expression of cytokines in the gastric mucosa of patients.
Concerning polymorphisms in TLR5, we did not observe any association between the 1174C > T SNP and gastroduodenal diseases, whereas one study in a Chinese population found a significant association between the TLR5 SNP rs5744174 (2487C allele) and risk for gastric cancer (GC), and we plan to study whether this SNP is also associated with gastric cancer (GC) in our population. One study in Brazilian children reported no association of the 1174C > T SNP with the development of duodenal ulcer, 19 which would agree with our findings in adults. A recent study in white people found that the TLR5 1775A > G (N592S) polymorphism was associated with decreased colorectal cancer survival; 20 however, this polymorphism was almost absent (0.76%) in the studied Mexican population, which included >561 patients, and suggest that the N592S polymorphism has low relevance for disease in our mestizo population. Although studies have demonstrated the importance of TLR5 and TLR9 for H. pylori recognition, their role in the evolution of gastric lesions is still unknown, and the importance of genetic variations in these receptors is poorly understood. Our results suggest that polymorphisms in both TLR5 and TLR9 might play no role in the development of gastric cancer, at least in our mestizo population, as has been reported for TLR9 polymorphisms in white populations. 21
In contrast, we found that the TLR9 2848G > A SNP was significantly associated with duodenal ulcer, which might be related to the observation that TLRs are important for healing of the injured intestinal epithelium and for the balance between epithelial proliferation and apoptosis.22,23 In silico analysis have suggested that the −1237C SNP in TLR9 creates a motif with increased binding affinity to NF-κB, which might be functionally relevant, leading to an increased risk for H. pylori-associated gastric diseases. 16 It might also be possible that the observed modifications in TLR5 and TLR9 may alter the capacity of the stomach epithelia to counteract H. pylori infection and predispose to the more severe forms of the gastroduodenal diseases associated with this bacterium. Deficient signaling after H. pylori recognition via TLR5 or TLR9 may reduce the immune responsiveness during the acute phase of infection, and may favor persistent infection.
We next examined whether the mucosal expression of cytokines and chemokines was affected in patients carrying the variants alleles in TLR5 and TLR9 compared with that in patients carrying the ancestral allele. We found that the expression of the inflammatory cytokines TNF-α, IL-1β, IL-6 and IL-10 was decreased in patients with the 1174T allele of TLR5, which makes sense if we consider that this SNP causes the insertion of an early stop codon in the ligand binding domain that seems to abrogate the signaling cytoplasmatic tail. 24 Our work also showed that both studied polymorphisms in TLR9 resulted in an impaired expression of the pro-inflammatory cytokines TNF-α and IL-1β but not the expression of IL-6 or IL-10. We suggest that the modification in the sequence of the extracellular domain of TLR9 with SNPs −1237C and 2848A probably modify the recognition of their ligands and the signaling pathways activated to induce the production of cytokines. 25 While the expression of IL-6 and IL-10 might be activated by a different pathway, it is not affected by the SNPs studied.26,27 An inappropriate inflammatory response and a lack of control over the infection may increase the risk to develop severe disease; in fact, it has been shown that MyD88-deficient mice (signaling molecule of TLRs) have an increased colonization with H. pylori compared with wild type mice, and generated an early and rapid advancement to gastric dysplasia post-infection.28,29 The TLR9 2848A allele was significantly associated with duodenal ulcer and to a reduced expression of IL-1β; IL-1β is a strong inhibitor of acid production and its impaired production would result in increased acid secretion and increased risk for duodenal ulcer. Thus, the risk for duodenal ulcer would be expected to be similar for SNPs 2848A, −1237C and 1174T. We think that this is a reflection of differences in frequency of the SNPs in the population; whereas the frequency of the TLR9 2848A allele was 64.2%, the frequency of −1237C was 12.82% and the frequency of TLR5 1174T was only 2.38%.
In conclusion, in the present study we found that in our population TLR5 1174C > T, 1775A > G and TLR9 −1237T > C polymorphisms were not found associated with gastroduodenal diseases. However, we found that the TLR9 2848G allele increases the risk of developing duodenal ulcer in H. pylori-infected patients twofold, showing for the first time the association between the 2848G SNP and duodenal ulcer. We also found that individuals with SNPs in TLR5 and TLR9 had decreased mucosal expression of inflammatory cytokines, which may impair the ability of the patient to counteract the infection and lead to an increased risk for disease. Together, these results suggest that TLR9 plays an important role in the outcome of H. pylori infection.
Footnotes
Funding
This work was supported by CONACYT-FOSISS, MEXICO, Grant SALUD-2007-01-69450, and by Instituto de Ciencia y Tecnología del Distrito Federal, Grant ICYTDF/234/2012 (PICSA 12-209). J. Torres is a recipient of an Exclusivity Scholarship from Fundación IMSS, Mexico. N. Sánchez-Zauco is a fellow of CONACYT Mexico for doctoral studies in science 2010 to 2014. N. Sánchez-Zauco is an ENCB-IPN doctoral student.
Conflict of interest
The authors do not have any potential conflicts of interest to declare.
Acknowledgements
We thank Dr. Julio Granados Arriola, División de Inmunogenética, Departamento de Trasplantes, Instituto Nacional de Ciencias Médicas y Nutrición Salvador Zubirán, for his critical comments.
