Abstract
Gestational diabetes mellitus is a growing public health concern due to its large disease burden; however, the underlying pathophysiology remains unclear. Therefore, we examined the relationship between 107 single-nucleotide polymorphisms in insulin signalling pathway genes and gestational diabetes mellitus risk using a nested case-control study. The SOS1 rs7598922 GA and AA genotype were statistically significantly associated with reduced gestational diabetes mellitus risk (ptrend = 0.0006) compared with GG genotype. At the gene level, SOS1 was statistically significantly associated with gestational diabetes mellitus risk after adjusting for multiple comparisons. Moreover, AGGA and GGGG haplotypes in SOS1 gene were associated with reduced risk of gestational diabetes mellitus. Our study provides evidence for an association between the SOS1 gene and risk of gestational diabetes mellitus; however, its role in the pathogenesis of gestational diabetes mellitus will need to be verified by further studies.
Introduction
Gestational diabetes mellitus (GDM), defined as glucose intolerance of various degrees that is first detected during pregnancy, is a common pregnancy complication that has been associated with adverse health outcomes for mothers and infants. 1 GDM affects approximately 5%–17% of all pregnancies worldwide, and the prevalence has increased upwards of 10%–100% during the past 20 years,2,3 leading to its emergence as a major public health concern. 4
Recent evidence suggests that insulin signalling pathways, which play a key role in energy metabolism, growth and cellular differentiation, are involved in the development of GDM. 5 The relationship between GDM risk and SNPs of insulin signalling pathway genes, including IRS1, INSR, ENPP1 and PIK3R1, has been evaluated in recent studies.6–8 Insulin signalling genes including GRB2, SOS, SHC, PRKAA1, PRKAR2B, PYGM, INPPL1, ACACA and PHKA1 may be involved in the activation of mitogen-activated protein kinase (MAPK), the phosphorylation for insulin receptor substrate (IRS) and phosphatidylinositol-3-kinase (PI3K), and then affect lipid and glucose homeostasis. In order to further clarify the potential role of SNPs in the insulin signalling pathway genes, we examined 107 SNPs in these 9 genes and the risk of GDM in a Chinese Han population.
Materials and methods
Subjects
The subjects were enrolled from an ongoing birth cohort at the First Affiliated Hospital of Shanxi Medical University between 1 March 2012 and 30 July 2014 in Taiyuan, China. The study included pregnant women waiting for delivery in the hospital whose gestational ages were 20 weeks or more, had no mental illnesses and were aged 18 years or older. All study procedures were approved by the Institutional Review Boards at the Shanxi Medical University. Written consent was obtained from eligible women prior to enrolment. Information on demographic factors, reproductive and medical history, smoking and alcohol consumption was collected using standardized questionnaires by trained interviewers.
Cases and controls selection
GDM was diagnosed using a 75-g oral glucose tolerance test (OGTT) during weeks 24–28 of pregnancy. Subjects who met at least one of the following criteria were diagnosed as having GDM: (1) fasting blood glucose was more than 5.1 mmol/L, (2) 1-h blood glucose >10.0 mmol/L and (3) 2-h blood glucose >8.5 mmol/L. A total of 334 subjects were identified as having GDM, and 334 control subjects without GDM were randomly selected through frequency matching to cases by age, month of conception and residence. In all, 13 cases and 18 controls were excluded due to missing genotyping per person or per SNP. Finally, 321 cases and 316 controls were included in the analysis.
Genotyping
Illumina Golden Gate Platform was used for genotyping. A total of 107 SNPs from the insulin signalling pathway genes including SOS1, PRKAA1, PRKAR2B, PYGM, INPPL1, SHC4, ACACA, GRB2 and PHKA1 were included in the study. The completion rate for all SNPs was over 99%. Chi-square test was used to test the Hardy–Weinberg equilibrium (HWE) for each SNP. Five SNPs were excluded from the final analysis due to violation of HWE (p < 0.05; as shown in Supplementary Table 1).
Statistical analysis
The odds ratios (ORs) and their 95% confidence interval (CI) were estimated by unconditional logistical regression as a measure of the associations between genotypes and GDM adjusting for maternal age, maternal body mass index (BMI), parity and family history of diabetes. The common homozygous genotypes served as reference. A linear trend test was conducted by assigning the ordinal values 1, 2 and 3 to the homozygous common, heterozygous and homozygous rare alleles, respectively. Minimum P (minP) tests were conducted to examine the association at the gene level using ‘minPtest’ R packages. Haplotype block structures were evaluated with HaploView version 4 using the method of Four Gamete Rule. Multiple comparisons were adjusted using the false discovery rate (FDR) method at a significant level of 0.2.
Results
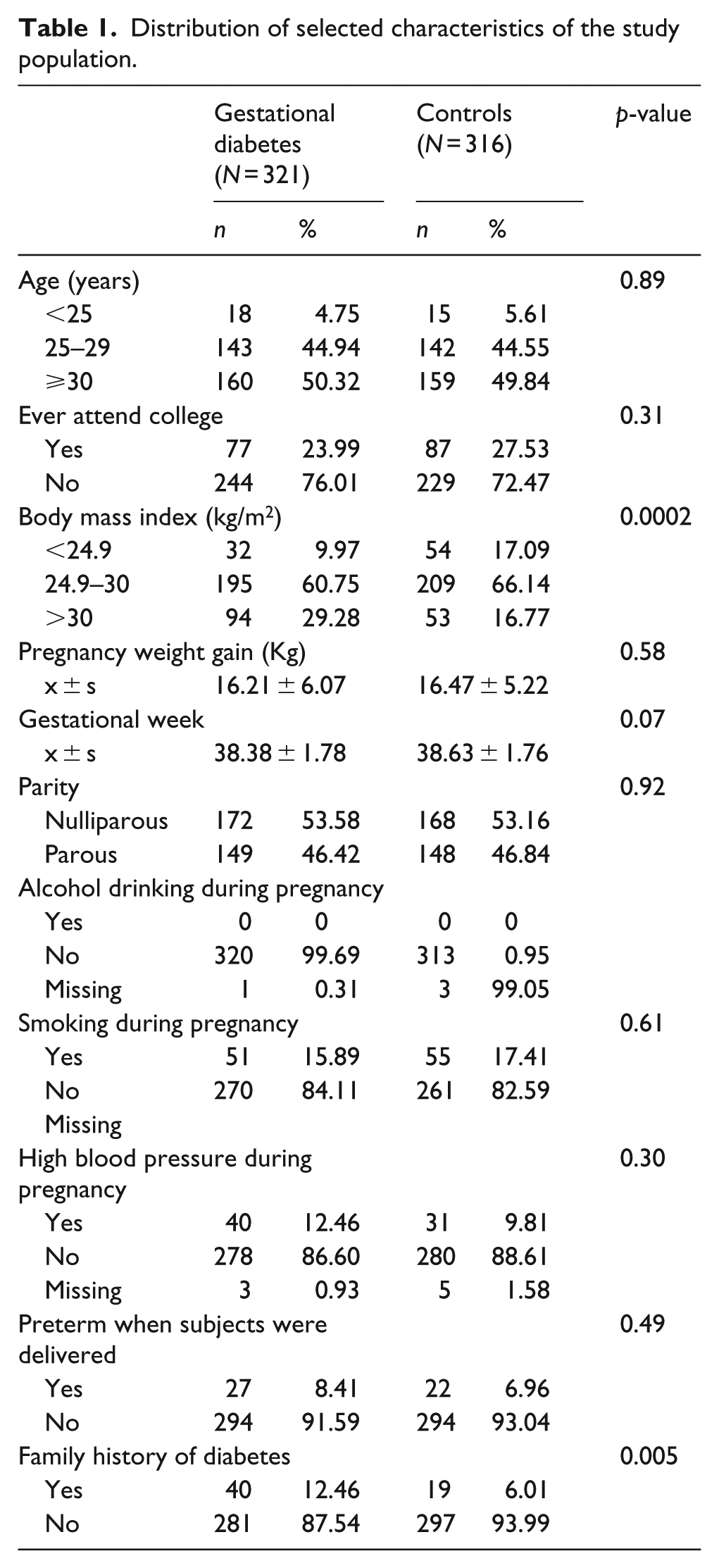
As shown in Table 1, there were more GDM cases with a higher BMI than control subjects (p = 0.0002). Overall, 12.5% of GDM cases and 6% of controls reported a family history of diabetes (p = 0.005). None of the cases or controls consumed alcohol during pregnancy. The distributions of maternal age, weight gain during pregnancy, parity, high blood pressure during pregnancy, smoking and gestational weeks between cases and controls were similar.
Distribution of selected characteristics of the study population.
As shown in Supplementary Table 2, reduced risk of GDM was observed in women who carried the SOS1 rs7598922 GA genotype (OR = 0.62, 95% CI = 0.43–0.88) or AA genotype (OR = 0.28, 95% CI = 0.09–0.73, ptrend = 0.0006), the rs3099968 AG genotype (OR = 0.69, 95% CI = 0.47–0.99, ptrend = 0.025) and the rs1010859 GG genotype (OR = 0.12, 95% CI = 0.01–0.66) compared with the homozygous common in each SNP. It was also found in women who carried the PHKA1 rs969927 AG genotype (OR = 0.58, 95% CI = 0.37–0.89) and AG or GG genotype (OR = 0.57, 95% CI = 0.37–0.87) compared with AA genotype (ptrend = 0.009).
Increased GDM risk was found in women who carried the PRKAA1 rs1002424 AA genotype, the rs10053664 AG genotype and the rs13361707 GG genotype, the PRKAR2B rs10276321 AA genotype, the rs4727681 GG genotype, the rs2712197 AA genotype, the rs1015411 GA genotype, the AA genotype, the PYGM rs10897540 GA genotype and GA or AA genotype compared with the homozygous common genotypes in these SNPs for the three genes.
Inconsistent associations with GDM were observed among SNPs in the SHC4 and ACACA genes. Reduced GDM risk was observed among women who carried the SHC4 rs7164585 AG genotype and the ACACA rs16237 AA genotype, and increased risk was observed among women who carried the SHC4 rs12906911 CC genotype, the ACACA rs12946701 AC genotype and the rs9893861 AC genotype compared with homozygous common in these SNPs.
At gene level, SOS1, PRKAA1, PRKAR2B and PHKA1 were associated with risk of GDM. Only the SOS1 gene was statistically significant after adjustment for the multiple comparisons using the FDR method (as shown in Supplementary Table 1).
Two blocks were defined in 9 SNPs of the SOS1 gene, including block 1 (rs3112188, rs13029406, rs3099968 and rs17023076) and block 2 (rs72797392, rs1010859, rs3923144 and rs4670899), by the haplotype analysis (as shown in Supplementary Table 3). Reduced risk of GDM was associated with the AGGA haplotype in block1 (OR = 0.68, 95% CI = 0.48–0.95, p = 0.025) and the GGGG haplotype in block 2 (OR = 0.62, 95% CI = 0.43–0.91, p = 0.013) as compared with the most common haplotype.
Gene-by-gene and gene-by-environment interaction analysis did not provide additional insight into the associations.
Discussion
In this study, SOS1 (son of sevenless homolog 1 in Drosophila) gene and two haplotypes in two blocks of this gene were found to be associated with decreased GDM risk. In previous studies, SOS1 was reported to be associated with a reduced risk of T2DM. 9 Similarities between the pathophysiology of GDM and T2DM may suggest that SOS1 could have a similar role in the pathophysiology of GDM. 10 GDM is thought to be a secondary reaction to maternal adipocytokine, placental and other hormones.1,11 However, this cannot fully account for GDM pathophysiology suggesting that genetics, the environment and their interaction may play an important role. Characterizing the mechanisms by which these factors enhance or interfere with the insulin signalling pathway is critical to understanding how these factors contribute to the pathogenesis of metabolic disorders. However, in our study, gene-by-gene and gene-by-environment interaction analysis did not provide additional insight into the associations. To our knowledge, this is the first study to report an association between the SOS1 gene and risk of GDM. However, the underlying mechanism remains unclear, warranting further investigation into the role this gene plays in GDM risk.
Our study has both strengths and limitations. First, the diagnosis of GDM was obtained by investigating the medical records and the diagnosis was based on the national guidelines of GDM diagnosis. This was likely to minimize potential misclassification. Second, information on potential confounders was collected using a standardized questionnaire. Third, the subjects were enrolled from hospital, which may have limited the generalizability of the result. Finally, the sample size in our study was moderate, and weak effects of SNPs on GDM maybe not have been observed.
In conclusion, our study suggests that the SOS1 gene and haplotypes in this gene may be involved in the pathogenesis of GDM. Further population-based studies with larger sample sizes among different ethnic groups are needed to verify these results.
Footnotes
Acknowledgements
The authors express their appreciation to the participants in the Taiyuan Birth Cohort Study for their enthusiastic support. Y.Z., S.W. and Q.C. conceived the idea and designed the study; Q.C. wrote the manuscript; H.Y., Y.F., P.Z., W.W., S.L., B.T., X.W., T.P., F.W., B.X., P.G., M.L., Y.W., N.Z., S.W. and Y.Z. interpreted the results and reviewed and edited the manuscript; and B.T. edited the manuscript. All authors provided intellectual input into the paper, and all authors read and approved the final manuscript.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship and/or publication of this article.
Funding
This work was supported by the National Institutes of Health grants (no. K02HD70324), National Natural Science Foundation of China grants (no. 81473061), Natural Science Foundation of Shanxi Province grants (no. 2013021033-2), ‘100 Talent Plan’ Award of Shanxi Province, Construction Project of Characteristic Key Disciplines for Universities of Shanxi Province and ‘10 Talent Plan’ Award of Shanxi Medical University.
