Abstract
Objectives
To investigate the association between toll-like receptor 9 (TLR9) single nucleotide polymorphisms (SNPs) and human papillomavirus (HPV) infection among Chinese Han women with cervical cancer.
Methods
TLR9 –1486 and 2848 SNPs were investigated in patients with cervical cancer and controls using polymerase chain reaction (PCR)-restriction fragment length polymorphism. HPV16 E6 and E7 infections were assessed using PCR.
Results
Of 120 patients with cervical cancer and 100 controls, there was a significant association between TLR9 2848 SNP and cervical cancer risk, but there was no such association with TLR9 –1486 SNP. Frequency of the TLR9 2848 GA genotype was significantly higher in patients with cervical cancer than in controls. There was no statistically significant between-group difference in presence of HPV16 infection. Presence of HPV infection with TLR9 2848 (rs352140) GA/AA genotype increased the risk of cervical cancer 13.8-fold compared with the GG genotype.
Conclusions
The TLR9 2848 G/A polymorphism in Chinese Han women was associated with increased risk of cervical cancer in the presence of HPV16 infection. Further studies are necessary to uncover the functional aspect of this TLR9 2848 polymorphism.
Keywords
Introduction
Cervical cancer is the third most commonly diagnosed cancer and the fourth leading cause of cancer death in females worldwide, with 85% of cases occurring in developing countries. 1 In China, it is estimated that 45 689 new cervical cancer cases occur annually, representing approximately one tenth of cases worldwide. 1 Human papillomavirus (HPV) infection is known to be the single most important risk factor for cervical cancer development, and improved screening techniques (including Papanicolaou testing, liquid-based cytology, and high-risk HPV testing) have significantly reduced cervical cancer deaths by >50%. 2 Infection with a high-risk HPV, such as the most prevalent HPV 16 and 18 subtypes, accounts for 76.7% of cervical cancer cases in China.3–5 DNA from these HPV subtypes is frequently integrated into the host cell genome in noninvasive squamous intraepithelial lesions and squamous cell carcinomas. 6 This integration is thought to be crucial in the development of invasive cervical cancer by increasing viral oncoprotein E6 and E7 expression levels. 7 E6 and E7 oncoproteins can inactivate the functions of the tumour suppressor proteins p53 and retinoblastoma protein, and can also inhibit inflammatory responses, in turn suppressing host defence mechanisms against infection and cancer.
Toll-like receptors (TLRs) have been identified as key components of the pathogen recognition process in human inflammatory responses against infectious disease. Thirteen genes of the TLR family have been identified in humans and mice, 8 and their proteins can recognize structurally conserved molecules derived from different micro-organisms, to activate immune cell responses. The TLR9 gene has been mapped to chromosome 3p21.3 and covers ∼5 kb with two exons, the second of which is the major coding region. 9 TLR9 appears to function as an innate immune pattern recognition protein for motifs that contain unmethylated CpG-rich bacterial and viral DNA, acting through the myeloid differentiation primary response (MYD88)-dependent signal transduction pathway. 9 At the molecular level, TLR9 is involved in nuclear factor of κ light polypeptide gene enhancer in B-cells 1 (NF-κB) transcription factor activation, proinflammatory cytokine production, and costimulatory molecule expression in human B cells and plasmacytoid dendritic cells, to eliminate harmful pathogens. 10 TLR9 has been shown to recognize CpG motifs from HPV16, and HPV16 E6 and E7 oncoproteins have been reported to inhibit TLR9 transcription in B lymphocytes and in cervical cancer cells. 11 Further investigations into the mechanics of HPV/ host TLR9 interactions would increase understanding of HPV infection and body defence mechanisms, and may facilitate the development of novel strategies for controlling HPV infection and reducing cervical carcinogenesis.
In the current study, the GenBank database (www.ncbi.nlm.nih.gov/genbank/) was searched for single nucleotide polymorphisms (SNPs) in TLR9 (rs352144, rs187084, rs352139, rs352140, and rs445676). Four of these have been reported to have higher frequencies in the human population, including –1486 (rs187084, located in the gene promoter region) and 2848 (rs352140, located in exon 2). 12 Such polymorphisms may affect TLR9 transcription or protein coding, and –1486 and 2848 polymorphisms have been shown to be associated with autoimmune disease and cancer. 13 Thus, the aim of the current study was to detect TLR9 gene polymorphisms at –1486 and 2848 in patients with cervical cancer and controls, to determine their association with HPV16 infection and clinicopathological characteristics of disease.
Patients and methods
Study population
This case–control study included Chinese Han patients with cervical cancer and Chinese Han control patients with uterine fibroids or endometriosis (matched with respect to age and ethnicity) who were recruited between June 2007 and January 2010 from West China Second Hospital, Sichuan University, Chengdu, China. Patients were excluded if they had previous history of atypical squamous cells of undetermined significance, low- or high-grade squamous intraepithelial lesions, or conization of the cervix.
Tissue samples from these patients were studied for genotyping of TLR9 gene polymorphisms. Well preserved paraffin wax-embedded tissue blocks were retrieved and cut into 10-µm thick sections for DNA extraction. Diagnosis was confirmed by histopathological examination of haematoxylin and eosin stained sections in all cases; such examinations were undertaken by one of the coauthors (L.S.), Demographic/clinicopathological data were obtained from medical records, including age, sex, cancer grade (cell differentiation level, size of tumour), and International Federation of Gynecology and Obstetrics (FIGO) staging data (http://www.medscape.com/viewarticle/722721). The study was approved by the Medical Ethics Committee of Sichuan University West China Second Hospital (Lun approval No. 019) and all study participants (or their legal proxies) provided written informed consent.
Genotyping of TLR9 polymorphisms
Genomic DNA was extracted from paraffin wax-embedded sections with phenol-chloroform and precipitated with ethanol using standard techniques. Polymerase chain reaction-restriction fragment length polymorphism (PCR-RFLP) was performed on the extracted DNA (100 ng/reaction) to detect the presence of two TLR9 SNPs, –1486 and 2848. Primers were designed using Primer 3 software (SimGene.com), and primers were obtained from Beijing Bo three Polygala Biotechnology Company Ltd. For the –1486 (T/C) SNP, PCR was performed using the QIAamp® DNA Mini Kit (Tektronix Biotechnology Company Ltd, Shanghai, China) and 5 µl each of the following primers: forward 5′-TGG GCA CTG TAC TGG ATC CTG G-3′ and reverse 5′-GCC TTG GGA TGT GCT GTT C-3′. The PCR conditions were 95℃ for 5 min, followed by 35 cycles of 95℃ for 45 s, 55–61℃ for 45 s, 72℃ for 30 s, and a final extension at 72℃ for 7 min. Following amplification, the PCR product was digested with Hpy 188III endonuclease (New England Biolabs, Ipswich, MA, USA), separated on a 3% agarose gel using electrophoresis, and stained with ethidium bromide. This generated three heterozygous DNA fragments of 327, 278, and 49 base pairs (bp) in length. For the 2848 (G/A) SNP, PCR was performed using the QIAamp® DNA Mini Kit (Tektronix Biotechnology Company Ltd) and 5 µl each of the following primers: forward 5′-AAG CTG GAC CTC TAC CAC GA-3′ and reverse 5′-TTG GCT GTG GAT GTT GTT-3′. The PCR conditions were 95℃ for 5 min, followed by 35 cycles of 95℃ for 45 s, 55–61℃ for 45 s, 72℃ for 30 s, and a final extension at 72℃ for 7 min. The PCR product was then digested with Aci I endonuclease (Fermentas International, Burlington, ON, Canada), separated on a 3% agarose gel using electrophoresis, and stained with ethidium bromide. Aci I digestion produced three fragments of 177, 137, and 40 bp. Gel images were obtained using the ATTO densitograph UV-image analyzer (ATTO Corporation, Tokyo, Japan).
Detection of HPV16 in tissue specimens
To detect HPV infection in cervical tissue specimens, genomic DNA from paraffin-wax embedded sections was extracted with phenol-chloroform and precipitated with ethanol, using standard techniques. Samples containing 100 ng of DNA were subjected to PCR amplification. PCR was performed using the QIAamp® DNA Mini Kit. For HPV16 E6, the forward primer was 5′-CCC ACA GGA GCG ACC CAG AAA GTT ACC-3′, and the reverse primer was 5′-CCC ATA TCT ATA TAC TAT GCA TAA ATC CC-3′. For HPV16 E7, the forward primer was 5′-ATG CAT GGC AGG CAC GTG ACC CTG AAG-3′, and the reverse primer was 5′-TTG GGG GCG CAG ATG GGG CAC ACG ATG TTC-3′. PCR reaction conditions were as previously described. 9 DNA fragments were visualized using 3.0% agarose gel electrophoresis with ethidium bromide staining, and gel images were obtained using the ATTO densitograph UV-image analyser.
Statistical analyses
Between-group differences in the distributions of TLR9 –1486 and 2848 genotype variants were evaluated using χ2-test. The risk of cervical cancer with each SNP genotype was estimated by computing the odds ratios (ORs) and their 95% confidence intervals (CIs), using logistic regression analyses. Comparison of clinical and laboratory parameters between patient subgroups was performed using Wilcoxon signed–rank test for continuous variables and χ2-test for categorical data. Genotype frequencies between HPV16(+) and HPV16(−) patient groups were compared using χ2-test. All statistical analyses using two-sided distribution were performed using SPSS® software, version 15.0 (SPSS Inc., Chicago, IL, USA) for Windows®. A P-value ≤ 0.05 was considered statistically significant.
Results
Data from a total of 220 Chinese Han patients were used in this study: 120 patients with cervical cancer and 100 age and ethnicity-matched controls with uterine fibroids or endometriosis. TLR9 –1486 and 2848 SNP genotypes in cervical cancer and matched control tissues were obtained using PCR-RFLP (Figure 1). Of the 220 patients analysed, two patients with cervical cancer and one control had the TLR9 –1486 SNP. Analysis of the TLR9 SNP gene frequencies showed a significant association between the TLR9 2848 SNP and the risk of cervical cancer (P = 0.002; OR = 7.04, 95% CI = 2.02, 24.52), but no association between the TLR9 –1486 SNP and the risk of cervical cancer (Table 1). The heterozygous variant genotype GA was associated with an increased risk of developing cervical cancer compared with homozygous (GG or AA) genotypes (P = 0.004, OR = 6.92, 95% CI = 1.53, 33.3; and P = 0.036, OR = 7.91, 95% CI = 0.97, 64.52; respectively, Table 1). There was no statistically significant association between either TLR9 –1486 or 2848 (G/A) SNPs and clinicopathological data (Table 2).
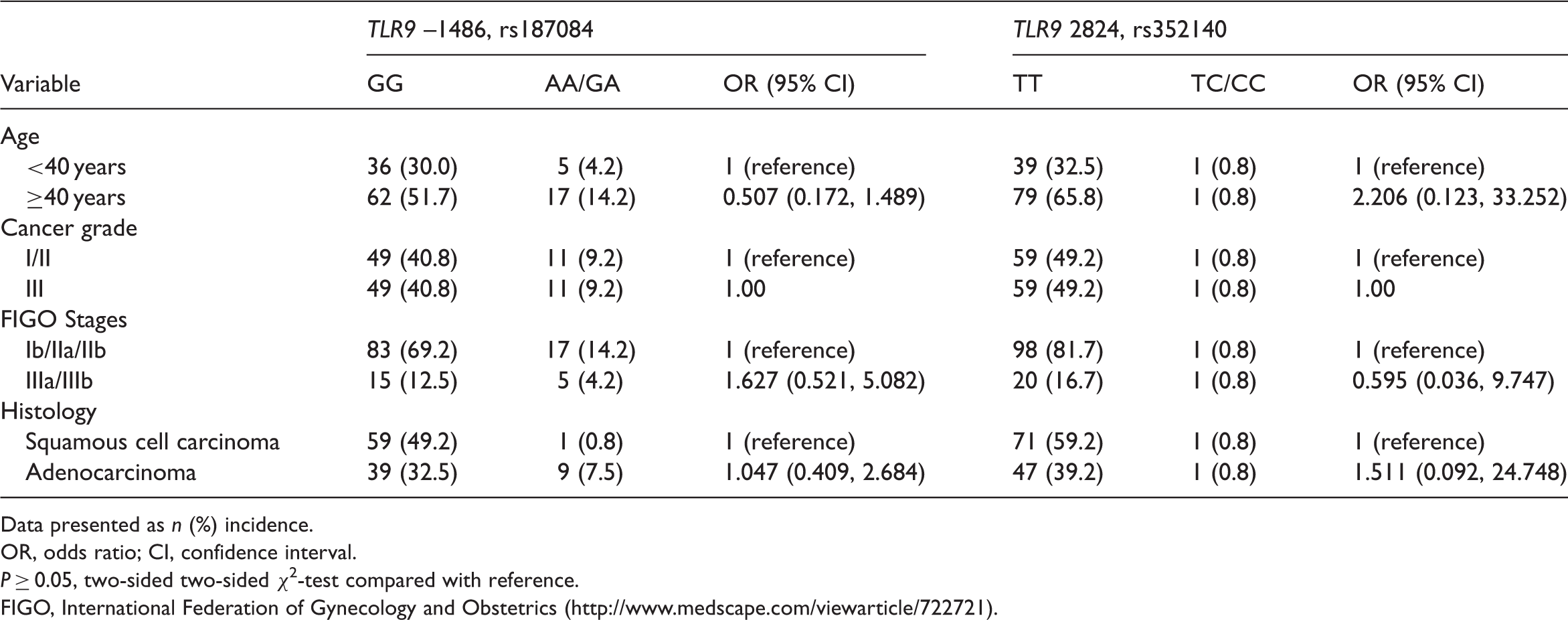
Representative gel images showing polymerase chain reaction-restriction fragment length polymorphism (PCR-RFLP) analysis of toll-like receptor 9 (TLR9) single nucleotide polymorphisms (SNPs) in cervical cancer tissue specimens, versus controls. (a) TLR9 –1486 (T/C) SNP. The PCR product was digested with Hpy 188III endonuclease and separated on a 3% agarose gel: lane 1, 100-base pairs (bp) DNA marker; lanes 5, 7 and 9, T/C heterozygous genotype; lane 8, C/C homozygous genotype; lanes 3, 4, and 6, T/T homozygous genotype. (b) TLR9 2848 (G/A) SNP. The PCR product was digested with Aci I endonuclease and separated on a 3% agarose gel: lane 1, 100 bp DNA marker; lanes 2, 4, and 7, G/A heterozygous genotype; lanes 3, 6, 8 and 9, G/G homozygous genotype; lane 5, A/A homozygous genotype. Associations between genotype or allele distributions of toll-like receptor 9 (TLR9) –1486 and 2848 polymorphisms and the risk of cervical cancer in Chinese Han women with cervical cancer, and age/ethnicity matched controls with uterine fibroids or endometriosis. Data presented as n (%) incidence. OR, odds ratio; CI, confidence interval. NS, no statistically significant differences (P ≥ 0.05), two-sided χ2-test compared with reference. Associations between genotype distributions of two toll-like receptor 9 (TLR9) single nucleotide polymorphisms (SNPs) and clinicopathological parameters in 120 Chinese Han women with cervical cancer. Data presented as n (%) incidence. OR, odds ratio; CI, confidence interval. P ≥ 0.05, two-sided two-sided χ2-test compared with reference. FIGO, International Federation of Gynecology and Obstetrics (http://www.medscape.com/viewarticle/722721).

Association between toll-like receptor 9 (TLR9) –1486 and 2848 single nucleotide polymorphisms (SNPs) and the risk of cervical cancer in the presence of human papillomavirus (HPV)16 infection in Chinese Han women with cervical cancer or age/ethnicity matched controls with uterine fibroids or endometriosis.

Data presented as n (%) incidence.
OR, odds ratio; CI, confidence interval.
NS, no statistically significant differences (P ≥ 0.05), two-sided two-sided χ2-test compared with reference.
Discussion
Innate immunity plays an important role in the suppression of cervical carcinogenesis in humans. 14 The TLR9 protein can recognize unmethylated CpG DNA and activated NF-κB channels, resulting in inflammatory factors that play an important role in tumour immunity. 15 CpG-oligodeoxynucleotides have been shown to enhance antitumour immunity in peripheral blood mononuclear cells from patients with lung cancer, through elevating cytokine secretion (e.g. tumour necrosis factor-α) and enhancing interferon-γ production. 16 Moreover, the combination of a G allele at position +1174 with a C allele at position –1486 was shown to have the ability to downregulate TLR9 expression and increase the risk of systemic lupus erythematosus. 17 The current case–control study associated a TLR9 gene polymorphism with HPV infection in Chinese Han patients with cervical cancer. A significant association was found between the TLR9 2848 SNP and risk of cervical cancer. No such association was found between the TLR9 –1486 SNP and risk of cervical cancer; however, only two patients with cervical cancer cases and one control possessed the TLR9 –1486 SNP in the current study. The frequency of the TLR9 2848 GA genotype was significantly higher in cervical cancer specimens than in controls. Although there was no between-group difference in terms of HPV16 infection, when HPV16 infection was present, the TLR9 2848 (rs352140) GA/AA genotype increased the risk of cervical cancer 13.8-fold compared with the GG genotype. These data suggest that the TLR9 2848 (G/A) polymorphism was associated with HPV16 infection and cervical cancer in Chinese Han women.
The increased frequency of TLR9 2848 GA genotype in patients with cervical cancer compared with controls also indicated an association between the TLR9 2848 polymorphism and the risk of cervical cancer. The region around 2848 is the major coding region of the TLR9 protein,18,19 and a polymorphism of the TLR9 2848 GA genotype has been reported to reduce TLR9 expression at the transcriptional level.8,19 Downregulated TLR9 could reduce the functions of innate immune reactivity against infection, in this case HPV infection, in turn prolonging infection and eventually progressing to precancerous and cancerous lesions. The current study did not, however, detect any association between the genetic variations of TLR9 and tumour differentiation, stage or histological type of cervical cancer. These data suggest that the TLR9 gene may play a role in cervical carcinogenesis but have a lesser (or no) role in tumour progression. Thus, further investigations of the TLR9 gene could be important in improving our understanding of cervical cancer pathogenesis.
In mammals, the TLR9 protein functions as a bacterial and viral CpG motif-specific receptor that triggers innate immune responses to eliminate foreign pathogenic micro-organisms, such as viruses.20,21 Studies have shown that TLR9 polymorphisms are associated with various viral infections.22,23 For instance, an association was revealed between a TLR9 promoter polymorphism and respiratory syncytial virus infection; 22 in addition, the TLR9 1635GG genotype was reported to be more frequently found in people with rapidly progressing HIV-1 infection. 23 To date, there have been no reports showing an association between genetic variations of TLR9 and HPV16 infection in Chinese patients. The current study found that TLR9 2848 GA/AA genotypes were more frequent in HPV16-positive patients with cervical cancer than in HPV16-negative patients.
The most prevalent type of HPV associated with cervical carcinogenesis is HPV16. The majority of HPV16 infections do not lead to cervical cancer, 24 however, which may be due to clearance of the HPV16 virus by the host innate immune system or suppression of HPV16 tumourigenic functions by body defence mechanisms. For example, HPV16 E6 and E7 recombinant retroviruses have been shown to inhibit TLR9 transcription and enhance functional loss of TLR9-regulated gene pathways. 11 However, TLR9 gene polymorphisms could alter HPV oncoprotein accessibility, thereby having an impact on HPV infection and tumourigenesis.
The current preliminary study suggested that the TLR9 2848 (G/A) polymorphism may be associated with increased risk of cervical cancer in the presence of HPV16 infection, in Chinese Han women. Further investigations should be conducted to confirm that the TLR9 2848 polymorphism is a risk factor for cervical cancer among these women. In addition, future studies should focus on detecting TLR9 expression in tumour tissue for the association with TLR9 polymorphisms, and a study with a larger sample size should be conducted to investigate multiple sites and the association between TLR9 polymorphisms and a wide range of HPV subtypes.
Footnotes
Declaration of conflicting interest
The authors declare that there are no conflicts of interest.
Funding
This work was supported in part by the Program for Natural Science Foundation of Sichuan University, Sichuan, China. Sichuan Provincial Health Department Research Projects, China (110438). Applied Basic Research Program of Sichuan Provincial Science and Technology Department, China (2012JY0009).
