Abstract
The severity of systemic meningococcal disease (SMD) correlates to plasma concentrations of LPS and IL-10, with the highest levels detected in non-survivors. Here, plasma from patients with SMD containing high and low concentrations of LPS were incubated with human monocytes before and after immunodepletion of IL-10 to study the effect of IL-10 on gene expression and cytokine release. Patient plasma containing IL-10 induced the expression of 1657 genes in human monocytes when compared with gene expression induced by low LPS plasma. After immunodepletion of IL-10, this number increased to 2260. By directly comparing the gene expression profiles induced before and after immunodepletion of IL-10, the presence of IL-10 differentially regulated 373 genes. Functional classes associated with these genes were cellular function and maintenance, cellular development, cellular growth and proliferation, cell–cell signaling and interaction and cellular movement. Immunodepletion of IL-10 resulted in down-regulation of genes of the leukocyte immunoglobulin-like receptor family, and up-regulation of genes of type I IFN signaling, TLR signaling, the inflammasomes, coagulation and fibrinolysis. Finally, immunodepletion of IL-10 increased the protein levels of IL-1β, IL-8, TNF-α, MIP-1α and MIP-1β. Data suggest that IL-10 in meningococcal sepsis plasma regulates a variety of genes and signaling pathways, likely leading to an overall inhibitory effect on the inflammatory response induced in meningococcal sepsis.
Keywords
Introduction
The initial phase of bacterial sepsis is characterized by a pro-inflammatory response. The intensity of the response is variable and related to the load and virulence of the bacteria. 1 In patients surviving this pro-inflammatory phase, studies suggest that a subsequent anti-inflammatory or inert phase is associated with morbidity and mortality owing to down-regulation or weakening of the immune response. 2 Meningococcal sepsis is characterized by its rapid development and overwhelming character, which is explained by the propensity of Neisseria meningitidis to multiply very rapidly in the circulation, reaching levels as high as 108 bacteria per ml plasma within 12–24 h.3,4 This massive bacterial growth generates high levels of LPS in plasma, which up-regulates the pro-inflammatory response.5–7 LPS is associated with fatality in a dose-dependent manner.8,9 The anti-inflammatory phase in meningococcal sepsis appears to closely follow the pro-inflammatory phase, as plasma concentrations of the anti-inflammatory cytokine IL-10 correlate with levels of LPS, pro-inflammatory cytokines and disease severity.9–11 A previous study by our group demonstrated that these high levels of IL-10 have a potent inhibitory activity on cytokine release and pro-coagulant activity in human monocytes. 12
In addition to cytokine release, extensive gene expression changes may occur during the pro- and anti-inflammatory phase of sepsis.13–16N. meningitidis and purified meningococcal LPS have the ability to alter the expression of more than 4000 genes. 17 IL-10 can regulate selected groups of these genes, potentially affecting cellular functions and signaling pathways important for the systemic inflammatory response. 18 We, and others, have developed experimental models to study the effects of purified LPS, whole bacteria and recombinant IL-10 on the gene regulation in monocytes/macrophages derived from humans and mice.19–22 In this study, we have investigated gene expression induced in human monocytes by plasma collected from patients with meningococcal sepsis containing high levels of biologically active LPS, cardinal pro-inflammatory cytokines and IL-10. By using an immunoprecipitation technique depleting IL-10 from plasma, we have been able to investigate gene expression and cytokine release induced in the presence and absence of IL-10. To our knowledge, this is the first detailed study of gene regulation in human monocytes using meningococcal sepsis plasma. It may contribute to our understanding of genomic changes associated to the pro- and anti-inflammatory response in overwhelming sepsis, and serve as a model to elucidate the effect of sepsis plasma on gene regulation of specific cell types.
Materials and methods
Reagents and solutions
All reagents and solutions were analyzed for the presence of LPS using the Limulus amebocyte lysate (LAL) assay (Pyrochrome; Associates of Cape Cod Incorporated, Falmouth, MA, USA). The lower detection limit was 0.16 endotoxin units (EU/ml). FCS (Biowhittaker; Lonza, Walkersville, MD, USA) was tested to contain <1 EU LPS, and was heat-inactivated to inactivate inhibitory serum-proteases and complement proteins. 23 RPMI medium 1640 was obtained from Gibco (Invitrogen, Carlsbad, CA, USA). Penicillin and streptomycin was obtained from Sigma-Aldrich (St. Louis, MO, USA).
Pooled human normal plasma
Heparinized (30 IU/ml) whole blood samples were collected from consenting, healthy donors (n = 10). After immediate centrifugation (1400 g for 10 min at 20℃), plasma was pipetted off, mixed, aliquoted and stored at −70℃ in Nunc cryotubes (Sigma-Aldrich).
Neisseria meningitidis
The reference strain N. meningitidis H44/76 (serogroup B:15:P1:7,16), originally isolated from a culture of blood from a Norwegian patient with lethal meningococcal sepsis, was provided by the Norwegian Institute of Public Health (NIPH; Oslo, Norway). The number of bacteria was quantified as described in our previous studies.4,18
Collection of patient plasma samples
The patient plasma samples (n = 6) were collected after informed consent from parents, relatives or patients, and in accordance with guidelines approved by the Regional Medical Ethics Committee of Health Region I in Norway (Biobank material access number 948; Oslo University Hospital, Oslo, Norway). Blood from patients 1, 2, 3 and 5 was collected in endotoxin-free heparinized tubes [30 IU sodium heparin/ml] (Endotube Chromogenix, Bedford, MA, USA). Blood from patients 4 and 6 was collected in endotoxin-free vaccum tubes (15 IU sodium heparin/ml) from Becton Dickinson (Franklin Lakes, NJ, USA). Blood samples were immediately centrifuged (1400 g for 10 min at 20℃), plasma pipetted off, aliquoted and stored at −70℃ in Nunc cryotubes (Sigma Aldrich).
LPS quantification in patient plasma
Quantification of LPS was performed by the LAL assay, as previously described. 9
Clinical description and laboratory findings of the patients
Clinical and laboratory data of patients included in the study.
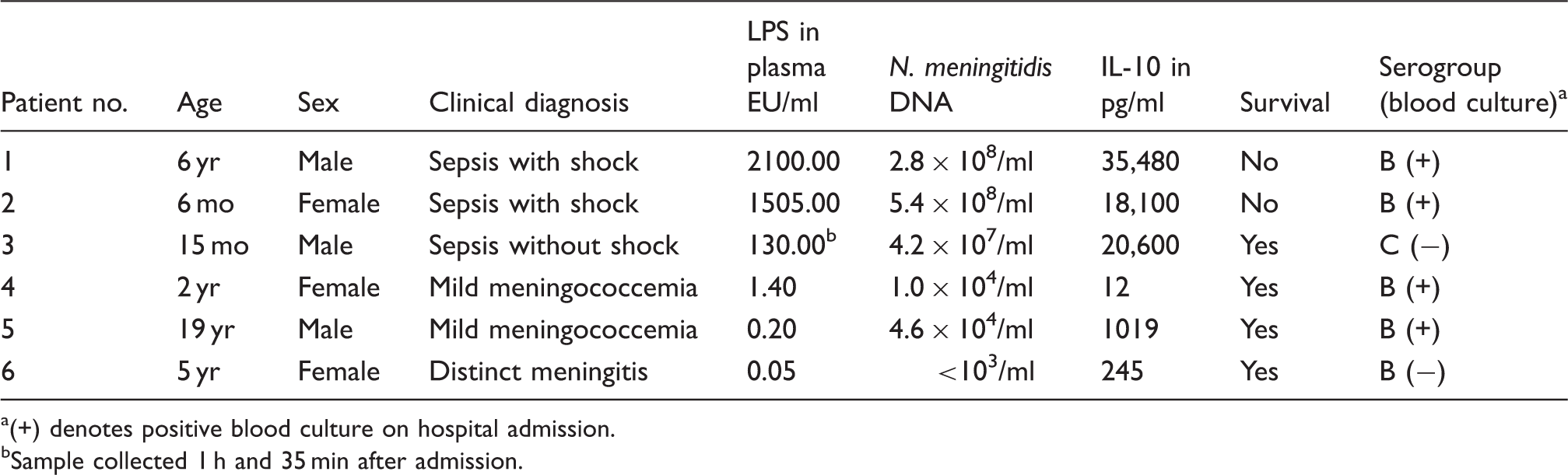
(+) denotes positive blood culture on hospital admission.
Sample collected 1 h and 35 min after admission.
Patient 1
A six-year-old boy was hospitalized 17 h after the first symptoms of fatigue and later fever. On admission, he was characterized as having severe sepsis that rapidly developed into septic shock. Petechiae and ecchymoses increased rapidly in number and size. Leukocytes were 1.5 × 109/l and platelets 53 × 109/l. Coagulation tests indicated severe disseminated intravascular coagulation (DIC). Metabolic acidosis with a of pH 7.33 and a base excess of −11.5 mmol/l was present. Creatinine concentration in the serum was pathologically elevated. He was treated at the pediatric intensive care unit (PICU) with benzylpenicillin and chloramphenicol, a large volume of fluids and norepinephrine, and was artificially ventilated. He died of progressive cardiovascular failure 5 h after admission. N. meningitidis serogroup B in blood culture was sensitive to benzylpenicillin and chloramphenicol. Heparin plasma collected on admission showed an LPS level of 2100 EU/ml and the copy number of N. meningitidis DNA was 2.8 × 108/ml. Final diagnosis was lethal meningococcal septic shock.
Patient 2
A six-month-old girl developed diarrhea 17 h before hospital admission, and subsequently experienced fever and hemorrhagic skin lesions. On admission, she had septic shock with petecchiae and ecchymoses. Leukocytes were 3.2 × 109/l (pathologically low), platelets were 124 × 109/l, and she had severe metabolic acidosis with pH 6.99 and a base excess of −20.9. Creatinine was pathologically elevated. She was treated at the PICU with benzylpenicillin and chloramphenicol, a large volume of fluids and norepinephrine, and was artificially ventilated. She died of progressive cardiovascular failure 5.5 h after admission. N. meningitidis serogroup B grew in blood culture and was sensitive to benzylpenicillin and chloramphenicol. Heparin plasma collected on admission showed an LPS level of 1505 EU/ml and the copy number of N. meningitidis DNA was 5.4 × 108/ml. Final diagnosis was lethal meningococcal septic shock.
Patient 3
A 15-month-old boy awoke in the morning with fever. His general condition deteriorated and, upon hospital admission 11 h later, he had petechiae but no ecchymoses. His consciousness was reduced, but he lacked marked neck and back stiffness. His blood pressure was normal for his age. He was classified as having severe sepsis, but without septic shock. Leukocytes were 3.2 × 109/l and platelets were 115 × 109/l (both pathologically low), and he had severe metabolic acidosis with a pH of 7.32 and a base excess of −16.0 mmol/l. Creatinine in serum was pathologically elevated. He was treated at the ICU in a local hospital with cefotaxime and a normal volume of fluids. He did not require vasoactive drugs or artificial ventilation. Blood culture was negative. PCR and ELISA studies at the NIPH showed N. meningitidis serogroup C in the patient’s blood. LPS in heparin plasma collected on admission was 214 EU/ml and, 100 min after initiation of cefotaxime, LPS declined to 130 EU/ml (sample used for analysis in this study). The copy number of N. meningitidis DNA on admission was 4.2 × 107/ml. Final diagnosis was severe meningococcal sepsis without septic shock, surviving without sequelae. Clinically, this patient represents an ‘outlier’, that is a patient apparently tolerating much higher levels of meningococcal LPS than the majority of such patients. He did not develop septic shock, multiple organ failure or DIC despite having 35-fold higher LPS level at admission than the previously defined septic shock level (7 EU/ml) based on studies of 135 patients with systemic meningococcal patients. 9 The sample used in the present study contained 130 EU/ml LPS, and was collected 1 h and 35 min after the first sample upon admission. A 52% reduction in LPS from the admission level is compatible with the established elimination curve of LPS in meningococcal sepsis patients.8,9
Patient 4
A two-year-old girl suddenly developed high fever and was hospitalized 2 h later with few petechiae but no ecchymoses. Her general condition was good and her blood pressure was normal for her age. She had no neck or back stiffness. Leukocytes were 15.8 × 109/l (pathologically increased) and platelets were 291 × 109/l. She had no metabolic acidosis, with a pH of 7.38 and a base excess of −1.6. Creatinine was normal for age. CSF was normal. She was treated at the PICU with benzylpenicillin and chloramphenicol and, after 48 h, with benzylpenicillin alone. She received normal maintenance fluid, but no vasoactive drugs or artificial ventilation. After 5 d of treatment with antibiotics she was discharged without sequelae. N. meningitidis serogroup B grew in blood culture and was sensitive to benzylpenicillin and chloramphenicol. Heparin plasma collected on admission showed an LPS level of 1.4 EU/ml and the copy number of N. meningitidis DNA was 1.0 × 104/ml. Final diagnosis was mild meningococcemia (very early stage of systemic meningococcal disease).
Patient 5
A 19-year-old man, recently enrolled in the military service, awoke with a fever and a sore throat, and later a headache and back pain. Eleven hours later he was admitted to hospital with a fever and multiple petechiae but no ecchymoses. Leukocytes were 23.0 × 109/l (pathologically increased), platelets were 200 × 109/l, and he had mild compensated metabolic acidosis with a pH of 7.40 and a base excess of −6.0 mmol/l. Creatinine was slightly increased above the upper normal level. CSF contained leukocytes (47 × 106/l) but no growth of N. meningitidis. His blood pressure declined but reacted positively to increased intravenous fluids. He was treated with benzylpenicillin and chloramphenicol, and, after 72 h, with benzylpenicillin alone. After initial rehydration, he later received normal volumes of maintenance fluid, no vasoactive drugs or artificial ventilation. After 5 d of treatment with benzylpenicillin he was discharged without sequelae. N. meningitidis serogroup B grew in blood culture and was sensitive to benzylpenicillin and chloramphenicol. Heparin plasma collected 9 h after antibiotic treatment started showed an LPS level of 0.2 EU/ml and the copy number of N. meningitidis DNA was 4.6 × 104/ml. Final diagnosis was mild meningococcemia without sequela.
Patient 6
A 5-year-old girl had gradually increasing symptoms of meningitis for more than 24 h. On hospital admission she was afebrile but with marked neck stiffness. She had reduced consciousness and petechiae but no ecchymoses. Her blood pressure remained normal. Leukocytes were 20.8 × 109/l (pathologically increased), platelets were 181 × 109/l, and she had no metabolic acidosis with a pH of 7.40 and a base excess of −2.4 mmol/l; creatinine was normal. CSF was cloudy with leukocytes >109/l, increased protein and growth of N. meningitidis serogroup C. Blood culture was negative. She was treated with benzylpenicillin and chloramphenicol, and, after 72 h, with benzylpenicillin alone for another 4 d. She received normal maintenance fluid, no vasoactive drugs or artificial ventilation. She was discharged after 10 d without sequelae. Heparin plasma collected on admission showed an LPS level of 0.05 EU/ml and the copy number of N. meningitidis DNA was <103/ml. Final diagnosis was meningococcal meningitis without shock and sequelae.
Definitions
Plasma samples from patient 1–3 contained high levels of LPS and IL-10, and are denoted as ‘patient plasma containing IL-10’. Plasma samples from patient 1–3 after immunodepletion of IL-10 are denoted as “IL-10 immunodepleted patient plasma”. Plasma samples from patient 4–6 contained low levels of LPS (<1.5 EU/ml) and are denoted as“low LPS patient plasma”.
Recombinant human IL-10 and anti-human IL-10 Abs
Stock solution (10 µg/ml) of recombinant IL-10 (rIL-10; R&D Systems, Abingdon, UK) was reconstituted by adding 1 ml PBS to the lyophilized powder, aliquoted and stored in working aliquots of 50 µl at −70℃.
Stock solution (1 mg/ml) of anti-human IL-10 Abs (anti-IL-10) (Sigma-Aldrich) was reconstituted by adding 1 ml of PBS to the lyophilized powder, aliquoted and stored in working aliquots of 100 µl at −70℃.
Immunoprecipitation of rIL-10 from normal pooled plasma
In order to test the ability of anti-IL-10 to remove IL-10 through immunoprecipitation, we boosted three tubes of normal pooled plasma with 25 ng/ml of IL-10 and subsequently added 160 µg/ml, 80 µg/ml and 40 µg/ml anti-IL-10 to the respective tubes. The tubes were incubated for 1 h at 37℃ with gentle agitation and then immediately incubated for 18 h without agitation at 4℃. After centrifugation (3000 g for 30 min at 4℃), the supernatant plasma was pipetted off. The IL-10 levels in the supernatant plasmas were measured by an ELISA (Life Technologies; Invitrogen, Carlsbad, CA, USA). We found that 160 µg/ml anti-IL-10 was needed in order to completely remove 25 ng/ml of IL-10 from the pooled normal plasma (results not shown). Similar concentrations of anti-IL-10 have previously been used to immunoprecipitate IL-10 from plasma samples. 24
Testing for LPS contamination of anti-IL-10 and for the stimulatory capacity of anti-IL-10/IL-10 immune complexes
In order to assess if anti-IL-10 activated human monocytes, we stimulated 7.5 × 105/ml monocytes, suspended in 500 µl 5% (v/v) FCS-RPMI and 500 µl pooled normal plasma for 3 h with 160 µg/ml, 80 µg/ml and 40 µg/ml anti-IL-10. Monocyte activation was estimated by measuring levels of TNF-α in the culture supernatants. Neither the Abs nor the immune complexes stimulated the release of TNF-α as compared with unstimulated monocytes (results not shown).
Immunoprecipitation of IL-10 in patient plasma
We incubated 300 µl of patient plasma from three patients with high levels of LPS (≥130 EU/ml) and from three patients with low levels of LPS (≤1.4 EU/ml) with 160 µg/ml anti-IL-10. Concentrations of IL-10 in these samples are presented in Table 1. Immunoprecipitation was performed as described above, and plasma depleted for IL-10 was immediately transferred to the monocyte target assay.
Monocyte target assay
Elutriation-purified, cryopreserved human monocytes (>90% purity) from a consenting, healthy donor (Biobank material access number 908) was thawed and re-suspended in RPMI 1640 in 5% v/v FCS containing 2% v/v penicillin/streptomycin and seeded (1.5 × 106 monocytes suspended in 250 µl 5% FCS-RPMI/well) in microtiter plates (Costar 3524; Corning Tewksbury, MA, USA), as previously described. 25 The monocyte suspensions were incubated with 250 µl patient plasma samples containing high (n = 3) and low (n = 3) levels of LPS and IL-10 prior to and after immunodepletion of IL-10. Four control wells with monocytes were incubated with 250 µl pooled normal plasma spiked with 1.0 × 106/ml N. meningitidis, 1.0 × 106/ml N. meningitidis and 25 ng/ml rIL-10, 25 ng/ml rIL-10 and vehicle. The microtiter plate was sealed off and incubated for 3 h (37℃, 5% CO2), and then centrifuged (47 g for 5 min at 37℃). After centrifugation, the supernatant was gently removed, aliquoted and stored at −70℃. Cells were subsequently harvested and stored in 700 µl Qiazol lysis buffer [supplied with the miRNeasy Mini Kit (Qiagen, Hilden, Germany] at −70℃ until subjected to total RNA isolation.
RNA preparation
Total RNA was extracted using the miRNeasy Mini Kit (Qiagen), and was further enriched using Microcon Centrifugal Filter Units (Millipore, Darmstadt, Germany), following manufacturer’s instructions. Total mRNA was quantified using a Nano Drop spectrophotometer (Saveen Werner, Limhamn, Sweden) and quality controlled using the RNA 6000 Nano assay (Agilent BioAnalyzer 2100 System; Agilent Technologies, Santa Clara, CA, USA). The RNA integrity numbers after enrichment ranged from 6.1 to 7.8.
Microarray analyses
Microarray analyses were performed using the Affymetrix GeneChip Human Gene 1.0 ST Arrays (Affymetrix, Santa Clara, CA, USA). Total RNA (150 ng) was subjected to the GeneChip HT one-cycle cDNA synthesis kit and GeneChip HT IVT labeling kit, following the manufacturer’s protocol for whole-genome gene expression analysis. Biotinylated and fragmented single-stranded cDNAs were hybridized to the GeneChips. The arrays were washed and stained using an FS-450 fluidics station (Affymetrix). Signal intensities were detected by a Hewlett Packard (Palo Alto, CA, USA) 3000 7G gene array scanner. The scanned images were processed using the Affymetrix GeneChip Command Console software, and the CEL files were imported into Partek Genomics Suite software (Partek, St. Louis, MO, USA). The robust multichip analysis algorithm was applied for the generation of signal values and normalization. Gene transcripts with maximal signal values of <32 across all arrays were removed to filter for low and non-expressed genes. For expression comparisons of different groups, profiles were compared using a two-way ANOVA model. The results were expressed as fold change (FC). Genes with FC ≥|1.5| and a P-value <0.01 were regarded as significantly expressed. Partek Genomics Suite software was used to generate heatmaps and principal component analysis (PCA) of the gene expression data. Final microarray data were deposited in Gene Expression Omnibus.
Ingenuity Pathway Analysis
Gene lists (Excel files) containing gene identifiers (probe set IDs) and corresponding expression values were uploaded to Ingenuity Pathway Analysis (IPA; Ingenuity Systems, Redwood City, CA, USA). A cut-off at FC ≥|1.5| and P < 0.01 was set to identify significantly differentially expressed genes. Several tools in IPA were used to analyze the gene expression data. Biological functions associated with the differentially expressed genes were identified by mapping each gene to its corresponding function in the Ingenuity Knowledge Base. IPA performs the Fisher’s exact test to calculate a P-value, determining the probability that each biological function assigned to the data set is due to chance alone.
In order to create gene networks, which display interaction between genes and pathways and their association with cellular functions, IPA overlaid the significantly expressed genes (called network eligible molecules) onto a global molecular network developed from information contained in the Ingenuity Knowledge Base. The Ingenuity Knowledge Base contains existing scientific information about the molecular relationships between the network eligible molecules and is used by IPA to generate gene networks. IPA computed a score for each network according to the fit of the genes in the data set in order to identify the most highly expressed networks. This score is generated using a P-value calculation determined by right-tailed Fisher’s exact test, and the score is displayed as the negative log of that P-value. This score indicates the likelihood that the fit of the set of focus genes in a network can be explained by chance alone. A score of 2 indicates that there is a 10−2 probability that the focus genes are together in a network by random chance.
Furthermore, we used functional analysis in IPA to predict causal effects between the observed gene expression data and changes in cellular functions. Expected causal effects between changes in gene expression and functions are derived from existing research findings in Ingenuity Knowledge Base. By identifying the functions associated with the differentially expressed genes and analyzing the genes’ direction of change (up- or down-regulated), IPA issues a prediction for the direction of change in function. IPA uses a z-score algorithm to calculate the significance of the prediction. The algorithm is designed to reduce the likelihood that random data generate significant predictions about the direction of change in function. A z-score ≥ 2.0 predicts a significant increase in the function, while a z-score ≤ −2.0 predicts a significant decrease in function.
Validation of mRNA data with RT-PCR
To validate the gene expression data analyzed by micro array analysis, selected differentially expressed genes were quantified by qRT-PCR [TaqMan gene expression assays and the Applied Biosystems ViiA7 sequence detection system (Applied Biosystems, Foster City, CA, USA]. Owing to small quantities of materials, cRNA from the Gene Chip HT one cycles cDNA synthesis kit was used for the validation. cRNA (20 ng) was reversed transcribed using random hexamers (Applied Biosystems) and Omniscript (Qiagen) in a total volume of 20 µl cDNA (2 µl), and 1 µl of RT-PCR primers (NLRP3 Hs 00366465-m1, LILRA3 Hs 00846590-s1, LILRB1 Hs 00366806-m1, IFIT3 Hs 00382744-m1, IFIT2 Hs 00533665-m1, CD14 Hs 02621496-s1, AIM2 Hs 00915710_m1, IL1b Hs 01555410_m1 and TNF-α Hs 99999043_m1) was added to 10 µl TaqMan universal PCR mastermix (Applied Biosystems) and 7 µl of H2O. The relative changes of each transcript using the mean of B2M (Hs 99999907_m1) and PPIB(Hs 00168719) as endogenous controls were calculated using ViiA 7 Software v1.1 (Life Technologies) and the ΔΔCT method. 26
Quantification of selected proteins in patient plasmas and supernatants before and after immunodepletion of IL-10
A microsphere-based multiplexing bioassay system with Xmap technology (Luminex, Austin, TX, USA) was used to quantify cytokines and chemokines in the patient plasmas prior to incubation with human monocytes and before and after immunodepletion of IL-10. The quantification was conducted according to the manufacturer’s instructions. The culture supernatants were examined for 30 cytokines [IL-1β, IL-1RA, IL-2, IL-2R, IL-4, epidermal growth factor (EGF), IL-5, IL-6, IL-7, IL-8, IL-10, fibroblast growth factor (FGF)-Basic, IL-12, IL-13, IL-15, IL-17, TNF-α, granulocyte colony-stimulating factor (G-CSF), IFN-α, IFN-γ, granulocyte macrophage (GM)-CSF, macrophage inflammatory protein (MIP)-1α, MIP-1β, hepatocyte growth factor (HGF), IFN-γ-induced protein 10 (IP-10), monokine induced by IFN-γ (MIG), Eotaxin, RANTES, monocyte chemotactic protein (MCP)-1 and vascular endothelial growth factor (VEGF)] using the Human Cytokine 30-Plex Panel from Invitrogen. All samples were run in duplicate.
Statistical analysis
For Affymetrix gene expression, a combination of FCs and two-way ANOVA was used to detect differentially expressed genes in the data set. Genes with a FC ≥|1.5| and a P-value <0.01 were regarded as significantly expressed. Pearson’s correlation test in SPSS 14.0 (IBM, Armonk, NY, USA) was used to calculate the correlation between FCs in Affymetrix and real time-PCR (RT-PCR).
Results and discussion
Immunodepletion of IL-10 from patient plasma increases cytokine release from human monocytes
Cytokine levels in patient plasma at admission, and after incubating human monocytes for 3 h with patient plasma containing IL-10 and IL-10 immunodepleted patient plasma.
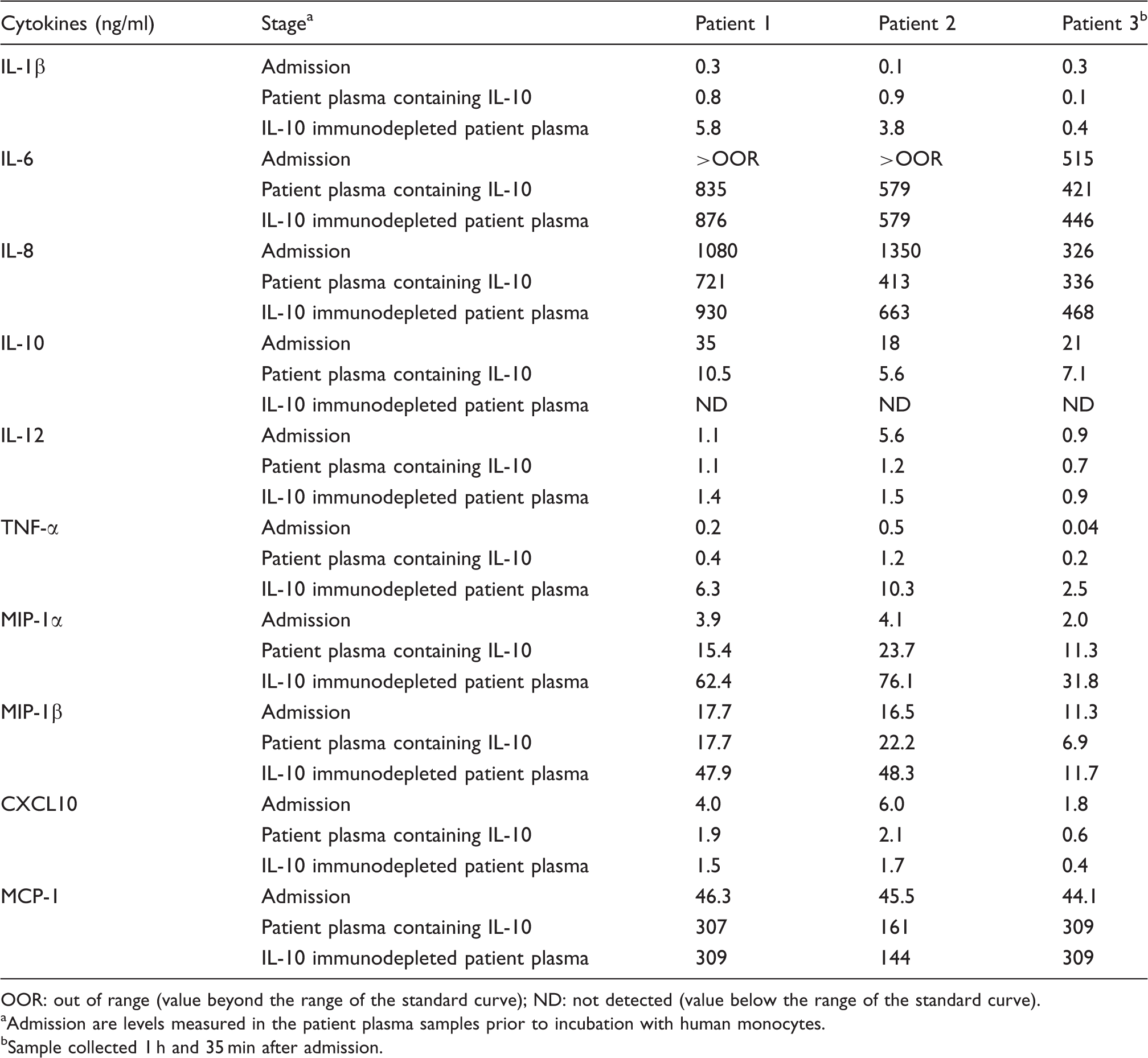
OOR: out of range (value beyond the range of the standard curve); ND: not detected (value below the range of the standard curve).
Admission are levels measured in the patient plasma samples prior to incubation with human monocytes.
Sample collected 1 h and 35 min after admission.
Overall, our results confirm findings from previous studies suggesting that the circulating levels of IL-10 have an inhibitory effect on cytokine release from human monocytes, which limit the pro-inflammatory response in patients with septic shock and multiple organ failure due to meningococcal sepsis.12,27
PCA clusters gene expression according to levels of LPS and IL-10 in patient plasma
We generated a PCA plot to obtain an overall picture of the gene expression induced by the patient plasma samples in our experimental model (Figure 1). Principal component analysis identifies the directions (principal components) in which the variation in the data is maximal, and enables us to visualize this variation in a plot. Each circle in the PCA plot represented gene expression data induced by each sample in our experimental model. The six circles (purple and orange) representing gene expression induced in human monocytes by plasma from patients with mild meningococcemia, containing low or undetectable levels of LPS and IL-10, clustered together in a “baseline cluster”. Closely located to the “baseline cluster” were the circles (pink and green) representing gene expression induced in IL-10-stimulated human monocytes and unstimulated controls. This indicated that the gene expression patterns induced by patient plasmas with low LPS levels were similar to gene expression induced by plasma from healthy donors and by IL-10 alone. The two other clusters in the PCA plot represented plasma samples from patients with meningococcal sepsis. One cluster of blue circles, furthest apart from the baseline cluster, consisted of gene expression profiles induced by patient plasma after immunodepletion of IL-10. Closely located to this cluster was a circle representing human monocytes stimulated by normal pooled plasma spiked with N. meningitidis. Taken together, the clustering suggests that N. meningitidis LPS is most likely the main activator of gene expression in human monocytes induced by meningococcal sepsis plasma after immunodepletion of IL-10.
PCA of the gene expression profiles induced in human monocytes. The PCA displays the clustering of samples inducing gene expression (FC ≥ 1.5, P-value < 0.01) in the experimental model. Each circle represents an individual sample, and the distance between them represent differences in the gene expression patterns. Blue circles are induced by meningococcal sepsis plasma after immunodepletion of IL-10; red circles are induced by meningococcal sepsis plasma containing LPS and IL-10; purple circles are induced by plasma from patients with mild meningococcemia containing low levels of IL-10 and LPS; orange circles are induced by plasma from patients with mild meningococcemia after immunodepletion of IL-10. In addition, the experimental model included gene expression induced by four control samples; the pink circle was induced by pooled normal plasma; the green circle was induced by recombinant IL-10; the brown circle was induced by pooled normal plasma containing N. meningitidis and recombinant IL-10; the light blue circle was induced by pooled normal plasma containing N. meningitidis. The percentage value describes the proportion of the total variance described by each principal component (PC) axis (PC1, PC2 and PC3). The first three PCs explain 63.6% of the variance in the data set.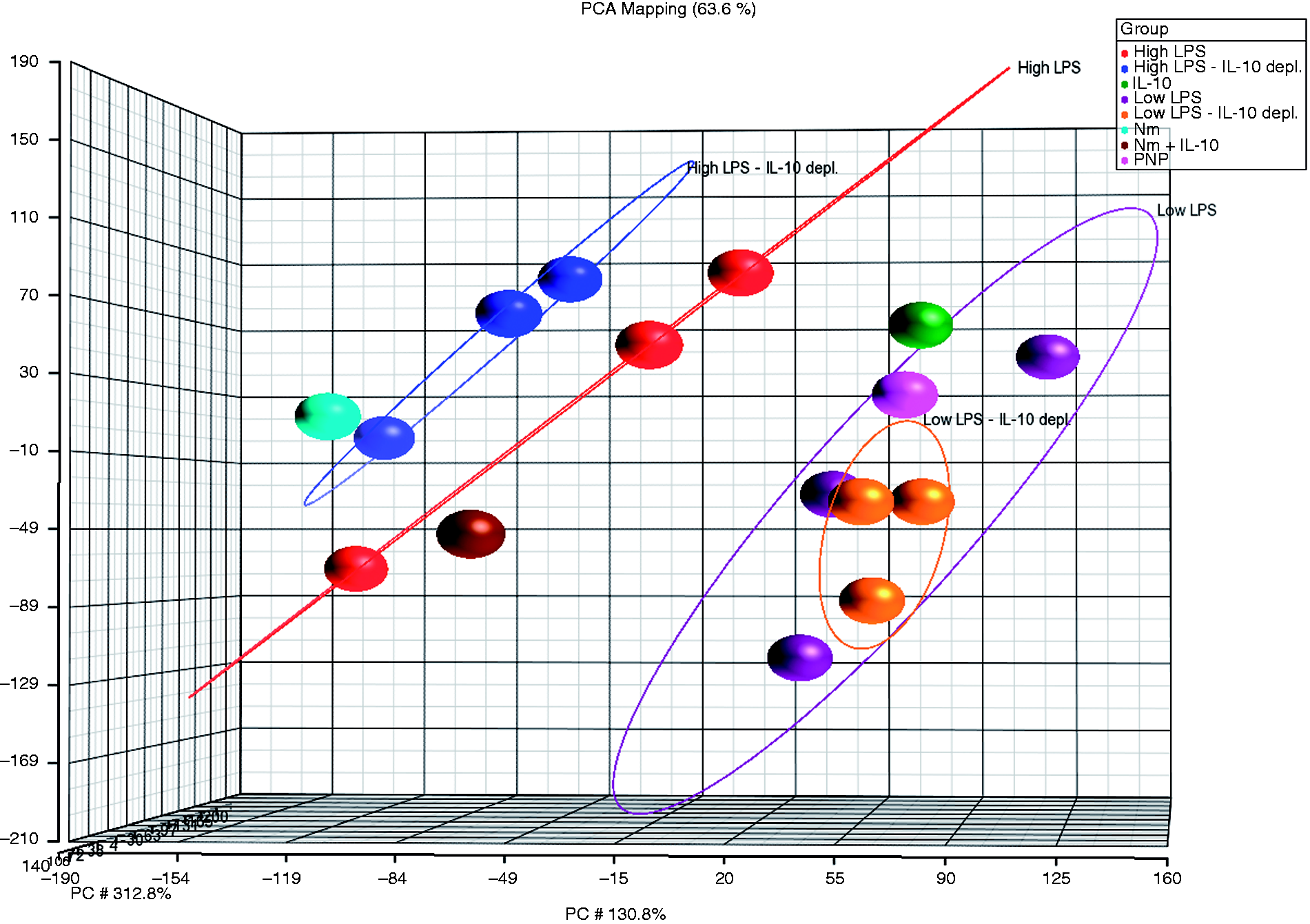
A second cluster of red circles, located closer to the baseline cluster, were samples inducing gene expression before immunodepletion of IL-10. Closely located to this cluster was a brown circle representing gene expression in human monocytes stimulated by normal pooled plasma spiked with N. meningitidis and IL-10. Thus, the expression pattern induced by patient plasma prior to immunodepletion of IL-10 closely resembled gene expression induced by the model system containing only N. meningitidis and IL-10. Taken together, this clustering suggests that in meningococcal sepsis plasma the dominant anti-inflammatory effect on gene expression is induced by IL-10. In these respective clusters, the location of one circle differed slightly from two other circles with corresponding levels of LPS and IL-10. These two circles, blue and red respectively, represented gene expression induced by the patient who survived despite high levels of meningococcal LPS. It suggests that plasma from this patient induces genes with slightly different expression patterns compared with plasmas from the non-surviving patients 1 and 2. One possible explanation might be the lower level of LPS in this plasma sample compared with the plasmas from the two patients who died from fulminant meningococcal septicemia. However, the PCA plot demonstrates that the gene expression pattern induced by this plasma, with high but not lethal LPS levels, is markedly different from the gene expression induced by the plasma from patients with mild meningococcemia, suggesting that this plasma has a strong pro-inflammatory content. Overall, the PCA plot demonstrates that global gene expression patterns induced in human monocytes cluster according to the levels of LPS and IL-10 in the plasma from patients with systemic meningococcal disease. Furthermore, the plots suggest that N. meningitidis LPS and IL-10 appear to be the principal inducers of gene expression in human monocytes incubated with meningococcal sepsis plasma.
Descriptive and functional analysis of the gene expression regulated in human monocytes by patient plasma containing IL-10 and IL-10 immunodepleted patient plasma
Genes differentially expressed in human moncytes by patient plasma containing IL-10 and IL-10 immunodepleted patient plasma. a
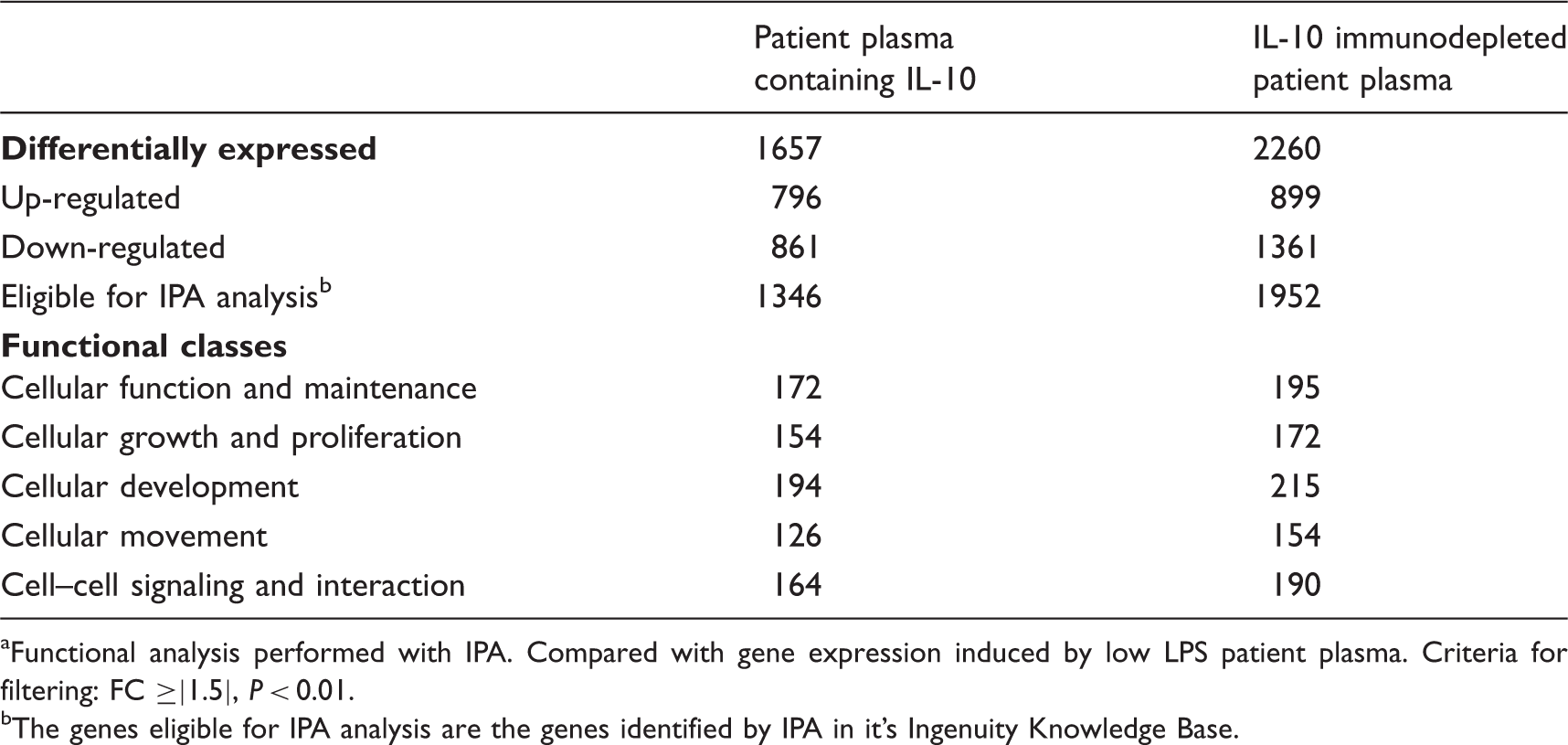
Functional analysis performed with IPA. Compared with gene expression induced by low LPS patient plasma. Criteria for filtering: FC ≥|1.5|, P < 0.01.
The genes eligible for IPA analysis are the genes identified by IPA in it’s Ingenuity Knowledge Base.
A total of 373 genes were regulated by FC ≥|1.5| after immunodepletion of IL-10, and gene network analysis identified several potential targets of the IL-10 anti-inflammatory response
By directly comparing the gene expression profiles induced before and after immunodepletion of IL-10, we identified 373 genes (121 up-regulated, 252 down-regulated) to be altered by FC ≥|1.5| (P < 0.01). This suggests that the presence of IL-10 in patient plasma down-regulated 121 genes and up-regulated 252 genes by FC ≥|1.5| (see online Supplementary Material). These 373 genes are displayed in a heatmap (Figure 2). IPA identified 322 (230 up-regulated, 92 down-regulated) of these to be eligible for analysis. The molecular and cellular functions most significantly associated with these genes were cellular function and maintenance (47 genes), cellular development (52 genes), cellular growth and proliferation (40 genes), cell–cell signaling (42 genes), and interaction and cellular movement (28 genes) (Table 4), demonstrating the breadth of functions potentially affected by IL-10 in human monocytes stimulated by meningococcal sepsis plasma. Using IPA, we generated gene networks displaying the interconnection between the genes and their associated functions. Two networks with the significant scores of 55 and 24 were generated (Table 5). Gene network 1 contained a total of 140 genes, and 60 of these were regulated by patient plasma containing IL-10 compared with IL-10 immunodepleted plasma. Functional classes associated with this network were cell–cell signaling and interaction, hematological system development and function, and immune cell trafficking (Table 5). Of particular interest was the regulation of genes encoding ligand receptors and transcription factors that could increase our understanding of the anti-inflammatory response mediated by IL-10. In network 1 we identified LILRA2, LILRA5 and LILRB3, which are genes belonging to the leukocyte immunoglobulin-like receptor (LILRs) family, to be up-regulated in patient plasma containing IL-10 compared with IL-10 immunodepleted plasma.30,31 These genes have also previously been identified to be up-regulated by IL-10, and we examined this group of genes in more detail (see below).18,22 Furthermore, we identified the differential expression of transcription factors, where several are known to regulate cytokines that are inhibited by IL-10. The gene REL, demonstrated by others to decrease expression of IL-12 (p35 sub-unit) and IL-23A when knocked out in mice, was down-regulated by the presence of IL-10 in meningococcal sepsis plasma (FC = −1.7).32,33 DUSP10, down-regulated (FC = −1.7), is a transcriptional regulator if knocked out in mice results in increased production of IL-1 and TNF-α.
34
While down-regulation of REL is consistent with the primary role of IL-10 as an inhibitor of cytokine release, down-regulation of DUSP10 is not. Other transcriptional regulators identified to be up-regulated by the presence of IL-10 were BATF (FC = 3.4), CITED2 (FC = −1.6) and VAV1 (FC = 2.9). BATF is a transcription factor required for differentiation of Th17 cells (IL-17-producing helper T cells) and T-follicular helper cells, while CITED2 knock-out attenuates the pro-inflammatory NF-kB response.35,36 VAV1 was identified to be up-regulated by IL-10 in a previous study.
18
Up-regulation of VAV1 by IL-10 in meningococcal sepsis plasma is consistent with a recent study that demonstrated VAV1 to be a down-regulator of IL-6 transcription in human macrophages.
37
The heatmap, generated by hierarchical clustering in Partek Genomics Suite, displays the 373 genes differentially regulated (FC ≥|1.5|, P < 0.01) by patient plasma containing IL-10 compared with IL-10 immunodepleted plasma. Red indicates up-regulation, while blue indicates down-regulation. The blue panels display genes induced in human monocytes by patient plasma with high levels of N. meningitidis LPS after immunodepletion of IL-10, green panels display genes induced by patient plasma with high levels of N. meningitidis LPS before immunodepletion of IL-10 and red panels display genes induced by patient plasma with low levels of N. meningitidis LPS. Heatmap cluster analysis revealed the gene expression profile induced by patients 4–6 after immunodepletion of IL-10 to be similar to expression patterns before immunodepletion of IL-10, and these panels are therefore not included in the heatmap. Functional classes associated with genes regulated in human monocytes by patient plasma containing IL-10 compared with IL-10 immunodepleted patient plasma.a Gene networks in human monocytes displaying the relationships between genes regulated in patient plasma containing IL-10 compared with IL-10 immunodepleted patient plasma.a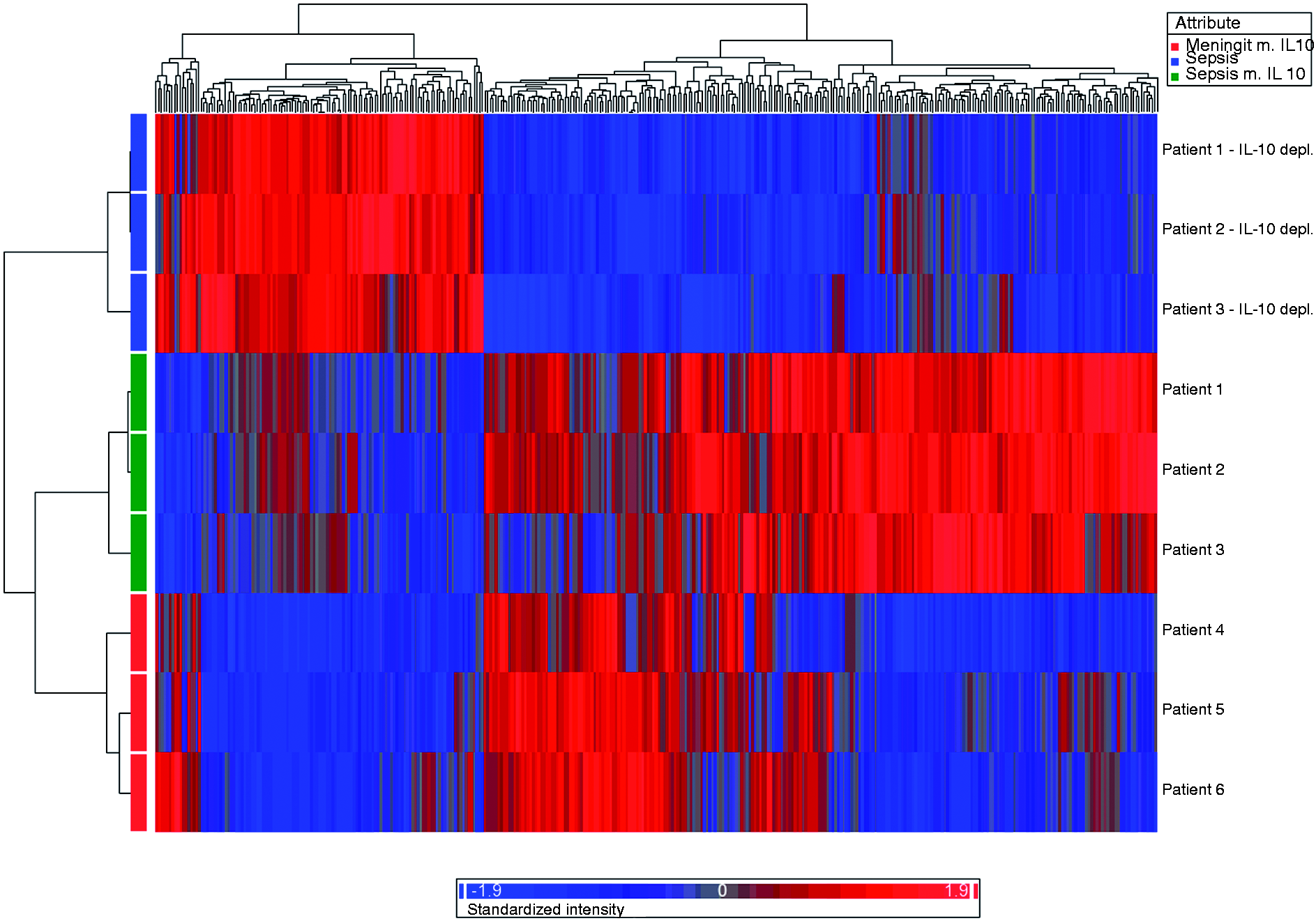
Network II contained 140 genes, and 38 of these were regulated by patient plasma containing IL-10 compared with IL-10 immunodepleted plasma. The network was associated with cellular development, cellular growth and proliferation, and hematological system development and function (Table 5). Transcription regulators with altered expression after immunodepletion of IL-10 in this network were ARID5A, CARM1, CITED2, E2F7, PBX3 and KAT2B. ARID5A is up-regulated in macrophages in response to LPS, and knockout of ARID5A results in reduced release of IL-6 from LPS-stimulated mice. 38 In our study, ARID5A was up-regulated by the presence of IL-10 (FC = 1.7), which is inconsistent with the primary role of IL-10 as an inhibitor of cytokine release. CARM1 was up-regulated by the presence of IL-10 (FC = 2.4), while E2F7 (FC = −2.8), KAT2B (FC = −2.5) and PBX3 (FC = −1.9) were down-regulated. The potential role of these transcription factors in the induction of genes and cytokines in response to LPS is, to our knowledge, not well described. Both Network I and Network II included the plasma membrane kinase RIPK2 (also known as RICK), which was down-regulated (FC = −2.8) in the presence of IL-10. Consistent with our previous study, this finding suggests that the presence of IL-10 in patient plasma decreases RIPK2-expression in human monocytes. 18 Previous research has identified RIPK2 as a target for regulating TLR signaling, as inhibition reduces the amount of cytokines released from human monocytes following TLR4 stimulation.33,34 Furthermore, RIPK2-deficient mice exposed to LPS prior to infection with Pseudomonas aeruginosa have reduced production of pro-inflammatory cytokines and reduced susceptibility to infection than wild type mice. 39 These studies suggest that inhibition of RIPK2 dampens the pro-inflammatory response, which may be one of many mechanisms by which IL-10 attempts to protect the host against the pro-inflammatory response induced by LPS.
Overall, the generated gene networks display the complex relationship between the genes regulated by the presence of IL-10 in meningococcal plasma, involving a number of functions and signaling pathways. The direction of altered gene expression was, for some regulators, such as REL, RIPK2 and VAV1, consistent with the role of IL-10 as an anti-inflammatory cytokine. However, for other genes, such as DUSP10 and ARIDA5, the direction of regulation was inconsistent with this role. This underscores the necessity of investigating such findings in different model system, and the carrying out of future functional experiments to determine the biological significance of the suggested regulation of these transcriptional regulators. One of the aims of this study was to investigate whether signaling pathways identified to be regulated by N. meningitidis and recombinant IL-10 in our previous in vitro model studies is regulated in human monocytes stimulated by meningococcal sepsis plasma containing IL-10.17,18 We therefore explored in more detail the differential expression of genes associated with TLR signaling, the type I IFN signaling pathway, the LILR family and the inflammasome. In addition, we investigated expression of genes related to coagulation and fibrinolysis owing to its critical role in the pathophysiology of meningococcal sepsis.
Immunodepletion of IL-10 from patient plasma results in down-regulation of CD14 and MD2
Regulation of genes related to the TLR signaling pathway in human monocytes stimulated by patient plasma containing IL-10 and IL-10 immunodepleted patient plasma.
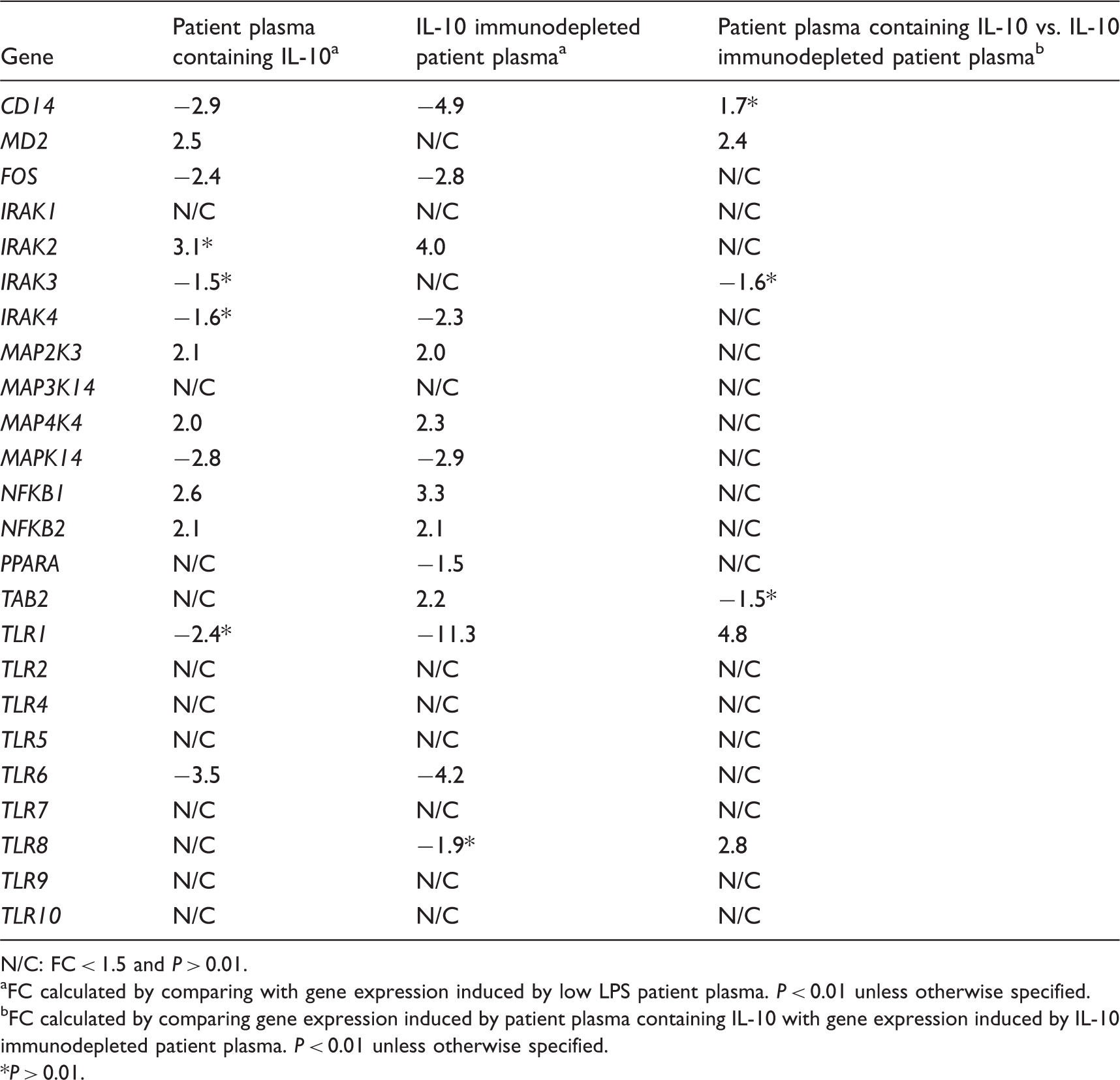
N/C: FC < 1.5 and P > 0.01.
FC calculated by comparing with gene expression induced by low LPS patient plasma. P < 0.01 unless otherwise specified.
FC calculated by comparing gene expression induced by patient plasma containing IL-10 with gene expression induced by IL-10 immunodepleted patient plasma. P < 0.01 unless otherwise specified.
P > 0.01.
Immunodepletion of IL-10 from patient plasma results in down-regulation of IFN-induced proteins with tetratricopeptide repeat (IFIT gene family)
Regulation of genes related to the type I IFN signaling pathway in human monocytes stimulated by patient plasma containing IL-10 and IL-10 immunodepleted patient plasma.
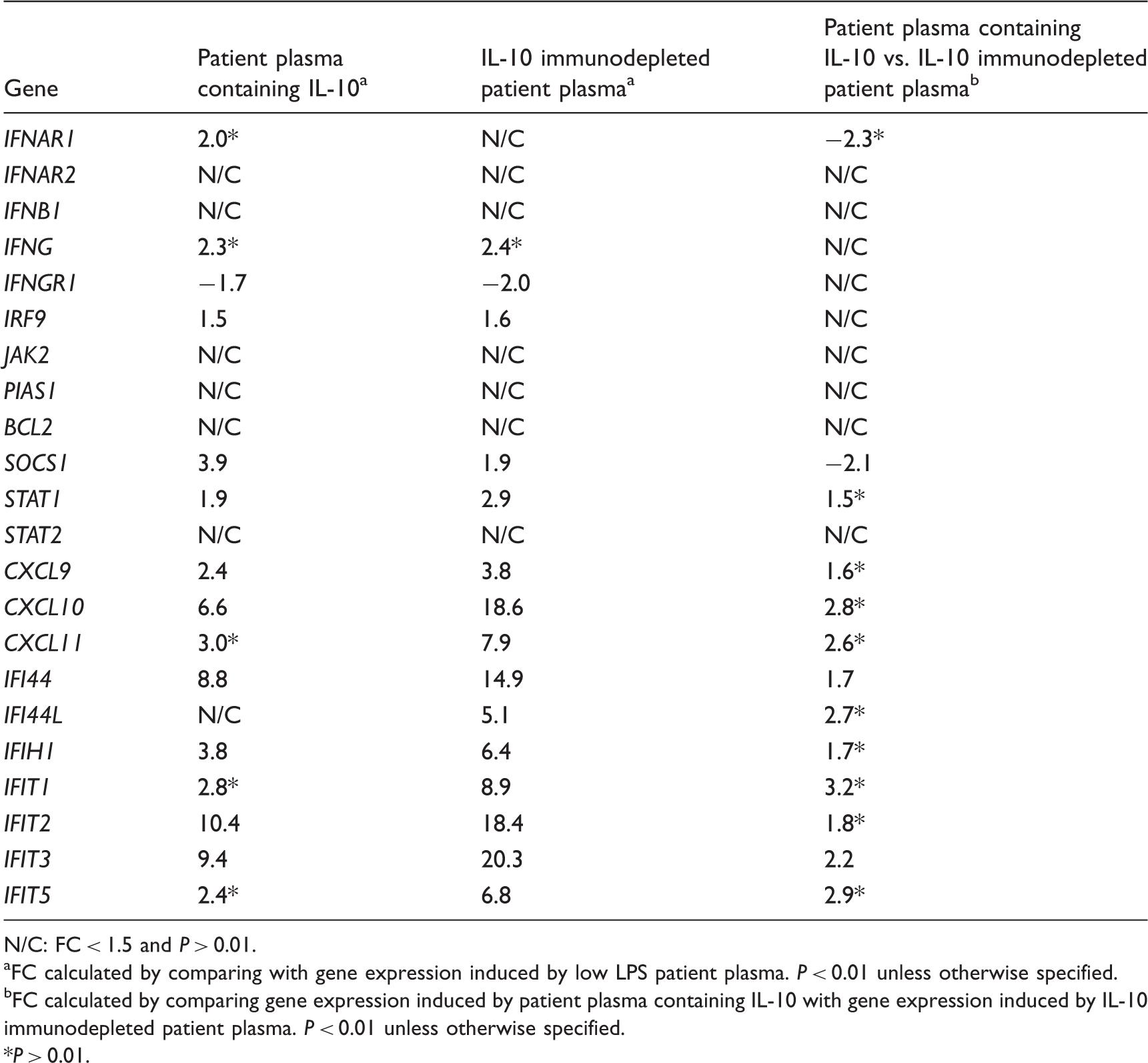
N/C: FC < 1.5 and P > 0.01.
FC calculated by comparing with gene expression induced by low LPS patient plasma. P < 0.01 unless otherwise specified.
FC calculated by comparing gene expression induced by patient plasma containing IL-10 with gene expression induced by IL-10 immunodepleted patient plasma. P < 0.01 unless otherwise specified.
P > 0.01.
We then investigated the regulation of the type I IFN inducible genes CXCL9, CXCL10, CXCL11, IFIT1, IFIT2, IFIT3, IFIT5, IFI44, IFI44L and IFIH1, which a previous study from our group had demonstrated to be up-regulated by wild type N. meningitidis and purified LPS. 17 In the present study, these genes were up-regulated by patient plasma containing IL-10. Immunodepletion of IL-10 resulted in increased up-regulation of all the genes, and IFIT3 and IFI44 were found to be significantly up-regulated (P < 0.01) when comparing the gene expression levels induced before and after immunodepletion of IL-10. IFIT1, IFIT2, IFIT3 and IFIT5 belong to the type I IFN-stimulated genes known as IFN-induced proteins with a tetratricopeptide repeat (IFIT gene family), which have an antiviral function. 45 Our results suggest that the presence of IL-10 in meningococcal sepsis plasma inhibits the IFIT gene family. Experiments in IFN-β-deficient mice suggest that IFN-β is an essential effector of lethality induced by LPS, as mice lacking IFN-β showed increased resistance to high doses of LPS. 46 Inhibition of IFN-β-induced gene expression may thus be one of the many anti-inflammatory mechanisms of IL-10 during sepsis. However, the relevance of this finding to the pathophysiology of meningococcal disease, as well as potential consequences for the pathophysiology of diseases caused by viral infections, requires further investigation in human cell models.45,47
Immunodepletion of IL-10 up-regulated genes in the LILR family
Regulation of genes related to the immunoglobulin-like receptor family in human monocytes stimulated by patient plasma containing IL-10 and IL-10 immunodepleted patient plasma.
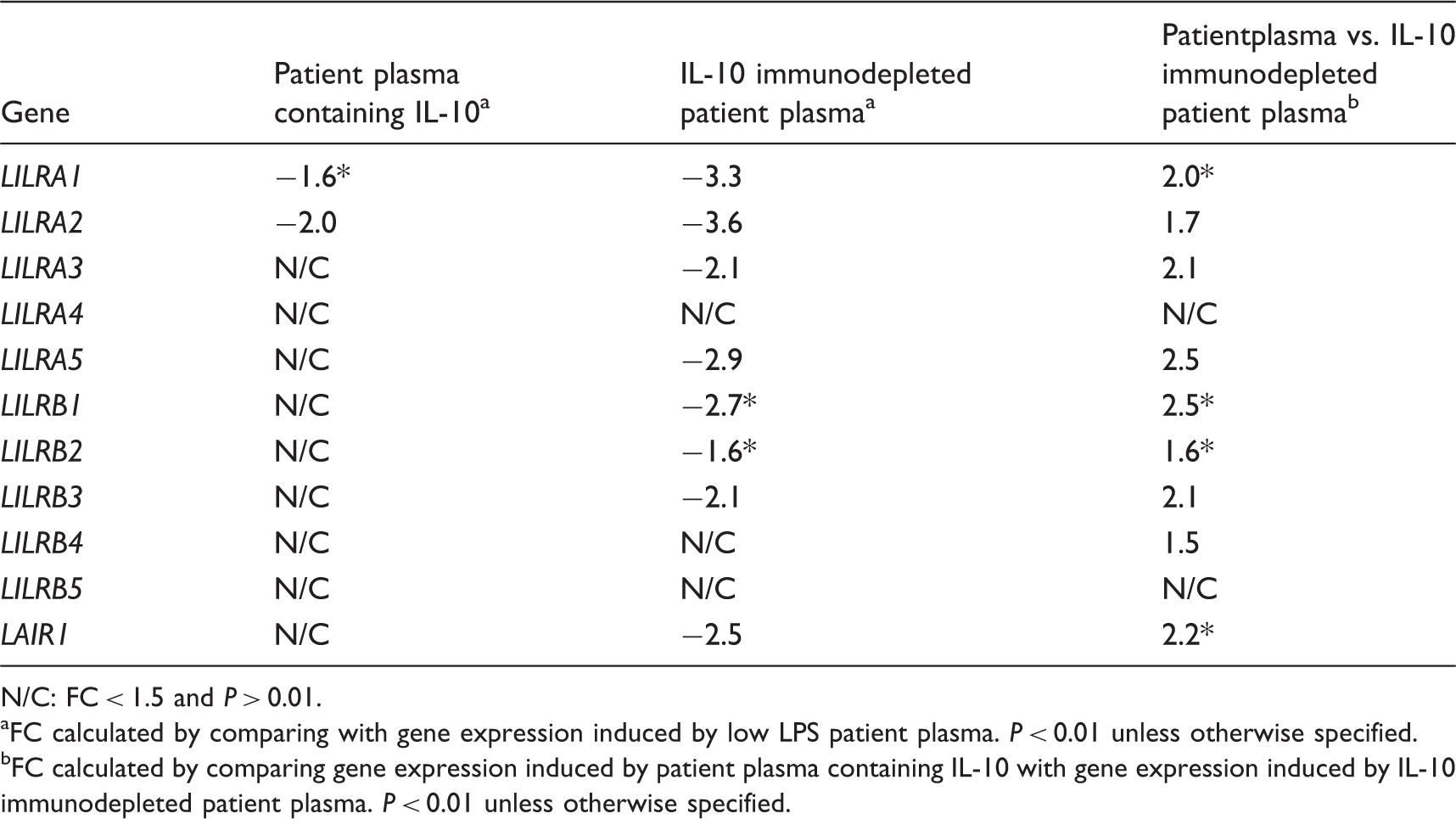
N/C: FC < 1.5 and P > 0.01.
FC calculated by comparing with gene expression induced by low LPS patient plasma. P < 0.01 unless otherwise specified.
FC calculated by comparing gene expression induced by patient plasma containing IL-10 with gene expression induced by IL-10 immunodepleted patient plasma. P < 0.01 unless otherwise specified.
Immunodepletion of IL-10 from patient plasma down-regulated NLRP3 and up-regulated NLRC4 and PYCARD
Regulation of genes related to the inflammasome in human monocytes stimulated by patient plasma containing IL-10 and IL-10 immunodepleted patient plasma.
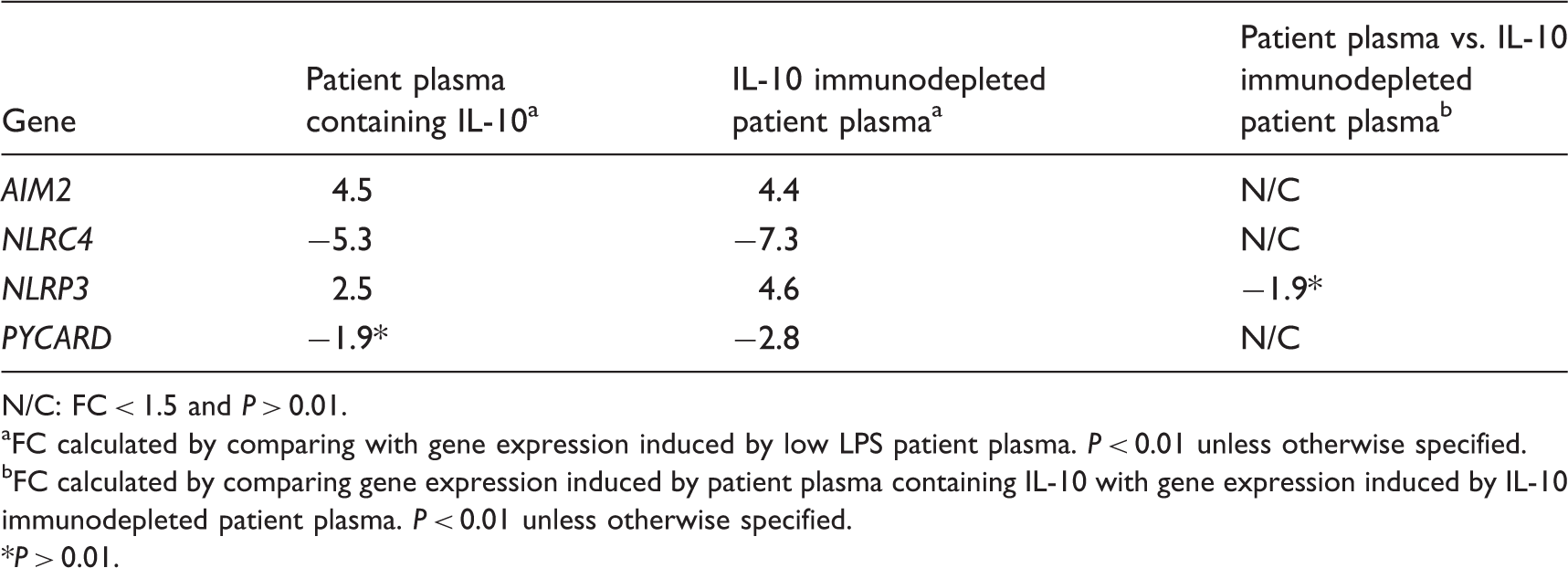
N/C: FC < 1.5 and P > 0.01.
FC calculated by comparing with gene expression induced by low LPS patient plasma. P < 0.01 unless otherwise specified.
FC calculated by comparing gene expression induced by patient plasma containing IL-10 with gene expression induced by IL-10 immunodepleted patient plasma. P < 0.01 unless otherwise specified.
P > 0.01.
Immunodepletion of IL-10 from patient plasma increased the expression of tissue factor and mediators of fibrinolysis
Genes related to the coagulation system and fibrinolysis induced in human monocytes by patient plasma containing IL-10 and IL-10 immunodepleted patient plasma.
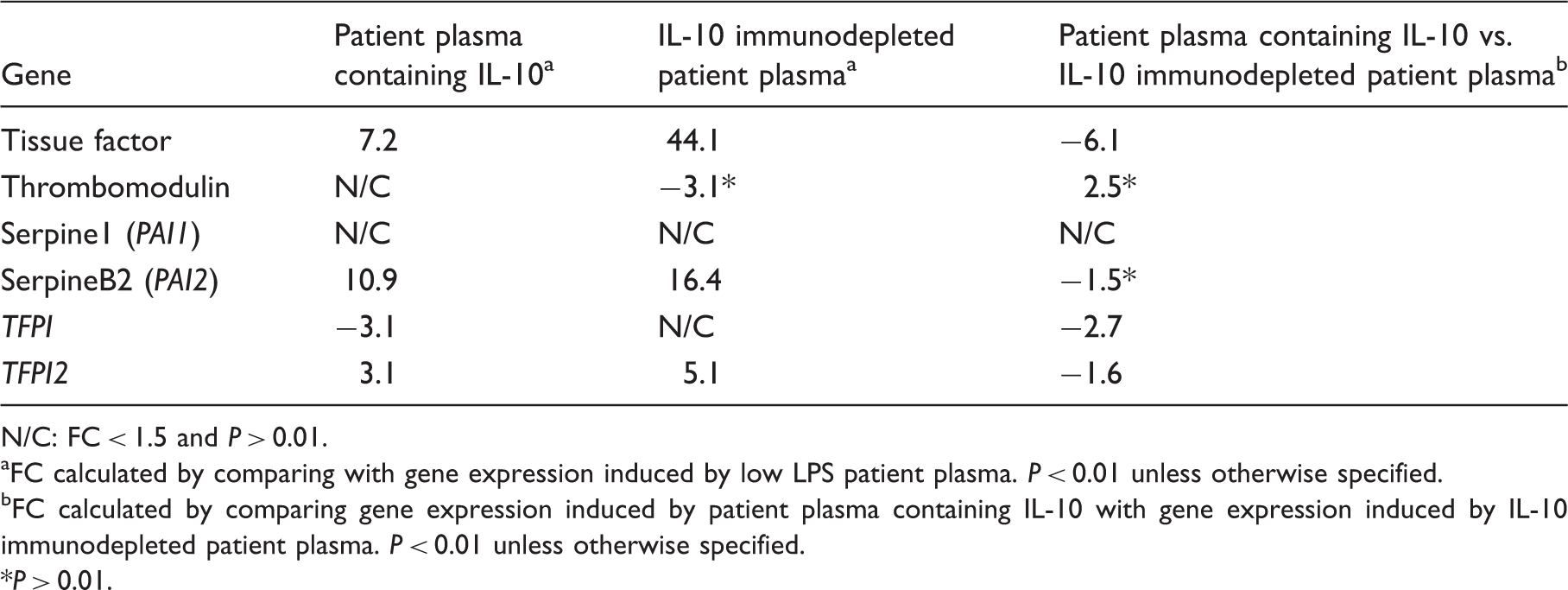
N/C: FC < 1.5 and P > 0.01.
FC calculated by comparing with gene expression induced by low LPS patient plasma. P < 0.01 unless otherwise specified.
FC calculated by comparing gene expression induced by patient plasma containing IL-10 with gene expression induced by IL-10 immunodepleted patient plasma. P < 0.01 unless otherwise specified.
P > 0.01.
Human monocytes are the primary immune cell exerting pro-coagulant activity. This pro-coagulant activity is largely mediated by tissue factor expressed on the cell membrane, as well on tissue factor-containing microparticles derived from the monocytes and may contribute to DIC in severe meningococcal disease.59,64,65 The strongly increased up-regulation of tissue factor after immunodepletion of IL-10 indicated that IL-10 in patient plasma had a potent inhibitory effect on tissue factor expression in human monocytes. IL-10-mediated inhibition of pro-coagulant activity of human monocytes has previously been demonstrated in other cell models, as well as by our own group.63,66–68 Our present gene expression findings considered together with the previous studies suggest that the presence of IL-10 in meningococcal sepsis plasma may partially inhibit the overwhelming pro-coagulant activity of human monocytes contributing to DIC, and that without IL-10 the patients would have suffered from an even more pronounced DIC.
Predicted decreased activation of mononuclear leukocytes and antigen presentation in patient plasma containing IL-10 compared with IL-10 immunodepleted plasma
Immune cell functions predicted by IPA to be decreased in human monocytes stimulated with patient plasma containing IL-10 compared with IL-10 immunodepleted patient plasma.a
Validation of gene expression results with real-time PCR
qRT-PCR and microarray data for selected differentially expressed genes in human monocytes incubated by patient plasma containing IL-10 and IL-10 immunodepleted patient plasma.a
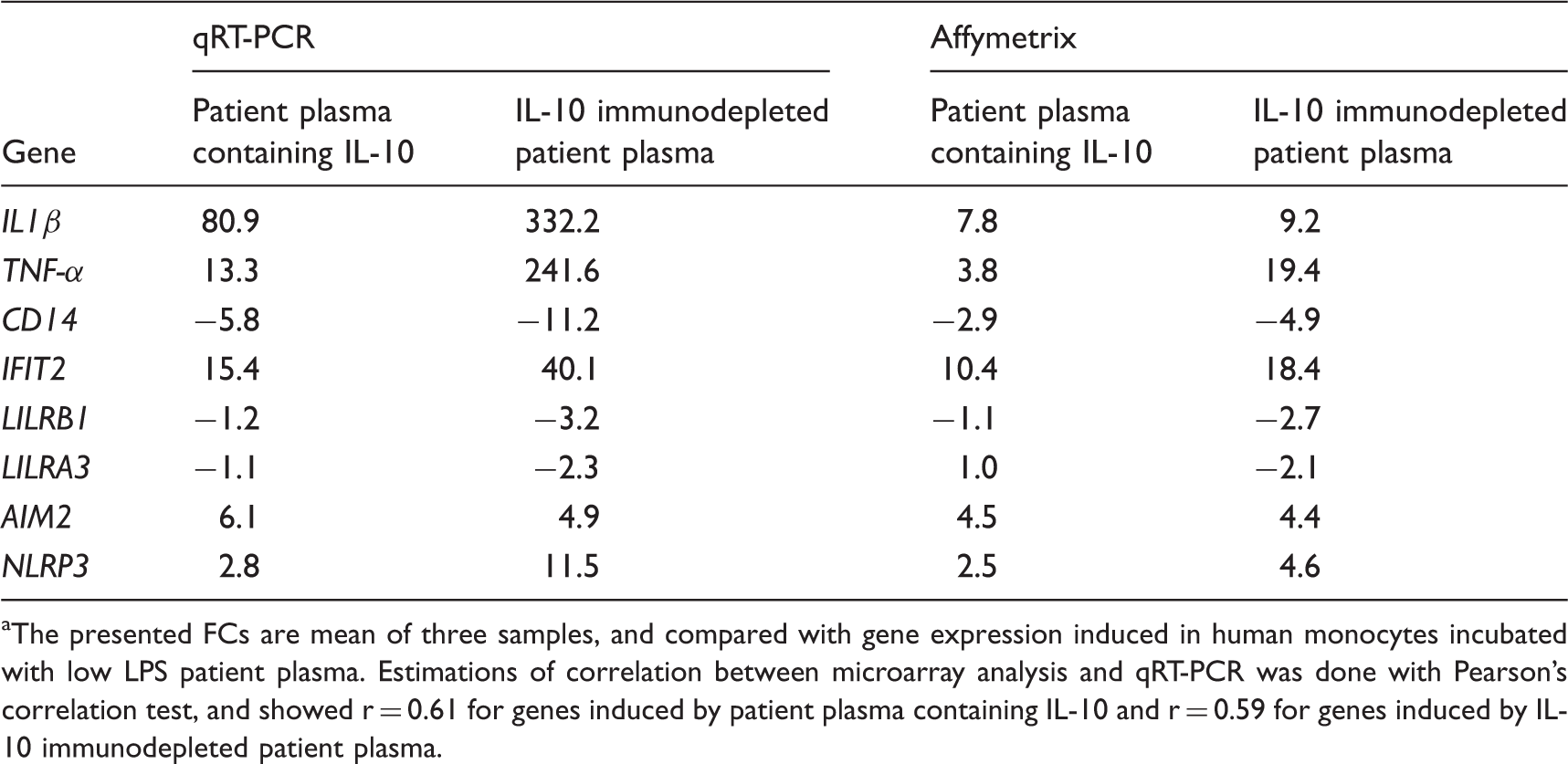
The presented FCs are mean of three samples, and compared with gene expression induced in human monocytes incubated with low LPS patient plasma. Estimations of correlation between microarray analysis and qRT-PCR was done with Pearson's correlation test, and showed r = 0.61 for genes induced by patient plasma containing IL-10 and r = 0.59 for genes induced by IL-10 immunodepleted patient plasma.
Conclusion
By using an immunoprecipitation technique to remove IL-10 from plasma from patients with meningococcal sepsis, we were able to study the induction of genes in human monocytes when incubated with such plasmas and to explore the effects exercised by IL-10. We suggest our in vitro patient plasma model more closely reflects the gene expression changes that occur during the initial stage of severe human endotoxemia than previously published studies.18–22 Thus, we believe that the present study adds knowledge to how the intense pro- and anti-inflammatory responses in a septic patient may alter gene expression. However, our data are descriptive, and future studies should investigate the functional significance suggested by these gene expression findings. We conclude that in human monocytes stimulated by meningococcal sepsis plasma containing high levels of LPS, the presence of IL-10 down- and up-regulated genes associated with a range of functions and signaling pathways, leading to an overall inhibitory effect on the inflammatory response induced by the patient plasma.
Footnotes
Funding
The research was carried out with financial support from the Research Council of Norway, as part of the Medical Student Research Program.
Conflicts of interest
The authors do not have any potential conflicts of interest to declare.
Acknowledgements
We thank the following persons for excellent technical assistance: Anne Marie Siebke Trøseid, Blood Cell Research Group, Section for Research, Department of Medical Biochemistry, for thawing human monocytes for the experiments; and Berit Nyland and Ernst Arne Höiby, National Institute of Public Health, Oslo, Norway, for providing cultures of the reference strain N. meningitidis 44/76.
