Abstract
Herpes simplex virus (HSV) is a human pathogen that can cause severe diseases such as encephalitis, keratitis and neonatal herpes. Control of HSV infection may be achieved by using small interfering (si)RNAs. We have designed and enzymatically produced pools of siRNAs targeting HSV. In addition to the target-specific effects, such siRNAs may induce innate immunity responses that may contribute to antiviral effects. HSV has versatile ways of modulating innate immunity, and it remains unclear whether HSV-specific antiviral treatment would benefit from the potential immunostimulatory effects of siRNAs. To address this, cell lines derived from epithelium and nervous system were studied for innate immunity reactions to HSV infection, to siRNA treatment, and to a combination of treatment and infection. In addition, the outcome of HSV infection was quantitated. We show that innate immunity reactions vary drastically between the cell lines. Moreover, our findings indicate only a minimal relation between the antiviral effect and the treatment-induced innate immunity responses. Thus, the antiviral effect is mainly sequence specific and the inhibition of HSV infection is not ascribed to the slight innate immunity induction.
Introduction
Herpes simplex viruses type 1 and 2 (HSV-1 and HSV-2) are ubiquitous human pathogens that cause clinically important diseases such as genital herpes, neonatal herpes, severe ocular infections, meningitis and herpes encephalitis. 1 HSV is capable of modulating innate immunity and escapes adaptive immunity. HSV infection induces the expression of IFN genes,2–5 and HSV is also able to evade and modulate innate immunity by several mechanisms.4,6,7 RNA interference (RNAi) using single-site, small interfering (si)RNAs has already been applied for treatment of HSV infection in cultured cells.8–11 A combination of two single-site siRNAs, targeting the UL29 gene (encoding the essential HSV DNA binding protein ICP8), has enhanced the survival of infected animals in a mouse model of genital herpes. 12 The possible induction of innate immunity by the siRNAs and the consequent effects on the infection control remain unknown.
siRNAs.
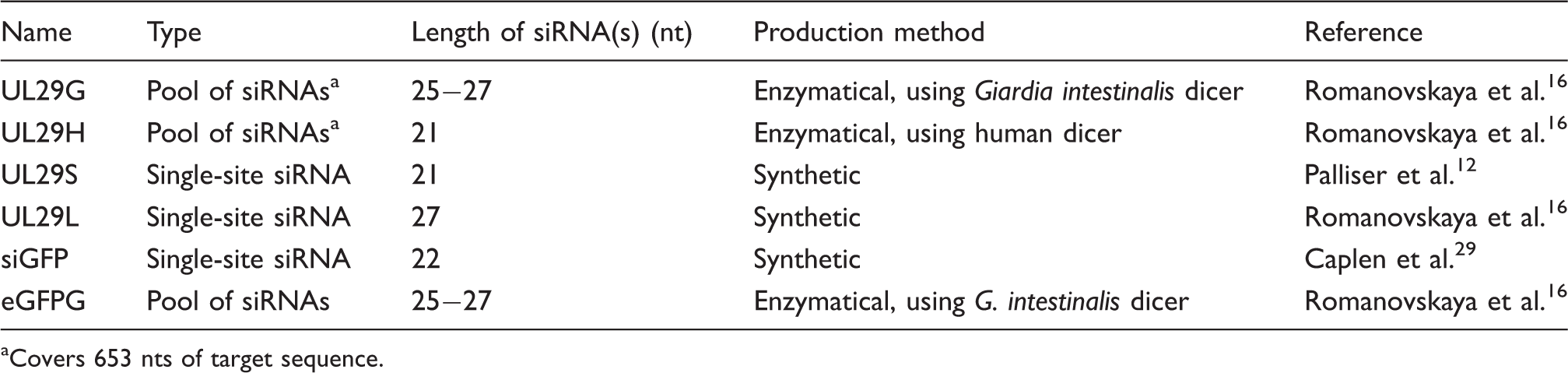
Covers 653 nts of target sequence.
To create pools of siRNAs, we have applied RNA polymerases of enterobacteria phage T7 and Pseudomonas phage phi6 for the enzymatic production of high-quality, double-stranded (ds)RNA for RNAi.14–16 The produced target-specific dsRNA molecules were cleaved in vitro to pools of canonical 21-nucleotide (nt)-long siRNAs or 27-nt-long dicer–substrate siRNAs using human or Giardia intestinalis dicers, respectively. Upon introduction into mammalian cells, the dicer–substrate siRNAs are further processed to 21-nt molecules by the endogenous dicer. 17 Consequently, their effects might differ from the canonical 21-nt siRNAs in RNAi applications. 17 Furthermore, we earlier utilized the enzymatically produced siRNA pools against coxsackie viruses and obtained a broader inhibition of related viral strains than with a single-site siRNA. 15 We have recently studied the inhibition of HSV infection in cell cultures using dicer–substrate and canonical siRNA pools, and found that the siRNA pools do not compromise cell viability in the efficient antiviral concentrations. 16 However, we did observe weak innate responses to the RNA species utilized. 16
Here, we investigated various siRNA preparations for the induction of innate immunity and for their efficacy in inhibiting HSV infection. We describe the innate immunity reactions induced by the siRNAs, and the immunomodulatory capabilities of HSV in cells originating from epithelium and the nervous system. Viral shedding and HSV gene expression profiles in siRNA-treated and untreated cells were characterized.
Materials and methods
Cell lines and virus
Human glioblastoma–astrocytoma U373MG cells (U-251 MG; ATCC, Manassas, VA, USA) were maintained in high-Glc DMEM (Gibco, Carlsbad, CA, USA) supplemented with 2 mM
dsRNA and siRNA molecules
Primers and siRNA sequences.
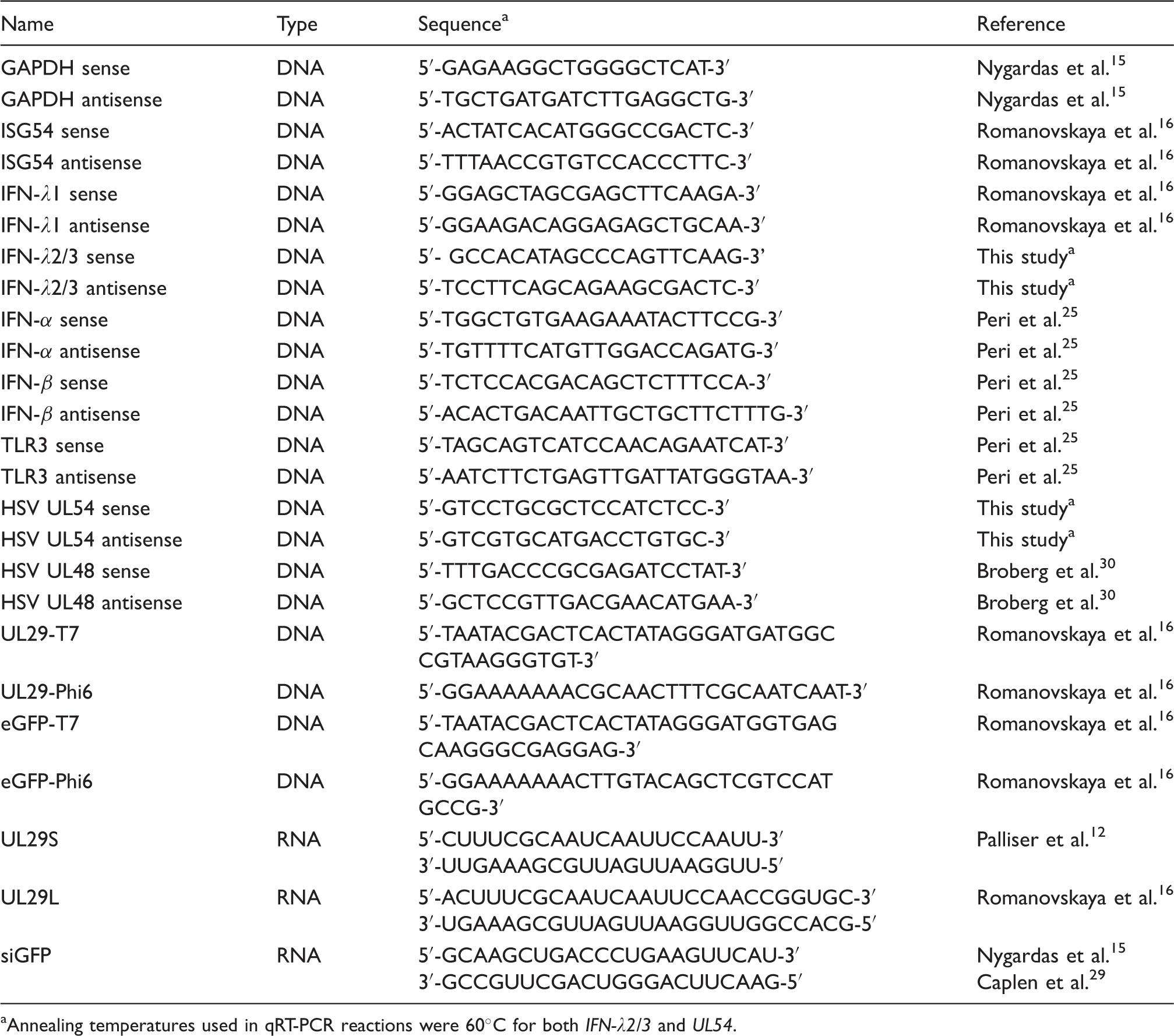
Annealing temperatures used in qRT-PCR reactions were 60℃ for both IFN-λ2/3 and UL54.
Transfection and infection
RNA or water transfections of U373MG and HaCaT cells, propagated in 96-well plates, were carried out using Lipofectamine RNAiMAX (Invitrogen) and the manufacturer’s forward transfection protocol. All the experiments were performed 2–5 times (n = 3–6 in each experiment).
Four h after transfection, the cells were challenged with a 100 -µl dose of HSV-1(17+) containing 1000 plaque forming units (pfu) per well. In the case of uninfected cells, they were subjected to the same treatment, excluding the virus. Prior to infection, the cells were washed twice with RPMI 1640 (Gibco) supplemented with 0.1% BSA. Cells were washed three times and covered with 200 µl of culture medium per well at 60–90 min p.i. At 48 h post-transfection (44 h p.i.) the cells were collected in TRI reagent (MRC, Cincinnati, OH, USA) for RNA extraction. Culture medium was collected prior to this and used to determine the amount of released virus by plaque titration on Vero cells.
Quantitative real-time PCR
Total cellular nucleic acid was isolated from cells using TRI reagent (MRC) according to manufacturer’s instructions. After treatment with DNase (Fermentas, Thermo Scientific), the total RNA was reverse transcribed into complementary DNA (cDNA) using RevertAid H Minus Reverse Transcriptase (Fermentas) and random hexamer primers (Fermentas). For the detection of selected mRNAs (viral or host) the produced cDNA were amplified using Maxima SYBR Green/ROX qPCR Master Mix (Fermentas). Quantitative real-time (qRT)-PCR was performed with a Rotor-Gene Q real-time instrument (Qiagen) as described previously. 20 The sequences of the primers are shown in Table 2. All results from the qRT-PCR were normalized to the housekeeping gene, GAPDH, copy number in the same sample.
Computational analyses
For statistical analyses, SPSS Statistics 20 (IBM, Armonk, NY, USA) software was used. Statistical significances were calculated with non-parametric Mann–Whitney U-test and the threshold was set to P < 0.05.
Results and discussion
In this study, we describe the effects of HSV infection on the cellular innate responses to enzymatically produced antiviral siRNA pools and chemically synthesized single-site siRNAs targeting the essential UL29 gene of HSV-1. We studied the possible correlation between the antiviral efficacy of siRNAs and their induction of innate immunity. We applied two size classes of siRNAs—21-nt-long and 27-nt-long (Table 1)—and used two cell lines, the astrocytoma-glioblastoma U373MG, representing a host cell of HSV infection in the central nervous system, and the HaCaT keratinocyte cell line, representing an epithelial type of host cell of HSV-1. Both cell lines produced progeny virus with similar efficiency (Figure 1).
Inhibition of viral shedding with siRNAs. U373MG and HaCaT cells, grown on 96-well plates, were challenged with 1000 pfu of HSV-1 (17+). The amount of infectious virus shed into the culture medium was determined by plaque titration at 44 h p.i. The titers are presented on a logarithmic scale. Mean + SD of six replicates are shown. Significances are shown for the groups of comparison. Non-parametric Mann–Whitney U-test was used to determine significances. **P < 0.01; and against all other treatments #P < 0.05, ##P < 0.01.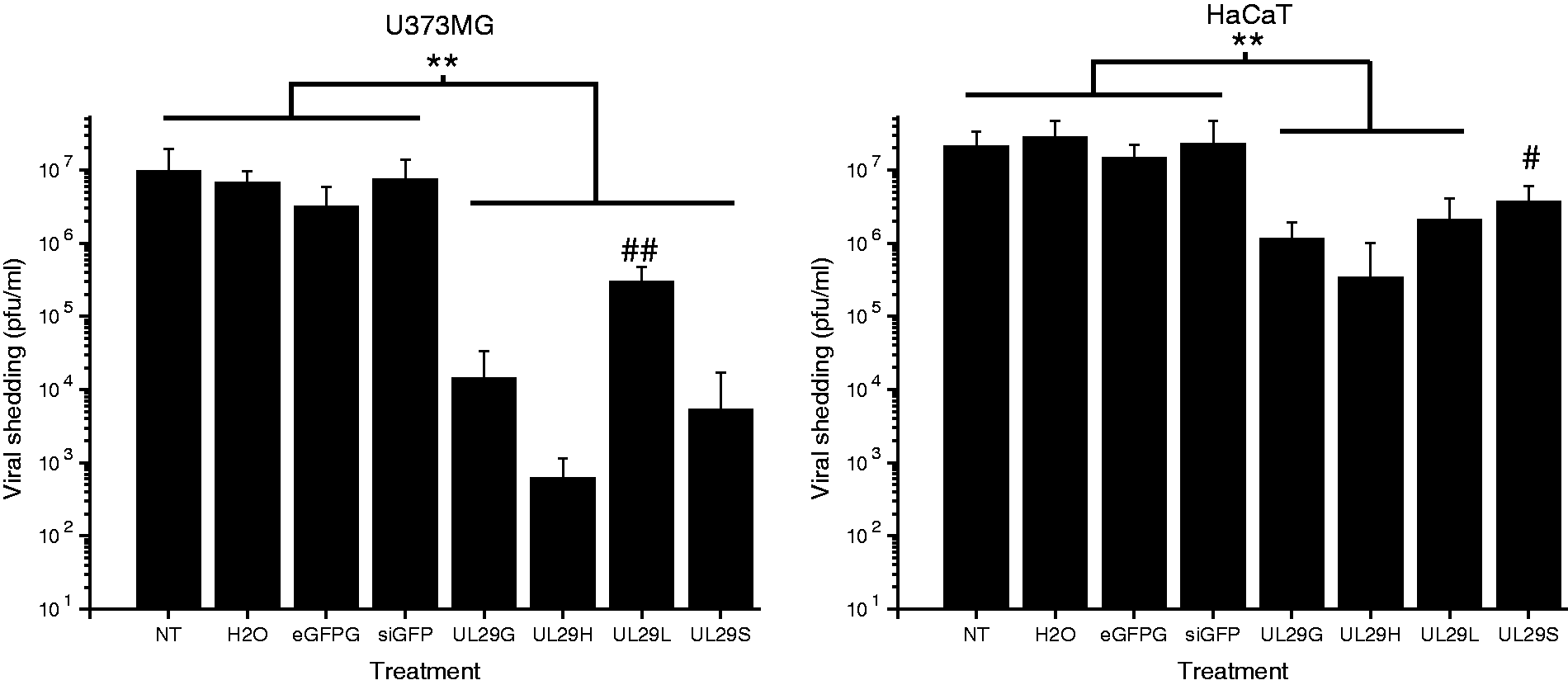
Anti-HSV potency of the siRNAs
The siRNA pools against the UL29 gene, prepared using human or Giardia dicer (UL29H and UL29G, respectively; Table 1), had a significant antiviral activity (P < 0.01; Figure 1) when compared with the nonspecific siRNAs targeting the eGFP gene (siGFP or eGFPG; Table 1) or control treatments in both HaCaT and U373MG cell lines. In accordance with previous studies, 16 both anti-HSV siRNA pools effectively inhibited HSV replication equally (Figure 1), reaching > 99% and 90% decrease in viral shedding in the astrocytic cell line U373MG and HaCaT keratinocytes, respectively, at 44 h p.i. In all cases, the antiviral effectiveness of the 21- and 27-nt single-site siRNAs (UL29S and UL29L, respectively) varied between cell lines but remained significant (P < 0.05).
Relative HSV late (γ) to immediate early (α) gene expression.
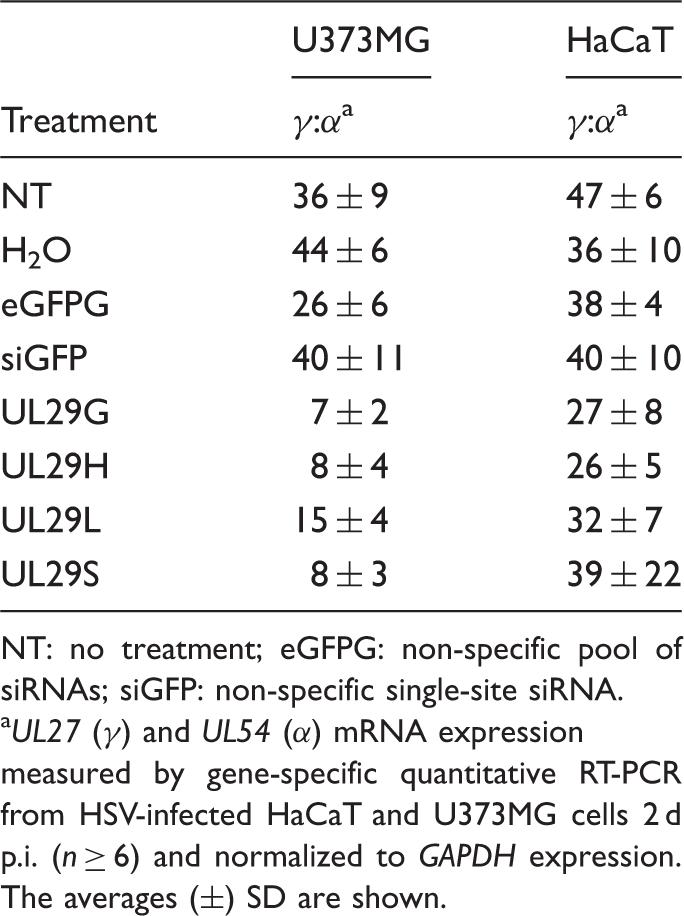
NT: no treatment; eGFPG: non-specific pool of siRNAs; siGFP: non-specific single-site siRNA.
UL27 (γ) and UL54 (α) mRNA expression measured by gene-specific quantitative RT-PCR from HSV-infected HaCaT and U373MG cells 2 d p.i. (n ≥ 6) and normalized to GAPDH expression. The averages (±) SD are shown.
Innate responses in siRNA-treated and HSV-infected U373MG and HaCaT cells
We focused on the cellular innate response, in particular on the changes in type I and III IFN gene expression, in cells treated with the siRNA preparations and infected with HSV-1. There is evidence that HSV can modulate or evade host innate responses.4,6,7 The effects of HSV infection on the studied innate responses in U373MG cells were limited (non- or water-transfected, infected and uninfected cells; Figure 2). This provided a platform to investigate the role that siRNA-induced IFN response play in antiviral efficacy. In contrast, HaCaT cells infected with HSV evoked robust innate responses (non- or water-transfected, infected and uninfected cells; Figure 3). This represented a combination of viral innate immunity modulation mixed with treatment-induced effects. In uninfected cells, siRNA treatments themselves caused some innate responses (Figures 2 and 3), in accordance with previous studies.
16
Nevertheless, these responses had minimal correlation with antiviral effectivity of a given siRNA preparation (Figure 1).
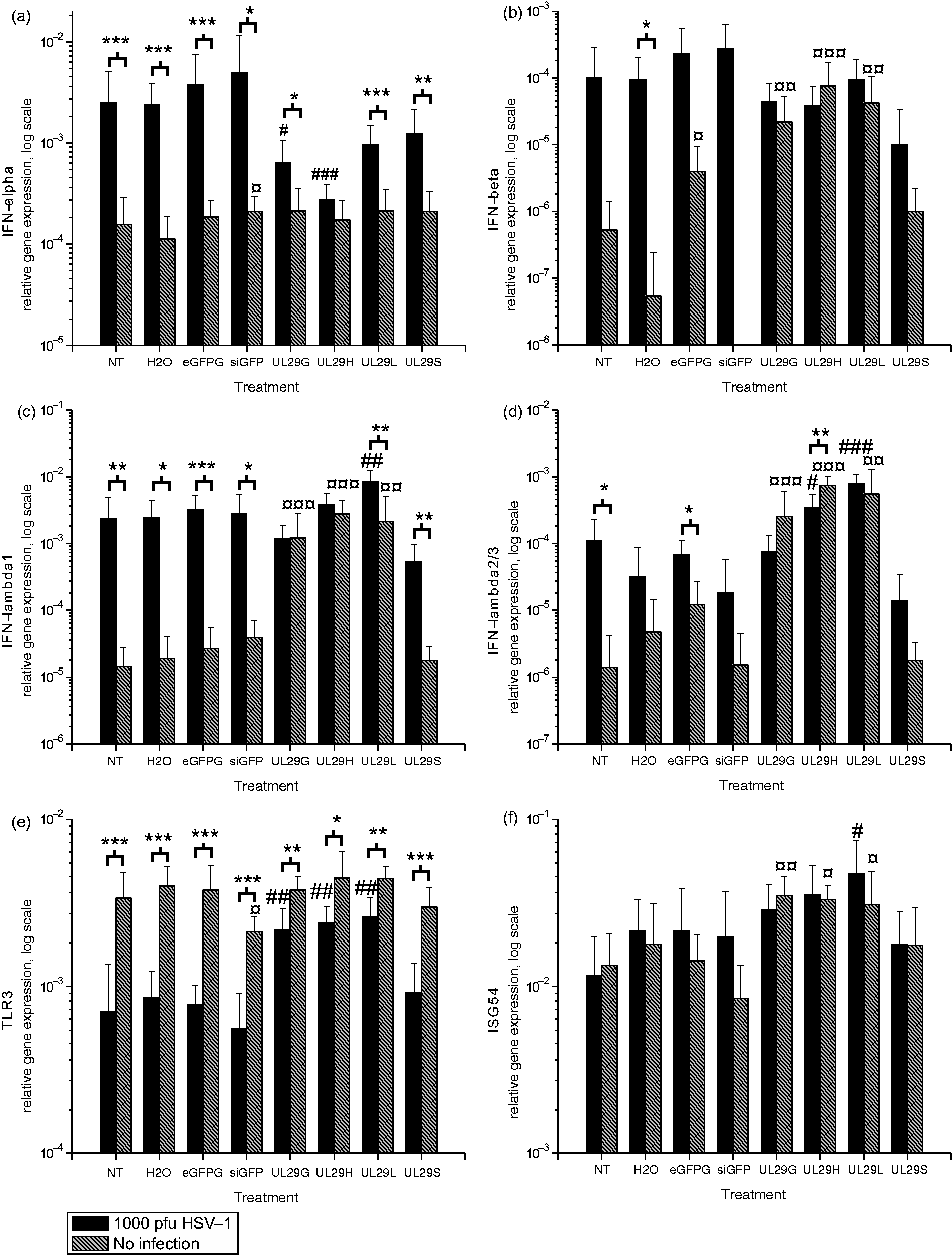
Effects of HSV-1 infection on the siRNA-induced innate responses in astrocytic cells. (a) IFN-alpha, (b) IFN-beta, (c) IFN-lambda1 (IL-29), (d) IFN-lambda2/3 (IL-28 A/B), (e) TLR3 and (f) ISG54 gene expression levels were measured by quantitative RT-PCR from uninfected (gray bars) and HSV-infected (black bars) U373MG cells at 48 h post-transfection (44 h p.i.). The relative expression, normalized to GAPDH gene expression, is shown on a logarithmic scale for each siRNA or control treatment. SDs of at least eight replicates are shown. A non-parametric Mann–Whitney U-test was used to determine significances. *P < 0.05, **P < 0.01, ***P < 0.001 (HSV-infected versus uninfected); #P < 0.05, ##P < 0.01, ###P < 0.001 as compared with infected controls (NT and H2O); ¤P < 0.05, ¤¤P < 0.01, ¤¤¤P < 0.001 as compared with non-infected controls (NT and H2O). Effects of HSV-1 infection on the siRNA-induced innate responses in HaCaT cells. (a) IFN-alpha, (b) IFN-beta, (c) IFN-lambda1 (IL-29), (d) IFN-lambda2/3 (IL-28 A/B), (e) TLR3 and (f) ISG54 gene expression was measured by quantitative RT-PCR from uninfected (gray bars) or HSV-infected (black bars) HaCaT cells at 48 h post-transfection (44 h p.i.). The relative gene expression, normalized to GAPDH gene expression, is shown on a logarithmic scale for each siRNA or control treatment. SDs of at least eight replicates are shown. Non-parametric Mann–Whitney U-test was used to determine significances. *P < 0.05, **P < 0.01, ***P < 0.001 (HSV-infected versus uninfected); #P < 0.05, ##P < 0.01, ###P < 0.001 as compared with infected controls (NT and H2O); ¤P < 0.05, ¤¤P < 0.01, ¤¤¤P < 0.001 as compared with non-infected controls (NT and H2O).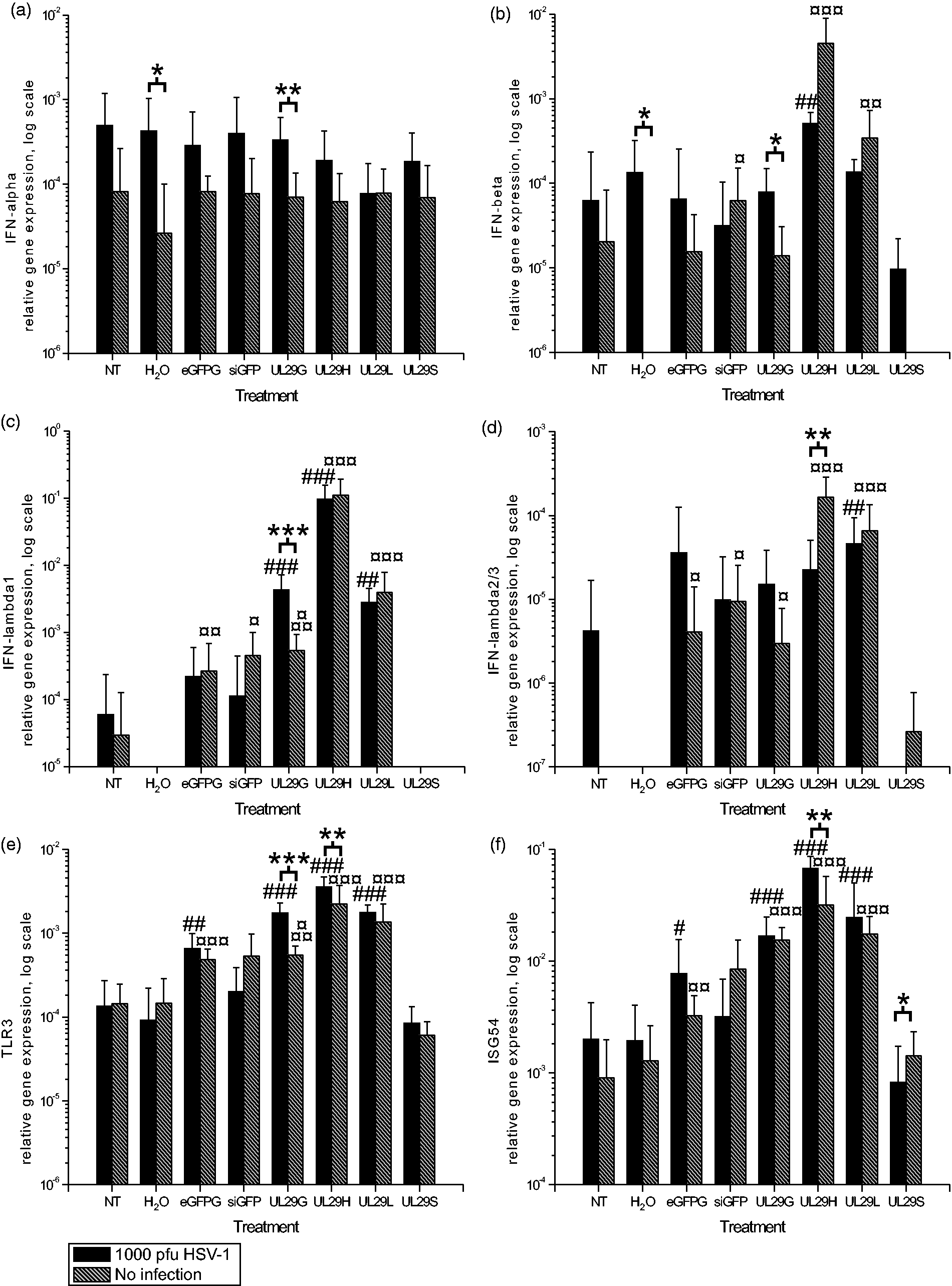
Innate cellular responses in U373MG cells
Astrocytic cells, including U373MG, express TLR3 and produce IFN type I and type III molecules.16,22,23 However, U373MG cells were unable to mount a distinct IFN response to the HSV infection (infected cells with non- and water transfection; Figure 2). The gene expression levels indicated a significant response to some of the siRNA treatments (Figure 2).
Transfection of U373MG cells with siRNAs caused only minimal alterations in IFN-α gene expression (Figure 2a). However, the UL29H siRNA pool and single-site UL29L siRNAs increased IFN-β and -λ expression (Figure 2b–d) and levels of TLR3- and ISG54-specific mRNAs (Figure 2e, f). Compared with the controls, UL29H- and UL29L-treated U373MG cells had significant IFN and IFN-stimulated gene expression responses irrespective of HSV infection (Figure 2b–f). Significant IFN responses to UL29L (Figure 2b–d) did not correlate with its anti-HSV activity (Figure 1).
Previous studies using astrocytes indicate that transfected and complexed dsRNA is recognized in these cell types via a melanoma differentiation-associated gene 5 cytosolic RNA sensor. However, the TLR3-dependent pathway is employed only when the dsRNA is naked in the culture media. 24 We observed a significant increase in TLR3 mRNA levels for all siRNA preparations (Figure 2e), except for short siRNAs. Although TLR3 is involved in the sensing of HSV infection,25,26 especially in astrocytes,22,23 we did not detect alterations in TLR3 gene expression after HSV infection (non- or water-transfected, infected and uninfected cells; Figure 2e). There was a significant upregulation of IFN-β, -λ1 and TLR3 in UL29G-treated and infected cells compared with their uninfected counterparts. The HSV infection modified the IFN-λ2/3, TLR3 and ISG54 gene expression in UL29H siRNA-treated cells (Figure 2d–f). The observed induction profile for IFN-stimulated gene 54 (ISG54) was similar to that of TLR3, which is also an ISG (Figure 2e, f) in different treatment groups.
The antiviral effect of the siRNA preparations (Figure 1) was not directly related to the type I or type III IFN gene induction. The UL29H siRNA pool induced the highest expression of IFN-β and IFN-λ, but the equally effective UL29G pool or single-site UL29S siRNA induced only low or undetectable expression of IFN-β or IFN-λ expression in U373MG cells (Figure 2b–d). Moreover, the longer siRNAs (27 nt), UL29G and UL29L (Table 1), had very similar effects on the gene expression of IFN type I and III, and on TLR3 and ISG54 gene expression in HSV infection, but their anti-HSV potency was significantly different (Figure 2 and Figure 1, respectively) in the U373MG cells. This suggests that the antiviral effects of the siRNAs are sequence-specific. The difference in the antiviral efficiency of the target-specific and nonspecific siRNAs, both single-site siRNAs and pools, support such a conclusion (Figure 1).
Innate cellular responses in HaCaT cells
In HSV-infected HaCaT cells, we observed type I and type III IFN responses (Figure 3a–d), consisting of distinct IFN-α and IFN-λ1 responses to HSV, but the IFN-β and IFN-λ2/3 responses were modest. The virus-induced effects overruled those caused by the siRNA pools or single-site siRNAs. The HSV infection-induced IFN-α gene expression was markedly suppressed when the cells were treated with anti-HSV siRNAs in comparison with the nontreated but infected controls (Figure 3a). This may reflect the significant reduction in HSV replication by these siRNA treatments (Figure 1), which, in turn, suppressed virus-induced IFN-α gene expression.
TLR3 is expressed in HaCaT cells (Figure 3e), 27 which are also capable of processing dicer–substrate RNAs to trigger RNAi. 28 In these cells, the HSV infection inhibited TLR3 gene expression both in untreated and siRNA-treated cells (Figure 3e). The downregulation of TLR3 gene expression correlated with the extent of HSV infection (Figure 1, Figure 3e). The highest expression occurred in cells treated with the effective anti-HSV siRNAs. This is in accordance with our earlier observation regarding the modulation of TLR3 expression by HSV infection in monocytic cells. 25 Only minor differences in ISG54 expression, in comparison with nontreated cells, were observed in HSV-infected or uninfected siRNA-treated cells (Figure 3f) at 48 h post-transfection. In general, the type I and III IFN gene expression, induced by the siRNA pools, was low or equal to the HSV-induced IFN expression in the keratinocyte cell line, supporting the concept that the inhibition of HSV infection by the siRNA pools was sequence-specific, and not a result of IFN induction by the siRNA pools.
As different cell lines give varying responses to HSV and the siRNA treatments, it remains to be studied what happens in an in vivo application. A slightly elevated immune response might enhance adaptive antiviral immunity, whereas too high a response to the treatment might cause adverse effects in a multicellular organism. Thus, the UL29G pool, causing only slight immune responses and having very effective antiviral properties, could be a prime candidate for an in vivo approach for the treatment of, for example, herpes keratitis.
Conclusions
The effects of HSV on the innate immunity responses depended on the studied cell line. Immunostimulation by the siRNA preparations had minimal effect against HSV. The antiviral effect of the siRNAs was mainly sequence specific. The inhibition efficacy varied significantly between the different anti-HSV siRNA molecules with similar innate responses. This was most evident in the U373MG cells, as the cells did not respond to HSV infection. In contrast, HaCaT cells reacted strongly to the HSV infection by mounting innate immunity responses. In addition, HSV was able to downregulate innate response in these epithelial cells as HSV-infected cells had diminished TLR3 gene expression compared with the baseline. The HSV replication inhibition correlated only weakly with the innate responses caused by the siRNA pools and single-site siRNAs of different length. In conclusion, HSV was capable of resisting a slight increase in intracellular innate immunity in HaCaT and U373MG cells, whereas the virus remained vulnerable to sequence-specific RNA interference.
Footnotes
Funding
This study was supported by the Academy of Finland (grant numbers 128915, 259725, 250113, 256069, 256197, 255342, 256518 and 272507); the Sigrid Juselius Foundation; the FinnPharma Doctoral Program – Drug Discovery section; the Instrumentarium Foundation; the Centre for International Mobility; the Turku Doctoral Programme of Biomedical Sciences; and the Finnish Cultural Foundation.
Acknowledgements
We thank R. Tarkiainen, T. Westerholm, M. Kleme, M.-L. Mattila, T. Laihinen, L. Lund and M. Laanti for technical assistance, and Dr. R.M. Badeau for the language checking of the manuscript.
