Abstract
Endotoxin is recognized as one of the virulence factors of the Bordetella avium bird pathogen, and characterization of its structure and corresponding genomic features are important for an understanding of its role in pathogenicity and for an improved general knowledge of Bordetella spp virulence factors. The structure of the biologically active part of B. avium LPS, lipid A, is described and compared to those of another bird pathogen, opportunistic in humans, Bordetella hinzii, and to that of Bordetella trematum, a human pathogen. Sequence analyses showed that the three strains have homologues of acyl-chain modifying enzymes PagL, PagP and LpxO, of the 1-phosphatase LpxE, in addition to LgmA, LgmB and LgmC, which are required for the glucosamine modification. MALDI mass spectrometry identified a high amount of glucosamine substituting the phosphate groups of B. avium lipid A; this modification was absent from B. hinzii and B. trematum. The acylation patterns of the three lipid As were similar, but they differed from those of Bordetella pertussis and Bordetella parapertussis. They were also found to be close to the lipid A structure of Bordetella bronchiseptica, a mammalian pathogen, only differing from the latter by the degree of hydroxylation of the branched fatty acid.
Introduction
Nine species have been described in the Bordetella genus, 1 with Bordetella pertussis and Bordetella parapertussis being the agents of whooping cough. The former is a strict pathogen for humans, 2 but strains causing pneumonia in sheep have been described for the latter. 3 Bordetella bronchiseptica infects many mammalian species, including humans, and causes respiratory diseases.4,5 Bordetella avium, 6 as its name indicates, is a bird pathogen. In addition to the major species described above, five others have more recently been identified and included in the genus: Bordetella hinzii, 7 Bordetella trematum, 8 Bordetella holmesii, 9 Bordetella petrii 10 and Bordetella ansorpii. 11
Involvement of LPS in Bordetella infections was first shown in B. pertussis, with LPS causing damage to ciliated tracheal cells by the induction of NO in synergy with tracheal cytotoxin. 12 Bordetella avium causes a mild upper respiratory tract disease called turkey coryza. This bacterium binds to ciliated tracheal epithelial cells, damaging them and thus favouring lethal secondary infections, often by Klebsiella pneumoniae, Escherichia coli and Pseudomonas spp. 6 Interest in the mechanism of infection is therefore driven, in part, by the economic stake in poultry production, especially turkey farming, but, more specifically, by the search for understanding bacterial evolution leading to the infectious disease process. Orndoff and collaborators have described different aspects of B. avium virulence. 13 Mutants were found with an altered LPS in SDS gel electrophoresis profiles and exhibited decreased binding to tracheal cells. 13 B. avium LPS has also been described as a virulence factor. 14 Another recent study involving a turkey tracheal explant culture model showed renewed interest in identifying LPS as a major immunomodulatory molecule. 14 Such experiments would benefit from the knowledge of well-characterized LPS structures, and the structural differences reported here for this bacterium underlines the importance of using species-specific LPS for these studies.
The use of a new technique of hydrolysis 15 and improved kinetics for selective ester-linked fatty acid removal 16 has allowed the detailed description of the lipid A moiety of B. avium [strain American Type Culture Collection (ATCC) 35086]. This structure is presented here for the first time, together with a characterization of the lipid A of B. trematum CCUG 13902, 17 —a pathogen associated with ear and blood infections in humans. Bordetella hinzii ATCC 51730 is another bird pathogen. 18 Like B. trematum, it has also been isolated from humans, including immunocompromised patients.8,19–21 The structure of B. hinzii lipid A 18 was reinvestigated in consideration of data relating to the absence of substitution at C-3, which we described a few years ago in B. parapertussis 22 and, more recently, in B. bronchiseptica, 23 subsequently confirmed by MacArthur et al. 24 Other important lipid A modifications were found first in B. bronchiseptica, 16 then in B. pertussis. 25 They consist of substitution of the phosphate groups with glucosamine, a substitution never described in other genera. We recently described the lgm (lipid A glucosamine (GlcN) modification) locus genes, lgmA, lgmB and lgmC, which are required for this modification 26 and the important effect on biological activities of this new structural element. 27 We will discuss the occurrence of the lgm locus in some of the species characterized herein. These data complete the polysaccharide structures we previously described for these three species.
Material and methods
Bacterial strains: culturing and LPS preparation
B. avium smooth-type strain ATCC 35806 28 B. hinzii ATCC 51730 1 8 and B. trematum CCUG 13902 17 were grown as described previously.17,18,28 The cells were killed in 2% cold phenol before harvesting. The LPS were extracted from lyophilized bacteria by the modified enzyme–phenol–water method, 29 and purified as described. 30
LPS cleavage for lipid A preparation
Three mild hydrolytic procedures developed in our laboratory were used for isolation of lipid A moieties from purified LPS.
Detergent-promoted hydrolysis. 31
LPS (100 mg) was cleaved by hydrolysis in 20 mM sodium acetate/acetic acid buffer (pH 4.5) containing 1% of SDS at 100℃ for 1 h at a concentration of 5 mg/ml. Lipid A was extracted from the hydrolysate after lyophilization and SDS removal by acidified ethanol washing.
Triethylamine–citrate micro-hydrolysis
This recently described method 32 allows direct analysis by MALDI-MS without salt removal. Briefly, LPS samples (50–100 µg) were suspended in 1.5-ml Eppendorf (Le Peck, France) tubes in 0.01 M triethylamine (TEA)–citric acid water solution (1:1 molar ratio, pH 3.6) at concentrations of 5 µg/µl for rough-type LPS and 10 µg/µl for smooth-type LPS. After agitation and homogenization using an ultrasonic bath, the tubes were incubated for 2 h at 1000 rpm and 100℃ in a thermomixer system (Eppendorf) under stirring. Cleaved lipid A was then analysed by MALDI-MS (see below) either directly from the hydrolysis samples or after lyophilization and extraction.
Alkaline β-elimination method. 33
LPS (50 mg) was first oxidized at the level of the Kdo extracyclic diol in 6 µM sodium periodate solution for 30 min (LPS concentration 4 µg/µl). Lipid A was then separated from the polysaccharide moiety by the β-elimination reaction effected at pH 9.5 for 2 h and followed by reduction with sodium borohydride (2.5 mg/ml) for 1 h. The whole procedure was performed at room temperature (20℃). After neutralization of the reactive mixture with acetic acid, insoluble lipid A was recovered in the pellet after ultracentrifugation, washed with water and extracted.
In all three methods, cleaved lipid A was finally extracted with an appropriate volume of chloroform/methanol/water mixture (3.0:1.5:0.25, v:v:v).
Sequential liberation of ester-linked fatty acids by mild alkali treatments
As previously described, 16 the following conditions were applied in the first-step liberation of primary ester-linked fatty acids: lipid A (30 µg) was suspended at 1 mg/ml in 28% ammonium hydroxide and kept under stirring for 5 h at 50℃. For liberation of the secondary ester-linked fatty acids, lipid A was suspended in 41% methylamine and kept under stirring for 5 h at 37℃. The solutions were dried with a stream of nitrogen, the residues were taken up in a mixture of chloroform/methanol/water (3.0:1.5:0.25, v:v:v) and analysed by MALDI-TOF-MS.
Alternatively, for kinetic analysis, more precise and selective removal of primary ester-linked fatty acids was necessary, so we applied a step-by-step kinetics of release described in Tirsoaga et al. 16 Lipid A was suspended in 28% ammonium hydroxide and kept under stirring at 50℃ for different time intervals (0, 15 min, 30 min, 1 h, 2 h, 4 h) before evaporation under a stream of nitrogen and MALDI-MS analysis.
MS
Two different techniques were used in this study.
Plasma desorption mass spectrometry
Plasma desorption mass spectra were obtained with a Depil TOF 21 mass spectrometer (IPN, Orsay, France) as described 34 and lipid A samples were prepared as before. 33 Spectra were recorded in the positive and negative ion modes.
MALDI-MS
MALDI-TOF mass spectra were obtained using a Perseptive Voyager STR (PE Biosystem, Les Ulis, France) TOF mass spectrometer (IBBMC, Orsay, France). A suspension of native or treated lipid A (1 µg/µl) in chloroform/methanol/water mixture (3.0:1.5:0.25, v:v:v) was desalted with a few grains of Dowex 50 W-X8 (H+); 0.5 µl was deposited on the target, mixed with 0.5 µl of the matrix solution and dried. Different ratios of sample and matrix were tested to optimize spectra. Matrix, 2,5-dihydroxybenzoic acid (Sigma Chemical, St. Louis, MO, USA), was dissolved at 10 µg/µl in 0.1 M citric acid solution in the same solvent. 35 Analyte ions were desorbed with pulses of a nitrogen laser (337 nm). Negative and positive ion mass spectra were recorded in the linear mode with delayed extraction and ion acceleration set at 20 kV. Fragmentation of negative precursor ions was studied in post-source decay (PSD). 25
TLC
Chromatography was performed on aluminium-backed silica TLC plates (Merck, Molsheim, France) and spots visualized by charring (145℃ after spraying with 10% sulphuric acid in ethanol) and/or by ninhydrin. The mobile phase was a mixture of chloroform/methanol/water/triethylamine (30.0:20.0:2.5:0.1, v:v:v:v). 31
Chemical analyses
Published methods were used to assay hexosamines 36 and phosphate. 37 GlcN phosphate substituents were released after aqueous hydrofluoric acid treatment as previously described 25 and characterized by GC/MS on a HP5 column (30 m × 0.32 mm; Hewlett Packard, Les Ulis, France) and by amino acid analysis on a Hitachi L-8800 device equipped with a 2620 MSC-PS column (ScienceTec, Les Ulis, France). Fatty acids were analysed after hydrolysis of the LPS or lipid A with 4 M HCl for 2 h at 100℃, followed by esterification using methanolysis, 22 extraction with hexane and identification by GC retention time on a BP10 column using a temperature gradient of 160–240℃, 4℃/min. Ester-linked fatty acids were likewise characterized by selective removal as described above.
Whole-genome sequencing and genetic analyses
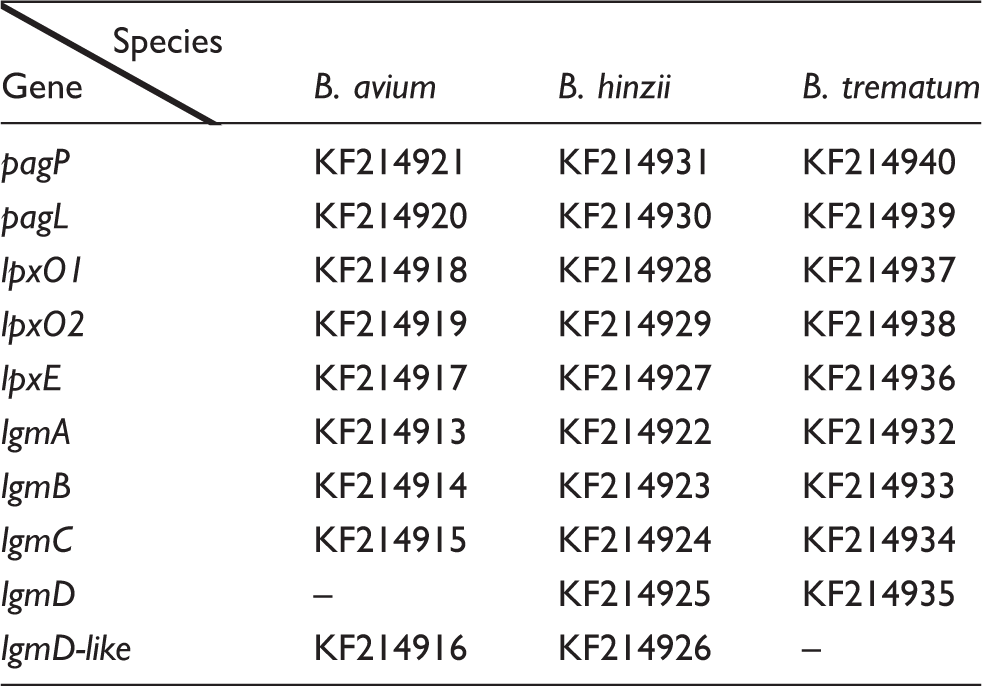
GenBank accession numbers for the Bordetella genes: pagP, pagL, lpxO1, lpxO2, lpxE, lgmA, lgmB, lgmC, lgmD and lgmD-like protein. Strains: B. avium ATCC 35086, B. hinzii ATCC 51730 and B. trematum CCUG13902.
Results
Comparison and selection of lipid A hydrolysis procedures
We previously described different hydrolysis methods to minimize lipid A structural modifications, especially at the level of the extremely labile glycosidic phosphate group. Three of these methods (SDS-promoted sodium-acetate hydrolysis, 31 triethylamine-citrate hydrolysis 32 and the alkaline β-elimination method 33 ) were tested on B. avium, B. trematum and B. hinzii LPS. In all experiments, the resulting MALDI mass spectra of the isolated lipids A were compared with those obtained by MALDI in-source-decay fragmentation of the corresponding LPS to confirm that no structural element was lost. Similar results were obtained with B. trematum and B. hinzii LPS following both SDS-promoted sodium-acetate and TEA–citrate hydrolyses. As the sample requirements for TEA–citrate hydrolyses were smaller than other methods, it was selected for further analysis. However, a higher level of monophosphoryl B. avium lipid A molecules was observed under TEA–citrate hydrolyses, especially in the low acylated molecular species. To avoid acid cleavage of the glycosidic phosphate, the alkaline β-elimination method was thus applied to B. avium LPS.
Chemical composition of the lipid A backbones
All three lipids A were shown to contain
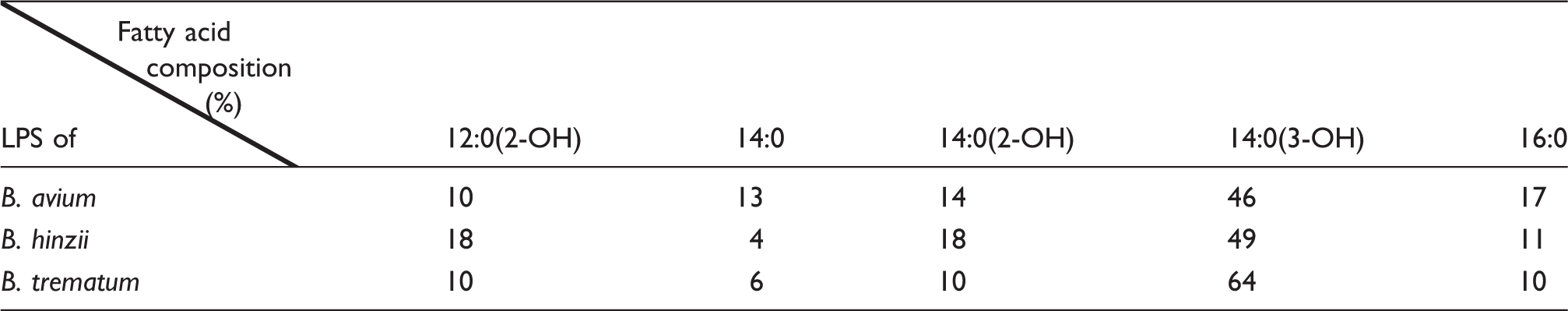
Total fatty acid composition
Relative amount of the different fatty acids present in the different LPS studied.
Comparison of the three lipid A negative ion MALDI mass spectra
Upon analysis by MALDI-MS, the three lipid A spectra were compared. B. avium (Figure 1a) was found to be more heterogeneous, in support of the TLC analysis, and presented additional higher mass molecular species than B. trematum and B. hinzii spectra. The latter two being similar, only B. trematum is presented on Figure 1b.
(a) Negative ion MALDI mass spectrum of B. avium lipid A and (b) Negative ion MALDI mass spectrum of B. trematum lipid A.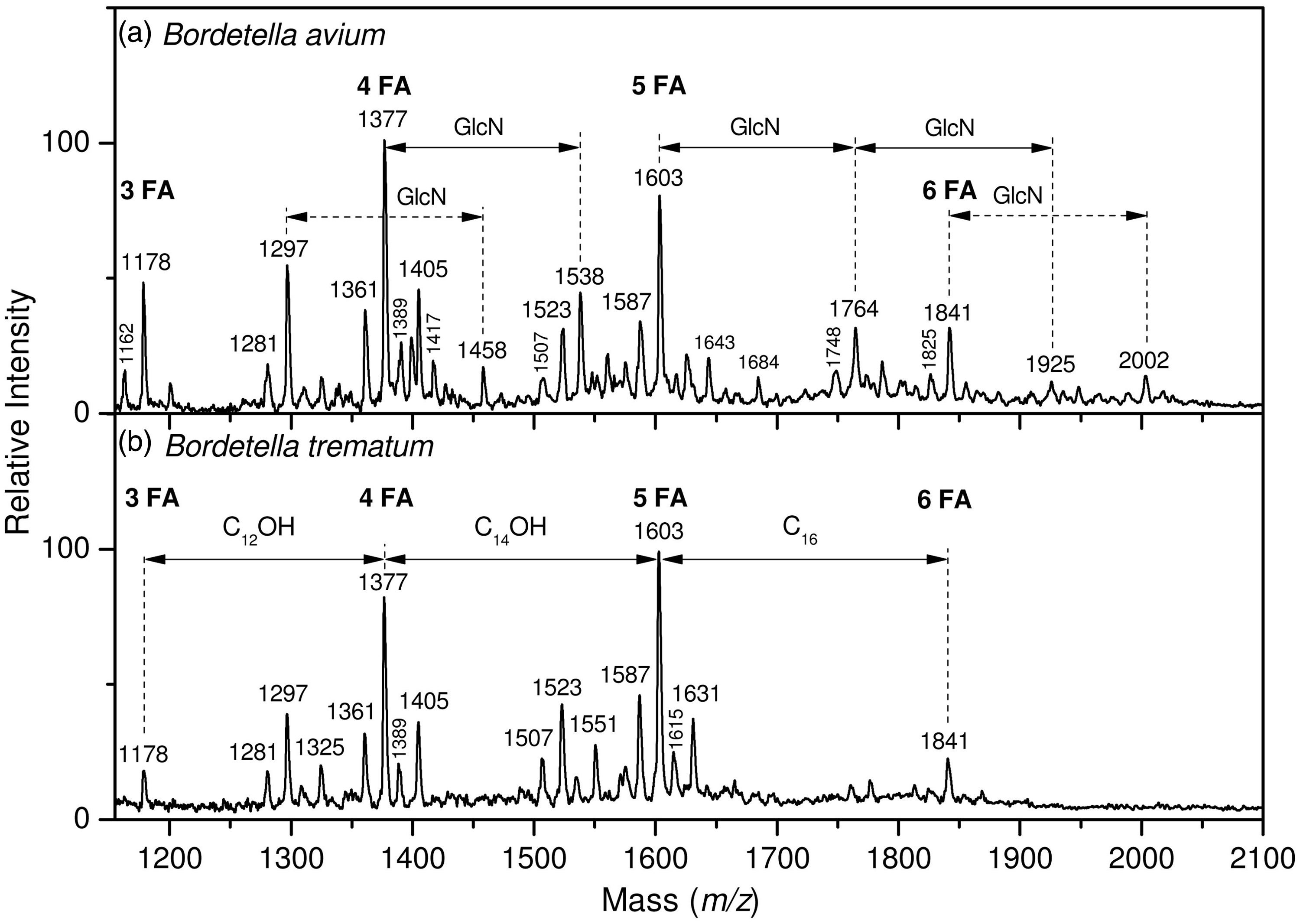
B. avium lipid A acylation
As shown in Figure 1a, major peaks of deprotonated [M-H]− lipid A molecular species were observed at 1178, 1377, 1603 and 1841 u. In accordance with the chemical composition and total fatty acid content data shown in Table 2, these peaks were attributed to tri-, tetra-, penta- and hexa-acyl molecular species in which the di-phosphorylated di-glucosamine backbone was substituted with the following fatty acids: three 14:0(3-OH) in tri-acyl species (1178 u), three 14:0(3-OH) and one 12:0(2-OH) in tetra-acyl molecular species (1377 u), three 14:0(3-OH), one 14:0(2-OH) and one 12:0(2-OH) in penta-acyl molecular species (1603 u), and three 14:0(3-OH), one 14:0(2-OH), one 12:0(2-OH) and one 16:0 in hexa-acyl molecular species (1841 u). These peaks were doubled by smaller ones observed at –16 u, i.e. at 1162, 1361, 1587 and 1825 u. They were attributed to the same structures in which either 14:0(2-OH) or 12:0(2-OH) were non-hydroxylated (14:0 or 12:0 respectively). Another type of microheterogeneity involved the length of fatty acids with peaks at plus 28 u (2 × CH2) from the major peaks, i.e. at 1405 u and 1631 u.
Lipid A sequential fatty acid liberation
The results obtained for the three lipids A were very similar, confirming the same acylation patterns for all three lipids A. B. hinzii lipid A is taken here as a model for all three lipids A, so B. trematum and B. avium kinetics are not shown.
The exact position of each fatty acid was determined by its liberation pattern following alkaline treatment in a sequential way, as described previously for MALDI experiments.
16
The transformation of native lipid A negative ion MALDI mass spectrum of B. hinzii lipid A during the kinetics of alkaline treatment is presented in Figure 2a. Under the conditions used, all substituents at the C-3 and C-3′ positions were liberated by ammonium hydroxide treatment. The B. pertussis lipid A deacylation pattern,
40
used as a reference, revealed almost total release of the fatty acid at position C-3 [10:0(3-OH)] in the first 15 min of treatment (data not shown). The first 15 min of treatment applied to the lipid A under study did not change the general aspect of the initial spectrum, which could be explained by an unsubstituted C-3 position and confirmed by the determination of all other fatty acid positions. After 2–5 h of treatment, 14:0(3-OH) and 14:0[3-0(16:0)] were completely released, which clearly indicated that these fatty acids were at the C-3′position in the native molecules. After treatment with methylamine, aimed at liberating secondary ester-linked fatty acids, 14:0(2-OH) and 12:0(2-OH) were completely released, which indicated that they were in secondary linkage at C-2 or C-2′ positions. According to the fragmentation data, C-2′ secondary position was substituted by 14:0(2-OH) and C-2 secondary position by 12:0(2-OH). Minor peaks corresponding to the native molecular species containing 14:0 and 12:0 fatty acids followed the same deacylation patterns, confirming that these fatty acids were present in alteration with their 2-hydroxylated counterparts, i.e. at secondary C-2′ and C-2 positions respectively.
(a) Kinetics of ester-linked fatty acids liberation upon alkaline treatment followed by negative-ion MALDI mass spectrometry. (b) Comparison of O-deacylated B. avium lipid A by (a) linear mode and (b) PSD-mode, showing the presence of GlcN residues on the lipid A core.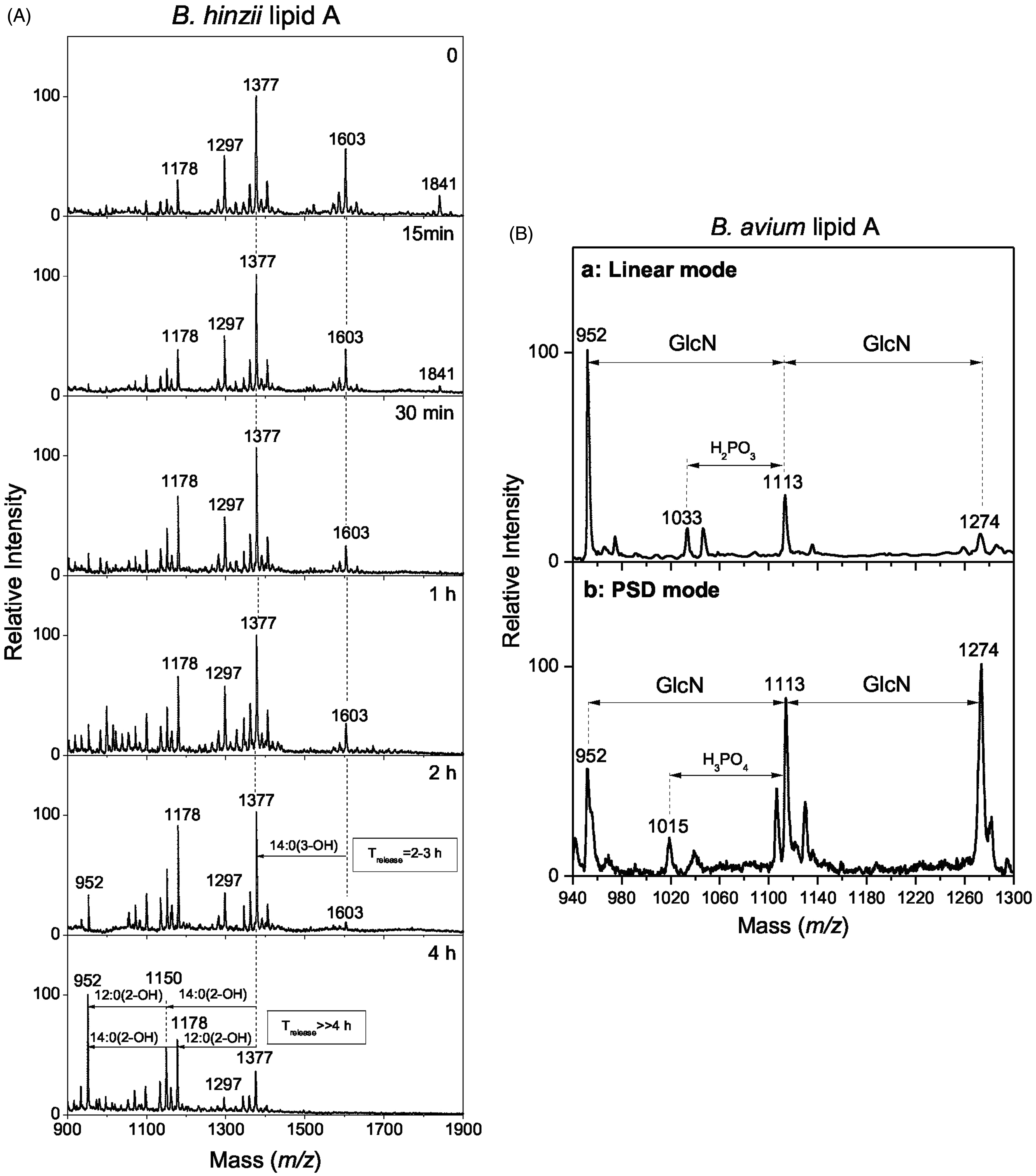
Phosphate group substitutions
The major difference observed in B. avium lipid A negative ion mass spectra from those of B. trematum and B. hinzii was the presence of a series of additional peaks at 161 or 322 u, higher than the mentioned common peaks (1538, 1764, 1925 and 2002 u). As the mass difference of 161 u corresponds to the molecular mass of one GlcN residue and according to the higher GlcN content found in B. avium lipid A, we hypothesized that they originated from GlcN residues present on the phosphate groups of this lipid A. These GlcN residues were effectively released by hydrofluoric acid treatment, giving confirmation that both phosphate groups were substituted.
To confirm the presence of extra GlcN on the phosphate groups, B. avium lipid A was O-deacylated by an alkali treatment with methylamine 16 and analysed by MALDI-TOF-MS in linear and PSD modes (Figure 2Ba, 2Bb). This methodology was previously used for the analysis of B. bronchiseptica strain 4650 lipid A, in which the substitution of lipid A phosphate groups with GlcN was described for the first time. 25 In a similar way as described for B. bronchiseptica, the direct MALDI negative ion mass spectrum of O-deacylated B. avium lipid A revealed a major peak at 952 u, which was attributed to di-phosphorylated di-glucosamine substituted with two 14:0(3-OH) fatty acids in amide linkages. Two other peaks were observed at 161 u and 322 u higher, i.e. at 1113 u and 1274 u, corresponding to substitution of the first molecular species with one and two GlcN respectively (Figure 2b). A PSD experiment on the 1274 u precursor ion revealed daughter ions at 1113 u and 952 u due to the release of one and two GlcN residues, respectively, as well as at 1033 u due to a consecutive release of one GlcN residue and one phosphate. GlcN was also characterized by amino acid analysis after hydrofluoric acid treatment, which also confirmed the presence of additional GlcN. 25
B. trematum lipid A acylation
MALDI-MS analysis of B. trematum lipid A is shown in Figure 1b. Some major peaks in common with B. avium lipid A were observed at 1178, 1377, 1603 and 1841 u. In accordance with the chemical composition and total fatty acid content data shown in Table 2, these peaks were attributed to tri-, tetra-, penta- and hexa-acyl molecular species in which the di-phosphorylated di-glucosamine backbone was substituted by the following fatty acids, as previously indicated for B. avium: three 14:0(3-OH) in tri-acyl species (1178 u), three 14:0(3-OH) and one 12:0(2-OH) in tetra-acyl molecular species (1377 u), three 14:0(3-OH), one 14:0(2-OH) and one 12:0(2-OH) in penta-acyl molecular species (1603 u), and three 14:0(3-OH), one 14:0(2-OH), one 12:0(2-OH) and one 16:0 in hexa-acyl molecular species (1841 u). These peaks were doubled by smaller ones observed at –16 u, i.e. at 1361 u and 1587 u. They were attributed to the same structures in which either 14:0(2-OH) or 12:0(2-OH) was replaced by their non-hydroxylated counterpart (14:0 or 12:0 respectively). Another type of microheterogeneity involved the length of fatty acids with peaks at plus 28 u (2 × CH2) at 1405 u and 1631 u from the major peaks.
No peaks were observed at + 161 u from the major molecular species, which indicated the absence of GlcN substitution on the phosphate groups.
Fatty acid distribution over di-glucosamine backbones lipid A
The position of each fatty acid in the lipid A structures were established by fragmentation analysis of native samples, which was completed by the analysis of sequential fatty acids liberation.
Distribution of fatty acids between the reducing (GlcN I) and distal (GlcN II) of the di-glucosamine backbone were established by native lipid A fragmentation analyses in the positive-ion MALDI-MS mode and the negative-ion plasma desorption MS (PDMS) mode.
41
The positive-ion MALDI mass spectra (Figure 3) of the three lipid As revealed prominent fragment peaks at 694, 920 and 1158 u doubled with smaller ones at 16 u lower, i.e. at 678, 904 and 1142 u respectively. In accordance with the fragmentation pattern previously established,
41
these peaks were attributed to B1 lipid A fragments made of GlcN II substituted with one phosphate group and different fatty acids: one 14:0(3-OH) and one 14:0 or 14:0(2-OH) for tri- and tetra-acyl molecular species, two 14:0(3-OH) and one 14:0 or 14:0(2-OH) for pentaacyl ones and two 14:0(3-OH), one 16:0 and one 14:0 or 14:0(2-OH) for hexa-acyl ones. Information about fatty acids substituting GlcN I were obtained from negative-ion PDMS mass spectra in which a group of three fragment peaks was observed for all three samples at 666, 682 and 711 u. According to Karibian et al.,
41
these peaks are attributed to Z1, Y1 and X1 fragment ions, respectively, related to the GlcN II carrying one phosphate group, one 14:0(3-OH) and one 12:0(2-OH) residues. Fragments containing additional GlcN substituting the phosphate group were not observed for B. avium lipid A because of the loss of this compound during the fragmentation process.
Positive ion MALDI mass spectrum of B. avium lipid A.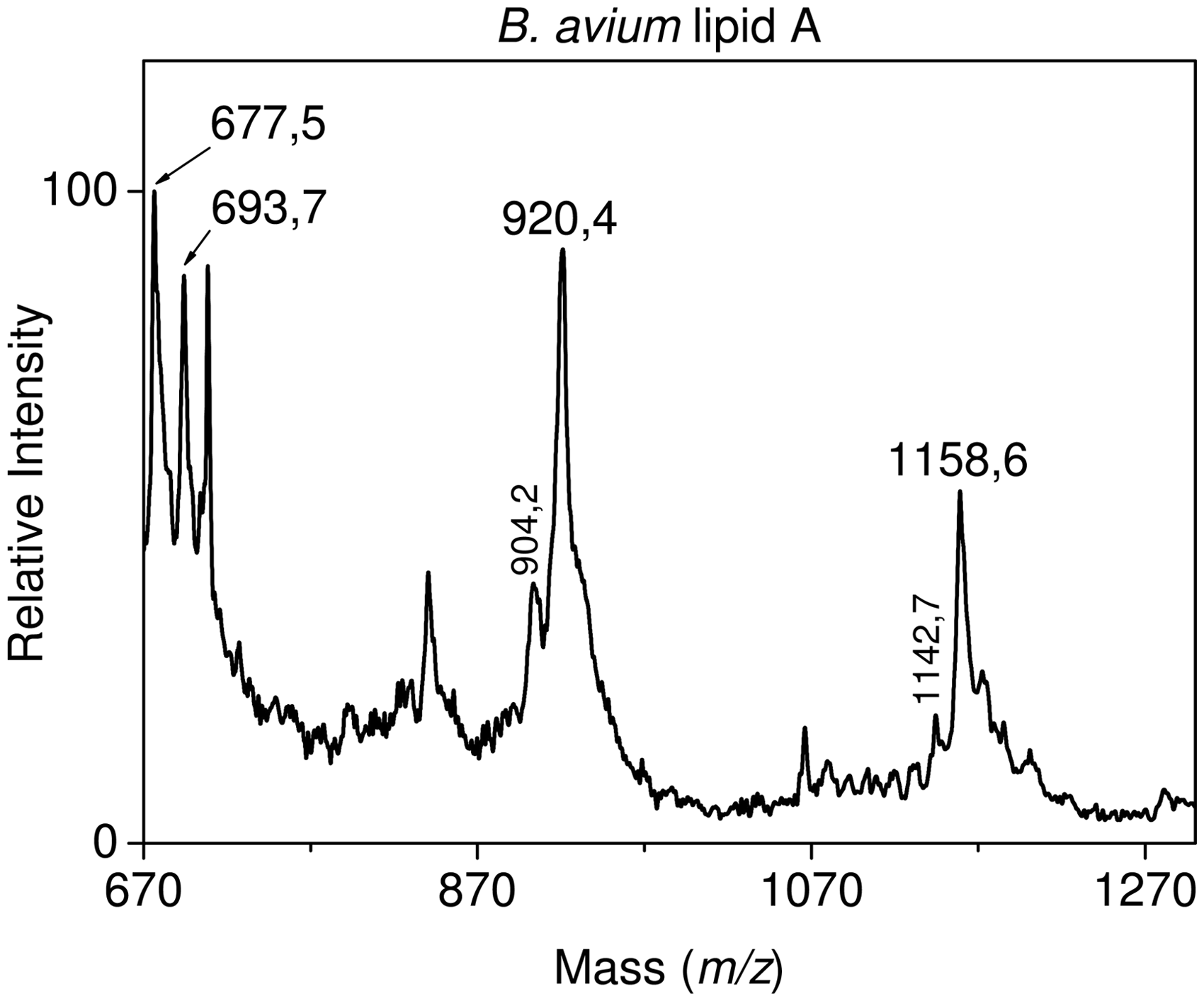
Comparison of the three lipid A structures
Taken together, these data established the lipid A structures of the B. avium, B. trematum and B. hinzii strains studied in this work. The structures of the major hexa-acyl molecular species are presented in Figure 4 (a,b); Figure 4 (c,d) displays B. pertussis and B. parapertussis lipid A structures, respectively, for comparison. The major difference distinguishing B. avium lipid A from that of B. trematum and B. hinzii was the presence of GlcN residues substituting one or both phosphate groups. This type of molecular species was completely absent from B. trematum and B. hinzii lipids A. As for the acylation patterns, they were shown to be very similar for all the three lipids A under study, with tri-, tetra-, penta- and hexa-acyl molecular species in the natural mixtures. The major hexa-acyl molecular species contained 14:0(3-OH) in primary amide C-2 and C-2′ linkages, as well as in a primary ester C-3′ linkage. The C-3 position was empty. Secondary C-2, C-2′ and C-3′ acyloxyacyl positions were occupied respectively by 12:0(2-OH), 14:0(2-OH) and 16:0. The major lower acylated molecular species corresponded to partial structures with the absence of 16:0 in penta-acyl molecular species, with an empty C-3′ position in tetra-acyl molecular species, and with an empty C-3′ position and the absence of 12:0(2-OH) in tri-acyl molecular species. Some micro-heterogeneity was observed due to alterations in hydroxylation of the 12:0(2-OH) and 14:0(2-OH) at secondary C-2 and C-2′ positions (replacing 12:0 and 14:0 respectively) and in the length of the 12:0(2-OH) at secondary C-2 position [replaced by 14:0(2-OH)].
Comparison of lipid A structures of (a), the highest mass molecular species of B. avium (b), the common molecular species of B. avium, B. hinzii and B. trematum (c), the main molecular species of B. pertussis and (d) the highest mass molecular species of B. parapertusssis.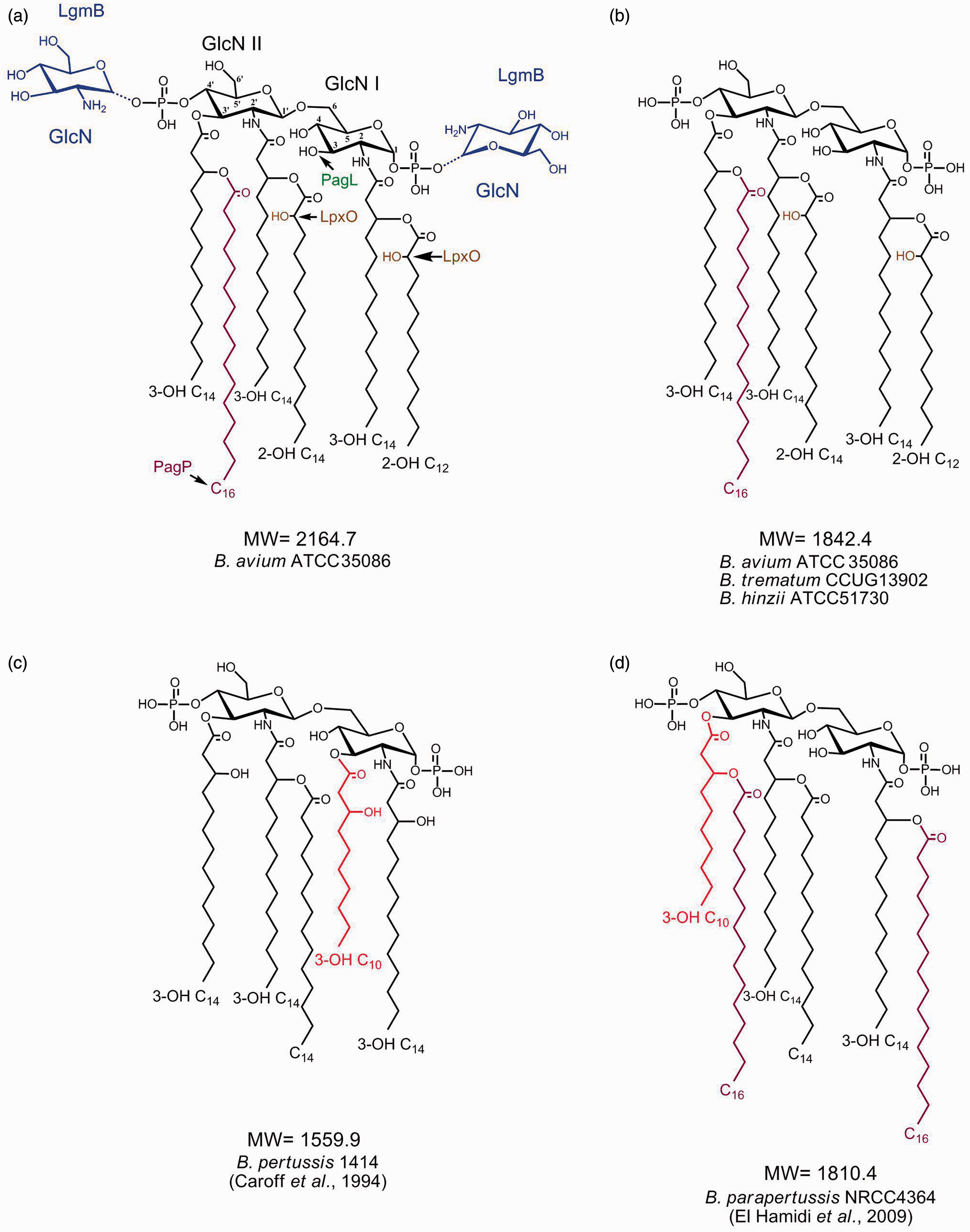
Genetic basis for the three lipid A structures
The lipid A structures of B. avium, B. hinzii and B. trematum suggest numerous modifications to the canonical lipid A structure. For example, the presence of a 16:0 at the secondary C3’ acyloxyacyl position in hexa-acyl species suggests the activity of PagP, an enzyme that functions to add a secondary 16:0 in B. bronchiseptica. 42 Furthermore, the absence of an acyl chain at the C-3 position and the hydroxylation of both the C-2 12:0(2-OH) and C-2′ 14:0(2-OH) suggest activity of PagL and LpxO homologs respectively.43,44 Whole-genome sequencing was performed (N. Shah et al., manuscript in preparation) to identify genes encoding lipid A-modifying enzymes. Sequence analysis of the genomes has revealed the presence of pagP, pagL and two lpxO homologues in each of these three Bordetella strains (Table 1), thus providing a possible genetic basis for these lipid A modifications. Interestingly, there are two sites displaying LpxO activity in these structures, corresponding to two lpxO homologs in the genomes, suggesting that each LpxO is responsible for hydroxylation of a particular secondary acyl chain.
We further analysed the genome sequences to explore the possible reasons as to why the B. hinzii and B. trematum strains lack GlcN-modified lipid A, and found that lgmA, lgmB and lgmC are present in all three strains (Table 1; Figure 5). The lgm locus of B. avium strain ATCC 5086 is almost identical to that of the previously sequenced strain B. avium 197N, except for minor polymorphisms in lgmA (data not shown). The lgm locus architecture for B. hinzii is similar to that of B. pertussis, B. bronchiseptica and B. parapertussis, all three of which have GlcN-modified lipid A. Although lgmD (previously shown to be dispensable in B. pertussis with regard to this GlcN modification
26
) is not found directly following lgmC in B. hinzii, as observed in these other three Bordetella strains: it is located elsewhere in the chromosome (Figure 5). Furthermore, B. avium, which possesses GlcN-modified lipid A, lacks an lgmD homolog, but does have an lgmD-like gene (BAV2926). BAV2926 is similar to lgmD and is primarily comprised of four predicted transmembrane helices. Interestingly, comparison of the B. avium lgm locus with other Bordetella species has revealed the presence of a novel open reading frame that encodes an LgmD-like protein in B. pertussis, B. bronchiseptica, B. parapertussis and B. hinzii in the same location as found in B. avium (Figure 5). Comparing the predicted Lgm proteins between different Bordetella species showed that, although LgmA and LgmB are well conserved among these species (data not shown), LgmC is more variable (Figure 6). In B. avium, the predicted LgmC protein is truncated at the N-terminus due to a stop codon in the 5’ region of lgmC. However, as we detect GlcN-modified lipid A in B. avium, this truncated lgmC must be expressed and functional.
Schematic representation of the lgm locus in Bordetella species: B. pertussis Tohama I, B. bronchiseptica RB50, B. parapertussis 12822, B. avium 197N and ATCC 35086, B. hinzii ATCC 51730, and B. trematum CCUG 13902. lgmA is black, lgmB is grey, lgmC is white, lgmD is vertically striped in light grey and lgmD-like protein is diagonally-striped in light grey. Double line represents the following region is in a different location of the chromosome. The directionality of lgmD in B. trematum is unknown in reference to lgmABC. Locus tags are given for B. bronchiseptica RB50, B. parapertussis 12822 and B. avium 197N; the open reading frame for lgmD-like protein has not been assigned a locus tag and therefore remains unlabelled. ClustalW alignment of LgmC for B. pertussis (Bpe), B. bronchiseptica (Bbr), B. parapertussis (Bpa), B. avium ATCC 35086 (Bav), B. hinzii (Bhi) and B. trematum (Btr). ‘*’: Identical amino acid residues; ‘:’: conserved amino acid residues; ‘.’: semi-conserved amino acid residues.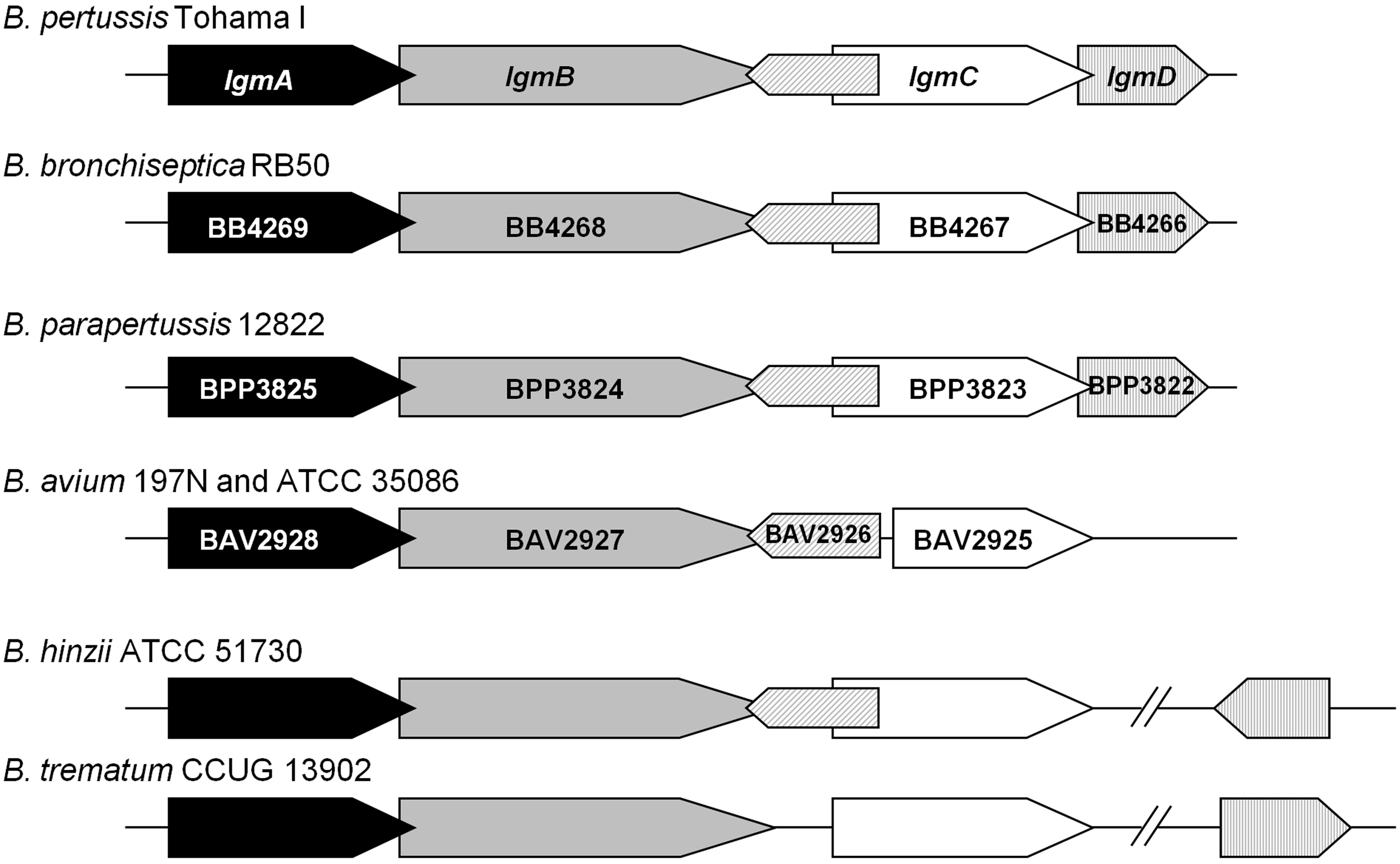

Discussion
Bordetella LPS are complex structures and since the beginning of their characterization they have displayed original traits.45–48 The first mass spectra of native LPS 34 were obtained with the aim of characterizing the apparent originality of the ‘LPS saga’ in this genus. 49 The polysaccharide moieties of B. avium 28 and B. hinzii48,50 have been described by our group, along with the two repetitive units structure present in B. trematum. 17
Owing to their extreme structural diversity and heterogeneity, several physico-chemical factors influence the efficiency of the hydrolytic methods used to split LPS between its lipid A and polysaccharide regions, as well as the integrity of native structures in both moieties. All three lipid As were isolated with the more adapted hydrolysis method. The non-acidic alkaline beta elimination 33 had to be tested with B. avium in order to better maintain the acido-labile glycosidic phosphate substituents. As some dephosphorylation was still observed, we concluded that the dephosphorylation could potentially result from the activity of LpxE, which is present in B. avium (Table 1).
It has been shown by us and others that B. pertussis, 25 B. bronchiseptica 16 and B. parapertussis 51 modify their lipid A with GlcN, and we have experimentally linked this function to the lgm locus (lgmA, lgmB and lgmC). 26 We have also reported that B. pertussis strain 18-323, which lacks this GlcN modification, had a frameshift mutation in lgmB and was also missing lgmC. 26 Here, we report that of the three Bordetella strains studied, only the B. avium lipid A structure shows GlcN-modified phosphates. However, we have found that the lgm locus is present in B. hinzii and B. trematum, with no frameshift modifications or absent genes, as was the case in B. pertussis 18-323. 26 Interestingly, polymorphism was observed in B. avium, as we found that LgmC is truncated, but from the presence of high levels of GlcN found in the lipid A of this strain, it is obviously functional. The polymorphisms observed in the genes comprising the lgm locus did not fall within the predicted active sites of any of the lgm gene products, and thus it is not known whether these differences are functionally relevant.
We propose that the lgm locus genes comprise the following pathway: LgmA, an ArnC ortholog, is predicted to be a glycosyl transferase that adds N-acetyl GlcN to the carrier lipid C55-P; LgmC is predicted to deacetylate this product; and LgmB, an ArnT ortholog, is predicted to transfer GlcN from C-55P-GlcN to lipid A. A flippase may also be involved in this pathway to transfer C-55-P-GlcN from the cytoplasmic side of the membrane to the periplasmic side of the membrane. Members of the gtrA-like Pfam family 52 have been shown to function in this manner, 53 and all the Bordetella strains we have analysed, with the exception of B. trematum, have a gene that encodes for an LgmD-like protein, which is a member of the gtrA-like Pfam family (Figure 5). The lack of an LgmD-like protein in B. trematum may be a possible reason as to why it also lacks the GlcN modification, but this cannot explain why B. hinzii has unmodified lipid A, as an LgmD-like protein is present in this strain. It is worth noting that in B. pertussis, the lgm locus is part of the Bordetella virulence gene (Bvg) regulon. 25 Genome analysis indicates that the bvgAS two-component regulatory system genes are present in B. avium, B. hinzii and B. trematum, suggesting that the lack of GlcN modification of lipid A is not due to the absence of the bvgAS genes. Culture conditions can also affect the expression of genes in the lgm locus; 25 however, in the present work all strains were grown under similar conditions. Therefore, the lack of GlcN modification in these strains may be due to the aforementioned differences in the lgm locus or to other variations in gene regulation and expression.
We previously described the detailed structure of the B. parapertussis lipid A 22 and showed that the GlcN backbone position was free at C-3. Further structural analysis of B. bronchiseptica lipid A from new human and animal bacterial isolates demonstrated that this C-3 position was also free in all the strains we characterized. 23 Removal of the C-3 position fatty acid is a function attributed to the enzyme PagL, and analysis of the corresponding genome sequence reveals the presence of pagL in both B. parapertussis 22 and B. bronchiseptica. 23 It is of note that this position was always found to be substituted with a 10:0(3-OH) fatty acid in B. pertussis strains, 49 indicating that PagL is not functional for this strict human pathogen. 24 Indeed, the presence of short-chain fatty acids in the C-3 position appears to be a signature for the two human Bordetella pathogens responsible for whooping cough: B. pertussis and a specific lineage of B. parapertussis.22,49 It is accepted that these two agents of whooping cough differentiated from B. bronchiseptica, 54 which expresses a high degree of heterogeneity at the level of fatty acid substitution,23,55 the result of relaxed enzyme specificity. 56 This might well be one of the structural characteristics conferring an advantage for infection of human hosts.
As shown in this work, another noticeable structural characteristic was found in B. avium, B. hinzii and B. trematum lipid A, which differed from that of B. bronchiseptica: the presence of a 14:0(2-OH) in acyl-oxyacyl position on the 14:0(3-OH) amide fatty acid at C-2′ replacing 14:0. This modification is due to another late-stage biosynthesis modification involving LpxO, 24 and earlier described in Salmonellae. 44 Such modifications are expected to affect the biological activities of lipid A, as suggested for Burkholderia pseudomallei 57 and for Pseudomonas isolates grown in biofilm medium. 58 The presence of these elements can be variable depending on the strain, natural niche or cultural conditions—something that is also seen in the GlcN substitution of the phosphate groups, as noted above. 25
Bordetella LPS are highly diverse and display a degree of species specificity. Whole-genome analyses has shown that, while the acyl chain modifications observed can be readily traced to the presence of genes encoding the enzymes responsible, the reasons for the glucosamine modifications are more elusive and prove to be much more complex than previously realized. Modifications in lipid A structures have been shown to play a role in modulating immune responses to allow bacteria to escape the immune system. 59 In this regard, a structure-to-activity relationship study is currently under investigation comparing B. hinzii and B. trematum LPS activities to those of B. bronchiseptica. Bordetella avium LPS is also being assessed for the influence of the presence of GlcN residues on its phosphate groups.
Footnotes
Funding
This work was supported by funding from CNRS, with a French–Canadian PICS (Program of International Scientific Collaboration) grant and by funding from the Canadian Institutes of Health Research (grant MOP-102706). N.R.S. was the recipient of a Natural Sciences and Engineering Research Council of Canada CGSD award. A.T. was a recipient of the Agence Universitaire de la Francophonie, S. AN of a Lebanese regional grant and I.T. from the ERASMUS program.
Acknowledgements
We are very grateful to JC Richards, L MacLean and P Fleming, as well as to all of Malcolm B. Perry’s colleagues at the NRC, Ottawa, Canada, who helped us finish this manuscript by providing the necessary bacterial growth cultures after our dear friend and colleague passed away.
