Abstract
The pyrin and HIN-domain (PYHIN) family member1 (pyhin1) is a member of PYHIN proteins and involved in transcriptional regulation of genes important for cell cycle control, differentiation and apoptosis. The regulatory action of mouse pyhin1 on LPS-induced inflammatory response was examined. LPS augmented the pyhin1 mRNA expression in murine RAW 264.7 macrophage cells and peritoneal macrophages. The augmentation of pyhin1 mRNA expression was abolished by parthenolide, a NF-κB inhibitor. Silencing of pyhin1 with small interfering RNA reduced the production of IFN‐β and NO. However, pyhin1 silencing did not affect the production of TNF-α, IL-6, IL-10 and prostaglandin E2. Reduced IFN-β production by pyhin1 silencing caused inactivation of STAT1 and reduced expression of IRF1. Pyhin1 silencing inhibited the expression of TRAF6, TBK1 and TRIF, which trigger IFN-β production in the MyD88-independent pathway. However, pyhin1 silencing did not affect the expression of MyD88, IRAK4 and several mitogen-activated protein kinases in the MyD88-dependent pathway. Taken together, mouse pyhin1 was suggested to be a NF-κB-responsible gene in response to LPS and positively regulate LPS-induced IFN-β and NO production through up-regulating the MyD88-independent signaling pathway.
Introduction
Members of the mammalian pyrin and HIN-domain (PYHIN) family have recently come to prominence as receptors mediating the detection of foreign DNA and initiating innate immune responses.1,2 The PYHIN family member 1 (pyhin1) is a member of PYHIN proteins that belongs to the HIN-200 family of IFN-inducible proteins.1–3 Pyhin1 is primarily a nuclear protein and is involved in transcriptional regulation of genes important for cell cycle control, differentiation and apoptosis.3,4 Down-regulation of the pyhin1 gene is associated with breast cancer 2 and the protein acts as a tumor suppressor by promoting ubiquitination and subsequent degradation of MDM2, which leads to stabilization of p53/TP53. 5 Recently, the functions of human PYHIN proteins, such as absent in melanoma 2 (AIM2) and IFI16, have been uncovered as receptors of intracellular microbial DNA.1,2,6 Thus, PYHIN proteins are involved in defense against infection through recognition of foreign DNA. Furthermore, some family members bind cytosolic DNA via their HIN domains and initiate inflammasome formation via their pyrin domains.4,7 Thus, human PYHIN proteins are involved in defense against some bacteria and viruses. However, the role of the mouse PYHIN family in innate immunity is still unclear. Moreover, the regulatory function of mouse PYHIN proteins in the LPS response has not yet been characterized.
TLRs are pattern recognition receptors that recognize the specific molecular patterns present in microbial components and induce a number of antimicrobial and inflammatory responses. 8 LPS is one of the components of the outer membrane of Gram-negative bacteria that is recognized by TLR4, and triggers the activation of series of signaling pathways, which leads to the production of a number of pro-inflammatory mediators, such as TNF-α, IFN-β, NO and ILs. 9 Excessive production of pro-inflammatory mediators causes systemic inflammatory response syndrome, endotoxic shock and multi-organ failure.10–12 In the present study, we investigated the involvement of mouse pyhin1 in the regulation of LPS-induced pro-inflammatory mediator production in murine RAW 264.7 macrophage cells and peritoneal macrophages. Here, we report that mouse pyhin1 is a NF-κB-responsive protein in response to LPS and positively regulates LPS-induced inflammatory response via augmentation of the MyD88-independent pathway of LPS signaling.
Materials and methods
Materials
LPS from Escherichia coli O55: B5 and poly I:C was purchased from Sigma Chemicals (St Louis, MO, USA). A series of Abs to p65 NF-κB, p38, stress-activated protein kinase/C-Jun N-terminal kinase (JNK), extracellular signal-regulated kinase (ERK) 1/2, IκB kinase (IKK)-α/β, Akt, IRF3, STAT1 and their phosphorylated forms, and HRP-conjugated anti-rabbit IgG were purchased from Cell Signaling Technology (Beverly, MA, USA). Abs to actin, IRF1, IRAK4, TRAF6 and MyD88 were purchased from Santa Cruz Biotechnology (Santa Cruz, CA, USA). Abs to TRIF and TBK1 ware obtained from Imgenex (San Diego, CA, USA) and Epitomics (Burlingame, CA, USA) respectively.
Cell culture
The murine macrophage cell line, RAW 264.7 was obtained from the Riken Cell Bank (Tsukuba, Japan) and maintained with minimum essential medium (MEM) (Sigma, St Louis, MO, USA) containing 5% FCS, non-essential amino acid (Invitrogen, Carlsbad, CA, USA) and antibiotic cocktail at 37℃ under 5% CO2. Peritoneal macrophages were collected by washing out the peritoneal cavity of male BALB/c mice with D-MEM medium (Sigma) and then spun down at 1000 rpm for 5 min. Cells were re-suspended and cultured overnight with D-MEM media containing 10% FCS and antibiotic. After removal of non-adherent cells with washing, adherent cells as macrophages were used for experiments. The animal experiments were carried out following the Guide for Care and Use of Laboratory Animals, Aichi Medical University.
Determination of nitrite concentration
Nitrite, the end product of NO metabolism, was measured using the Griess reagent as described elsewhere. 13 Briefly, RAW 264.7 cells were transfected with pyhin1 or control siRNA and then stimulated with LPS (50 ng/ml) for 24 h. The nitrite concentration in the culture supernatant was determined with reference to the standard curve.
Determination of cytokines concentrations
RAW 264.7 cells were transfected with pyhin1 or control siRNA and then stimulated with LPS (100 ng/ml) for 4 h. The concentrations of TNF-α, IL-6 (R & D System; Minneapolis, MN, USA), IL-10 (Thermo Scientific, Rockford, IL, USA) and IFN-β (PBL Interferon Source, Piscataway, NJ, USA) in the culture supernatant were determined by an ELISA kit.
Immunoblotting
Immunoblotting was performed as described previously. 14 Briefly, the whole cell lysates were extracted by the lysis buffer. The protein concentration of each sample was determined by the bicinchoninic acid protein assay reagent (Pierce, Rockford, IL, USA). Equal amounts of protein (20 µg) were subjected to analysis by SDS-PAGE under reducing conditions. The proteins were transferred electrically to a polyvinylidene fluoride membrane (Millipore, Billerica, MA, USA) and the membranes were treated with a series of appropriately diluted Abs. The immune complexes were detected with HRP-conjugated second Ab at 1:2000 for 1 h. The protein bands were visualized using a chemiluminescence reagent (Pierce). The chemiluminescence was detected by a light capture system analyzer AE6955 (Atto, Tokyo, Japan). For re-probing, the membranes were stripped with the restore Western blot striping buffer (Thermo Scientific) for 15 min and treated with the corresponding Abs. The molecular sizes of the Ags were determined by comparison with a pre-stained protein size marker kit (Invitrogen). To quantify the expression of each molecule, an equal area of each band image on an immunoblotted membrane was gated and calculated as the intensity of band by ImageJ Software (National Institutes of Health, Bethesda, MA, USA).
Semi-quantitative RT-PCR and real-time PCR
Semi-quantitative RT-PCR was performed as described previously. 15 Briefly, RNA was extracted from the cells with a RNeasy mini kit (Qiagen, Valencia, CA, USA). RT-PCR was carried out by using the Access Quick RT-PCR system (Promega, Madison, WI, USA). Primers with the pyhin1 sequence forward 5′-AGATGCTGGAGTGGACCAACTGA-3′ and reverse 5′-TCCGTTTCCCTGTCCGTGTCCT-3′, GAPDH sequence forward 5′-ATGGGGAAGGTGAAGGTCGGAGTC-3′ and reverse 5′-GCTGATGATCTTGAGGCTGTTGTC-3′ were obtained from Invitrogen, and TRIF sequence forward 5′-ATGGATAACCCAGGGCCTT-3′ and reverse 5′-TTCTGGTCACTGCAGGGGAT-3′ from Rikaken (Nagoya, Japan). GAPDH was used as an equal loading control. Optimized RT-PCR conditions were 45℃ for 45 min followed by 95℃ for 2 min, and 32 cycles at 95℃ for 4 s, 62℃ for 30 s and 70℃ for 30 s. The PCR products were analyzed by electrophoresis on 2% agarose gel. The gels were stained with CYBR gold nucleic acid gel stain (Molecular Probes, Invitrogen) and visualized under an ultraviolet transilluminator. The 100-base pair DNA size marker (Invitrogen) was also run to determine the approximate size of the product.
Quantitative real-time PCR was performed as described elsewhere. 16 RNA was reverse-transcribed in RT buffer (Toyobo, Osaka, Japan) according to the manufacturer’s instructions, and quantitative PCR was carried out using SYBR green real-time PCR master mix (Toyobo). PCR was performed with the ABI PRISM 7700 sequence detection system (Applied Biosystems, Hamilton, New Zealand). The relative quantitative expression of pyhin1 was normalized by the expression levels of the reference gene GAPDH. The expression levels of pyhin1 were presented as fold increase to the mean value of the control.
Transfection of small interfering RNA
Transfection of small interfering (si)RNA was done as described previously. 14 Briefly, pyhin1-specific siGENOME SMART pool and a non-targeting siRNA were obtained from Dharmacon (Chicago, IL, USA). Around 2 × 105 cells were seeded in a 24-well culture plate in complete growth medium with 5% FCS and antibiotics. After 12 h incubation under normal growth conditions, the cells were transfected with the Hiperfect transfection reagent (Qiagen, Hilden, Germany), according to the manufacturer’s protocol. Briefly, pyhin1 or control siRNA (400 ng) was diluted in 100 μl/well OPTI–MEM medium (Gibco–BRL, Gaithersburg, MD, USA) and cationic lipid complexes were prepared by incubating for 10–15 min with 6 μl/well Hiperfect transfection reagent. After 8 h incubation, an additional 400 μl of culture medium containing 5% FCS and antibiotics was added to each well and incubated for 48 h. Finally, transfected and controls cells were stimulated with LPS (100 ng/ml) for 4 or 6 h, and the efficiency of pyhin1 silencing was evaluated by PCR.
NF-κB-dependent luciferase assay
RAW 264.7 cells were transfected with pyhin1 or control siRNA and incubated for 48 h. The cells were further transfected with 500 ng/well of NF-κB-Taluc luciferase reporter gene (Invitrogen) and an equal amount of pRL-TK plasmid (Promega) by FuGene HD transfection reagent (Roche Applied Science, Mannheim, Germany) and incubated for 48 h. The transfected and control cells were stimulated with LPS (100 ng/ml) for 6 h. After treatment with a lysis reagent, luciferase activity was determined with the dual luciferase assay kit (Promega). The NF-κB-dependent luciferase activity in the cell lysates was determined with a luminometer. The fold increase was calculated based on the untreated control.
Statistical analysis
Experimental values are represented as the mean ± SD in triplicate. Experiments were carried out at least three times independently. The significance of differences between experimental and control groups was determined by the Student’s t-test. A value of P < 0.01 was considered statistically significant. A typical experimental result of at least three independent experiments is shown in immunoblotting and RT-PCR analysis.
Results
LPS induces the expression of pyhin1 mRNA in macrophages
The effect of LPS on the expression of pyhin1 mRNA was examined in mouse RAW 264.7 macrophage cells and peritoneal macrophage cells (Figure 1). RAW 264.7 cells were stimulated with LPS (100 ng/ml) for various h and the expression of pyhin1 mRNA was determined by semi-quantitative RT-PCR (Figure 1A). Pyhin1 mRNA expression was detected 3 h after LPS stimulation and increased up to 6 h. The high level of pyhin1 mRNA was still expressed 9 h after LPS stimulation. The expression of pyhin1 mRNA in response to various concentrations of LPS was examined (Figure 1B). LPS at 10 ng/ml was effective for the induction of pyhin1 mRNA expression, and LPS at 100 ng/ml induced the maximal expression. Moreover, LPS-induced pyhin1 mRNA expression between RAW 264.7 cells and peritoneal macrophages was compared (Figure 1C). LPS induced the expression of pyhin1 mRNA in mouse peritoneal macrophages, as well as RAW 264.7 cells, although peritoneal macrophages expressed a lower level of pyhin1 mRNA compared with RAW 264.7 cells.
Expression of pyhin1 mRNA in macrophages in response to LPS. (A) RAW 264.7 macrophage cells were stimulated with LPS (100 ng/ml) for various lengths of time. (B) RAW 264.7 cells were stimulated with various concentrations of LPS for 6 h. (C) RAW 264.7 cells and peritoneal macrophages were stimulated with various concentrations of LPS for 6 h. The expression of pyhin1 mRNA was determined by semi-quantitative RT-PCR (A, B) or real-time PCR (C). *P < 0.01 versus untreated control.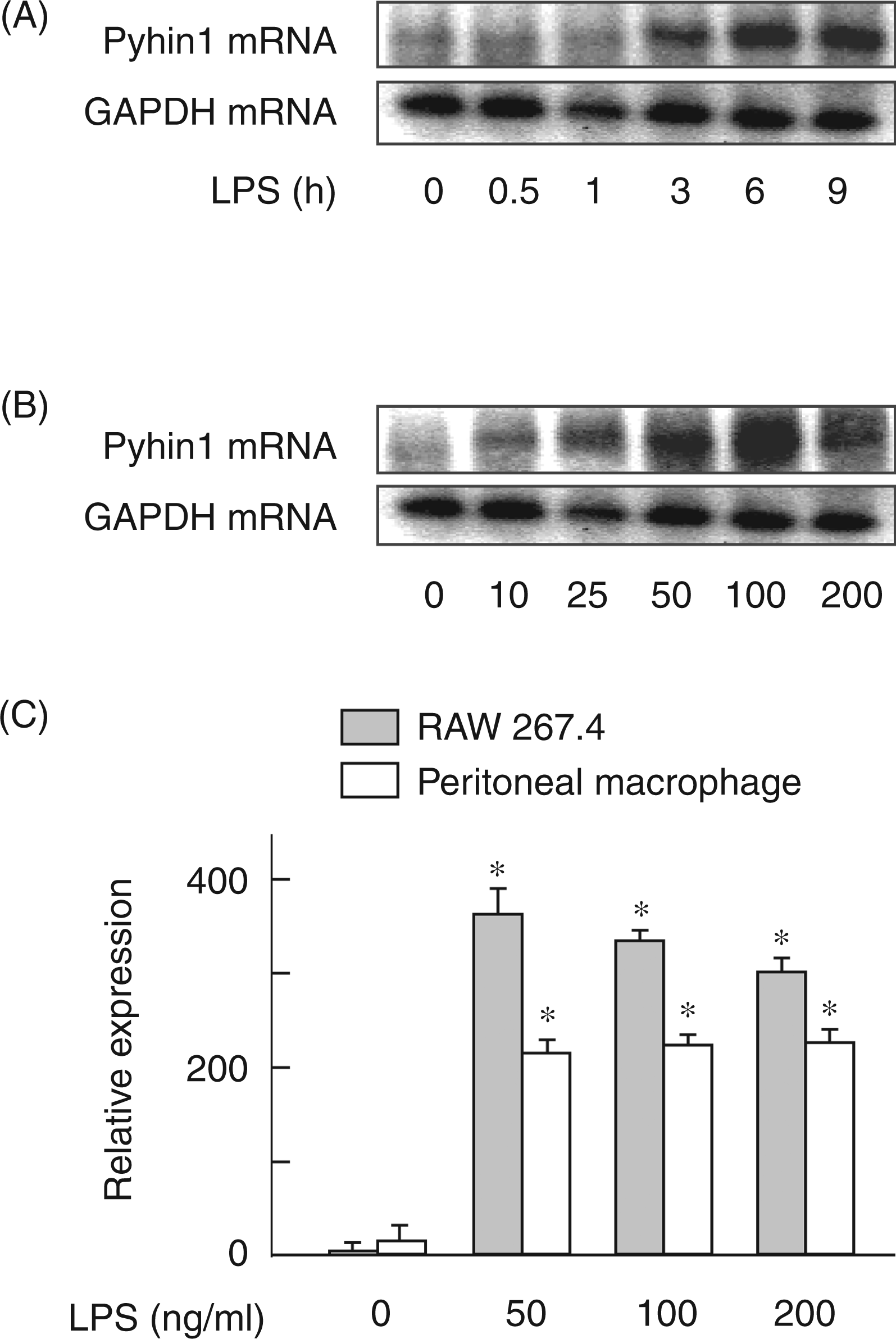
LPS-induced pyhin1 mRNA expression is mediated by early activation of NF-κB
In order to clarify the signaling molecule(s) involved in LPS-induced pyhin1 mRNA expression, the effect of a series of pharmacological signaling inhibitors was examined (Figure 2). RAW 264.7 cells were pretreated with SB203580 (a p38 inhibitor), JNK inhibitor-II, PD98059 (a ERK1/2 inhibitor) and parthenolide (a NF-κB inhibitor) for 30 min, and then stimulated with LPS (100 ng/ml) for 6 h. LPS clearly induced the expression of pyhin1 mRNA in untreated control cells. However, parthenolide as a NF-κB inhibitor almost completely inhibited pyhin1 mRNA expression. However, PD98059, SB203580 and JNK inhibitor-II did not affect the pyhin1 mRNA expression in response to LPS. In addition, anti-IFN-β Ab did not affect LPS-induced pyhin1 mRNA expression (data not shown).
Effect of various signaling inhibitors on LPS-induced pyhin1 mRNA expression. RAW 264.7 cells were pretreated with PD98059 (5 µM), parthenolide (5 µM), SB203580 (10 μM) or JNK inhibitor II (5 μM) for 30 min, and then stimulated with LPS (100 ng/ml) for 6 h. The expression of pyhin1 mRNA was determined by RT-PCR.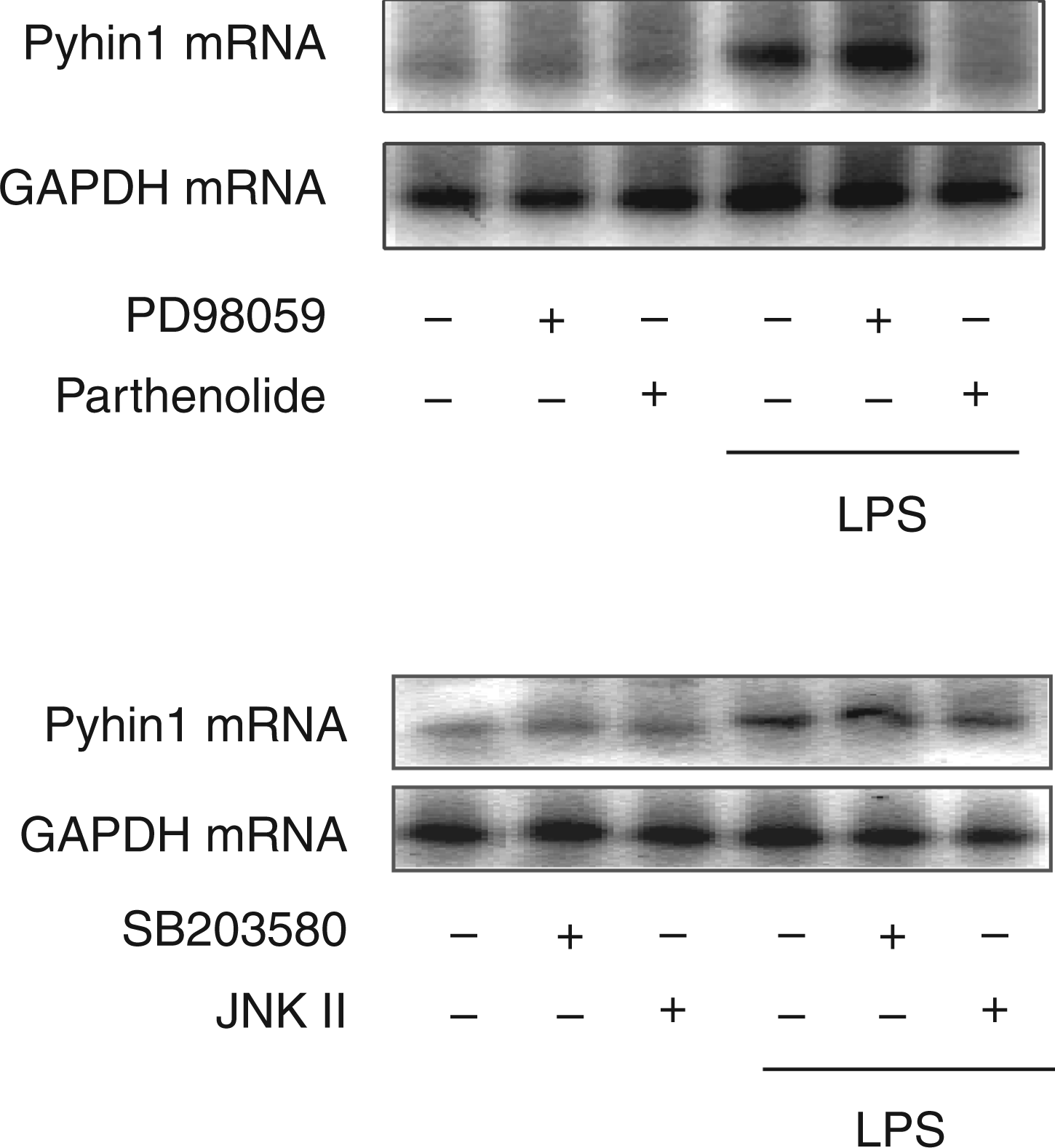
Silencing of pyhin1 reduces LPS-induced IFN-β production
The binding of LPS on TLR4 triggers production of various pro-inflammatory mediators, such as TNF-α, ILs, IFN-β and NO via the MyD88-dependent or independent pathway.12,17 The effect of pyhin1 silencing on the production of LPS-induced pro-inflammatory mediators was examined with siRNA (Figure 3). First, pyhin1 siRNA was confirmed to specifically inhibit the expression of pyhin1 mRNA (Figure 3A). The cells were transfected with pyhin1 or control siRNA and stimulated with LPS (100 ng/ml) for 4 h. The levels of IFN-β, TNF-α, Il-6 and IL-10 in the culture supernatant were determined by ELISA (Figure 3B). Silencing of pyhin1 exclusively reduced the production of IFN-β in response to LPS, whereas it did not affect the production of TNF-α, IL-6 or IL-10. The reduced intracellular IFN-β expression with pyhin1 silencing was confirmed with immunoblotting (Figure 3C). As parthenolide inhibited LPS-induced pyhin1 mRNA expression, the effect of parthenolide on LPS-induced IFN-β production was examined (Figure 3D). Parthenolide, as well as pyhin1 silencing, inhibited LPS-induced IFN-β production.
Effect of pyhin1 silencing on LPS-induced cytokines production. (A) Pyhin1 or control siRNA-transfected cells were stimulated with LPS (100 ng/ml) for 6 h. The efficiency of pyhin1 siRNA was determined by the expression of the mRNA. (B, C) Pyhin1 or control siRNA-transfected RAW 264.7 cells were stimulated with LPS (100 ng/ml) for 4 h. The expression levels of various cytokines were determined with ELISA (B) or immunoblotting (C). (D) Cells transfected with pyhin1 siRNA or treated with parthenolide (5 µM) for 30 min were stimulated with LPS (100 ng/ml) for 4 h. The concentration of IFN-β in the culture supernatant was determined by ELISA. *P < 0.01 versus LPS alone.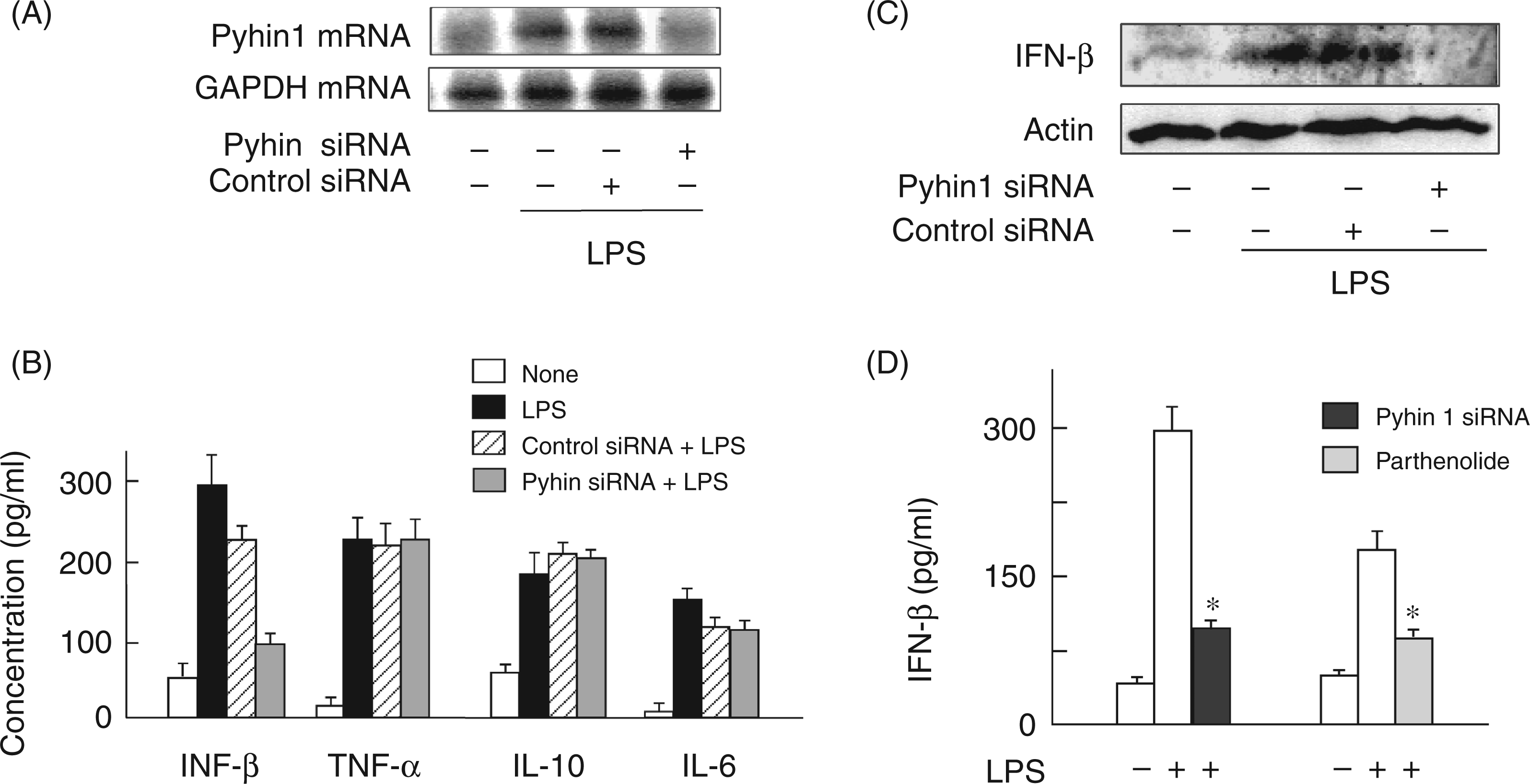
Silencing of pyhin1 reduces iNOS-dependent NO production in response to LPS
Silencing of pyhin1 exclusively inhibits LPS-induced IFN-β production, which is a key event in the MyD88-independent pathway of LPS signaling.
18
Therefore, the effect of pyhin1 silencing on IFN-β-dependent gene expression in response to LPS was examined. As LPS-induced IFN-β production is known to lead to the iNOS-dependent NO production,19,20 the effect of pyhin1 silencing on the iNOS-dependent NO production in response to LPS was examined (Figure 4). Pyhin1 or control siRNA-transfected cells were stimulated with LPS (50 ng/ml) for 24 h. LPS clearly induced the NO production in control siRNA-treated cells, whereas it induced lower level of NO production in pyhin1 siRNA-treated cells (Figure 4A). The effect of pyhin1 silencing on LPS-induced iNOS expression was also examined. Silencing of pyhin1 markedly inhibited LPS-induced iNOS protein expression (Figure 4B), although significant iNOS expression was detected in control cells.
Effects of pyhin1 silencing on iNOS-dependent NO production in response to LPS. Pyhin1 or control siRNA-transfected cells were stimulated with LPS (50 ng/ml) for 24 h (A) and LPS (100 ng/ml) overnight (B). The concentration of nitrite from culture supernatant was determined by Greisse reagent (A) and the expression of iNOS protein was determined by immunoblotting (B). *P < 0.01 versus LPS alone.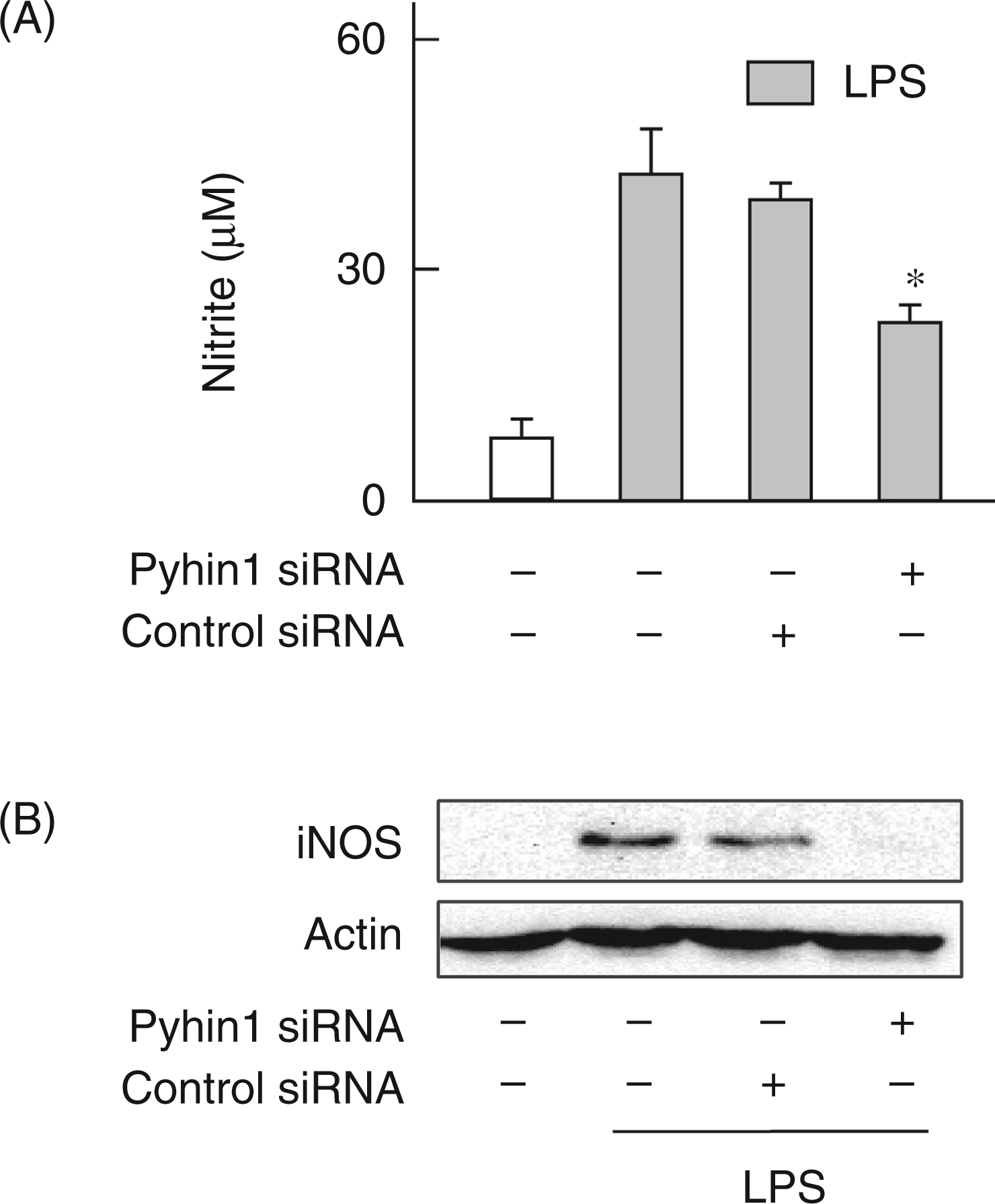
Silencing of pyhin1 inhibits the activation of STAT1 and IRF1
IFN-β is reported to induce NO production via activation of STAT1 and expression of IRF1 in the MyD88-independent pathway.
19
The effect of pyhin1 silencing on the activation of STAT1 and IRF1 in response to LPS was examined (Figure 5). LPS induced the phosphorylation of STAT1 (Figure 5A) and augmented the expression of IRF1 protein (Figure 5B). However, they were significantly prevented by the silencing of pyhin1.
Effect of pyhin1 silencing on LPS-induced STAT1 activation and IRF1 expression. Pyhin1 or control siRNA-transfected RAW 264.7 cells were stimulated with LPS (100 ng/ml) for 4 h. The phosphorylation of STAT1 (A) and the expression of IRF1 (B) were determined by immunoblotting.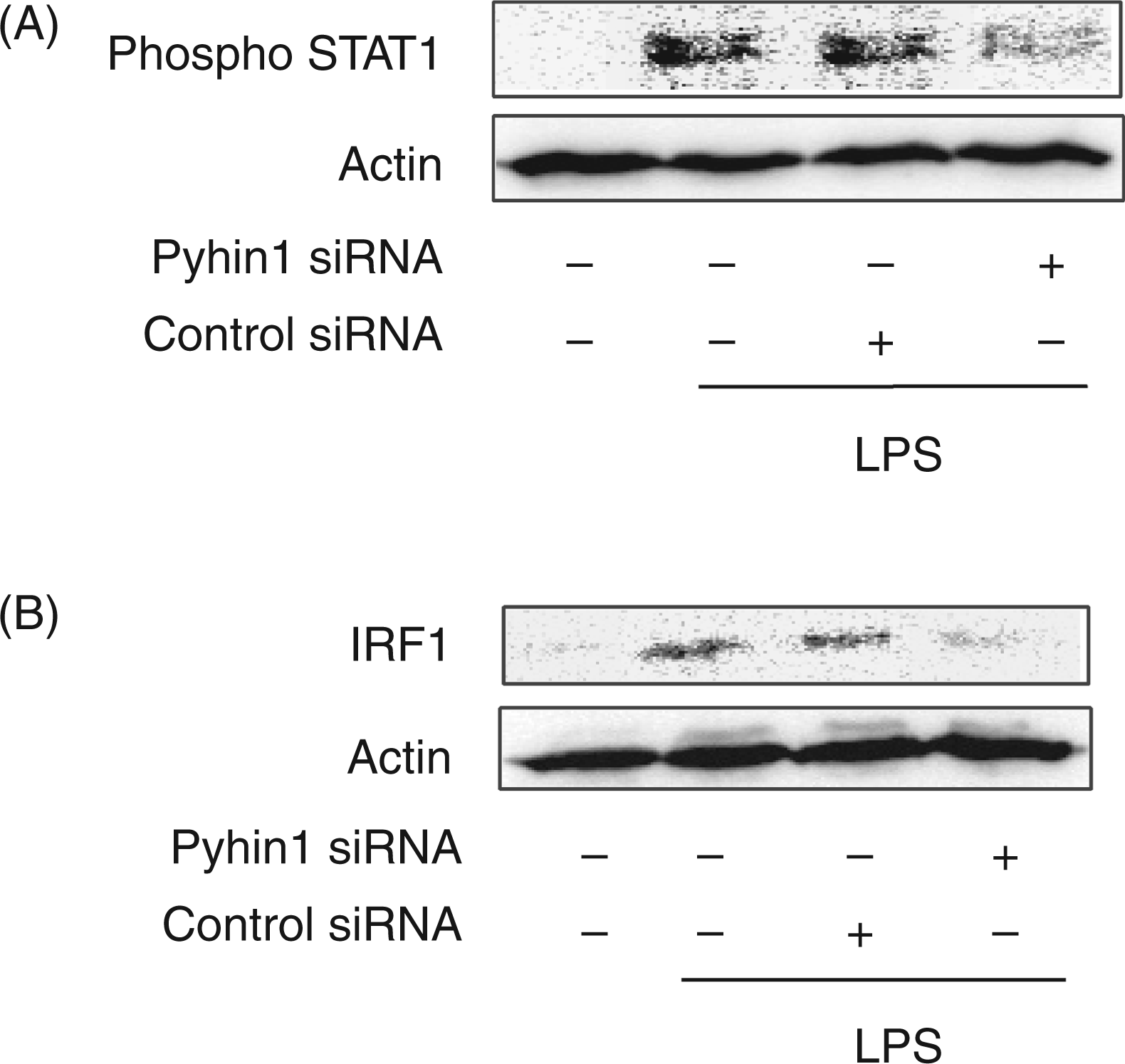
Silencing of pyhin1 down-regulates the activation of IRF3 and NF-κB, which is required for LPS-induced IFN-β production
The late phase activation of NF-κB and IRF3 in the MyD88-independent pathway triggers LPS-induced IFN-β production.21,22 Therefore, the effect of pyhin1 silencing on the phosphorylation of IRF3 and NF-κB in response to LPS were examined (Figure 6). Pyhin1 or control siRNA-transfected cells were stimulated with LPS (100 ng/ml) for 45 min and 4 h, and then phosphorylation of p65 NF-κB was determined by immunoblotting. Although LPS induced the late stage phosphorylation of p65 NF-κB, silencing of pyhin1 profoundly inhibited it at 4 h (Figure 6A). However, pyhin1 silencing did not affect the early NF-κB activation 45 min after LPS stimulation (Figure 6A). Similarly, silencing of pyhin1 reduced the NF-κB-dependent luciferase activity 4 h after LPS stimulation (Figure 6B). The effect of pyhin1 silencing on the activation of IKKα/β as an upstream molecule of NF-κB was examined (Figure 6C). Silencing of pyhin1 inhibited the phosphorylation of IKKα/β at 4 h in response to LPS. Furthermore, the effect of pyhin1 silencing on the phosphorylation of IRF3 was examined (Figure 6D). Silencing of pyhin1 also inhibited LPS-induced IRF3 phosphorylation at 4 h.
Effect of pyhin1 silencing on LPS-induced NF-κB and IRF3 activation. Pyhin1 or control siRNA-transfected RAW264.7 cells were stimulated with LPS (100 ng/ml) for 45 min (A) or 4 h (A, C, D), or 6 h (B). Mock (C) was transfected with empty vector. The phosphorylation of p65 NF-κB, IKKα/β and IRF3 was determined by immunoblotting (A, C, D). The activation of NF-κB was determined by a luciferase reporter gene assay (B). *P < 0.01 versus control siRNA.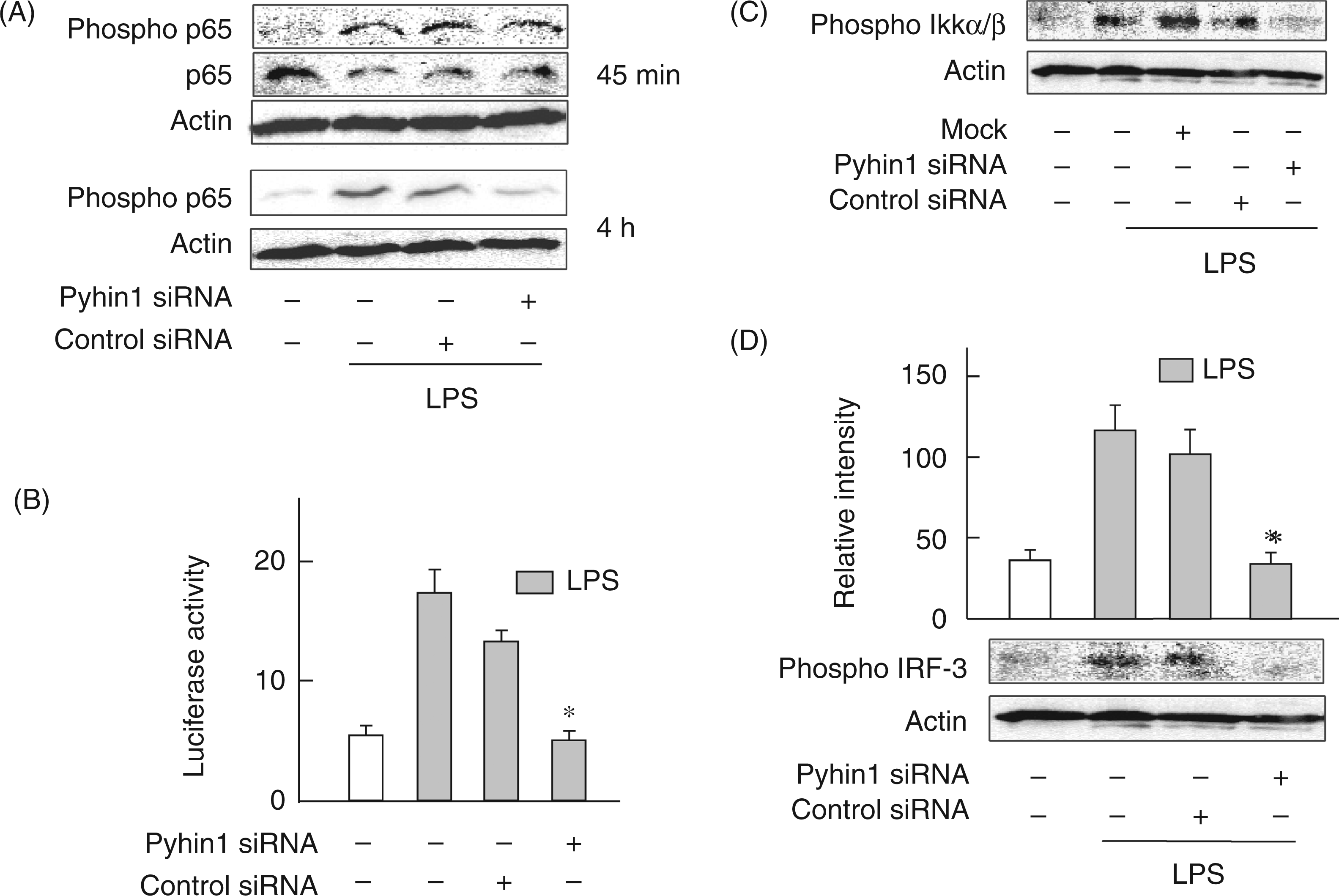
Silencing of pyhin1 inhibits the expression of TRIF in the MyD88-independent pathway
In the preceding section, the reduction of LPS-induced IFN-β production by pyhin1 silencing was suggested to be mediated by the inactivation of upstream molecules of IRF3 and NF-κB at a late stage, which regulate IFN-β production. Therefore, the effect of pyhin1 silencing on the activation of the upstream molecules in MyD88-independent pathway was examined (Figure 7). LPS-induced TBK1, TRAF6 and TRIF expression in pyhin1 or control siRNA-transfected cells were analyzed by immunoblotting (Figure 7A). The expression of TBK1, TRAF6 and TRIF in response to LPS was reduced by pyhin1 silencing. Transfection with siRNA had some cytotoxic action against cells. Therefore, control siRNA slightly reduced the expression of TRIF and TRAF6. Furthermore, the mRNA expression of TRIF, the most upstream molecule among them, was examined (Figure 7B). Silencing of pyhin1 clearly inhibited the TRIF mRNA expression, whereas control siRNA did not affect it.
Effect of pyhin1 silencing on the expression of various signaling molecules in response to LPS. (A–C) Pyhin1 or control siRNA-transfected RAW264.7 cells were stimulated with LPS (100 ng/ml) for 4 h. The expression of a series of indicated signaling molecules were determined by immunoblotting (A, C) and RT-PCR (B) respectively.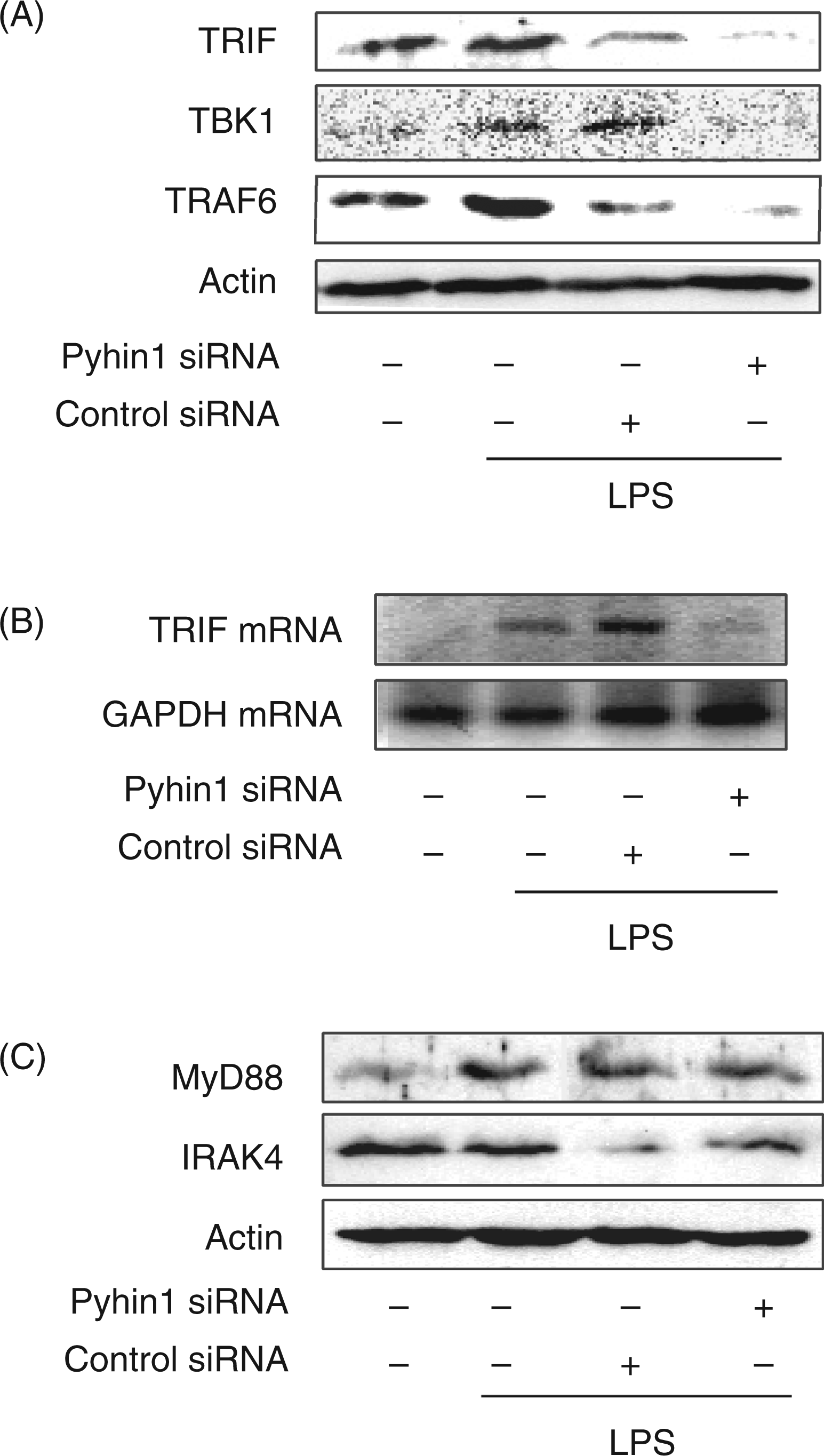
In order to confirm no effect of pyhin1 silencing on the MyD88-dependent pathway, the effect of pyhin1 silencing on the expression of IRAK4 and MyD88 was examined (Figure 7C). Immunoblotting analysis demonstrated that silencing of pyhin1 did not affect the expression of IRAK4 and MyD88 in the MyD88-dependent pathway.
Silencing of pyhin1 reduces poly I:C-induced IFN-β production
The effect of pyhin1 silencing on the production of poly I:C-induced IFN-β production was examined (Figure 8). The cells were transfected with pyhin1 or control siRNA, and stimulated with poly I:C (100 µg/ml) for 4 h. Silencing of pyhin1 reduced the production of IFN-β in response to poly I:C, as well as LPS.
Effect of pyhin1 silencing on poly I:C-induced IFN-β production. Pyhin1 or control siRNA-transfected RAW 264.7 cells were stimulated with poly I:C (100 µg/ml) for 4 h. The level of IFN-β was determined with ELISA.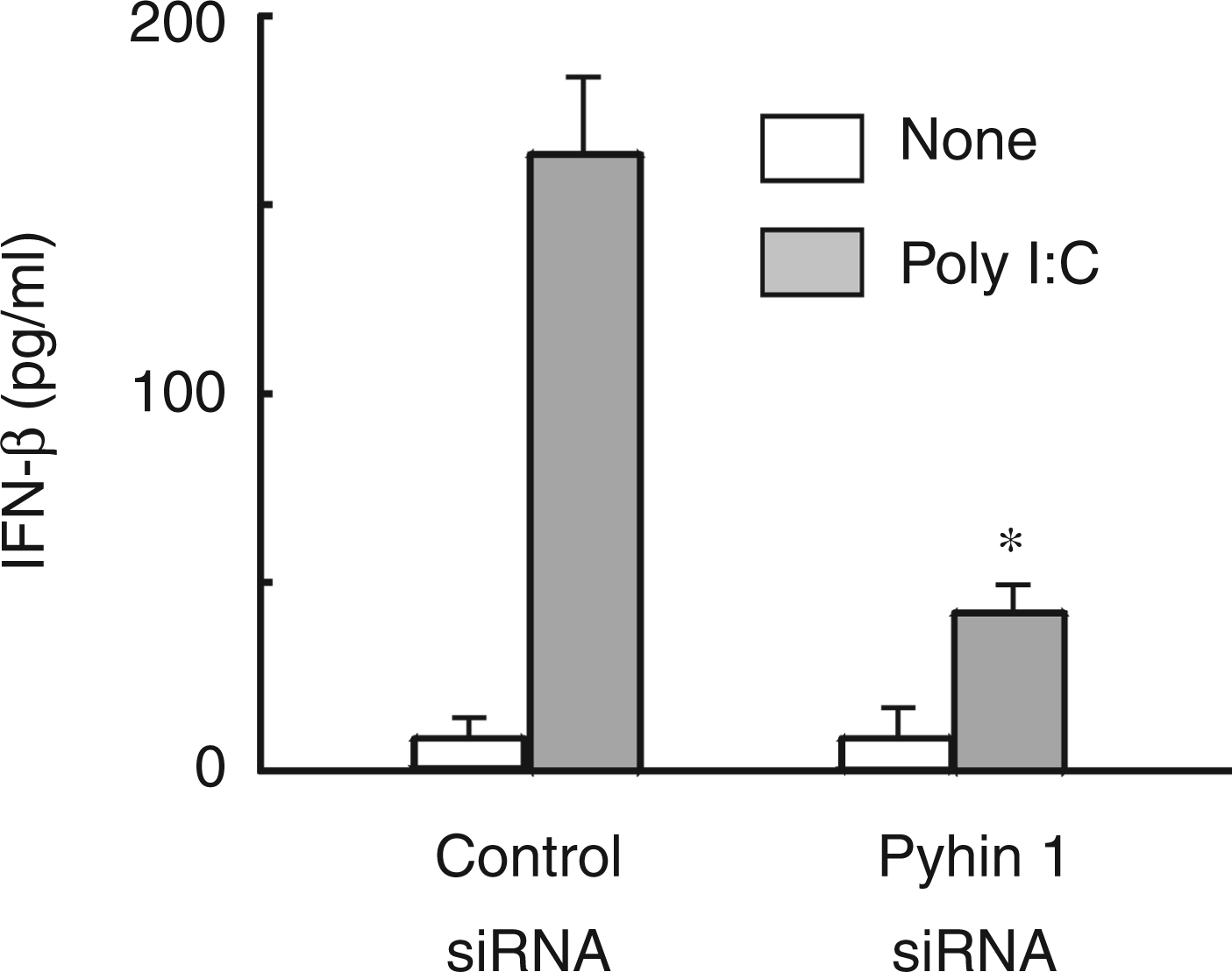
Discussion
In the present study, we have demonstrated that the expression of mouse pyhin1 is induced by LPS in macrophages and that it augments LPS-induced NO production. A putative schematic diagram is shown in Figure 9. Several lines of evidence suggest that mouse pyhin1 is a positive regulator of the MyD88-independent pathway, but not the MyD88-dependent one; first, silencing of pyhin1 reduces LPS-induced IFN-β production, characteristic of the MyD88-independent pathway; second, it down-regulates the expression of TRIF consisting of the MyD88-independent pathway, but not the expression of MyD88; third, it does not affect the activation of NF-κB and MAPKs in the MyD88-dependent pathway; finally, pyhin1 inhibits the IFN-β production in response to poly I:C, which exclusively triggers MyD88-independent signal pathway. Once again, mouse pyhin1 is suggested to exclusively up-regulate the MyD88-independent pathway in response to LPS.
A putative schematic diagram of LPS-induced pyhin1 expression.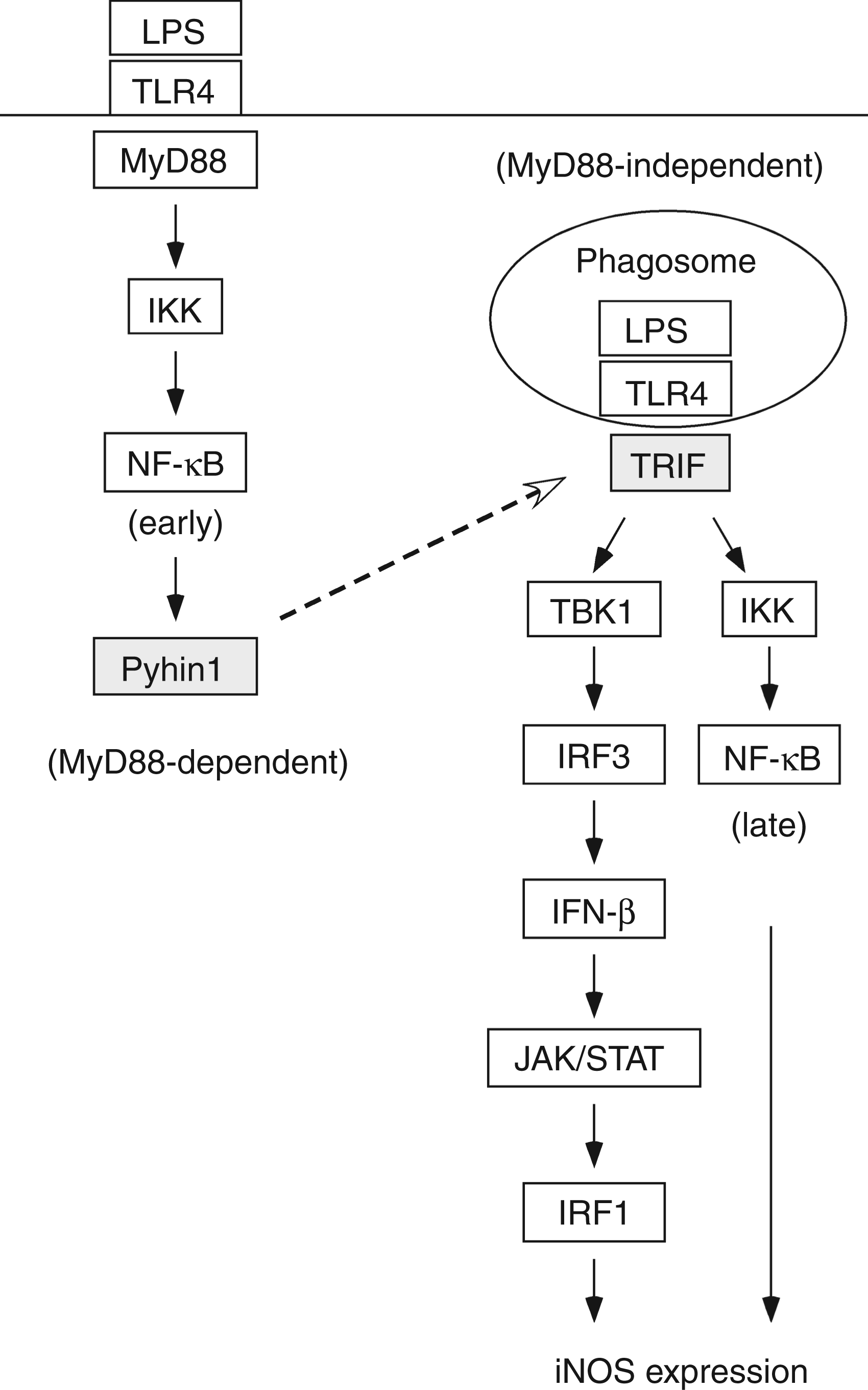
It is certain that pyhin1 does not affect the MyD88-dependent pathway as the expression of pyhin1 mRNA cannot be detected until 3 h after LPS stimulation. The MyD88-dependent pathway would be already triggered before pyhin1 is expressed. It is also supported by the fact that pyhin1 silencing does not affect the production of TNF-α, IL-6 and IL-10, which is dependent on the MyD88-dependent pathway. LPS induces the expression of pyhin1 via the MyD88-dependent pathway, and, once expressed, pyhin1 regulates the MyD88-independent pathway. Mouse pyhin1 might be an important regulator of LPS signaling.
In the present study, we, for the first time, demonstrated that mouse pyhin1 as a member of PYHIN protein family is a NF-κB-responsive protein in response to LPS, although human PYHIN proteins, such as IFI16 and IFIX, are known as IFN-inducible proteins.2,3 LPS appears to induce pyhin1 expression via the early NF-κB activation in the MyD88-dependent pathway as a NF-κB inhibitor prevents LPS-induced pyhin1 expression. Interestingly, pyhin1 siRNA inhibits LPS-induced NF-κB activation at 6 h, but not 45 min. Therefore, once expressed, pyhin1 is suggested to up-regulate the late NF-κB activation in the MyD88-indepedent pathway. In response to LPS, mouse pyhin1 is a NF-κB responsive protein and a positive feedback regulator of NF-κB activation.
A putative target molecule of pyhin1 might be TRIF. As pyhin1 augments LPS-induced IFN-β production, a putative target of pyhin1 must be the upstream signal molecule triggering LPS-induced IFN-β production. Pyhin1 silencing reduces the expression of TRIF, TBK1 and TRAF6, and TRIF is the most upstream signal molecule among them. Therefore, TRIF is likely to be a target of pyhin1 in response to LPS. If so, augmented expression of TRIF with pyhin1 could certainly enhance the NF-κB activation and IFN-β production in the MyD88-independent pathway in response to LPS.
LPS induces the expression of pyhin1 in physiological macrophages, as well as RAW 264.7 cells. Why does LPS induce the expression of pyhin1 in macrophages? Human IFI16 and its murine ortholog p204 is the first PYHIN protein shown to be involved in IFN-β induction as a sensor of DNA. 5 IFI16 containing the similar functional domain, like pyhin1, functions as an innate immune sensor for cytosolic DNA,2,6 as well as nuclear DNA 23 to initiate different innate immune responses and induce IFN-β production. Therefore, macrophages might promote the ability of sensing intracellular bacterial DNA in response to LPS. Pyhin1 might be involved in the immunosurveillance against microbial DNA in bacterial infection. LPS-stimulated macrophages may also sense DNA released from damaged host cells in endotoxic shock via pyhin1. It is still a matter for speculation that mouse pyhin1 induced by LPS may function as a DNA sensor in macrophages.
AIM2, a protein of the mammalian PYHIN (IFI200/HIN-200) family, binds cytosolic DNA via its HIN domain and initiates inflammasome formation via its pyrin domain. 24 Therefore, LPS might enhance the formation of inflammasome via mouse pyhin1 expression and lead to inflammatory responses. Mouse pyhin1 may positively augment LPS-induced inflammatory response via regulation of inflammasome formation, although it must await further characterization.
The present study demonstrates that the inhibition of pyhin1 expression prevents the production of LPS-induced pro-inflammatory mediators, such as IFN-β and NO. It suggests that pyhin1 might be a potential target for control of endotoxic shock. It is of particular interest to clarify the clinical significance of pyhin1 expression in endotoxin-related diseases.
In summary, mouse pyhin1 is induced by LPS in macrophages and is suggested to positively regulate LPS-induced IFN-β and NO production via MyD88-independent pathway. Mouse pyhin1 might be a positive regulator of LPS response in macrophages.
Footnotes
Acknowledgements
We are grateful to K. Takahashi for the technical assistance.
Funding
This work was supported, in part, by a Grant-in-Aid for Scientific Research from the Ministry of Education, Science, Sports and Culture of Japan, and a grant of Strategic Research Foundation Grant-aided Project for Private Universities from Ministry of Education, Culture, Sports, Science, and Technology, Japan (MEXT).
Conflicts of interest
The authors do not have any potential conflicts of interest to declare.
