Abstract
The role played by prolactin (PRL) in fish immunity is scant. We report here that stimulation of the Atlantic salmon monocytic cell line SHK-1 with native salmon PRL resulted in activation of the respiratory burst and induction of the expression of the genes encoding the phagocyte NADPH oxidase components p47phox, p67phox and gp91phox, and the transcription factor IFN regulatory factor-1 (IRF-1). Interestingly, the pharmacologic inhibition of the Jak/Stat signaling pathway with AG490 blocked reactive oxygen species (ROS) production, and the induction of genes encoding the NADPH oxidase components and IRF-1 in PRL-activated SHK-1 cells. In addition, PRL promoted the phosphorylation of Stat and induced the DNA binding activity of IRF-1. These results, together with the presence of several consensus target motifs for Stat and IRF-1 in the promoter of the tilapia p47phox gene, suggest that PRL regulates p47phox gene expression in fish through the activation of these two key transcription factors. Taken together, our results demonstrate that PRL induces the expression of the genes encoding the major phagocyte NADPH oxidase components and ROS production in fish macrophages via the JAK2/Stat/IRF-1 signaling pathway.
Introduction
The immune and neuroendocrine systems are intimately linked and are involved in bidirectional communications. Prolactin (PRL) is a 23-kDa polypeptide that is synthesized primarily in the pituitary and is secreted not only by the anterior pituitary gland, but also by many extra-pituitary sites, including immune cells. An even greater correlation between PRL and immune regulation is observed in animal models, where circulating PRL levels can be altered by hypophysectomy, bromocriptine treatment or genetic deletions. For example, hypophysectomized animals are deficient in mounting various B- and T-cell-mediated immune responses, which are restored by PRL injections.1,2
In fish, various immunologic functions have been attributed to PRL, such as regulation of the phagocytic activity of fish leukocytes,3–5 the proliferation of leukocytes in chum salmon (Oncorhynchus keta) and the circulating levels of IgM in rainbow trout (Oncorhynchus mykiss).6,7 However, data regarding the specific effects of PRL on fish immunity are still not conclusive, particularly in regard to innate immunity and, more specifically, NADPH oxidase—the enzyme responsible for reactive species oxygen (ROS) production. ROS refers to a group of highly reactive molecules that includes oxygen anions and free radicals, such as superoxide anion (O2−), hydroxyl radical (OH−) and hydrogen peroxide (H2O2), 7 and are generated endogenously in response to hyperoxia, hypoxia, growth factors and cytokines. 8 In this regard, we have demonstrated recently that PRL is able to induce the production ROS in head kidney leukocytes from the teleost fish gilthead seabream (Sparus aurata) via protein kinase C (PKC)-mediated phosphorylation of the NADPH oxidase component p47phox.9,10 Interestingly, PRL is also able to induce the expression of two major pro-inflammatory cytokines secreted by activated monocytes and macrophages, namely IL-1β and TNF-α, through the NF-κB signaling pathway. 9 Therefore, we hypothesized that the activation of NADPH oxidase might also be mediated through the induction of the genes encoding the NADPH oxidase components.
IFN regulatory factor-1(IRF-1), which belongs to a small family of 11 IRF proteins, 11 regulates the expression of a number of genes important for mediating antiviral and antibacterial responses, apoptosis, cytokine signaling, T-helper 1 immune responses, NK-cell differentiation, and dendritic cell and macrophage functions.12,13 It has been reported that the tyrosine residues Y580 and Y309/Y382 of the PRL receptor (PRLR) are important for the induction of IRF-1 via STAT5 and STAT1, respectively, in Nb2 lymphoma cells.14,15 In fact, PRL stimulates IRF-1 gene expression in normal rat leukocytes derived from bone marrow and spleen, and in human granulocytes.16,17 However, the transcription of the gp91phox gene, a key component of NADPH oxidase, is tightly modulated by inflammatory mediators, such as LPS, TNF-α and IFN-γ, in myeloid cells18,19 by the cooperation of STAT1, PU.1 and IRF-1 transcription factors. 20 IRF-1 binds to highly conserved DNA motifs, known as IFN-stimulating response elements (ISREs), found at the promoter of gp91phox gene. 20 Interestingly, infection of neutrophil precursors with Anaplasma phagocytophilum, the causative agent of human granulocytic ehrlichiosis, results in down-regulation of the gp91phox gene, a phenomenon that is associated with reduced expression of IRF-1 in nuclear extracts of A. phagocytophilum-infected cells. 21 Therefore, IRF-1 is a key player in the regulation of NADPH oxidase components.
Little is known about the nucleotide sequence and activity of IRF-1 in teleosts. In fact, this molecule has only been characterized in Japanese flounder (Paralichthys olivaceus), 22 pufferfish (Fugu rubripes), 23 rainbow trout (O. mykiss), 24 turbot (Scophthalmus maximus), seabream, 25 snakehead (Channa argus) 26 and large yellow croaker (Pseudosciaena crocea). 27 Although the ISRE motif has been identified in the IRF-1 promoter of rainbow trout, 24 Atlantic salmon 28 and snakehead, 26 no functional studies have been performed to date.
In the present study, we found that PRL was able to prime the production of ROS and to induce the expression of IRF-1 and several NADPH oxidase components, including p47phox, p67phox and gp91phox, in Atlantic salmon and gilthead seabream leukocytes. In addition, by using pharmacological inhibitors, Western blot, RT-qPCR, electrophoretic mobility shift assay (EMSA) and a complementary bioinformatics approach of the p47phox gene promoter of tilapia (Oreochromis niloticus), we obtained data that strongly suggest that PRL stimulates the expression of genes encoding NADPH oxidase components through a Jak2/Stat/IRF-1 signaling pathway.
Materials and methods
Cell culture and treatments
SHK-1 is a cell line from Salmo salar head kidney with macrophagic properties.
29
The cells were cultured at 18°C in 75 cm2 tissue culture-treated flasks (Corning B.V. Life Sciences, Amsterdam, The Netherlands), in L-15 medium (500 ml with 300 mg/l
ROS detection assay
O2− production by SHK‐1 cells was measured using NBT reduction.31,32 Approximately 106 cells per well were plated, grown for 24 h and stimulated for 4 h. The medium was then removed and the cells were washed with 5 ml PBS. Filtered 0.25% p-Nitro-Blue tetrazolium (NBT) (5 ml) was added and the cells were incubated at 18°C for 60 min to allow the O2− to reduce the NBT to blue formazan. The supernatants were removed, and the wells were washed thoroughly with methanol and then allowed to air dry. The insoluble blue formazan was solubilized by adding 120 µl/well 2 M potassium hydroxide and then 140 µl/well DMSO (Sigma–Aldrich). The contents of the wells were then mixed immediately to complete solubilization and give a final solution with an intense turquoise colour. The OD630 of this solution was read on an ELISA reader, using as blank a well without cells, which was incubated with the NBT solution and subjected to the same fixing, washing and solubilization steps.
Analysis of gene expression
Total RNA was extracted from cell pellets with TRIzol reagent (Life Technologies), following the manufacturer’s instructions, and treated with DNase I, amplification grade (1 U/µg RNA; Invitrogen). The SuperScript III RNase H ReverseTranscriptase (Life Technologies) was used to synthesize first-strand cDNA with oligo-dT18 primer from 1 µg total RNA at 50°C for 50 min. Real-time PCR was performed with an ABI PRISM 7500 instrument (Life Technologies) using SYBR Green PCR Core Reagents (Applied Biosystems). Reaction mixtures were incubated for 10 min at 95°C, followed by 40 cycles of 15 s at 95°C, 1 min at 60°C, and, finally, 15 s at 95°C, 1 min at 60°C and 15 s at 95°C. For each mRNA, gene expression was normalized to the ribosomal protein S11 (rps11) content in each sample using the comparative Ct method (2 −ΔΔ Ct). The primers used were p47phox (FJ594437): forward 5’-gaggagcctgaagaagctga-3’, and reverse 5’-tcc agcagcttgtgaatgac-3’; p67phox (FJ594436): forward 5’-tgtgtccacaatggtcgtct-3’ and reverse 5’-gcactgtccctcaaaccagt-3’; gp91phox (FJ594435): forward 5’-gtgttcggctatgaggtggt-3’ and reverse 5’-ggtctcaggacacagccaatt-3’; IRF-1 (EF067841.1): forward 5’-taaacgacaaggggttccag-3’ and reverse 5’-cgcagctctatttccgtttc-3’; RPS11 (NM001141447) forward 5’-acagaaatgccccttcactg-3’ and reverse 5’-gcctcttctcaaaacggttg-3’; PRLR (DQ508436.1) forward 5’-tcaacatcttcagcctgcac-3’ and reverse 5’-ggatccacatggattgttcc-3’. In all cases, each PCR was performed with triplicate samples and repeated with at least two independent samples.
Immunoprecipitation and Western blotting
SHK-1 cells were lysed in ice-cold NP-40 lysis buffer containing 1% Nonidet P-40, 25 mM Tris-HCl (pH 7.5), 150 mM sodium chloride, 1 mM EDTA, 5 mM sodium fluoride, 1 mM sodium orthovanadate, 1 mM leupeptin and 1 mM phenylmethylsulfonyl fluoride.
For the immunoprecipitation studies, cell lysates were incubated with the Stat5 polyclonal Ab (1:1000, #sc‐835) or Jak2 (1:1000, #sc-294) (both from Santa Cruz Biotechnology, Heidelberg, Germany) for 4 h. 9 Cell lysates were then mixed with protein A/G–agarose beads and stirred for 1 h at 4°C. The beads were washed three times with ice-cold NP-40 lysis buffer, and the precipitated proteins were boiled for 5 min and eluted with SDS-PAGE sample buffer.
Western blot analysis was carried out according to a standard protocol. 33 Briefly, leukocytes were lysed at 4°C and the BCA protein assay kit (Pierce, Rockford, IL, USA) was used to determine the protein concentrations of the cell lysates. Equal amounts of protein were resolved by SDS-PAGE and transferred onto nitrocellulose membranes (Schleicher & Schuell, Keene, NH, USA). After transfer, the membranes were blocked overnight in 5% dry milk and probed for 1 h with 4G10-Platinum, anti-phosphotyrosine mouse monoclonal cocktail (1:2500, #05-1050; Millipore, Madrid, Spain) or anti-p47phox (1:1000, #sc-14015; Santa Cruz Biotechnology) or anti-actin (1:4000, #sc-25778; Santa Cruz Biotechnology) or with rabbit polyclonal Abs produced against an internal epitope mapping from the amino acid residue 116 to 137 (1:1000, KPNNKRSKANNVKKNNKGSQIK) of salmon IRF-1 (ABO38869) or against an internal epitope (1:1000, KENSETVYECPDYHT) of salmon PRLR (ABF57668), and then for 1 h with a HRP-coupled secondary Ab (1:5000) (Sigma-Aldrich). The reactive bands were visualized via an enhanced chemiluminescence detection system (GE Health Care Life Sciences, Barcelona, Spain) according to the manufacturer’s procedures. The intensity of the signals was measured with a digital imaging system (IMAGE J).
EMSA
Nuclear and cytoplasmic extracts were prepared from SHK-1 cells, according to the method of Sadowski et al. 34 The double-stranded oligonucleotides used as probes in the EMSA were IRFe-1: 5′-actggactgacacggtttcactttgcagtgtctactatatt-3′/5′-aatatagtagacactgcaaagtgaaaccgtgt- cagtccagt-3′. Binding reactions and electrophoresis were performed as described previously.9,35 In the supershift assays, 1:10 anti-IRF-1 polyclonal Ab was added 20 min before or after adding the probe. 9
In silico analysis
Potential motifs and domains were searched using the Tfsitescan (http://www.ifti.org/cgi-bin/ifti/Tfsitescan.pl) and further improved manually by comparison with human or mouse NADPH oxidase components (http://www.ensembl.org/index.html). Finally, the MEME/MAST system version 3 (http://meme.nbcr.net/meme/cgi-bin/mast.cgi) for motif discovery and search was used to predict conserved sequence domain STATx and ISRE in the p47phox promoter region.
Statistical analysis
All data are shown as means ± SE. Differences were evaluated using ANOVA followed by a Student’s t-test. Statistical significance was defined as P < 0.05.
Results
PRL activates the respiratory burst of SHK-1 cells
RT-PCR and Western blot were used to confirm the expression of PRLR in SHK-1 cells at transcript and protein levels respectively (Figure 1A, B). Then, the time course of ROS production was analyzed in SHK-1 cells treated with PRL (25–1000 ng/ml) and LPS (positive control
36
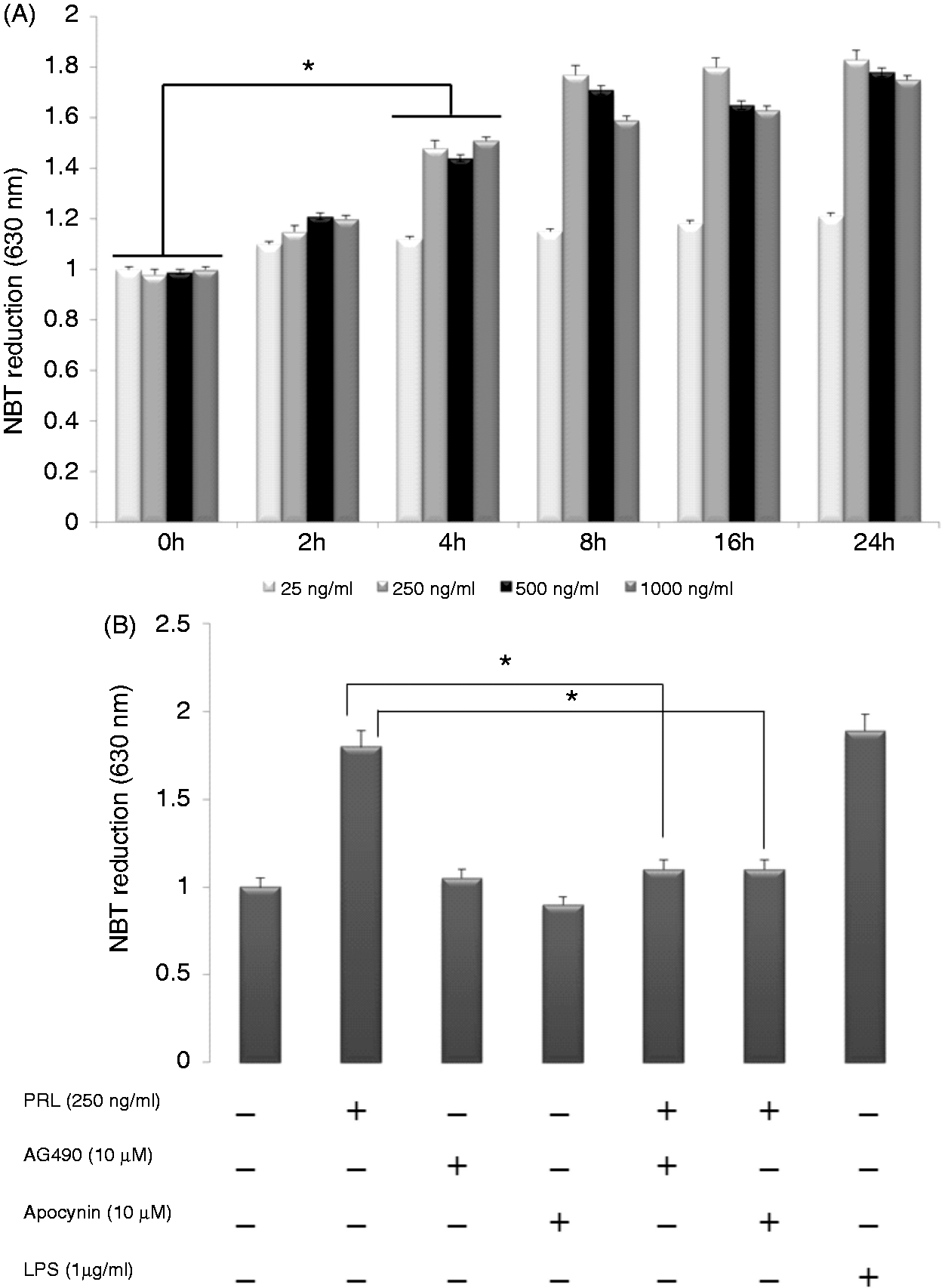
), as measured by NBT reduction. All the doses used are within the known physiological ranges.37,38 As shown in Figure 2A, 250 ng/ml PRL increased the amount of ROS by about 55% and 83% (P < 0.05) at 4 h and 8 h, respectively, as seen by when the maximal activity reached. Interestingly, 500 ng/ml and 1000 ng/ml failed to further increase ROS production, but resulted in a lower activity than 250 ng/ml at most time points. We next examined the effect of apocynin, a NADPH oxidase inhibitor, to confirm the involvement of this enzymatic complex in superoxide anion production. The PRL-mediated increase in ROS observed at 4 h was completely inhibited by 10 µM apocynin (Figure 2B). Additionally, the ROS production induced by PRL was attenuated by 10 µM AG490, a Jak2 inhibitor, which inhibits the phosphorylation of Jak2 in these cells.
10
We have shown previously that Jak2 and PKC are required for the leukocyte-mediated production of O2− in PRL-activated seabream leukocytes.9,10 Taken together, these data, indicate that PRL-induced ROS production in two distant phylogenetic species, i.e. S. salar and S. aurata, involves the Jak2/PKC-dependent activation of NADPH oxidase.
PRLR expression in SHK-1 cells. (A) The presence of PRLR was evaluated by RT-PCR amplification. Lane 1: negative control without template; lane 2: PRLR amplicon of 144 bp and (B) Western blot analysis of SHK-1 cell extracts with anti-PRLR Ab. Lane 1: total protein extract; lane 2: control Ab; lane 3: specific signal of PRLR using the anti-PRLR Ab. PRL activates the respiratory burst of SHK-1 cells. (A) SHK-1 cells were treated with PRL at the indicated concentrations for 0, 2, 4, 8, 16 and 24 h. The reduction of NBT in SHK-1 was then measured. Values are normalized to unstimulated cells and represent the mean ± SE of three independent experiments. *P < 0.05 vs unstimulated cells and (B) SHK-1 cells were treated for 4 h with PRL (250 ng/ml) alone or in the presence of the indicated concentrations of the pharmacological inhibitors AG490 (Jak) or apocynin (NADPH oxidase). As a positive control, LPS (1 µg/ml) was added to SHK-1. Values are normalized to unstimulated cells and represent the mean ± SE of three independent experiments. *P < 0.05 vs cells stimulated with PRL and in the presence of AG490.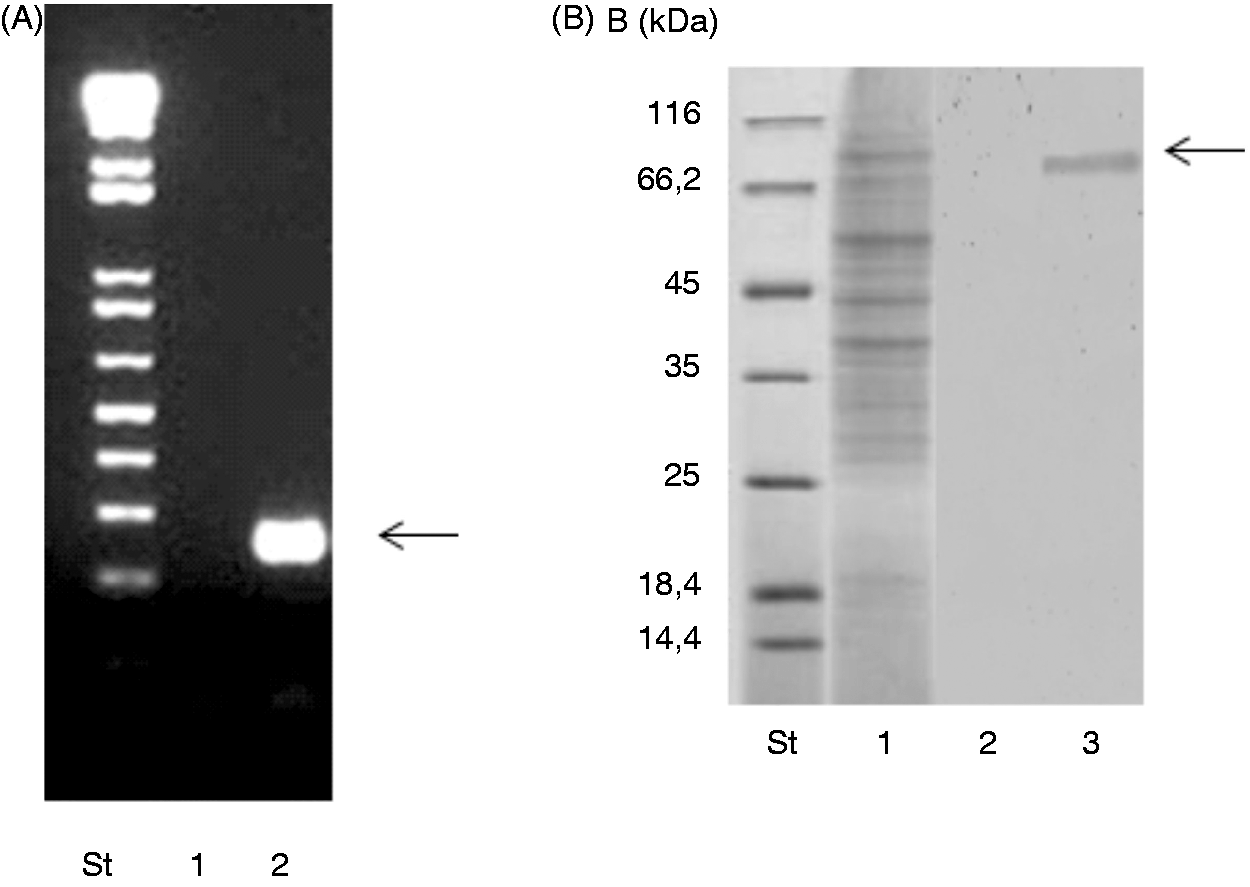
PRL induces p47phox, p67phox and gp91phox through the Jak/Stat pathway
The NADPH oxidase consists of a membrane-bound flavocytochrome b558 (gp91phox) and the four cytosolic subunits p47phox, p67phox, p40phox and Rac1/2.39–41 In the sets of experiments described in the following, we evaluated the effect of PRL on p47phox, p67phox and gp91phox gene expression by measuring their mRNA levels. PRL treatment resulted in a time-dependent increase in the expression of p47phox, p67phox and gp91phox, with the highest expression being observed at 8 h post-stimulation (Figure 3A). Quantitative analysis indicated maximum increases in p47phox, p67phox and gp91phox mRNA levels of 4.55, 3.05 and 2.05-fold (Figure 3A), respectively, in SHK-1 cells treated with 250 ng/ml PRL compared with untreated control cells (P < 0.05). To determine whether the increase in p47phox mRNA was associated with an increase in p47phox protein in SHK-1, Western blot analysis was performed on cell lysates obtained from PRL-treated and untreated SHK-1, taking the advantage of the availability of an Ab against salmon p47phox. A time-dependent increase in p47phox protein was also observed, following exposure to 250 ng/ml PRL (Figure 3B). The strongest immunoreactivity for p47phox protein was observed at 8 and 16 h. Semi-quantitative evaluation of p47phox protein was measured by scanning densitometry and using actin as housekeeping marker (Figure 3B). The results showed that p47phox synthesis had increased 3.7-fold 16 h post-treatment with 250 ng/ml PRL (P < 0.05).
Expression patterns of p47phox, p67phox and gp91phox genes in SHK-1 cells incubated with PRL (250 ng/ml). (A) The expression of p47phox, p67phox and gp91phox was evaluated by real-time RT-PCR at different times. Gene expression is normalized against rps11 and is shown as relative to the mean of unstimulated SHK-1 cells. Each bar represents the mean ± SE of triplicate samples. (B) Cell extracts from SHK-1 cells stimulated with 250 ng/ml PRL for the indicated times was evaluated by Western blot with an Ab directed against p47phox (1:1000). The signals for p47phox were quantified using Image J software and the ratio of p47phox/actin was calculated after densitometry detection. The results shown are representative of three independent experiments. (C) SHK-1 cells were treated for 8 h with PRL (250 ng/ml) alone or in the presence of the indicated concentrations of AG490. Values are normalized to unstimulated cells and represent the mean ± SE of three independent experiments. *P < 0.05 vs cells stimulated with PRL and in the presence of the lowest concentration of the inhibitor. (D, E) Cell extracts from SHK-1 stimulated with 250 ng/ml PRL for 30 min alone or in the presence of the Jak2 inhibitor AG490 were immunoprecipitated (IP) with an Ab directed against Jak2 (1:1000) or Stat5 and then probed with anti-Jak2 or anti-Stat5 (1:1000), respectively, or the anti-phosphotyrosine 4G10-Platinum (1:2500) Abs. The signals for p-Jak2 and total Jak2, or for p-Stat5 and total Stat5 were quantified using Image J software and the ratio of phosphorylated p-Jak2/Jak2 and p-Stat5/Stat5 were calculated after densitometry detection. The results shown are representative of three independent experiments.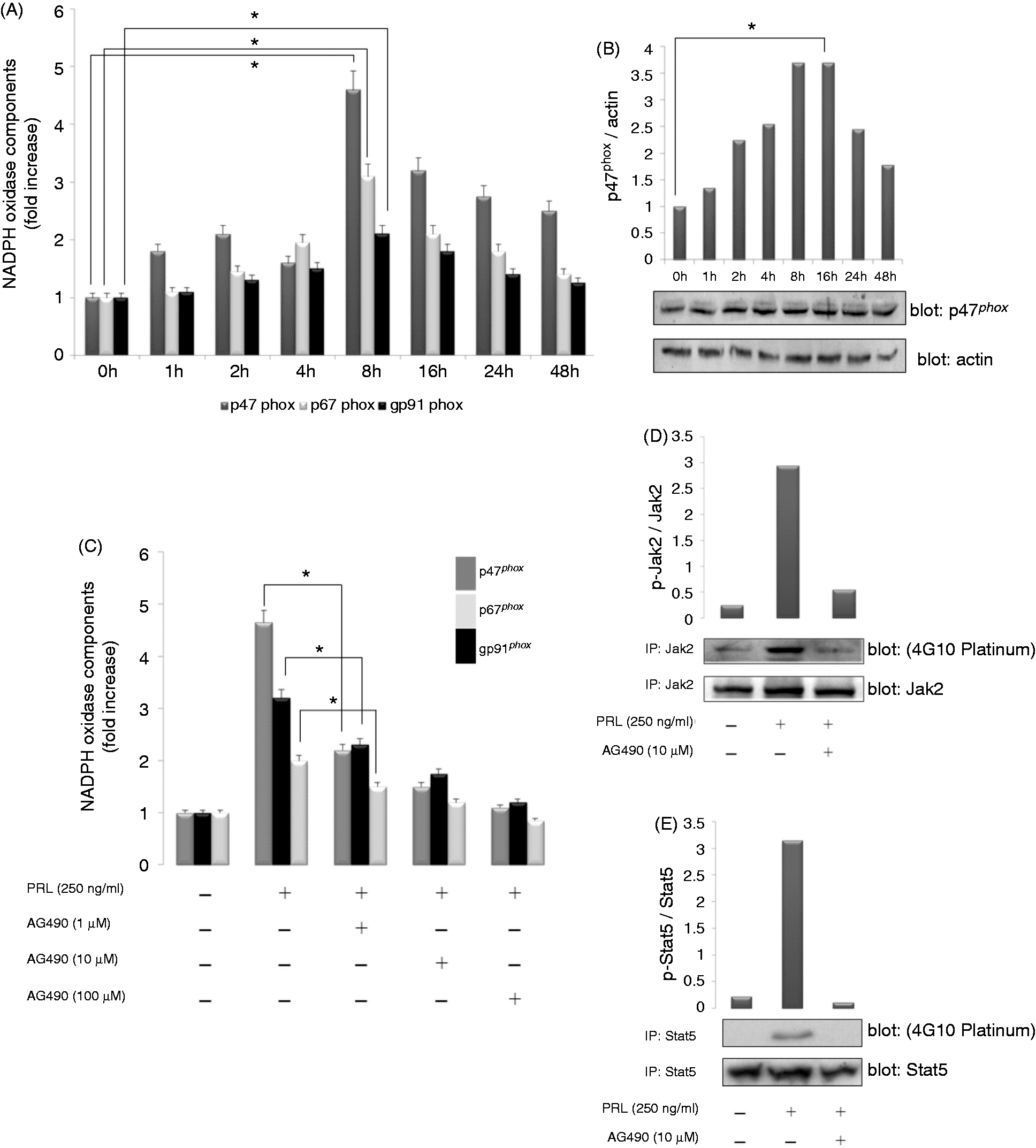
To assess whether the effect of PRL on the induction of p47phox, p67phox and gp91phox in SHK-1 was dependent on PRLR, SHK-1 cells were pretreated for 1 h with the Jak2 inhibitor AG490 (1, 10 and 100 µM) before adding 250 ng/ml PRL and leaving for 8 h. As seen in Figure 3C, pretreatment with AG490 strongly diminished induction of the genes coding for these three NADPH oxidase components in SHK-1 cells, compared with SHK-1 cells treated only with PRL (P < 0.05).
In light of these results and in order to clarify the mechanism underlying PRL-induced transcription of the genes encoding NADPH oxidase components, we investigated whether Stat5 became phosphorylated following PRL stimulation. The results showed that a 30-min exposure to PRL increased the phosphorylation of Jak2 (Figure 3D) and Stat5 (Figure 3E) 2.95 and 3.15-fold, respectively, compared with unstimulated cells. Notably, PRL-induced Jak2 and Stat5 phosphorylation was inhibited by AG490 (10 µM) (Figure 3D, E), whereas apocynin had no effect (data not shown). Collectively, these data indicate that PRL induced the expression of the genes encoding p47phox, p67phox and gp91phox in SHK-1 cells via the Jak2/Stat signaling pathway.
Blockage of Jak/Stat signaling pathway attenuates the gene expression and activity of IRF-1
As PRL is able to stimulate IRF-1 gene expression in rat leukocytes
16
and in human granulocytes,
17
and the transcription of the gp91phox gene is modulated by inflammatory mediators though the cooperation of STAT1, PU.1 and IRF-1 transcription factors,
20
we hypothesized that the PRL-mediated induction of genes coding for the NADPH components in SHK-1 cells is orchestrated by IRF-1. Real-time RT-PCR analysis demonstrated that PRL treatment of SHK-1 cells resulted in increased IRF-1 mRNA levels (∼9.5-fold increase) within 2 h, peaking (∼11.5-fold increase) 8 h post-stimulation (Figure 4A). Western blotting analysis also confirmed the induction of IRF-1 at protein level (Figure 4B). Consistent with previous results,9,10 Jak2 inhibition with 1 and 10 µM AG490 blocked PRL-induced IRF-1 gene expression in SHK-1 cells (Figure 4C). Finally, when an EMSA was performed using a probe containing an ISRE to evaluate IRF-1 activity in PRL-stimulated SHK-1 cells, protein binding to the ISRE probe was apparent within 1 h of stimulation with PRL (Figure 4D). The binding of this protein complex to the probe was specific because it was abolished by an excess of homologous competitor (Ho) (Figure 4D). Additionally, supershift assays with the anti-IRF-1 Ab and pre-incubation of SHK-1 cells with 10 µM of AG490 (Figure 4D) confirmed that PRL was able to activate Jak2/Stat signaling pathway in SHK-1 and that this led to the induction and activation of IRF-1.
Blocking of Jak2 attenuates PRL-induced IRF-1 expression and activity. (A) The expression of IRF-1 was evaluated by real-time RT-PCR at different times. Gene expression is normalized against rps11 and is shown as relative to the mean of unstimulated SHK-1 cells. Each bar represents the mean ± SE of triplicate samples. (B) Cell extracts from SHK-1 cells stimulated with 250 ng/ml PRL for the indicated times was evaluated by Western blot with an Ab directed against IRF-1 (1:1000). The signal for IRF-1 was quantified using Image J software and the ratio of IRF-1/actin was calculated after densitometry detection. The results shown are representative of three independent experiments. (C) SHK-1 cells were treated for 2 h with PRL (250 ng/ml) alone or in the presence of the indicated concentrations of the AG490. Values are normalized to unstimulated cells and represent the mean ± SE of three independent experiments. *P < 0.05 versus cells stimulated with PRL and in the presence of the lowest concentrations of the inhibitor and (D) EMSA was performed using nuclear extracts obtained from SHK-1 cells stimulated with 250 ng/ml PRL for 45 min and the oligonucleotide probe containing ISRE. Three identical EMSA experiments were performed to confirm reproducibility using nuclear extracts from SHK-1 cells with 10 µM AG490, as well as a cold competitor probe (Ho) (5- and 50-fold molar excess) and the Ab to IRF-1 added 20 min before (b) or immediately after (a) adding the probe. The arrows indicate the protein–DNA complexes or protein–DNA–Ab (supershift) observed in SHK-1 cells stimulated with PRL. Free: probe only.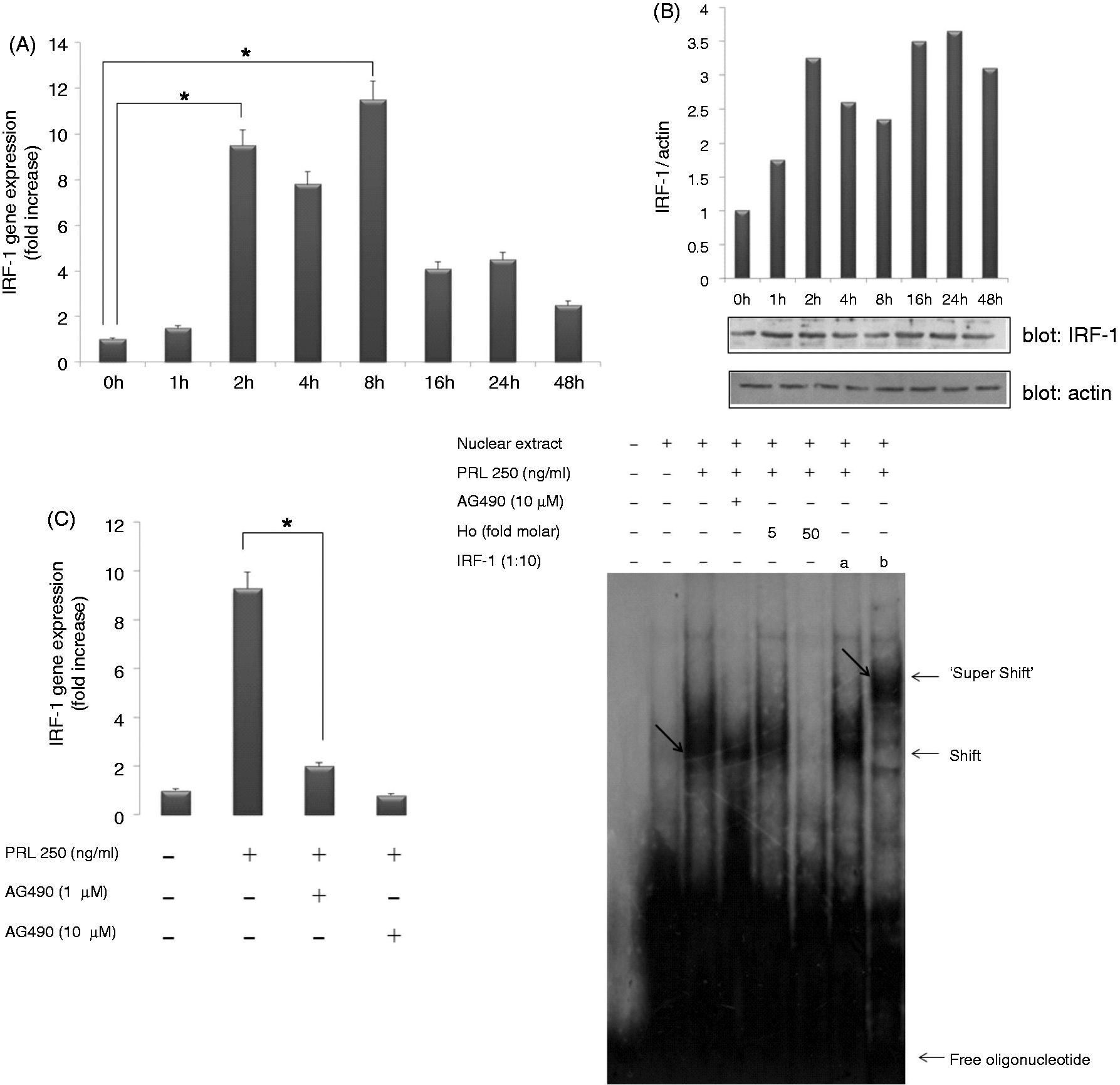
Discussion
PRLR is a single membrane-bound protein that belongs to class 1 of the cytokine receptor superfamily. 42 The intracellular domain is a key player in the initiation of the signal transduction mechanisms associated with PRLR. 43 The intracellular domains differ in length and composition among the various PRLR isoforms and show little sequence similarity to other cytokine receptors. 43 However, there are two relatively conserved regions, termed box 1 and box 2. 44 Box 1 is a membrane-proximal, proline-rich motif necessary for the consensus folding of the molecule recognized by the transducing molecules, whereas Box 2 is less conserved.43,45 Interestingly, both motifs are well conserved in the PRLR of several teleost fish species, despite the low degree of similarity that exists in the intracellular domain of the receptor.46,47 This suggests that PRL may display very specific functions in this group of animals. The membrane proximal region of the intracellular domain is constitutively (i.e. not induced by ligand binding) associated with Jak2, 48 and there are two major prerequisites for it to be phosphorylated: (i) the presence of box 1 motif in the intracellular domain of the PRLR and (ii) homodimeric stoichiometry of the ligand-induced PRLR dimerization. 49 All this is vital, considering that PRL is apparently responsible for the activation of several signal transduction pathways, including Jak/Stat and NF-κB, involved the regulation of innate immunity, particularly ROS production and pro-inflammatory gene induction, in the teleost fish gilthead seabream.9,10 In the present work, we demonstrated, additionally, an increase in the expression of NADPH oxidase components triggered by the PRL in SHK-1 monocytic cells from Atlantic salmon, thereby extending our previous observations to a very distant phylogenetic species. Furthermore, our results showed that PRL is a potent stimulator of the production of ROS through (i) the direct PKC-mediated phosphorylation and activation of p47phox; 10 and (ii) the induction of a new enzymatic complex (this study).
In general, the engagement of PRLR by PRL in target cells results in the activation of Jak2 and different Stat proteins, including Stat1 and Stat5.9,43 Activated Stat proteins subsequently translocate to the nucleus and up-regulate the transcription of target promoters. Thus, Stats, PKC and NF-κB were all involved in the PRL-mediated polarization of gilthead seabream macrophages to an activated cellular phenotype characterized by the production of ROS and pro-inflammatory cytokines.9,10 Additionally, in this study we report that PRL is able to induce the gene expression and activation of IRF-1 in SHK-1 cells. The specific role and target genes of the PRL-induced IRF-1 in fish leukocytes remain to be ascertained, as this transcription factor is pleiotropic in other biological models. Thus, IRF-1 influences the transcription of numerous target genes that facilitate cellular growth, apoptosis, adhesion, neoplastic transformation, viral resistance and differentiation.50,51 Therefore, it is tempting to speculate that PRL in fish leukocytes, probably via IRF-1, might regulate the expression of an array of target genes that would fine-tune a multitude of functions associated with the establishment of immunity and disease resistance. Despite these interesting and promising results, the paucity of Atlantic salmon and gilthead seabream genomic information hampers the confirmation of the role played by IRF-1 in the regulation of the genes encoding the NADPH oxidase components. However, we used a bioinformatics approach with the tilapia (O. niloticus) genome to elucidate a possible molecular connection between PRL, Jak/Stat, IRF-1 and the regulation of NADPH oxidase component genes. Figure 5A shows several putative recognition motifs for STATx (GAS) and IRF-1 (ISRE) transcription factor located upstream of the sequence coding for p47phox. Interestingly, several cis-regulatory elements, putative targets of key transcription factors, cluster in a 300-bp proximal promoter region of the p47phox gene. In particular, we found a binding motif for PU.1, which is a pivotal transcription factor of human p47phox
52
and gp91phox genes.
53
Interestingly, human IFN-γ has been found to induce the transcription of the p47phox, p67phox and gp91phox genes through the cooperative activation of PU.1, STAT-1 and IRF-1 via binding to the PU.1 binding sequence, GAS and ISRE respectively.20,52,54 We envisage a similar model in fish (Figure 5B), where PRL would coordinately induce the activation of NF-κB, STAT and IRF-1 transcription factors in leukocytes that, in turn, would promote the induction of genes encoding pro-inflammatory cytokines (IL-1β and TNF-α) and NADPH oxidase subunits, as well as the direct activation of NADPH oxidase through the PKC-mediated phosphorylation of p47phox9,10 and present study. This model will be further investigated once the genomic sequence of Atlantic salmon and/or gilthead seabream promoters of the genes encoding the NADPH oxidase subunits are available.
Model proposed for the PRL-Statx-IRF-1-p47phox signaling pathway in teleosts. (A) The promoter region of p47phox (ENSONIG00000000357.1). Numbering is relative to putative translation star and the first codon of ORF is indicated (+1, bold/underlined). Potential IFN regulatory motifs (ISRE), IFN-γ-activated site (GAS), PU.1 box and TATA box are in bold and underlined. (B) Model showing the signaling pathways involved in the regulation of NADPH oxidase activation by PRL in bony fish. (i) Binding of PRL to its receptor induces the phosphorylation of Jak2, which activates Stat1/Stat5 and PKC. (ii) PKC stimulates the phosphorylation and activation of p47phox, NADPH assembly and the generation of ROS. (iii) Phosphorylated Stat proteins dimerize, translocate to the nucleus and bind to GAS of the p47phox promoter (and likely to p67phox and gp91phox). (iv) Activated Stats also bind to GAS of the IRF-1 promoter and induce IRF-1. (v) IRF-1 binds to ISRE of the p47phox promoter and activates it in cooperation with Stat.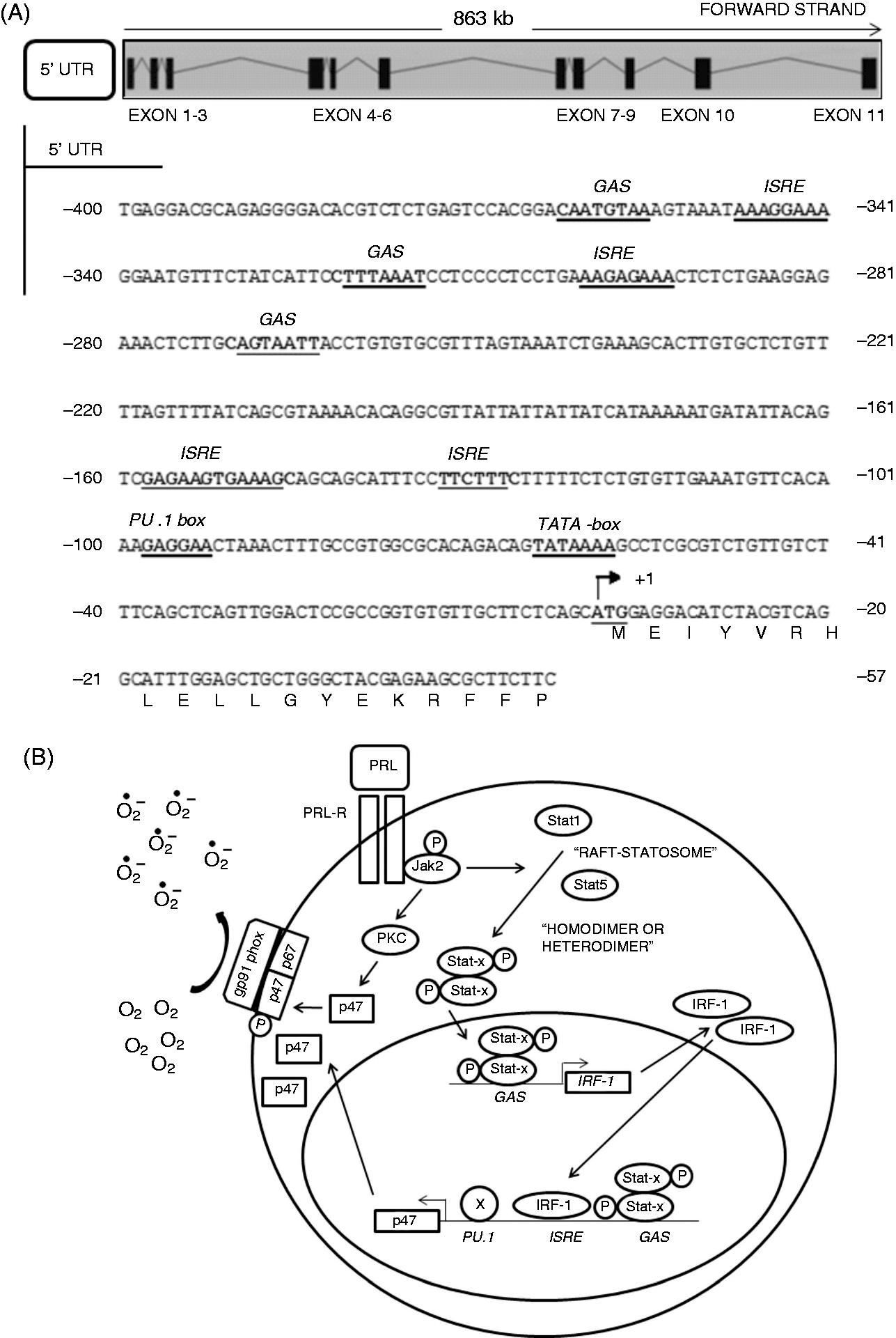
In conclusion, this study demonstrates the PRL-mediated induction of the genes encoding the NADPH oxidase components and the transcription factor IRF-1 via the Jak/Stat signaling transduction pathway in salmon monocytic SHK-1 cells. Moreover, for the first time, IRF-1 is confirmed as a PRL-responsive gene in fish leukocytes. Although the exact role of IRF-1 in fish immunity and its target genes remain to be elucidated, the study paves the way for illumination of the PRLR/Jak/Stat/IRF-1 signaling pathway that regulates NADPH oxidase activity in vertebrates.
Footnotes
Acknowledgments
We thank I. Fuentes for her excellent technical assistance and the ‘Servicio de Apoyo a la Investigación’ (University of Murcia) for assistance with cell culture.
Funding
This work was supported by the Spanish Ministry of Science and Innovation (grants BIO2008-01379 and BIO2011-23400), Dirección de Investigación y Desarrollo de la Universidad Austral de Chile (DID-UACh) and Fundación Marcelino Botín, Spain.
