Abstract
Chemotherapy-induced peripheral neuropathy (CIPN) is the most common side-effect of anti-cancer therapy. To date, there are no clinically effective analgesics that could prevent and treat CIPN. However, the exact pathogenesis of CIPN is still unclear. In the present study, we use the paclitaxel-induced peripheral neuropathy (PIPN) model, aiming to better understand the transcriptomic level of the Dorsal root ganglia (DRG) neurons in rats with PIPN. mRNA from each DRG sample was reverse transcribed to cDNA and sequenced using next-generation high throughput sequencing technology. Quantitative RT-PCR verification was used to confirm the identified Differentially expressed genes (DEGs) in the DRG of PIPN rats. RNAseq results have identified 384 DEGs (adjusted P-value < 0.05; fold change ≥ 2) in the DRG of rats 14 days after paclitaxel injection in total, including 97 up-regulated genes, and 287 down-regulated genes. GO analysis revealed that these DEGs were majorly involved in neuropeptide activity, chemokine receptor activity, defense response, and inflammatory response. Kyoto Encyclopedia of Gene and Genomes analysis showed that neuroactive ligand-receptor interaction and cytokine-cytokine receptor interaction were involved in sensory neurons of rats with PIPN. Besides, comparison analysis identified that 11 DEGs in the PIPN model are shared with either inflammatory pain (Ces1d, Cfd, Retn, and Fam150b) or neuropathic pain (Atf3, Csrp3, Ecel1, Gal, Sprr1a, Tgm1, and Vip). Quantitative RT-PCR results also confirmed the validation of the RNAseq data. These results suggested that neuroactive ligand-receptor interaction and cytokine-cytokine receptor interaction are majorly involved in sensory neurons of rats with PIPN. Immune, inflammatory responses and neuron functional changes are the major pathogenesis of PIPN. Paclitaxel-induced peripheral neuropathy has shared characteristics with both inflammatory pain and neuropathic pain.
Keywords
Introduction
Chemotherapy-induced peripheral neuropathy (CIPN) is one of the major complications of anti-cancer agents. 1 Among anti-cancer agents used clinically, paclitaxel is a first-line used chemotherapy agent that causes classical peripheral neuropathy. 1 Paclitaxel is mainly used for the treatment of breast cancer, ovarian cancer, and non-small cell lung cancer. Its unique and effective curative effect is irreplaceable in the treatment of cancer. To date, paclitaxel ranked first in the sales list of anti-cancer agents. Paclitaxel-induced peripheral neuropathy (PIPN) is representative of CIPN and affects a wide range of people. Clinical studies have shown that up to 81% of patients have numbness and/or pain in their hands and/or feet within 6-24 months after paclitaxel use, of which 25% have severe symptoms. 2 Paclitaxel-caused peripheral neuropathic symptoms in humans include cold-triggered acral paresthesias, and chronic distal sensory neuropathy. 3 Chemotherapy-induced peripheral neuropathy symptoms initially manifest as symmetrical numbness at the end of the limbs, gloves, socks-like deep sensation, shallow sensory disorder, needle-like, and burning-like sensory abnormalities, and then develop into sensory and motor disorders, and eventually develop to the far end of the limb muscle loss, movement difficulties, limb burning sensation, and severe pain. Chemotherapy-induced peripheral neuropathy severely limits the use of chemotherapy agents and is a common cause of chemotherapy termination. 4 Different from other side effects, CIPN in some patients cannot be relieved even after chemotherapy. To date, there are no clinically effective analgesics that could prevent and treat CIPN.
Behavioral changes in CIPN result in a series of changes in neuronal hyperexcitability, perineuronal homeostasis, gene expression, inflammation, and immune response in peripheral sensory neurons.5,6 The pathogenesis of CIPN induced by paclitaxel in peripheral sensory neurons mainly includes 1. Abnormal expression and activation of cell membrane receptors and ion channels; 2. Alterations in intracellular signal transduction; 3. Microtubule damage, oxidative stress, and mitochondrial damage; 4. Glial cell activity and neuroinflammation eventually lead to the sensitivity of sensory neurons and damage to peripheral neurons and nerve fibers. 7 Besides, clinical and animal experimental studies have shown that chemotherapy agent-induced CIPN is associated with peripheral nerve fiber atrophy, demyelination and nerve fiber loss, axon regeneration failure, 8 and reducing the damage of peripheral nerve fiber alleviates the pain behaviors associated with the paclitaxel-induced CIPN.9,10 However, the exact peripheral pathogenesis of paclitaxel-induced CIPN is still unclear. The full transcriptomic profiles in dorsal root ganglia (DRG) that correlated to CIPN, could be important for a better understanding of the peripheral pathogenesis of paclitaxel-induced CIPN.
Therefore, this study aims to better understand the transcriptomic profiles of the DRG neurons in animals with PIPN. The present study examined the transcriptomic changes in DRG neurons of rats with PIPN and summarized the general transcriptomic changes of DRG in PIPN animals by comparing them with either inflammatory pain or neuropathic pain models. The results revealed that 97 genes were up-regulated, and 287 genes were down-regulated in the DRG of rats 14 days after paclitaxel injection. GO and KEGG analysis demonstrated that these altered genes were mainly involved in hormone activity, neuropeptide activity, chemokine receptor activity, defense response, inflammatory response, and neuropeptide signaling transduction. Besides, compare with the data from inflammatory pain and neuropathic pain models, we identified that there are 11 Differentially expressed genes (DEGs) shared in the PIPN model with either inflammatory pain or neuropathic pain. Therefore, we concluded that neuroactive ligand-receptor interaction and cytokine-cytokine receptor interaction are mainly involved in sensory neurons of rats with PIPN. Paclitaxel-induced peripheral neuropathy has shared characteristics with both inflammatory pain and neuropathic pain.
Materials and methods
Animals
Sprague-Dawley rats (200-250 g) used in this study were purchased from Guangdong Laboratory Animal Center, Guangzhou, China. Rats were kept in a temperature-controlled room (22 ± 1°C) under a light/dark (12/12h) cycle, with free access to food and water. All experiment procedures in this study have been performed by the Basel Declaration and approved by the Ethics Committee of Animal Care and Use of Shenzhen University Health Science Center.
Experiment design and drug treatment
Rats were randomly divided into a vehicle group (6 rats) and a paclitaxel treatment group (6 rats). After acclimation to the environment, behavioral tests were conducted before or after being injected with saline or paclitaxel three times (days 0, 7, and 14). A rat model of PIPN was established as we previously described. 11 In brief, paclitaxel (Dalian Meilun, China) dissolved in DMSO (20 mg/ml) and serially diluted in 0.9% saline solution was intraperitoneal (i.p.) injected at a dosage of 2 mg/kg (using an injection volume of 10 ml/kg) on alternative days (day 1, 3, 5, and 7), with a final cumulative dose of 8 mg/kg paclitaxel injection in each rat as shown in Supplementary Figure 1. Each rat was sacrificed on day 15 and the ipsilateral lumbar (L) 4, L5, and L6 DRGs were collected for RNA-sequencing.
Behavioral assays
Mechanical allodynia (von Frey test)
Animal behaviors were tested in the daytime before and after paclitaxel injection. Animals were acclimated in a plastic box on a stainless steel mesh floor for 30 min before testing. An electronic von Frey (IITC Life Sciences Inc., Woodland Hills, USA) is used to assess mechanical withdrawal thresholds by applying the von Frey hairs (0.5-mm-diameter polypropylene rigid tip) to the plantar surface of the hind paw, upon paw withdrawal, the thresholds will be displayed and stored in grams as we previously reported.11,12 Each measurement was repeated 3 times at a 5-min interval.
Thermal and cold hyperalgesia (hot/cold-plate test)
Thermal and cold hyperalgesia was examined using a hot/cold plate analgesia meter (BIO-CHP, Bioseb, France) as previously reported. 13 Rats were kept on a hot (53 ± 1°C) or cold (4 ± 1°C) plate. Response latency was measured 3 times, with a 10 min interval between every measurement. Lifting for normal locomotion was excluded.
Cold hyperalgesia (tail immersion test)
Cold hyperalgesia was evaluated using the cold water tail immersion method according to previous literature. 14 The rat was kept in a rat holder and the tail (2 - 3 cm) was immersed in cold water (4 ± 1°C). The time taken to flick the tail was recorded as the response time.
RNA extraction and sequencing
The L4-6 DRGs were dissected and collected using Trizol reagent (Sigma-Aldrich, USA) for RNA extraction. RNA quality and quantity were examined using a NanoDrop ND-1000 (NanoDrop, USA). The construction of the cDNA library was performed by Beijing Genomics Institute (BGI, China) according to the manufacturer’s instructions. In brief, poly-(A) mRNA was isolated from total RNA with the Oligo d(T) magnetic beads. Then, mRNA was fragmented into small pieces with the action of fragmentation buffer under an elevated temperature. The first single-stranded (ss) cDNA was synthesized using mRNA fragments as templates. The second-strand cDNA was made by use of DNA polymerase I and RNase H. The cDNA ends were then repaired by polymerase and untemplated A residues were added to the 3’ ends of double-stranded cDNA. Illumina special adapters were ligated onto the cDNA pools and suitable fragments were selected for PCR amplification. After clustering of index-coded samples using the TrueSeq PE Cluster Kit v3-cBot-HS (Illumina, USA), pair-end reads were sequenced on an illumine HiSeq 2500 platform (Illumina, USA).
Data processing and bioinformatics analysis
The raw short reads were obtained after image analysis, base calling, and error estimation using Illumina/Solexa Pipeline version 1.8 (Off-Line Base caller software, version 1.8). FastQC was used for the raw data quality control. Trim Galore was adapted to remove poor-quality reads. The trimmed reads were mapped to the reference genome with Tophat, and cufflinks were applied to calculate the fragments per kilobase of exon per million fragments mapped. FeatureCount was utilized to count read numbers for each gene, and then the count matrix was submitted to the DEseq2 R package for differential expression analysis, in which Benjamini and Hochberg’s approach was applied to control the false discovery rate. 15 Genes with adjusted p-values < 0.05, and fold change ≥ 2 were considered as DEGs. Then gene Ontology (GO) enrichment analysis was performed for DEGs using BiNGO, 16 and Kyoto Encyclopedia of Gene and Genomes (KEGG) pathway enrichment was determined with the new enrichment detection mode implemented in the online STRING database. 17 Using the whole genome as background, the functional enrichment is based on the straightforward over-representation analysis using hypergeometric tests. The protein-protein interaction (PPI) network encoded by the DEGs was also searched in STRING and then visualized in a Cytoscape plugin Network Analyzer. Hierarchical cluster analysis of the top 50 strongest interactive genes was performed by TBtools software. 18 The sequencing data from PIPN rats were submitted to the Gene Expression Omnibus (GEO) database (GSE160543).
Quantitative RT-PCR for mRNA validation
Sequences of the primer for quantitative real-time RT-PCR.
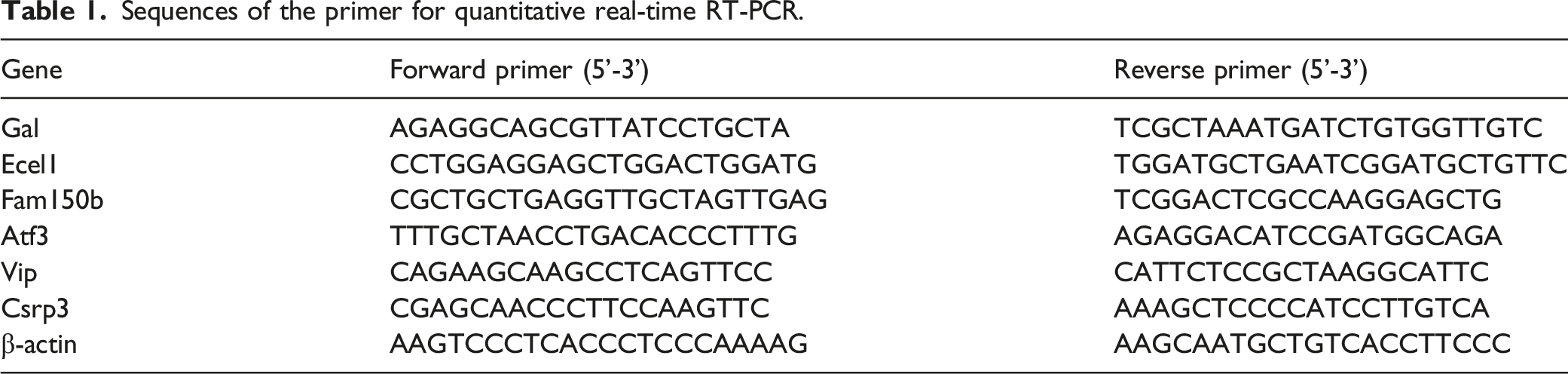

Statistical analysis
Data are represented as means ± SEM and analyzed with R studio. Statistical differences between the two groups were analyzed using two-way repeated-measures ANOVA with Bonferroni correction or unpaired student's t-test. p < 0.05 was considered a significant difference.
Results
Mechanical allodynia, cold and thermal sensitivity in PIPN rats
As shown in Figure 1, the neuropathic pain rat model was established by paclitaxel administration (i.p.). Mechanical allodynia, thermal and cold sensitivity were examined in all PIPN model rats at 0, 7, and 14 days after paclitaxel injection, respectively. Our results demonstrated that PIPN rats exhibited higher sensitivity to mechanical, thermal, and cold stimuli (either cold-plate test or tail immersion) from days 7 to 14. Paw withdrawal thresholds in response to mechanical and thermal stimuli were significantly decreased on the 7th and 14th day in paclitaxel-induced PIPN rats (p < 0.05 or p < 0.01, n = 6, Figure 1(a) and (b)). Besides, the paw withdrawal threshold in response to cold stimuli and latency of tail immersion was dramatically reduced on day 7 and day 14 as well (p < 0.05 or p < 0.01, n = 6, Figure 1(c) and (d)). These results indicated that the paclitaxel-induced CIPN model was established in rats successfully. Administration of paclitaxel induces mechanical allodynia, hot hypersensitivity, and cold hypersensitivity in rats. 
Differential gene expression and GO enrichment analysis
The summary of raw RNA sequencing data set.
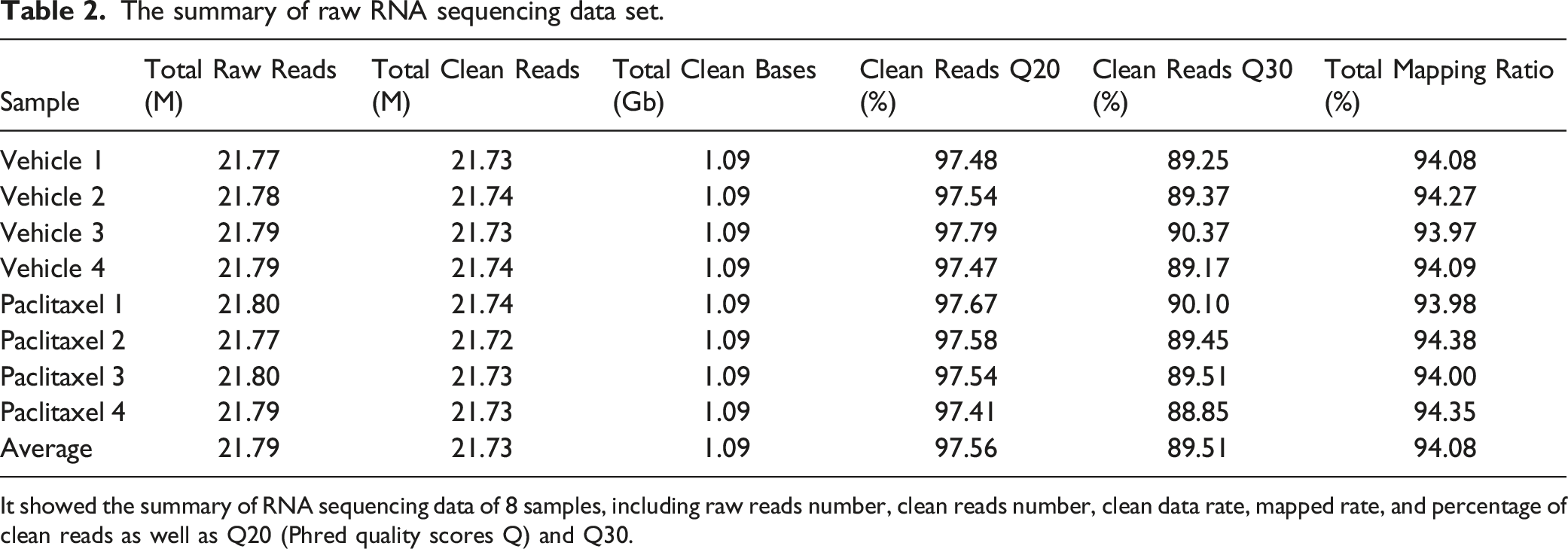
It showed the summary of RNA sequencing data of 8 samples, including raw reads number, clean reads number, clean data rate, mapped rate, and percentage of clean reads as well as Q20 (Phred quality scores Q) and Q30.
Differentially expressed genes screening and further functional analysis were then carried out. Normalized expression levels were compared to PIPN rats with control. Using the adjusted p-value of less than 0.05, we identified a total of 384 DEGs between the PIPN and the control group after 14 days of treatment with paclitaxel. These 384 genes included 97 up-regulated genes (e.g., Atf3, Npy, Sp) and 287 down-regulated genes (e.g., Camp, Ccl8) as shown in Figure 2(a). Besides, Table 3 listed the top 20 up- and down-regulated genes in the DRG of PIPN rats. Functional analysis showed that the up-regulated DEGs were mostly enriched in cellular components (CC) terms such as extracellular region (Figure 2(b)), molecular function (MF) terms related to hormone activity, receptor binding, and neuropeptide receptor activity (Figure 2(c)), and biological processes (BP) related to behavior, neuropeptide signaling pathway, and regulation of response to extracellular stimulus (Figure 2(d)). Meanwhile, the down-regulated DEGs were enriched in CC terms such as extracellular region, plasma region, and troponin complex (Figure 2(b)), MF terms such as carbohydrate-binding, serine-type endopeptidase activity, and glycosaminoglycan binding (Figure 2(c)) and BP terms such as defense response, immune system process and response to stress (Figure 2(d)). Functional analyses of DEGs by Gene Ontology and KEGG classifications in the DRG of PIPN rats. (a) Volcano plot showing the log2 fold change (x-axis) against adjusted P-value (y-axis) for differentially expressed genes between vehicle and paclitaxel treatment. Bar plot showing the number of DEGs. Genes with significantly downregulated are shown in green, while the genes with significantly upregulated are marked in red. (b) GO terms significant enrichment (p < 0.05) in cellular components with genes significantly decreased (green) and increased (red). The X-axis represents the enrichment score (-log10 P-value) for each GO term. (c) GO terms significant enrichment (p < 0.05) in molecular function with genes significantly decreased (green) and increased (red). The X-axis represents the enrichment score (-log10 P-value) for each GO term. (d) GO terms significant enrichment (p < 0.05) in biological process with genes significantly decreased (green) and increased (red). The X-axis represents the enrichment score (-log10 p-value) for each GO term. (e) Histogram shows the distribution of enriched KEGG pathways. The X-axis represents the number of genes, and the y axis represents the name of the enriched pathway. The color indicates the enrichment score (adjust p-value). DEGs: Differentially expressed genes; DRG: Dorsal root ganglia; KEGG: Kyoto Encyclopedia of Gene and Genomes; PIPN: paclitaxel-induced peripheral neuropathy. The top 20 up- and down-regulated genes in the DRG of paclitaxel-induced peripheral neuropathy rats Note: DRG: Dorsal root ganglia.

KEGG pathway enrichment analysis of DEGs
The DEGs between PIPN and control rats were subjected to KEGG pathway enrichment analysis using the database STRING. 17 According to the results of the KEGG pathway analysis, the DEGs were enriched in 7 signaling pathways significantly, including neuroactive ligand-receptor interaction, cytokine-cytokine receptor interaction, and chemokine signaling pathway, which were related to immune and inflammatory responses (Figure 2(e)). We further performed functional analysis of the neuroactive ligand-receptor interaction pathway and cytokine-cytokine receptor interaction pathway as shown in Supplementary Figure 3(a) and (b). The results demonstrated that the DEGs involved in the neuroactive ligand-receptor interaction pathway were mainly GPCRs, peptides, hormone protein, and ion channels (Supplementary Figure 3(a)). Besides, the DEGs enriched in the cytokine-cytokine receptor interaction pathway as shown in Supplementary Figure 3(b) were majorly enriched in chemokines (CC and CXC subfamily), and cytokines (IL-4, IL-1, IL-17, TNF family, and TGF-β family). These results further demonstrated that neuroactive ligand-receptor interaction and cytokine-cytokine receptor interaction are involved in sensory neurons of rats with PIPN.
Protein-protein interaction network analysis
The PPI network of DEGs in PIPN rats was constructed using a Cytoscape plugin Network Analyzer to investigate the interaction and hub genes of DEGs involved in the pathogenesis of PIPN. The predicted PPI in PIPN rats was driving the complex interaction network 14 days after paclitaxel treatment. The results demonstrated that the established PPI network consisted of 62 nodes (hub genes) and 422 edges (interactions). Fifty of these top genes with high connectivity were highlighted (Figure 3(a)). To elucidate the role of DEGs in the PIPN model, these 50 highly connectivity genes were hierarchically clustered (Figure 3(b)). The most prominently up-regulated genes consisted of families of chemokines (Cxcl10 and Cxcl11), neuropeptide (Npy, Sp, and Vip), and G protein-coupled receptors (Ghsr). This clustering analysis of RNAseq data revealed that the high connectivity genes in the PIPN model are closely associated with the development of neuropathic pain. Many novel DEGs that we screened may play a critical role in PIPN, but future in-depth studies are required. Protein-protein interaction (PPI) network and hierarchical clustering analysis of DEGs in the DRG of PIPN rats. (a) The PPI network of the DEGs. Circles represent genes, and lines represent the interactions between genes. The size of the circle represents the strength of interaction pairs. The bigger circle size indicates the larger interaction degree. (b) Heatmap showing hierarchical clustering of top 50 strongest interactive genes. Columns represent individual arrays and rows represent specific genes. The Z-score is used to normalize the expression value. The relative expression value is marked as color intensity, with green indicating down-regulated and red indicating up-regulated genes. DEGs: Differentially expressed genes; DRG: Dorsal root ganglia; PIPN: Paclitaxel-induced peripheral neuropathy.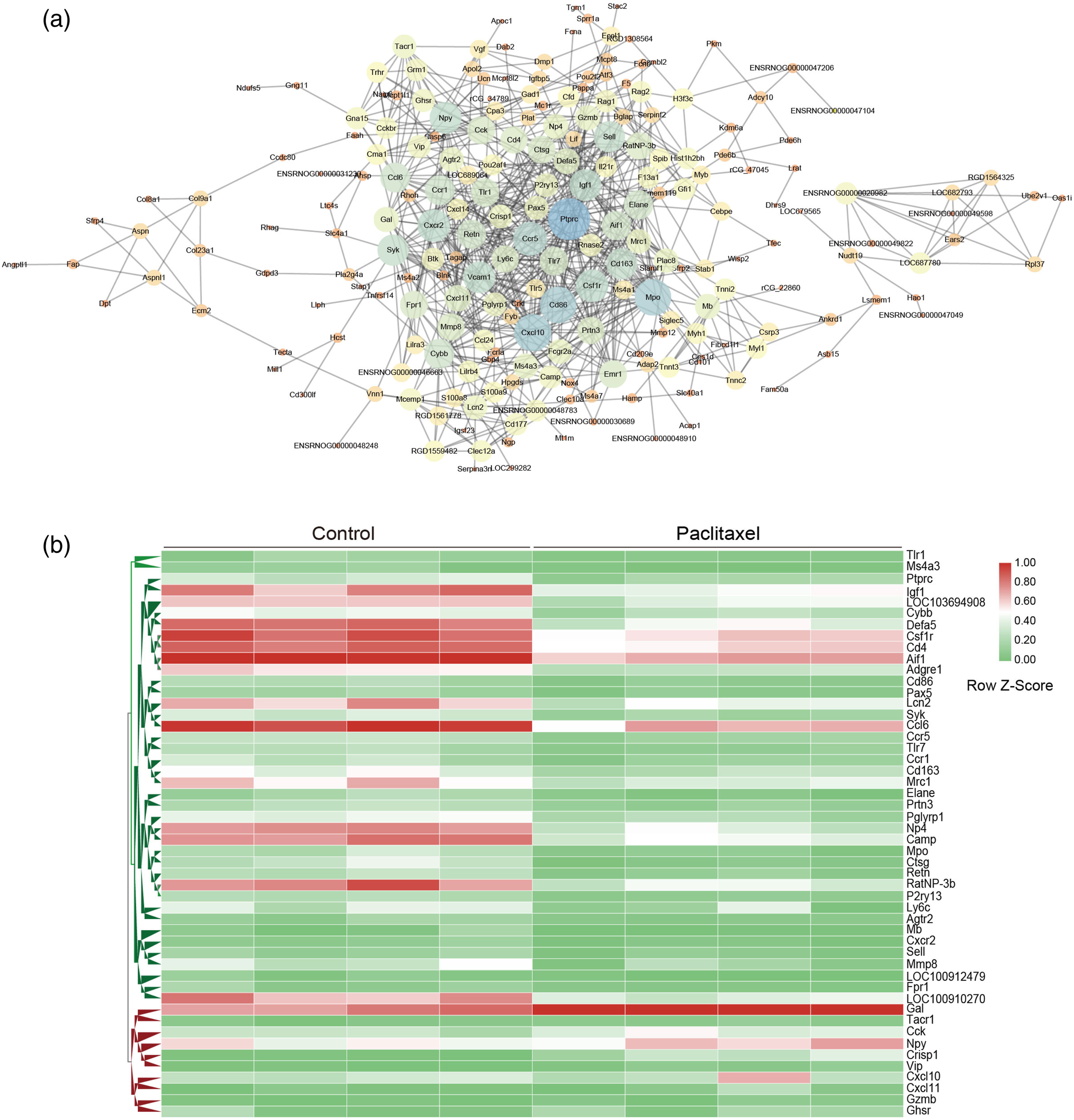
Transcriptomic comparison with inflammation pain and peripheral nerve injury models
To further elucidate the characteristics of PIPN, we compared the DRG transcriptomic data of PIPN rats with CFA and neuropathic pain models (spared nerve injury, chronic constriction injury).19,20 Transcriptomic data (GSE111216,
19
and GSE30691
20
) were downloaded from the GEO database. GEO2R, an interactive web tool, was used to compare the transcriptomic data from different pain models in a GEO series to identify DEGs across experimental groups. As shown in Figure 4(a), there were shared 4 DEGs in the PIPN model and CFA-induced inflammatory pain animals. Besides, compared with the SNI and CCI model, 7 of the shared DEGs among PIPN, SNI, and CCI models were identified (Figure 4(b)). We, therefore, listed the value of the RPKM of these 11 DEGs in PIPN rats, as shown in Figure 4(c), they were Ces1d, Cfd, Retn, and Fam150b altered in both PIPN and inflammatory pain models with the same direction. Besides, Atf3, Ecel1, Gal, Sprr1a, and Tgm1 in either PIPN, SNI, or CCI models were altered in the same direction but not Csrp3, and Vip (Figure 4(c)). Interestingly, DEGs shared in PIPN and CFA animals were down-regulated except Fam150b. Whereas DEGs altered in either PIPN, SNI, or CCI models were all up-regulated. These results suggested that PIPN has shared characteristics with both inflammatory pain and neuropathic pain. Transcriptomic comparison of the RNAseq data from PIPN with other pain models. (a) Venn diagram exhibits the number of unique or shared DEGs in PIPN and CFA-induced inflammatory animals. (b) Venn diagram presents the number of DEGs that are unique or shared among PIPN, SNI, and CCI models. (c) Value of the RPKM of each DEG selected from a and b. DEGs: Differentially expressed genes; PIPN: Paclitaxel-induced peripheral neuropathy.
Quantitative RT-PCR verification of the mRNA expression of DEGs shared in different pain models
We further validated a few of the key RNAseq findings using quantitative RT-PCR. Therefore, the mRNA expression of Atf3, Gal, Ecel1, Vip, Csrp3, and Fam150b in DRG of PIPN rats was examined. Our results demonstrated that the mRNA expression of Atf3, Gal, Ecel1, Vip, and Fam150b were up-regulated significantly except for Csrp3 (p < 0.05 or p < 0.01, n = 6, Figure 5). These results confirmed the validation of RNAseq data and suggested the consistency between RNAseq and quantitative RT-PCR results. Validation of DEGs by quantitative RT-PCR. 
Discussion
Paclitaxel is a first-line agent and is widely used for the treatment of various kinds of cancer clinically. However, the underlying mechanism involved in PIPN is still unclear so far. Therefore, in this study, the paclitaxel-induced CIPN model was used to analyze the transcriptomic profile changes in peripheral ganglia in response to chemotherapeutic treatment, which could help to better understand the molecular mechanisms of PIPN. Our data revealed that there were identified 384 DEGs in DRG, including 97 up-regulated genes and 287 down-regulated genes, from PIPN rats.
GO analysis revealed that the up-regulated DEGs in DRG from PIPN rats were mainly enriched in receptor binding and neuropeptide receptor activity, and neuropeptide signaling pathway. Meanwhile, the down-regulated DEGs were enriched in defense response, immune system process, and stress response. Besides, KEGG pathway enrichment analysis uncovered that the DEGs were significantly enriched in neuroactive ligand-receptor interaction, cytokine-cytokine receptor interaction, and chemokine signaling pathway, which were mostly corresponding to immune, inflammatory responses, and neuron functional changes. These results suggested that PIPN is mainly involved in immune and inflammatory responses. These results consisted of a previous microarray assay demonstrating inflammatory and/or immune responses were mainly involved in the DRG transcriptomic changes after paclitaxel treatment in rats. 21 Moreover, previous transcriptomic analysis results demonstrated that altered neuronal responses and immune and inflammatory responses were predominant biological events in the spinal cord of PIPN rats as well.22,23 Further pathway analysis demonstrated that DEGs involved in the neuroactive ligand-receptor interaction pathway were mainly GPCRs, ion channels, chemokines, and cytokines. These data demonstrated that neuroactive ligand-receptor interaction and cytokine-cytokine receptor interaction are majorly involved in sensory neurons of rats with PIPN. This is quite similar to the transcriptomic profile changes in DRG of the neuropathic pain model rats with sciatic nerve injury. 24
Besides, network analysis established PPI network consisted of 62 nodes (hub genes) and 422 edges (interactions). The most prominently high connectivity degrees genes consisted of families of chemokines, neuropeptides, and G protein-coupled receptors. This clustering analysis results revealed that the high connectivity genes in the PIPN model are closely associated with the pathogenesis of neuropathic pain. Many novel DEGs that we screened may play a critical role in the regulation of protein expression in PIPN, but future in-depth studies would be required.
A previous report has shown the signatures of CIPN induced by various chemotherapeutic agents in mice. 25 Vincristine predominantly caused dysregulation of genes associated with immune processes, whereas oxaliplatin caused dysregulation of neuronal function, and cisplatin resulted in a mixed inflammatory and neuropathic pathology. 25 Compare with the data from CIPN induced by vincristine, cisplatin, and oxaliplatin, our results indicated that paclitaxel causes dysregulation immune, inflammatory responses, and neuron functional changes. Therefore, paclitaxel-induced CIPN has more similar characteristics to the data from cisplatin-induced CIPN, but not vincristine and oxaliplatin. A further detailed comparison among paclitaxel, cisplatin, vincristine, and oxaliplatin is warranted.
In addition, to further elucidate the characteristics of PIPN, we compared the DRG transcriptomic data of PIPN rats with CFA and neuropathic pain animals.19,20 Four DEGs (Ces1d, Cfd, Retn, and Fam150b) were found in both PIPN and CFA-induced inflammatory animals. Meanwhile, compared with SNI and CCI models, 7 DEGs among PIPN, SNI, and CCI models were found, including Atf3, Csrp3, Ecel1, Gal, Sprr1a, Tgm1, and Vip. Interestingly, DEGs shared in PIPN and CFA animals were down-regulated except Fam150b. Whereas DEGs altered in either PIPN, SNI, or CCI models were all up-regulated. Atf3 is known to be induced by paclitaxel and involved in neuronal stress/injury.26,27 Gal 28 and Vip 29 have been reported to be involved in neuropathic pain. Vip is a neuropeptide exhibiting up-regulation in the nociceptive neurons after peripheral nerve injury. 29 Fam150b, also known as Alkal2, is a kind of receptor tyrosine kinase. Fam150b converts the extracellular signal into an intracellular response in the form of receiving specific ligands and induces the activation of tyrosine kinase. 30 Ecel1 is an endopeptidase with high neuron-specific expression in the nervous system. The expression of Ecel1 is increased in the damaged neurons and plays a protective role in injured neurons. 31 Importantly, we also examined the mRNA up-regulation of Atf3, Gal, Vip, Fam150b, and Ecel1, suggesting the involvement of these genes in PIPN. Csrp3 is a muscle-specific cysteine-rich protein, which is critical in maintaining the structure and function of muscle. 32 However, we did not find the upregulation of Csrp3 in PIPN. Quantitative RT-PCR results further confirmed the validation of RNAseq data, suggesting that PIPN has shared characters with either inflammatory pain or neuropathic pain.
Conclusions
The present study revealed that neuroactive ligand-receptor interaction and cytokine-cytokine receptor interaction are involved in sensory neurons of rats with PIPN. Immune, inflammatory responses, and neuron functional changes are the major pathogenesis of PIPN. Paclitaxel-induced peripheral neuropathy has shared characteristics with both inflammatory pain and neuropathic pain.
Supplemental Material
Supplemental Material - Transcriptome analysis reveals dysregulation of inflammatory and neuronal function in dorsal root ganglion of paclitaxel-induced peripheral neuropathy rats
Supplemental Material for Transcriptome analysis reveals dysregulation of inflammatory and neuronal function in dorsal root ganglion of paclitaxel-induced peripheral neuropathy rats by Wuping Sun, Shaomin Yang, Songbin Wu, Xiyuan Ba, Donglin Xiong, Lizu Xiao and Yue Hao in Molecular Pain
Footnotes
Authors’ contributions
The authors’ contributions were as follows: Y Hao and W Sun were responsible for the concept and design of the study; W Sun, S Yang, S Wu, D Xiong, and L Xiao were involved with experimental and analytical aspects of the manuscript; Y Hao and W Sun performed data interpretation, presentation and writing of the manuscript.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
This work was supported by grants from the National Natural Science Foundation of China (No. 82171378), Shenzhen Municipal Science, Technology and Innovation Commission (No. JCYJ20210324112202006), and Shenzhen Nanshan District Scientific Research Program of China [No. NS007].
Ethics approval
All experiment procedures in this study were approved by the Ethics Committee of Animal Care and Use of Shenzhen University Health Science Center.
Data availability
The sequencing data from PIPN rats were available in the GEO database (GSE160543).
Informed consent
The paper has been read and approved by all authors. All authors approved the submission of this paper to “Molecular Pain” for publication. All authors confirmed that neither the manuscript submitted nor any part of it has been published or is being considered for publication elsewhere.
Supplemental Material
Supplemental material for this article is available online.
Abbreviations
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
