Abstract
Background
Extrahepatic cholangiocarcinoma is a malignancy that originates from bile duct epithelium with an unfavorable prognosis. Epithelial membrane protein 1 was first discovered in 1995, functioning as an oncogene or anti-tumor gene in various cancers. However, the clinical role of epithelial membrane protein 1 extrahepatic cholangiocarcinoma remained unclear.
Methods
Differentially expressed genes were identified using Gene Ontology and the Kyoto Encyclopedia and Genomes pathway analysis. Out of 183 extrahepatic cholangiocarcinoma patients and 61 healthy controls, the expression level of epithelial membrane protein 1 was detected and compared using reverse transcription-quantitative polymerase chain reaction analysis and western blot assay. Meanwhile, the diagnosis and prognosis of EMP1 in ECCA were measured by receiver operating characteristic and Kaplan–Meier analysis. Finally, the relationship between epithelial membrane protein 1 expression and clinicopathological indexes were compared to further verify the clinical role of epithelial membrane protein 1 in extrahepatic cholangiocarcinoma.
Results
After analyzing data from GSE76297, GSE89749, and GSE26566GO, we found 1554 down-regulated and 1065 up-regulated genes. Through Gene Ontology and Kyoto Encyclopedia and Genomes analysis, extracellular matrix organization, extracellular structure organization, cholesterol metabolism, interleukin-17 signaling pathway, and vitamin digestion and absorption were significantly enriched and involved in targeted differentially expresses genes. Epithelial membrane protein 1 messenger ribonucleic acid was notably decreased in serum samples from extrahepatic cholangiocarcinoma patients, compared with that in healthy controls. Receiver operating characteristic analysis revealed that the area under the curve of epithelial membrane protein 1 messenger ribonucleic acid for the diagnosis of extrahepatic cholangiocarcinoma was 0.9281 (95% CI = 0.8967–0.9595). Moreover, the correlation analysis presented that epithelial membrane protein 1 expression was negatively correlated with lymph node metastasis, tumour node metastasis stage, cancer antigen 19-9 level, and carcinoembryonic antigen level.
Conclusion
Aberrant expression of epithelial membrane protein 1 contributed to distinguishing extrahepatic cholangiocarcinoma patients and healthy controls, and a low expression level of epithelial membrane protein 1 indicated an unfavorable prognosis. Hence, epithelial membrane protein 1 was a feasible and credible biomarker for extrahepatic cholangiocarcinoma diagnosis and prognosis, with high accuracy, sensitivity, and specificity.
Keywords
Introduction
Extrahepatic cholangiocarcinoma (ECCA) is a malignant tumor of bile duct epithelial cells derived from the biliary tract, which is highly prevalent in developing countries such as Southeast Asia.1,2 Worldwide, the incidence and mortality of ECCA are increasing, accounting for about 15% of primary liver cancer, 3 with poor prognosis and difficult diagnosis. 4 Clinical methods for early detection of ECCA included nuclear magnetic resonance imaging (MRI), magnetic resonance cholangiopancreatography, and ultrasound. 5 However, these diagnostic methods are expensive and invasive. The levels of tumor markers, such as cancer antigen 19-9 (CA19-9) and carcinoembryonic antigen (CEA), were recommended for clinical diagnosis. 6 However, these biomarkers were distinctively expressed in gastrointestinal cancer tumors as well. 7 Hence, how to improve accuracy, sensitivity and specificity remained challenging to clinical diagnosis. Furthermore, due to the difficulty of accurate early diagnosis of ECCA, many patients are not diagnosed until the disease's late stage. 8 Because of the limited choice of treatment, the overall survival rate of patients is low as well. Hence, it was crucial to identify and establish a reliable biomarker for the diagnosis of ECCA.
The differential expression of genes may affect several cellular processes, including cell proliferation, cell differentiation, and signal transduction.9,10 Recent studies showed that many genes are related to human cancers’ occurrence and development, including ECCA. For instance, Long et al. 11 discovered 1970 differentially expressed genes (DEGs) in lung cancer using bioinformatics analysis, suggesting their potential as candidates for lung diagnosis, monitoring, and treatment. Yin et al. 12 reported that Tigger transposable element-derived 1 was overexpressed in pancreatic, lung, liver, gastric, and colorectal cancers, serving as a potential biomarker in cancer progression and prognosis. As for ECCA, Zhou et al. 13 elucidated 159 DEGs, which may function as potential diagnostic and therapeutic parameters for ECCA. Epithelial membrane protein 1 (EMP1) was first discovered and named by Taylor et al. 14 in 1995, consisted of 157 amino acid residues. A growing number of reports claimed that EMP1 may be associated with tumor cell progression and antitumor drug chemoresistance.15,16 However, the clinical role of EMP1 in ECCA remained unknown.
In the present study, we addressed the impact of EMP1 in ECCA, discovering that ECCA was prominently decreased in serum samples of ECCA patients with high accuracy concerning discriminating ECCA from healthy controls. Meanwhile, low EMP1 level was negatively correlated with CA19-9, CEA and poor prognosis, suggesting the potential of EMP1 in ECCA detection and treatment.
Materials and methods
Subjects
A total of 183 patients with ECCA first admitted and 61 healthy participants who underwent routine medical examinations at Chongqing University Three Gorges Hospital from April 2013 to December 2014 were recruited in the present study. The exclusion criteria included: immune diseases, systematic inflammatory diseases, metabolic diseases, patients with gallbladder or intrahepatic CCA, recurrent or metastatic tumors. Written informed consent was obtained from each participant. The present study was approved by the Ethics Committee of Chongqing University Three Gorges Hospital in accordance with the Declaration of Helsinki.
Measurement of CA19-9 and CEA
The expressions of serum CA19-9 and CEA were determined by chemiluminescence immunoassays with corresponding reagents (JK-(a)-E00672, JK-(a)-3218; Jingkang, Shanghai, China).
Reverse transcription-quantitative polymerase chain reaction
According to the manufacturer's recommendations, total RNA was extracted from serum samples or exosomes using the Total RNA Extraction Kit (R1200-50T, Solarbio, Beijing, China). Then, complementary deoxyribonucleic acid (cDNA) was reverse transcribed into cDNA using a PrimeScript cDNA synthesis kit (Takara, Japan) following the instructions. Finally, reverse transcription-quantitative polymerase chain reaction (RT-qPCR) was performed using SYBR Green Master Mix (Roche, USA) on an ABI Prism 7500 PCR machine (Applied Biosystems; Thermo Fisher Scientific, Inc., Waltham, USA) following the protocol. A total of 40 ng cDNA was used for the PCR reaction. The expression level of EMP1 was normalized to the internal control, glyceraldehyde 3-phosphate dehydrogenase (GAPDH), calculated with the 2−△△Cq method. The average △△Cq of the healthy control group was set as 1, and the relative expression of EMP1 in each sample was then calculated.
All the primers used in the present study were detailed as follows: EMP1, forward 5'-CCCTCATTAAGCTGCCTATCAGT-3' and reverse 5'-CACACAGGTATGCCCTGGTT-3'; GAPDH, forward 5'-AGAAGGCTGGGGCTCATTTG-3' and reverse 5'-AGGGGCCATCCCACAGTCTTC-3'.
Western blotting analysis
Total protein was extracted from serum samples by the radio-immunoprecipitation assay lysis buffer (Beyotime, Shanghai, China) and quantified by the bicinchoninic acid (BCA) Protein Assay Kit (A53225; Pierce; Thermo Fisher Scientific, Inc., Waltham, USA) following the manufacturer's protocol. Then, the extracted proteins (50 μg) were separated by 10% sodium dodecyl sulfate-polyacrylamide gel electrophoresis (P1200-50T; Solarbio, Beijing, China) and transferred to polyvinylidene fluoride membranes (IPVH00010; Millipore, Solarbio, Beijing, China). After blocking with skimmed 5% non-fat milk for 50 min, the membranes were incubated with primary antibodies at 4 °C overnight. The primary antibodies were detailed: rabbit anti-EMP1 antibody (ab202975; 1:300; Abcam, Shanghai, China) and rabbit anti-GAPDH antibody (AB9485; 1:1000; Abcam, Shanghai, China). Afterward, secondary antibodies goat anti-rabbit immunoglobulin G H&L (horseradish peroxidase) preadsorbed (ab7090; 1:1000; Abcam, Shanghai, China) were added into membranes and incubated for another 1 h at room temperature. Finally, an enhanced electro chemi luminescence (ECL) Chemiluminescent Substrate kit (YEASEN, Shanghai, China) was applied to visualize the protein bands while the band was quantified via the National Institutes of Health ImageJ software. The average gray value of EMP1/GAPDH in the healthy control group was set as 1, and the gray value of EMP1/GAPDH in each sample was determined.
Follow-up analysis
Overall survival (OS) and disease-free survival (DFS) were evaluated in the present study. DFS was defined as the time from randomization to disease recurrence or death due to disease progression, while OS referred to the time of death from any cause. For subjects who had been lost to follow-up before death, the last follow-up time was deemed the time of death. The patients’ follow-up investigation was conducted every 3 months after the first admission for surgery in 5 years.
Isolation of exosomes
The exosomes were extracted from serum samples of 10 patients with ECCA. Briefly, 5 ml blood samples were centrifuged for 60 min at 2400 g, and then supernatants were collected and centrifuged for 10 min at 800 g. Next, the supernatants were collected, and the samples containing the exosomes were transferred to Ultra-Clear tubes and centrifuged at 70,000 g for 60 min. After centrifugation, the supernatants (exosome-free serum) were collected and transferred into a new tube; meanwhile, the precipitates (containing the exosomes) were resuspended in phosphate buffered saline (PBS). The exosome-free serum samples as well as the PBS containing the exosomes were stored at −80 °C until further analysis.
Statistical analysis
All the data were displayed as the mean ± standard deviation (SD). SPSS 17.0 software (IMB Corp., Armonk, NY, USA) and GraphPad Prism 6.0 (GraphPad Software, Inc., La Jolla, CA, USA) were used to analyze the data. Student's t-test followed by Tukey's post-hoc test were applied to distinguish between two groups. Pearson's χ2 test was employed to determine the differences between EMP1 expression and clinicopathological features among ECCA patients and healthy participants. OS rate and DFS rate were evaluated using the Kaplan–Meier analysis along with log-rank test analysis. A P value of <0.05 was deemed statistically significant.
Results
24DEGs in ECCA
After processing the raw data in GSE76297, GSE89749, and GSE26566GO, 2609 genes were verified. Among them, 1544 were down-regulated, while 1065 were up-regulated in ECCA samples compared with healthy controls. We then chose 24 DEGs, comprising 14 down-regulated genes and 10 up-regulated genes. The heatmap and volcano plot of DEGs associated with ECCA are shown (Supplementary 1). According to Supplementary 1, we listed 24 DEGs in Supplementary 2 as well.
Gene Ontology (GO) and the Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analysis of EMP1 in ECCA
GO results in Supplementary 1 displayed that targeted 24 DEGs’ biological processes were notably implicated in the extracellular matrix organization and extracellular structure organization. Moreover, KEGG analysis showed that targeted genes were mainly enriched for cholesterol metabolism, interleukin-17 signaling pathway and vitamin digestion and absorption (Supplementary 1).
The eccentric expression of EMP1 in ECCA serum samples
The serum levels of EMP1 in patients with ECCA and healthy participants were determined using RT-qPCR and western blot assays. According to the results in Figure 1(A) and (B), the mRNA and protein expression levels of EMP1 were significantly decreased in ECCA patients compared with healthy controls.
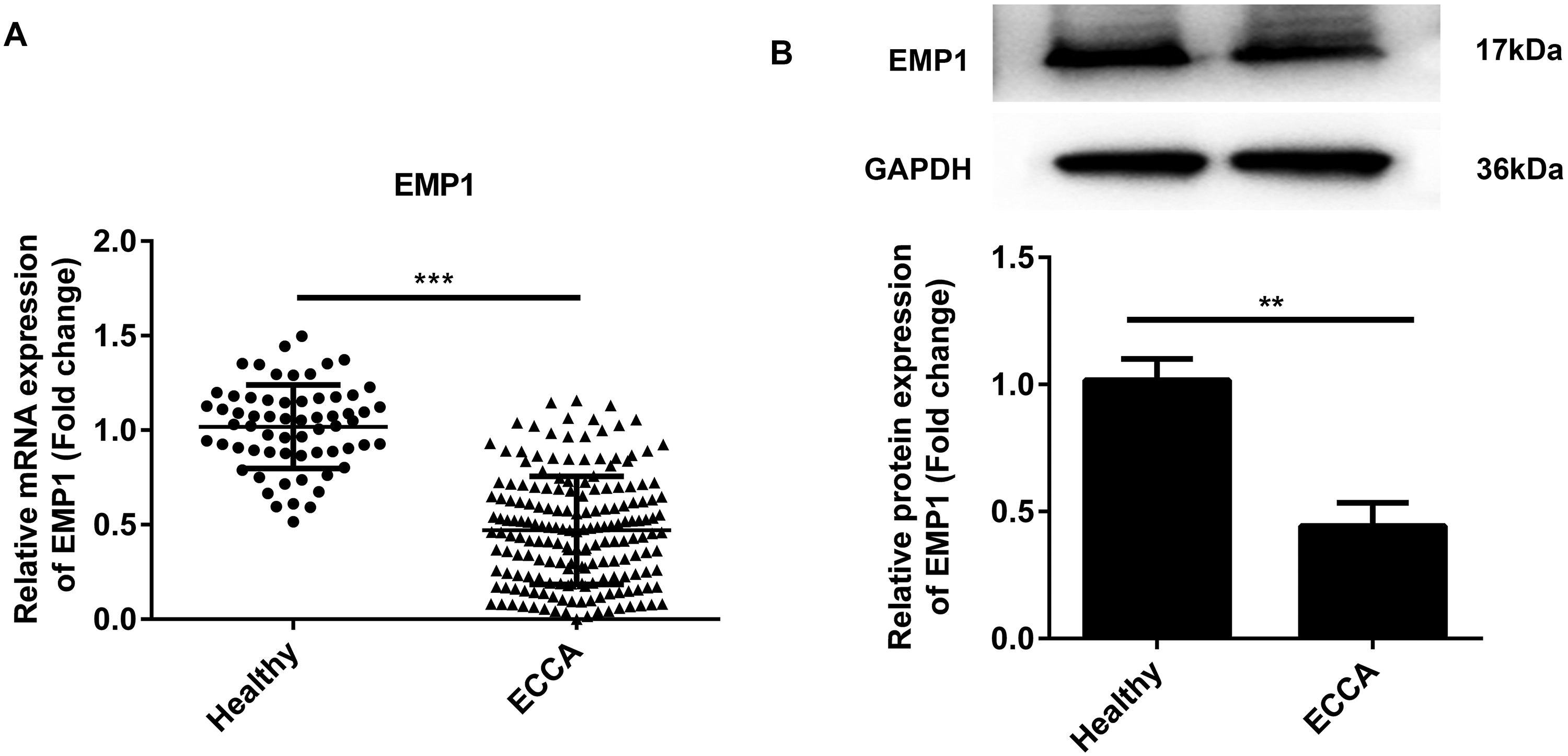
RT-qPCR analysis. (a) The mRNA expression level of EMP1 in serum samples from ECCA patients and healthy controls at admission, and (b) the protein expression level of EMP1 in serum samples from ECCA patients and healthy controls at admission.
The discriminatory power of EMP1 in ECCA diagnosis
The diagnostic value of EMP1 in ECCA was evaluated by generating a receiver operating characteristic (ROC) analysis (Figure 2). As shown, the area under the curve (AUC) of EMP1 mRNA was 0.9281, with 95% CI = 0.8967–0.9595. Moreover, the correlation between EMP1 mRNA mRNA expression and gastrointestinal cancer-related indicators, CA19-9 and CEA, was measured subsequently. As shown in Figure 3, EMP1 mRNA expression was negatively correlated with CA19-9 (r = −0.1990, P = 0.0069) and CEA (r = −0.1892, P = 0.0103), further suggesting the diagnostic significance of EMP1.
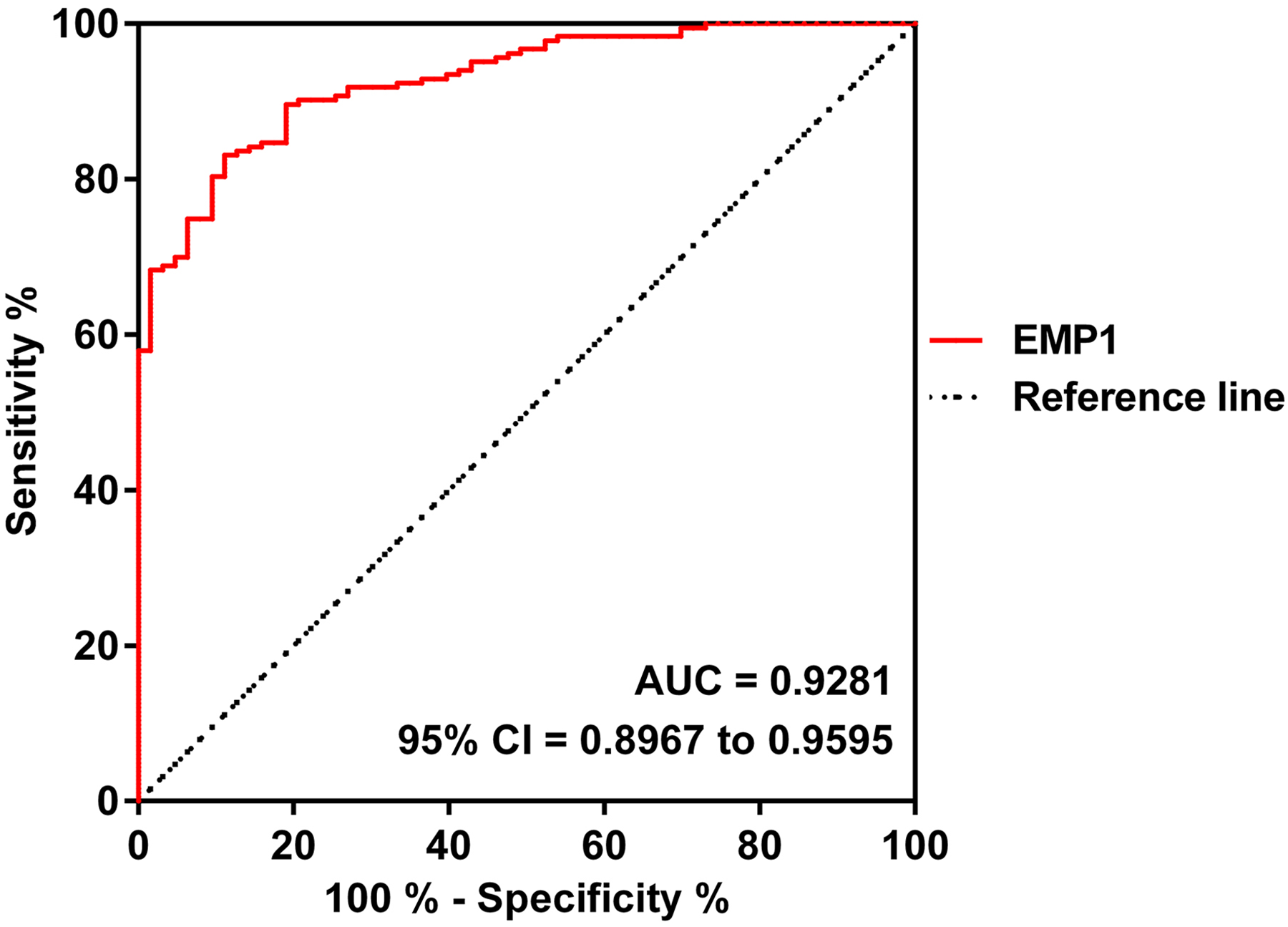
ROC analysis. The diagnostic value of EMP1 mRNA in terms of discriminating ECCA from healthy controls. AUC = 0.9281, 95% CI = 0.8967–0.9595.
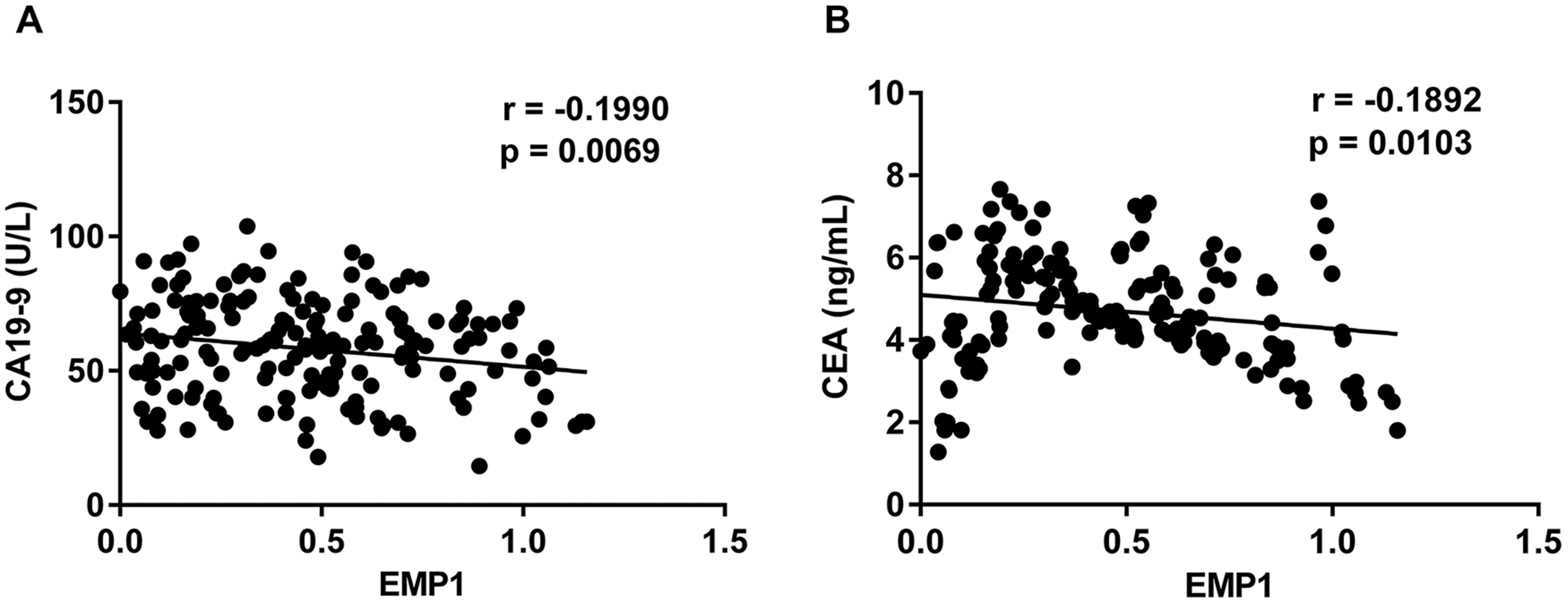
Spearman's correlation analysis. (a) The correlation between EMP1 mRNA expression and CA19-9 and (b) the correlation between EMP1 mRNA expression and CEA.
Clinicopathologicial features in ECCA patients with eccentric EMP1 mRNA expressions
According to the average expressions of EMP1 mRNA in ECCA patients, we divided them into two groups: EMP1 mRNA low-expression group and EMP1 mRNA high-expression group (cut off value 0.4600). As shown in Supplementary 3, there were significant differences in lymph node metastasis, TNM stage, CA19-9 level and CEA level between the two groups (P < 0.01); however, there were no differences in gender, age, differentiation, tumor size, and invasion.
Elevated EMP1 expression indicated unfavorable outcomes in ECCA
As to evaluate the prognostic significance of EMP1 in ECCA, the OS and DFS rates were measured after a 5-year follow-up investigation. From the results in Figure 4(A) and (B), both OS and DFS were notably different between the two groups. Moreover, ECCA patients with lower EMP1 mRNA expressions exhibit unfavorable prognosis compared to those with higher EMP1 mRNA expressions.
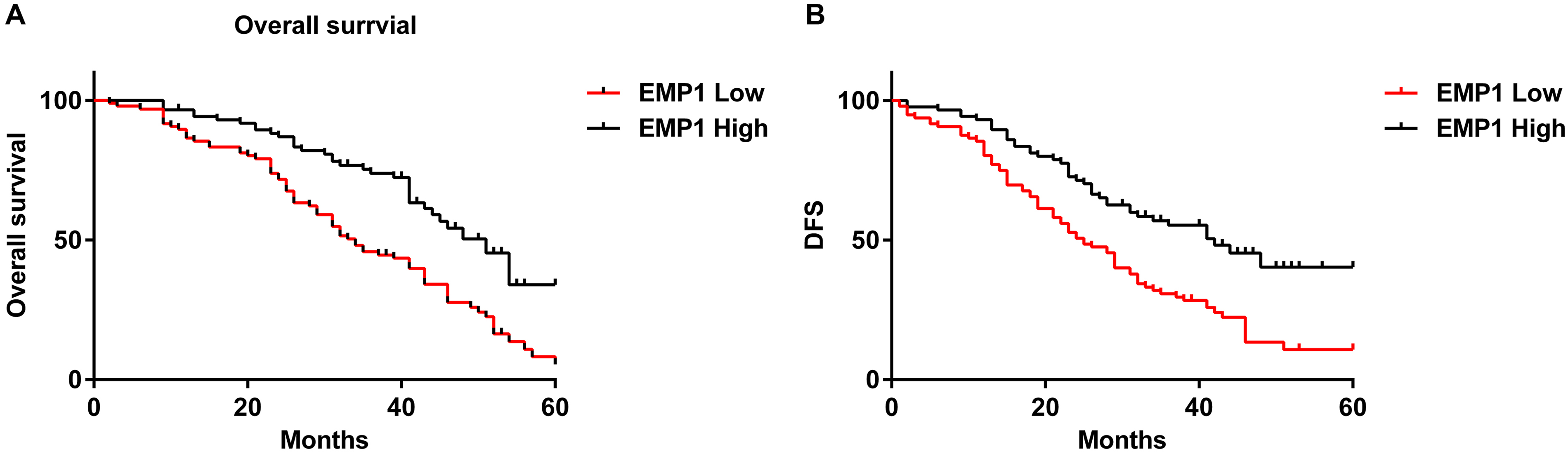
Kaplan–Meier survival analysis of EMP1 mRNA in ECCA. (a) Overall survival analysis and (b) disease-free survival analysis.
Expression of EMP1 in exosomes derived from serum of ECCA patients
Finally, to determine the origin of serum EPM1, serum samples of 10 ECCA patients were collected, and the exosomes were collected by centrifugation; then RNAs were isolated from exosomes and exosome-free serum samples, and the expression of EMP1 in exosome-free serum and exosomes were compared. EMP1 was highly expressed in exosomes derived from the serum of patients with ECCA, and the expression of EMP1 was nearly absent in exosome-free serum samples of cholangiocarcinoma patients (Supplementary 4, P < 0.0001).
Discussion
Worldwide, morbidity and mortality rates in ECCA are still on the rise. Epidemiological reports showed that the main risk factors for ECCA included biliary abnormalities, intrahepatic stones, hepatobiliary shunt, and cirrhosis.3,17 According to clinical information, the main clinical manifestations of ECCA were jaundice, loss of appetite, nausea and vomiting, gallbladder enlargement, liver injury, biliary bleeding, and infection. 18 Since the clinical manifestations of ECCA were nonspecific symptoms or asymptomatic, there were no sensitive screening criteria, and only a few cases were diagnosed at an early stage. Most ECCA patients were diagnosed at an advanced stage and the tumor at that point had invaded the adjacent structure or metastasized to a distance. 19 Even for patients with early diagnosis, cirrhosis complications may increase the difficulty of treatment. 20 In only about 30% of patients ECCA can be surgically removed, and the recurrence rate remained high. 21 Although clinical studies improved patients’ management with ECCA, the prognosis was still low, with a 3-year survival rate of 30% and a 5-year survival rate of 18%. 22 Therefore, finding credible potential diagnostic and prognostic biomarkers for ECCA remains essential. Our study aimed to investigate the expression of EMP1 in human ECCA serum and whether its expression was associated with clinicopathological data in ECCA patients. Subsequently, the possibility of EMP1 as an ECCA diagnostic and prognostic marker was further studied and compared with the level of gastrointestinal tumor markers such as CA19-9 and CEA to confirm the clinical value of EMP1.
After being discovered and named by Taylor, EMP1 was expressed in the heart, placenta, lung, skeletal muscle, kidney, spleen, thymus, prostate, testis, ovary, and small intestine, and colon of adults. 14 A variety of studies recently revealed that EMP1 may be implicated in cell adhesion, apoptosis, proliferation, drug resistance, metastasis, cancer detection, and prognosis. For instance, EMP1 was reported to be overexpressed in prednisolone-resistant pediatric acute lymphoblastic leukemia patients, presenting an unfavorable prognosis compared to those with lower expressions. 23 Meanwhile, EMP1 was reported to function as an oncogene in various cancers, such as lung cancer, 24 human endometrial and ovarian cancers, 25 glioma, 26 and osteosarcoma. 27 On the contrary, EMP1 was also proved to be down-regulated in many cancers, functioning as an anti-tumor gene. In gastric cancer, colorectal cancer, prostate cancer, and breast cancer, EMP1 was deemed a negative tumor regulator associated with survival and prognosis.28–31 In the present study, we used bioinformatic analysis to demonstrate that EMP1 was significantly down-regulated in ECCA; meanwhile, a decreased level of EMP1 was found in serum samples from ECCA patients. Moreover, through ROC and Kaplan–Meier analysis, the potential of EMP1 in ECCA diagnosis and prognosis was examined. From the results, we verified the high accuracy, sensitivity, and specificity of EMP1 mRNA in ECCA; besides, ECCA patients with lower EMP1 mRNA expression presented poorer prognosis than those with higher EMP1 mRNA expression.
CA19-9 and CEA were verified to be markers in gastrointestinal tumors, such as pancreatic, gallbladder, colon, and gastric cancers. 7 As broad-spectrum tumor markers, they cannot be used as specific indicators for diagnosing certain malignant tumors due to their low specificity or sensitivity. However, they still have important clinical value in diagnosis, condition monitoring, and evaluation of malignant tumors’ curative effect. In ECCA, CA19-9 and CEA were also reported to have potentials in predicting tumor diagnosis, progression, and outcome.6,32,33 In our results, we discovered that EMP1 mRNA was negatively correlated with CA19-9 and CEA expressions, further confirming the value of EMP1 in ECCA diagnosis and development.
EMP1 is known as a surface protein, and it is unclear how EMP1 was secreted to the serum. Previous studies suggested that exosomes may be an important transporter for the delivery of the proteins from cells.34,35 Interestingly, our data also suggested that EMP1 was highly expressed in exosomes derived from patients with ECCA, and the expression of EMP1 was nearly absent in exosome-free serum samples of ECCA patients. EMP1 is expressed in exosomes derived form the serum of ECCA patients. However, the underlying mechanism still requires further investigation.
In summary, EMP1 might be a feasible biomarker in ECCA diagnosis and contribute to predicting prognosis. However, a larger number of samples are needed to be studied in the future.
Supplemental Material
sj-docx-1-jbm-10.1177_17246008211035142 - Supplemental material for Serum epithelial membrane protein 1 serves as a feasible biomarker in extrahepatic cholangiocarcinoma
Supplemental material, sj-docx-1-jbm-10.1177_17246008211035142 for Serum epithelial membrane protein 1 serves as a feasible biomarker in extrahepatic cholangiocarcinoma by Xiang Li, Lang Yan and Hao Xue in The International Journal of Biological Markers
Supplemental Material
sj-docx-2-jbm-10.1177_17246008211035142 - Supplemental material for Serum epithelial membrane protein 1 serves as a feasible biomarker in extrahepatic cholangiocarcinoma
Supplemental material, sj-docx-2-jbm-10.1177_17246008211035142 for Serum epithelial membrane protein 1 serves as a feasible biomarker in extrahepatic cholangiocarcinoma by Xiang Li, Lang Yan and Hao Xue in The International Journal of Biological Markers
Supplemental Material
sj-docx-3-jbm-10.1177_17246008211035142 - Supplemental material for Serum epithelial membrane protein 1 serves as a feasible biomarker in extrahepatic cholangiocarcinoma
Supplemental material, sj-docx-3-jbm-10.1177_17246008211035142 for Serum epithelial membrane protein 1 serves as a feasible biomarker in extrahepatic cholangiocarcinoma by Xiang Li, Lang Yan and Hao Xue in The International Journal of Biological Markers
Supplemental Material
sj-jpg-4-jbm-10.1177_17246008211035142 - Supplemental material for Serum epithelial membrane protein 1 serves as a feasible biomarker in extrahepatic cholangiocarcinoma
Supplemental material, sj-jpg-4-jbm-10.1177_17246008211035142 for Serum epithelial membrane protein 1 serves as a feasible biomarker in extrahepatic cholangiocarcinoma by Xiang Li, Lang Yan and Hao Xue in The International Journal of Biological Markers
Footnotes
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
