Abstract
Background:
Several prognostic factors were proposed to improve early detection of recurrence after liver resection of metastases of colorectal cancer. Circulating tumor cell-related transcripts were evaluated in colorectal cancer patients with conflicting results. The aim of this study was to investigate usefulness of carcinoembryonic antigen CAM5, epidermal growth factor receptor, and ERCC1 transcripts in the bloodstream as predictive factors of recurrence in patients who underwent liver resection for metastases of colorectal cancer.
Methods:
Peripheral blood was collected from 29 patients at the time of the colorectal cancer liver metastasis resection, and from 25 normal controls. Follow-up draws (FUDs) were also performed at 30 days, and 3 and 12 months since surgery. On each sample, carcinoembryonic antigen CAM5, ERCC1, and GAPDH mRNAs were examined by quantitative reverse transcription (qRT).
Results:
Carcinoembryonic antigen transcript levels were linearly correlated to the number of spiked cells (qRT analytical limit = five cells). Among 29 patients (20 M/9 F; mean age 63 years (range 32–79), highly significant levels of carcinoembryonic antigen, if compared to the baseline, were detected in those relapsing after surgery (P <0.05). The main differences were between the 1st- and 12th-month FUDs. Significantly higher levels of carcinoembryonic antigen were also detected in patients who died from disease progression during the follow-up (as evaluated at 30 days and 90 days FUDs).
Conclusions:
Blood carcinoembryonic antigen-mRNA absolute copy number overtime variation can represent a valid early predictor of relapse after liver resection in colorectal liver metastases patients. Prospective studies, in the context of large clinical trials, will provide further data to also qualify ERCC1 as a predictive biomarker for decisions on therapeutic strategies.
Introduction
Colorectal cancer (CRC) is the third most common cancer worldwide with 1.2 million cases diagnosed every year.1–2 Improvements in surgical technique and increased effectiveness of modern chemotherapy have led to good results after liver resection, with a mean of 5-years overall survival of 40% to 45%. Recurrence after radical liver resection still represents the main issue, with 60% of patients being at risk of relapse, and 40% relapsing within 12 months after surgery.3–5
Conventional circulating biomarkers (such as carcinoembryonic antigen (CEA) and Ca19.9) are not sufficient to predict a patient’s response after surgical treatment or chemotherapy; therefore, there is an urgent need to identify new early biomarkers above all in CRC. During the last decade, several studies focused on the role of tumor cells in blood (circulating tumor cells (CTCs)) as a new prognostic factor in patients with metastatic breast, colon, and prostate cancer.6–11
Along with CTCs, circulating free nucleic acids (ctDNAs, mRNAs, and non-coding RNAs) can be used as indirect markers of the presence of CTC in the bloodstream. Therefore, polymerase chain reaction (PCR)-based methods are widely used for the detection of CTCs, for example to target tumor-specific genes mRNAs. Once released in the bloodstream from malignant cells, mRNA is relatively unstable, with an unpredictable half-life; nevertheless, once detected, such transcripts can be indicative of the presence of viable tumor cells in the patient’s blood. 12 Traditional reverse transcription polymerase chain reaction (RT-PCR) can provide qualitative information about blood cancer cell load, being sometimes semiquantitative.13–15 Moreover, quantitative RT-PCR or digital PCR (qRT-PCR; dPCR) allow for quantification tumor load in peripheral blood. The assessment of specific RNA copy number cut-off can allow clinicians to distinguish between those relapsing and those not. The setting of these methods needs to fit with specific requirements above all for the normalization of data.16–17
The carcinoembryonic antigen (CEA or CEACAM 5) holds a special position among overall tumor markers. In fact, it is released by dysregulated tumor tissue into the blood and lymphatic vessels through intercellular spaces. CEA is overexpressed in many tumors, mainly in CRC and its metastases. In many patients, the blood level of CEA rises; therefore, pre- and post-operative CEA levels provide independent prognostic information. There are many factors influencing the dynamics of circulating CEA, mainly related to tumor size and staging18–19 and some of these affect the usefulness of CEA protein assay in the clinical setting.
Very recently it has been reported that ERCC1 is the most promising predictive marker of resistance to oxaliplatin; although currently there is no standard test available, ERCC1 assay could be implemented as a routine test in CRC patients in the near future.20–22
Furthermore, the epidermal growth factor receptor (EGFR) also has been reported as detectable in a wide variety of cancers (e.g. lung, breast, ovarian, bladder, esophageal, cervical, and head and neck tumors) where its overexpression was suggested as a progression factor associated with poor prognosis.23–27
The aim of the present study was to verify the usefulness of CEACAM5, EGFR, and ERCC1 transcripts in the bloodstream as prognostic factors for recurrence in patients who underwent liver resection for metastases of CRC. To this end, the circulating levels of these three specific mRNAs were assayed by qRT-PCR before and after surgery. Finally, ERCC1 variations were evaluated during the follow-up of patients treated with FOLFOX at 3 and 6 months after the start of adjuvant chemotherapy.
Materials and methods
Patients and controls selection
From November 2013 to February 2016, 29 patients who underwent liver resection for colorectal metastases at the Hepatobiliary Surgery Unit in our institution were considered eligible for this study. Any eligible patient was enrolled only after having signed the specific informed consent approved by our Ethical Committee.
The inclusion criteria were patients: (a) with both synchronous or metachronous liver metastases (colorectal liver metastases (CRLM)); (b) with both single or multiple CRLM; (c) primary tumor already resected; and (d) without extrahepatic disease who were radically resected.
Blood samples were collected at the time of surgery (0) and 30 days, and 3 and 12 months after resection.
During surgery, we collected three different blood samples: the first from a peripheral vein, the second from the central vein catheter (CVC) and the third from the portal vein. Blood draws were taken at basal conditions, before liver mobilization and manipulation.
Every patient underwent a strict clinical–radiologic follow-up that included a computed tomography (CT) scan and/or magnetic resonance imaging (MRI) and/or ultrasound. The first radiologic examination was undertaken 3 months after surgery.
Not all patients received chemotherapy after surgery, according to both surgeon and medical oncologist decisions. One month after surgery, a new peripheral blood sample was collected, before starting the first chemotherapy scheme. Another sample was collected 3 months after liver resection, mostly simultaneously to chemotherapy. The last blood draw was obtained 12 months after surgery.
Follow-up was performed by telephone calls every 3 months for each patient. Some patients were followed by the Clinical Oncology Unit of our hospital, where each case was routinely discussed within multidisciplinary meetings. To ensure a closer follow-up, we offered an additional liver ultrasonography procedure to further evaluate patients between the CT or MRI that was already routinely scheduled. Imaging was evaluated according to the RECIST criteria.34
In order to try to define a possible cut-off for each marker investigated exclusively at the peripheral blood vein, we compared the results of the 29 patients with 25 controls. The age range of the control individuals was between 25 and 60 years. Inclusion criteria were: non-smokers and people not suffering from conditions related to increased CEA levels (pancreatitis, inflammatory bowel diseases, liver cirrhosis, etc.).
Sample processing, RNA extraction and cDNA synthesis
Total RNA from 3 mL of blood samples (derived from 9 mL of blood into a tube with EDTA) was extracted and treated with DNAse using Perfect pure RNA blood kit (5 Prime, Eppendorf s.r.l., Italy) according to the manufacturer’s recommendation. RNA integrity and quantification were electrophoretically determined using high-sensitivity Experion-chip (Biorad, Hercules, CA). All RNA samples were stored at −80°C until used. All samples were reverse-transcribed and amplified. Reverse transcription was carried out on about 250 ng of total RNA according to the protocol of Transcriptor First Strand cDNA synthesis kit (Roche Applied Science, Indianapolis, IN). The intact cDNAs were stored at −20°C or immediately used for subsequent amplification reactions.
Real-time amplification
The amplification in qRT-PCR was performed on Roche Light Cycler 480 by using primers and probes for CEA-CAM5, ERCC1, and GAPDH (TaqMan or SyB Green-based platforms). Assay detection limit was performed by mixing serial dilutions of HCT 116 with donor-derived peripheral blood. Standard curves were generated by making recombinant clones; the expression of each marker was normalized by the GAPDH housekeeping gene as a ratio. The CEA mRNA level was linearly correlated with numbers of spiked cells (qRT analytical detection limit = five cells).
Statistical analysis
Data were analyzed using IBM SPSS version 20 (IBM Corporation, Armonk, NY, USA). Differences in continuous variables were tested using an independent sample Student t test and a Chi square test for contingency tables.
Results
Patients
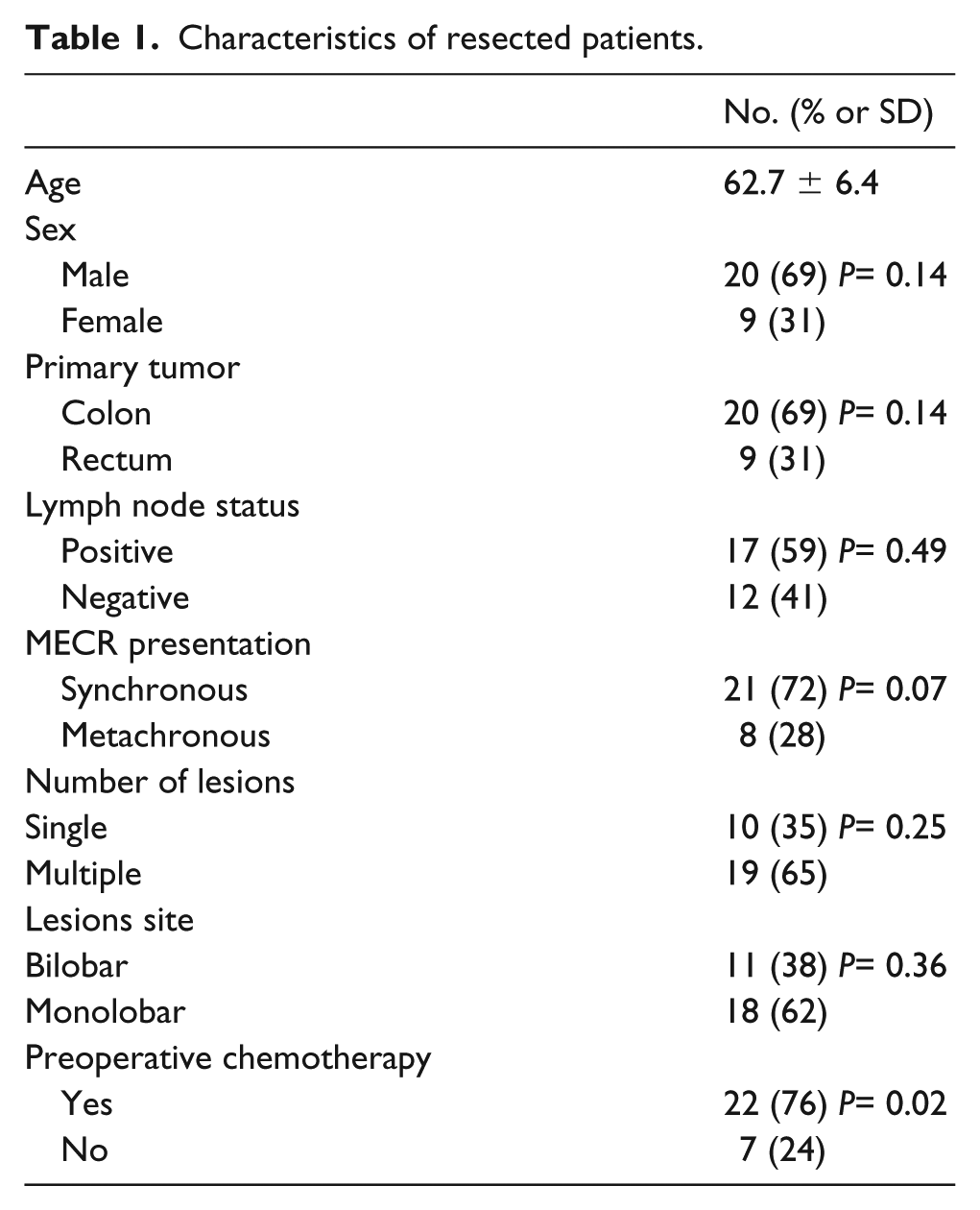
Between November 2013 and February 2016, we enrolled 29 patients resected for colorectal liver metastases at the Hepatobiliary Surgery Unit in our hospital. The mean age was 62.7 years (range 32–79). Twenty out of 29 patients were male (68.9%): 21 showed synchronous metastases (72.4%), while 19 (65.5%) presented with multiple metastases. In detail, the mean number of metastases per patient was 6 (range 2–17) with bilobar localization in 11/29 cases (37.9%). In twenty patients (68.9%) the site of the primary tumor was the colon, while in 9 patients it was the rectum (60.9%). Lymph node involvement of the primary was present in 17 patients (58.6%). Most of the patients underwent preoperative chemotherapy (n=22; 75.8%), that was interrupted at least 3 weeks prior to surgery. Biological agents have been administered in 14 cases (63.6%). Patients’ characteristics are summarized in Table 1. No significant results were found regarding the clinical characteristics of our patients, with the exception of preoperative chemotherapy (P< 0.02); 76% of our patients received this therapy before surgical treatment.
Characteristics of resected patients.
Comparison of markers expression between cancer patients and controls
To determine cut-off values of RT-PCR, we compared mRNA copy numbers between CRLM patients and healthy subjects. Our analysis showed no significant differences of baseline CEACAM5, ERCC1, and EGFR blood levels between CRLM and controls. For this reason, we couldn’t set a cut-off value for these markers in order to identify patients with worse prognosis.
Furthermore, EGFR mRNA blood levels were always negative or close to 0 in both groups; therefore, we excluded this transcript from the statistical analysis.
The copy numbers of circulating transcripts are summarized in Table 2.
Comparison of markers expression between CRLM and controls at baseline.
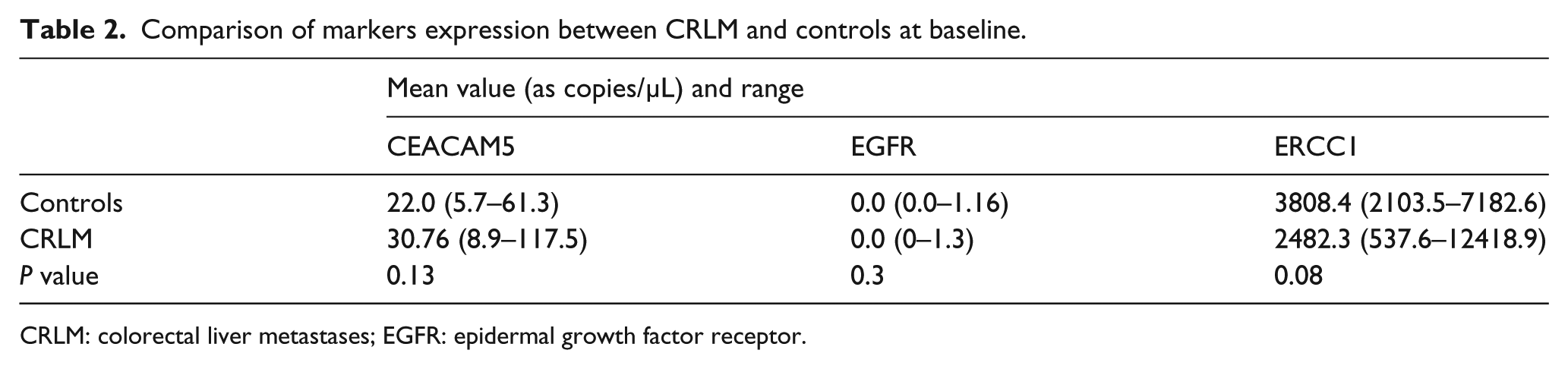
CRLM: colorectal liver metastases; EGFR: epidermal growth factor receptor.
Comparison of markers expression according to the site of blood draw
At the time of surgery, from the first 15 patients enrolled in our study, we collected three blood draws from three different venous accesses before any manipulation and mobilization of the liver: (a) peripheral blood; (b) portal vein; and (c) central venous circulation by using a CVC. By comparing the expression of CEACAM5 and ERCC1 with the housekeeping GHPDH gene, no significant difference was found regarding the three sites of the blood draws that were analyzed. For this reason, for the remaining 14 patients, only a peripheral blood sample was drawn at the baseline.
Association of CEACAM5 and ERCC1 expression and clinic data
Among the 29 patients included in this study, 15 (51.7%) developed recurrence within 12 months after surgery. In particular, liver metastasis was present in 10 patients (66.6%), while other recurrence sites involved lung (n=5, 33.3%), abdominal lymph nodes (n=2, 13.3%), brain (n=1, 6.6%), bone (n=1, 6.6%), and peritoneum (n=1, 6.6%). Six patients presented with multiple localizations of recurrence (40%). Recurrence appeared after a mean disease-free survival (DFS) of 6.6 months (range 3–12 months).
During the period of enrollment, five patients died: four died from disease progression, and one from heart failure. A second blood draw was performed on all five patients during the first month after surgery, while only two completed the check-point scheduled at the third month. None of them were able to entirely complete the blood draws scheme schedule.
Twenty-three patients (79.3%) underwent post-operative chemotherapy. Oxaliplatin based regimens were administered in 13 cases (56.5%). One month after surgery no patients had started chemotherapy; however, at the third month of blood draw, 22 CRLM (84.6%) were receiving systemic chemotherapy (1 out of the 23 had died by then). At the 12th month of the scheduled check-points, only 4 out of 21 patients still alive were undergoing treatment (19%). The number of patients receiving treatment at 3 months was significantly higher than at 1 and 12 months (P<0.001).
We stratified our CRLM patients into two groups: the first showed early recurrence (n=15) and the second had slower progression (n=14). By analyzing the behaviors of our transcripts, we found that CEACAM5 was significantly differentially expressed between the two groups. Particularly, patients with disease recurrence showed an increasing trend of CEACAM5 expression during the 12-month follow-up (Figure 1). Surprisingly, similar (and not significantly different) levels of CEACAM5 between the two groups were found at the third month of observation. These results were confirmed after correction for preoperative chemotherapy by univariate analysis (P< 0.05).
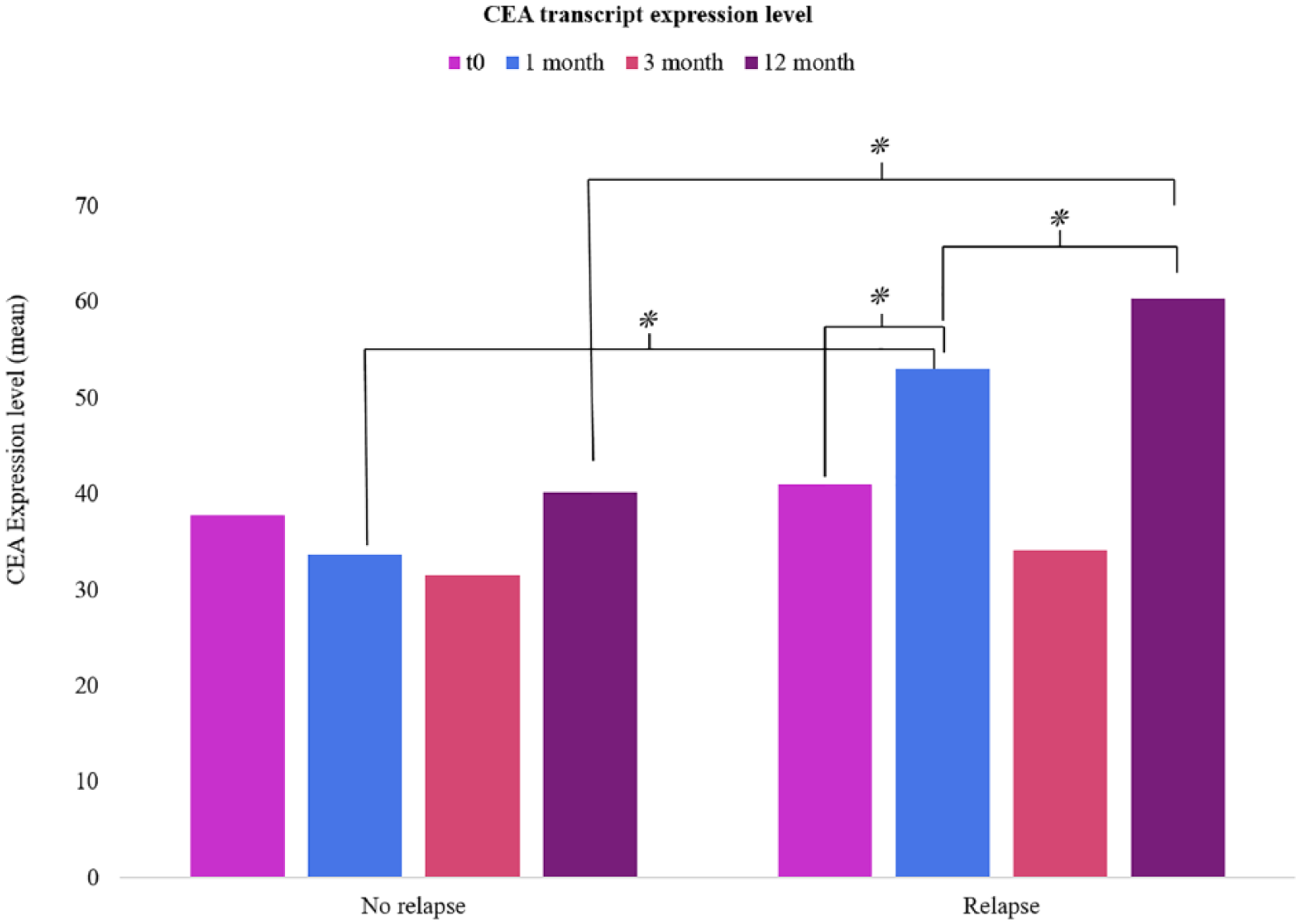
Differential expression level of CEA transcript among patients with or without recurrence at each follow-up draw.
Patients who died during the 12-month follow-up showed impressively higher CEACAM5 levels at the baseline and after 1 and 3 months from surgery, compared to the other patients (Figure 2).
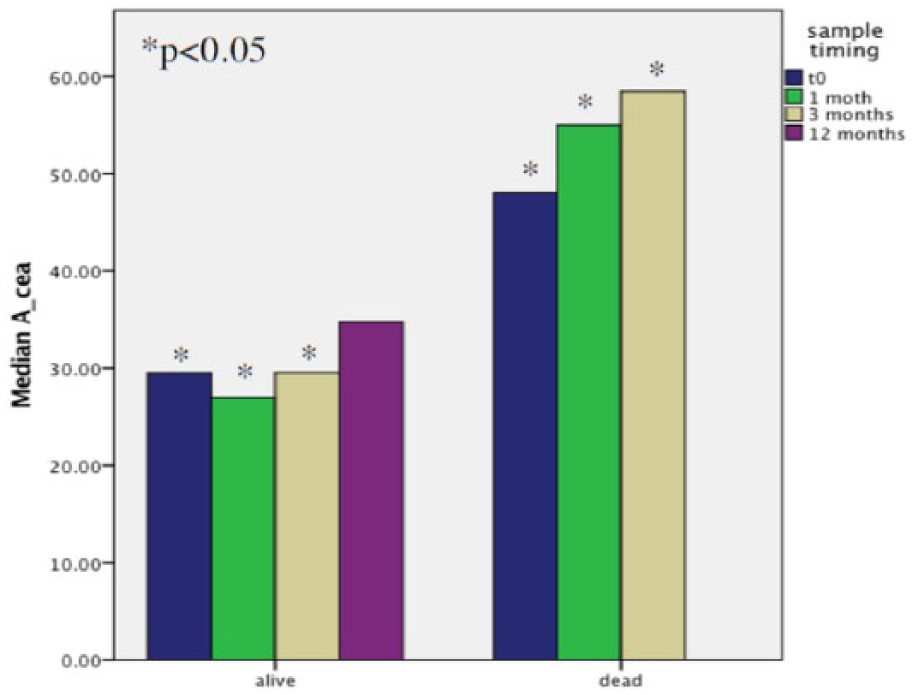
Comparison of CEACAM5 copy amounts between patients who died during the study period and those who did not.
Patients who underwent post-operative oxaliplatin-based chemotherapy showed reduced ERCC1 mean copy levels compared to those untreated. Moreover, this difference was particularly evident at 1 and 3 months after surgery, while it was not confirmed after 12 months (Figure 3).
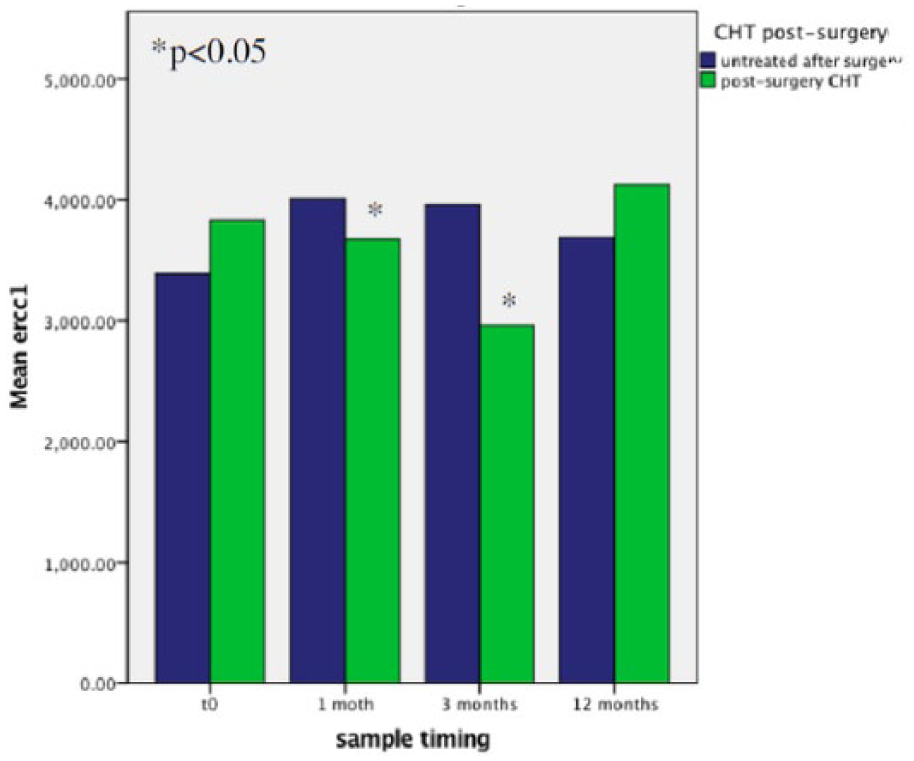
ERCCI expression overtime, according to post-operative chemotherapy.
Discussion
In this study we considered a group of 29 patients resected for CRLM. These patients were heterogeneous in terms of both number of lesions and presentation of the metastatic disease (synchronous or metachronous). The inclusion of patients with this variety of clinical presentations reflects the case mix of the patients routinely admitted at our Policlinico A. Gemelli Hepatobiliary Surgery Unit. Seventy-six percent of patients underwent preoperative chemotherapy, while 79% received post-operative treatment.
The aim of our study was to explore the role of CEACAM5, EGFR, and ERCC1 transcript blood levels as prognostic markers for recurrent disease after liver resection for metastases of CRC. In particular, we attempted to evaluate the possible validity of CTC-related transcripts as potential prognostic factors for recurrence.
Therefore, mRNA expression levels of the above three CTC-related transcripts were evaluated and normalized by the GAPDH housekeeping gene as a ratio.
Regarding the quantitative molecular method, our qRT-PCR method was set-up in order to detect one colorectal carcinoma cell within 107 leukocytes (1–10 mL of blood), which is three times higher than immunocytochemistry. 13
For each patient, four blood drawings were scheduled: the first at basal conditions, and the remainder at 1, 3 and 12 months after surgery. Since in the first 15 patients’ blood collected from peripheral, portal, and CVC did not show any difference in terms of copy numbers for the three transcripts, we decided to evaluate only the peripheral blood.
Due to the absence of any significant difference in marker expression between controls and metastatic liver patients at the baseline, we were not able to set a cut-off to identify patients with the worst prognosis. Furthermore, since EGFR was close to zero at any detection time scheduled, it was not considered for the statistical analysis. We underline that data regarding EGFR mRNA evaluation on the blood of CRC metastatic patients are not reported in the literature; unfortunately, our paper is not able to provide significant results about this point of investigation. Nevertheless, by reviewing literature data, 28 we found that EGFR genomic gains were the most frequent in the primary CRC tissues (11%) and (37%) of metastatic lesions. Therefore, although it would be conceivable that at least one-third of metastatic patients might also show increased levels of circulating RNA, in our setting, both the three blood sources used (peripheral vein, CVC, and portal vein) at the T0 and the following T1 and T2 peripheral blood samples did not show any difference in the levels of EGFR. These results are not so surprising if we consider data recently reported by Vojtechova et al., 29 where they were not able to enrich CTCs from the blood of stage 3 and 4 CRC patients. In fact, only one out of three patients was positive for CTCs: this stage 4 CRC patient was positive for EGFR mRNA only in the first of five samples (before surgical treatment), while the remaining ones (during the surgery, 1 and 7 days after surgery, and finally 3 months later), were positive only for CEA mRNA. Similar results were also obtained by Pesta et al. 30 These findings completely overlap with those obtained in our study; therefore, we can agree with the conclusions of Vojtechova et al. 29 who stated that as there are numerous reports in the literature showing that the presence of CTC in colorectal patients (especially in advanced metastatic disease stages) it can hardly be questioned.8,29 In fact, the successful CTC recovery naturally relies on: (a) proper timing and methodology applied during the sample acquisition; (b) leukocyte contamination whose genetic material could saturate the RT-PCR reaction, thus hampering detection of EGFR, CEA, and GA733-2 transcripts originating from CTCs. 29 Interestingly, 52% of patients developed recurrence within 12 months after surgery. Not surprisingly, these patients showed increased CEACAM5 copy numbers, with a mean maximum peak of expression at the 12th-month check-point (P<0.05). Circulating levels of CEACAM5 were significantly higher at 1 and 12 months in relapsing patients compared to those who did not. Although no difference was found at 3 months between the above reported groups, we found decreasing CEACAM5 transcript levels from 1 to 3 months, which was probably due to the fact that at that time of observation 85% of patients were receiving chemotherapy. In the absence of any clinically validated cut-off for CEACAM5 circulating levels, we mainly focused on the dynamics of its circulating levels, which, when increasing, raised the clinical prognostic relevance. Although preliminary, this study showed that the post-surgery increase of CEACAM5 copy numbers was associated with disease recurrence. The limit of the CEACAM5 transcript is that it is not able to work as an early marker, with the significant difference detectable only at 12 months. Nevertheless, in our setting, changes between baseline and 1 month after surgery could become a useful and easily achievable marker if validated on larger cohorts of patients.
Additionally, patients who died during our 12 months enrollment showed significantly higher CEACAM5 levels at each time point. Although these findings could represent a promising result, they need to be further validated. To this end, we would like to emphasize that such a follow-up is not easy, especially when liquid-biopsy-based studies are set; in fact, in order to guarantee a homogeneous path (from patients’ blood collection to the biomarker assays) and to reduce pre-analytical biases, we enrolled only patients for whom it would be possible to follow-up in our hospital. Therefore, although the number of enrolled patients may appear to be low, we were able to set a highly standardized setting.
Finally, also Shirota et al. 20 showed how patients receiving FOLFOX with decreasing level of ERCC1 are associated with better overall survival. In keeping with these literature data, our patients showed significantly lower levels of ERCC1 at both 1 and 3 months after surgery, especially between those undergoing chemotherapy. This feature is difficult to understand for three reasons: (a) there is an imbalance between the two groups of patients, with 79% of them being treated with post-operative chemotherapy; (b) oxaliplatin, which is correlated to ERCC1 function, was administered in 56% of cases; and (c) 1 month from surgery, ERCC1 levels were still significantly lower among patients who had not yet started chemotherapy.
In conclusion, CEACAM5 mRNA seems to play a promising role as an early predictor of recurrence after liver resection for CRLM. Perspective studies in the context of large clinical trials will provide further data to also qualify ERCC1 as a predictive biomarker for selection therapy.
Footnotes
Author contributions
Felice Giuliante and Elena Panettieri are co-first authors.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
