Abstract
Introduction:
Glioma is the most common primary brain tumor. The small nucleolar RNA host gene (SNHG) SNHG6 is a potential oncogene in the development of several types of cancers.
Methods:
In this study, we investigated the functional role of long non-coding RNA (lncRNA) SNHG6 in the malignancy of glioma in cell lines and transplanted nude mice.
Results:
We found that the expression of lncRNA SNHG6 was higher in glioma tissues and cells than in normal brain tissues and cells. The expression of lncRNA SNHG6 was positively correlated with the malignancy and poor prognosis of glioma patients. microRNA (miR)-101-3p expression was decreased in glioma tissues and cells and was negatively correlated with the malignancy and poor prognosis of glioma patients. In glioma tissues, the expression of lncRNA SNHG6 was negatively correlated with the expression of miR-101-3p. SNHG6 contained a binding site of miR-101-3p. Knockdown of SNHG6 expression resulted in a significant increase of miR-101-3p expression. miR-101-3p mimic markedly decreased the luciferase activity of SNHG6. Knockdown of SNHG6 inhibited glioma cell proliferation, migration, and epithelial-mesenchymal transition (EMT), and increased apoptosis. miR-101-3p mimic enhanced knockdown of SNHG6-induced inhibition of cell proliferation, migration, and EMT, and an increase of apoptosis. Anti-miR-101-3p reversed the the effects of si-SNHG6 on cell malignancy. Knockdown of SNHG6 remarkably reduced the increase of tumor volumes in xenograft mouse models. In tumor tissues, knockdown of SNHG6 increased the expression of miR-101-3p and reduced EMT biomarker expression.
Conclusions:
Our study provides novel insights into the functions of lncRNA SNHG6/miR-101-3p axis in the tumorigenesis of glioma.
Introduction
Glioma is the most common primary tumor in the adult central nervous system and is characterized by a high malignancy, rapid progression and early metastasis. The treatment and prognosis of glioma patients have only been slightly improved by surgical resection followed by radiation therapy and chemotherapy. The average survival of malignant glioma patients is only 12–15 months (1-4). Therefore, identification of new biomarkers and novel therapeutic strategies is crucial for effective treatment.
It is well known that although more than 90% of the DNA sequences are actively transcribed, only 2% encode protein. Most of the transcripts are considered to be non-coding RNAs (ncRNAs) (5, 6). Originally, those ncRNAs are believed to be junk RNA with no function, including long non-coding RNAs (lncRNAs) (7). Emerging evidence has supported that lncRNAs function as critical regulators of physiological processes and pathological conditions, including cancer (8). A variety of lncRNAs play oncogenic or tumor suppressive roles, or both (9, 10). In recent years, it has been found that small nucleolar RNAs (snoRNAs) and their host genes (SNHGs) have various functions in the development of several types of human cancers (11-13). In particular, SNHG6 has been identified to be a potential oncogene in the development of gastric cancer and hepatocellular carcinoma (14-16). However, the functional role of lncRNA SNHG6 in glioma is still not clear.
In the current study, we investigated the functional role of lncRNA SNHG6 in the malignancy of glioma. We showed that lncRNA SNHG6 expression was increased in glioma tissues and cells, and was positively correlated with the malignancy and poor prognosis of glioma patients. The expression of microRNA (miR)-101-3p was decreased in glioma tissues and cells, and was negatively correlated with the malignancy and poor prognosis of glioma patients. The expressions of lncRNA SNHG6 and miR-101-3p in glioma tissues were negatively correlated. lncRNA SNHG6 possessed a direct binding site of miR-101-3p. Knockdown of SNHG6 inhibited proliferation, decreased migration ability, and increased apoptosis in glioma cells, and inhibited the growth of tumor volume in transplanted nude mice. miR-101-3p mimic enhanced, but anti-miR-101-3p inhibited, the reduction of glioma malignancy induced by knockdown of SNHG6.
Materials and methods
Patients and tissue specimens
A total of 71 glioma tissues and 10 normal brain tissues were obtained from Tangdu Hospital, Fourth Military Medical University between October 2015 and December 2016. The informed consent was obtained from all the involved patients, and the study was approved by the Research Ethics Committee of Tangdu Hospital, Fourth Military Medical University. The patients did not receive any radiotherapy or chemotherapy before surgery. The age- and gender-matched normal brain tissues were obtained from patients with cerebral trauma or cerebral hemorrhage, and were cut and used as controls. Tissue samples were stored in liquid nitrogen immediately after surgery for the extraction of total RNA. Two independent experienced clinical pathologists confirmed the pathological diagnosis of all glioma patients according to the World Health Organization (WHO) classification. The patients in our study had 29 low grade (grade I–II) and 42 high grade (grade III–IV) glioma. From the date of surgical resection, all patients had experienced a 60-month follow-up period. Overall survival (OS) was defined as the time from the date of initial surgical operation to the date of death or the date of the last follow-up (if death did not occur).
Cell culture
Human brain astrocyte, HEB and human glioma cell lines, including U87, U251, LN229, and T98G, were purchased from the American Type Culture Collection (ATCC, Rockville, MD, USA). Normal human embryonic kidney 293T (HEK293T) cells were purchased from the Cell Bank of the Chinese Academy of Science (Shanghai, China). Cells were cultured in Dulbecco’s Modified Eagle’s Medium (DMEM; Gibco, USA) supplemented with 10% fetal bovine serum (FBS; Gibco, USA). Cells were maintained in a humidified incubator at 37°C with 5% CO2.
Plasmid construction and cell transfection
To construct si-SNHG6 vector, the self-complementary hairpin DNA oligonucleotides (5′-CTGCGAGGTGCAAGAAAGCCT-3′) targeting the transcript of SNHG6-203 (ENST00000520944.5) and a negative control (named as si-NC vector: 5′-GCTTCGCGCCGTAGTCTTA-3′) were annealed and ligated into the Bal II and Hind III sites of linearized pSuper RNA system shRNA expression vector (Oligoengine). miR-101-3p mimic (5′-UACAGUACUGUGAUAACUGAA-3′; 5′-CAGUUAUCACAGUACUGUAUU-3′) and control mimic (NC-mimic) (5′-UUCUCCGAACGUGUCACGUTT-3′; 5′-ACGUGACACGUUCGGAGAATT-3′) were used to overexpress miR-101-3p. In parallel, anti-miR-101-3p (5′-CAGUACUUUUGUGUACAA-3′; 5′-UUCAGUUAUCACAGUACUGUA-3′) was used to downregulate miR-101-3p expression with anti-control (anti-NC) as a control. miR-101-3p mimics and inhibitors and their respective controls were synthesized by RioBio Company (Guangzhou, China). Twenty nM si-SNHG6, si-NC, miR-101-3p mimic, NC-mimic, anti-miR-101-3p, and anti-NC were transfected into U87 and LN229 cells with Turbofect reagent (Thermo Fisher Scientific, Rockford, IL, USA). Stable cell lines were established by G418 selection (0.5 mg/mL, Sigma–Aldrich, St Louis, MO, USA).
Luciferase reporter assay
The full SNHG6 cDNA were amplified by polymerase chain reaction (PCR) from human glioma brain tissues, and then cloned into the pGL3-Basic luciferase reporter vector (Promega, Madison, WI, USA) to generate pGL3-SNHG6 (SNHG6-WT) (RLuc-SNHG6-WT-F 5′-CTA GTC TAG AGA GGT GAA GGT GTA TGA AAG-3′; RLuc-SNHG6-WT-R 5′-CCG GAA TTC CAT CAT TTG CTT CTG TTC ATG-3′). In addition, the mutant miR-101-3p binding sites were replaced as indicated to generate pGL3-SNHG6-MUT (SNHG6-MUT) (RLuc-SNHG6-MUT-F 5′-CTA GTC TAG AGA GGT GAA GGT GTA TGA AAG-3′; RLuc-SNHG6-MUT-R 5′-CCG GAA TTC CAT CAT TTG CTT CTG TTC ATG-3′). The luciferase assays were performed using the Dual-luciferase reporter assay system (Promega, Madison, WI, USA). Briefly, 4.5×104 HEK293T cells were planted into 48-well plates. When the cells were 50% confluent, they were co-transfected with 500 ng constructed SNHG6 plasmid vector and 30 ng miR-101-3p mimic or NC-mimic by Turbofect reagent (Thermo Fisher Scientific). Cells were harvested 48 h after transfection for luciferase reporter assay with a Dual-Luciferase Assay Kit (Promega, Madison, WI, USA). The relative luciferase activity was normalized to the Renilla luciferase activity.
Cell viability
In brief, after the treatment, the cell culture medium was removed. Cell viability was evaluated by a cell counting kit (CCK)-8 (Beyotime, Shanghai, China) according to the manufacturer’s instructions. After incubation for 2 h at 37°C, the absorbance was recorded at 450 nm with a microplate reader.
Apoptosis
Apoptotic cell death was measured using a TUNEL apoptosis detection assay kit (Beyotime). After treatment, the cells were trypsinized and then suspended in a reaction solution and stained with TUNEL. After washing with PBS, cells were analyzed using a FACScan flow cytometer (BD Biosciences, USA). The results were expressed as a percentage of apoptotic cells.
Invasion assays
Cell invasion assay was carried out using the 24-well Transwell chamber with 8 mm pore size (BD Biosciences). The cells were trypsinized, suspended with DMEM medium, and then 1×105 cells was added into the upper chamber with 100 μl serum-free DMEM. The bottom chamber was filled with DMEM containing 10% FBS. Undergoing migration for 24 h, non-migrated cells in the upper chamber were removed using a cotton swab, then the filters were individually fixed with 4% paraformaldehyde for 0.5 h and stained with hematoxylin and eosin. The number of invasive cells on the bottom surface was counted under a light microscope. The experiments were performed in triplicate and the cell numbers were counted in five random fields of each chamber under the microscope.
Quantitative real-time PCR
Cells were lysed with Trizol (Life Technologies, Carlsbad, CA, USA) and total RNA was isolated from cell lines. RNA was reversely transcribed into cDNA using the PrimeScript RTTM according to the manufacturer’s instructions. miRNA from total RNA was reverse transcribed using the Prime-Script miRNA cDNA Synthesis Kit (TaKaRa). Quantitative real-time PCR was used to precisely quantify target genes using gene-specific primer pairs on a ABI PRISM 7500 real-time PCR system (Applied Biosystems, Foster City, CA, USA) using SYBR Premix Ex Taq II (Takara). GAPDH was used as the endogenous control for the detection of mRNA expression levels, while U6 was used as the endogenous control for miRNA expression analysis. The conditions of the reactions were: an initial step at 94°C for five min, followed by 40 cycles of denaturation at 94°C for 30 s, annealing at 63°C for 30 s, and then extension at 72°C for 10 s. Data were collected and analyzed using 2−ΔΔCt method for quantification of the relative mRNA expression levels. The primers used were: SNHG6: Sense: 5′-CCT ACT GAC AAC ATC GAC GTT GAA G-3′; Antisense: 5′-GGA GAA AAC GCT TAG CCA TAC AG. E-cadherin: Sense: 5′-CTG CTG CAG GTC TCC TCT TG-3′; Antisense:5′-TGT CGA CCG GTG CAA TCT TC-3′. Vimentin: Sense: 5′-AAG GCG AGG AGA GCA GGA TT-3′; Antisense: 5′-GGT CAT CGT GAT GCT GAG AAG-3′. GAPDH: Sense: 5′-GGG AGC CAA AAG GGT CAT-3′; Antisense: 5′-GAG TCC TTC CAC GAT ACC AA-3′. miR-101-3p: 5′-GCGCGTACAGTACTGTGATAACTGAA-3′. U6: Sense: 5′-CTC GCT TCG GCA GCA CA-3′; Antisense: 5′-AAC GCT TCA CGA ATT TGC GT-3′.
Transplanted tumor nude mice
Twenty nude mice were obtained from the Animal Center of Fourth Military Medical University. Animal experiments were performed according to the animal experiment protocol approved by Tangdu Hospital, Fourth Military Medical University. In brief, 4 × 106 U87 cells were stably transfected with si-SNHG6 and si-NC were subcutaneously transplanted into the left flank of the nude mice. Tumor volumes were measured every five days using a Vernier caliper. Tumor volume (mm3) = length ×width2/2. The experimental period was 40 days. After the animal experiment, the mice were killed and tumor tissues were collected for further determination.
Statistical analysis
Data were exhibited as mean ± SD of three independent experiments and processed using SPSS 17.0 statistical software (SPSS Inc., Chicago, IL, USA). The differential expression of SNHG6 between glioma and normal brain tissues was analyzed by an independent two-sample t test. OS was evaluated by the Kaplan–Meier analysis, and the log-rank test was used to compare the difference between groups. Correlations between the expression of SNHG6 and miR-101-3p were performed by Pearson’s correlation. P values less than 0.05 were considered statistically significant.
Results
Upregulation of lncRNA SNHG6 is correlated with poor prognosis and is negatively correlated with miR-101-3p level in glioma patients
To examine the pattern of lncRNA SNHG6 expression in glioma, we compared the expression level of lncRNA SNHG6 in normal brain tissues and glioma tissues. We found that the expression level of lncRNA SNHG6 was higher in glioma tissues than in normal brain tissues (Fig. 1A). In addition, we investigated the relationship between lncRNA SNHG6 expression and histological malignancy in 29 cases of low-grade (I–II) and 42 cases of high-grade (III–IV) glioma tissues. In Figure 1B we showed that the expression of lncRNA SNHG6 of the high-grade glioma cases was markedly higher than the low-grade cases. We also compared the expression level of lncRNA SNHG6 in human brain astrocytes, HEB cells, and glioma cell lines, including T98G, U87, U251, and LN229 cells. We showed that compared with the HEB cells, the expression level of lncRNA SNHG6 in T98G, U87, U251, and LN229 cells had notably increased (Fig. 1C). A Kaplan–Meier analysis and a log-rank test were performed to examine the correlation of lncRNA SNHG6 expression with the prognosis of glioma patients. As shown in Figure 1D, patients with higher lncRNA SNHG6 expression had shorter OS time than those with lower lncRNA SNHG6 expression (HR=1.652; 95% confidence interval (CI) 1.102, 3.321; P=0.0076). To examine the pattern of miR-101-3p expression in glioma, we compared the expression level of miR-101-3p in normal brain tissues and glioma tissues. We found that the expression level of miR-101-3p was lower in glioma tissues than in normal brain tissues (Fig. 1E). In addition, we investigated the relationship between miR-101-3p expression and histological malignancy in low-grade (I–II) and high-grade (III–IV) glioma tissues. In Figure 1F we showed that the expression of miR-101-3p in high-grade glioma cases was markedly lower than in low-grade cases. We also compared the expression level of miR-101-3p in human brain astrocyte and glioma cell lines. We showed that the expression level of miR-101-3p in T98G, U87, U251, and LN229 cells was notably lower than that in HEB cells (Fig. 1G). The results of the Kaplan–Meier analysis and the log-rank test showed that patients with higher miR-101-3p expression had longer OS time than those with lower miR-101-3p expression (HR=1.912; 95% CI 1.135, 3.681; P=0.0069) (Fig. 1H). Furthermore, we found that the level of lncRNA SNHG6 was negatively correlated with the miR-101-3p level in tumor tissues in glioma patients (Fig. 1I).
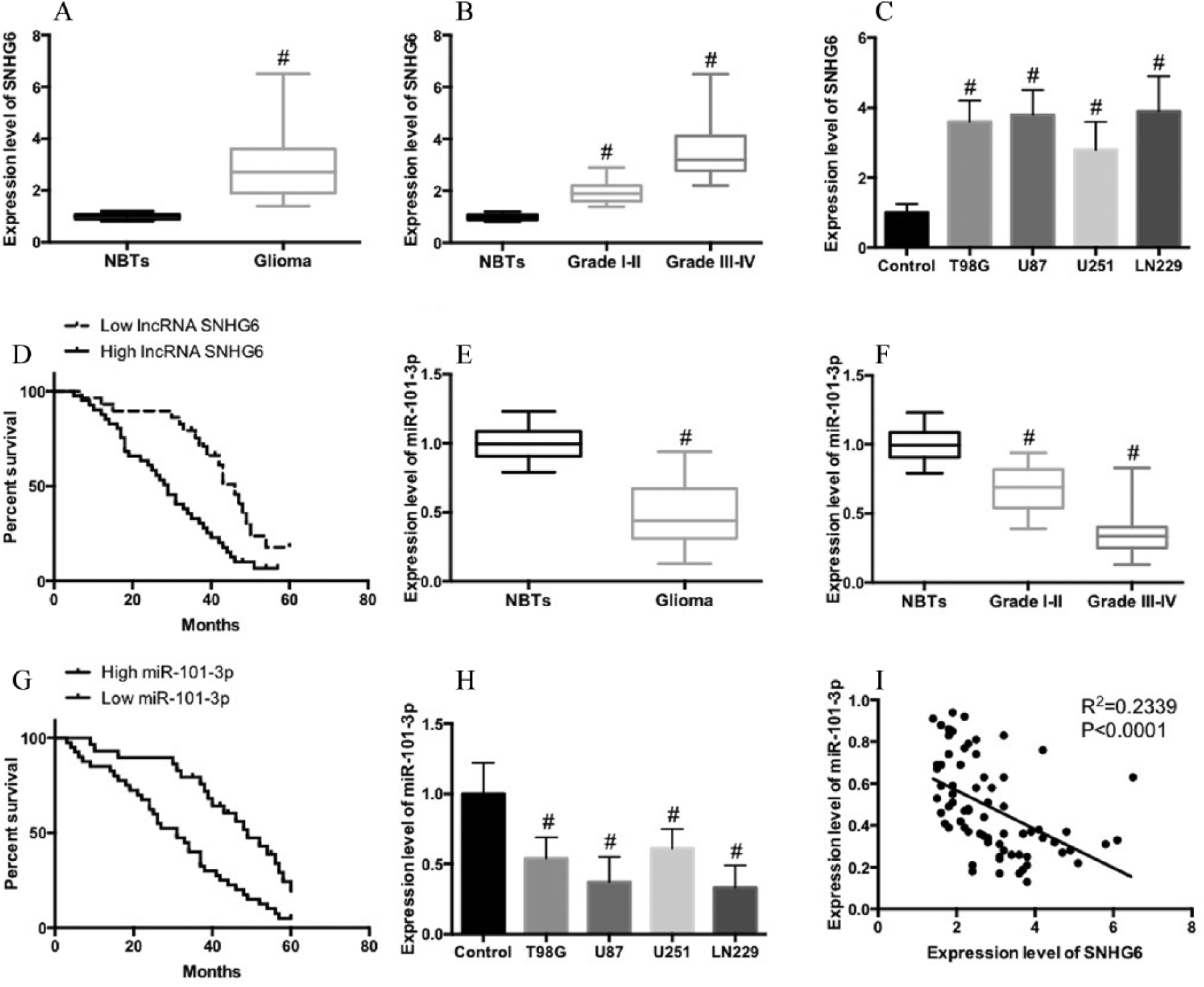
Upregulation of lncRNA SNHG6 is correlated with poor prognosis and is negatively correlated with miR-101-3p level in glioma patients. A) lncRNA SNHG6 expression in 10 normal brain tissues and 71 human glioma tissues were measured by qRT-PCR. B) lncRNA SNHG6 expression in low-grade (I–II) (29) and high-grade (III–IV) (42) glioma tissues. C) lncRNA SNHG6 expression in normal brain and four glioma cell lines as determined by qRT-PCR. D) The patients with glioma were divided into lncRNA SNHG6 high-expression and low-expression group and Kaplan–Meier analysis was performed to evaluate the relationship of lncRNA SNHG6 expression level with prognosis of glioma patients. E) miR-101-3p expression in 10 normal brain tissues and 71 human glioma tissues were measured by qRT-PCR. F) miR-101-3p expression in low-grade (I–II) (29) and high-grade (III–IV) (42) glioma tissues. G) miR-101-3p expression in normal brain and four glioma cell lines as determined by qRT-PCR. H) The patients with glioma were divided into miR-101-3p high-expression and low-expression groups and Kaplan–Meier analysis was performed to evaluate the relationship of miR-101-3p expression level with the prognosis of glioma patients. I) Pearson’s correlation analysis of the relationship between lncRNA SNHG6 and miR-101-3p.
SNHG6 is a target of miR-101-3p
Previous studies have shown that SNHG6 contained a binding site of miR-101-3p (Fig. 2A) (14, 16). In U87 and LN229 cells, knockdown of SNHG6 expression resulted in a significant increase of miR-101-3p expression (Fig. 2B–E). In addition, using the dual-luciferase reporter assay, we found that miR-101-3p mimic markedly decreased the luciferase activity of SNHG6-WT (Fig. 2E and F). In contrast, miR-101-3p mimic did not alter the luciferase activity of SNHG6-MUT (Fig. 2E and F). The results indicated that miR-101-3p can bind directly to SNHG6 within the recognition site.
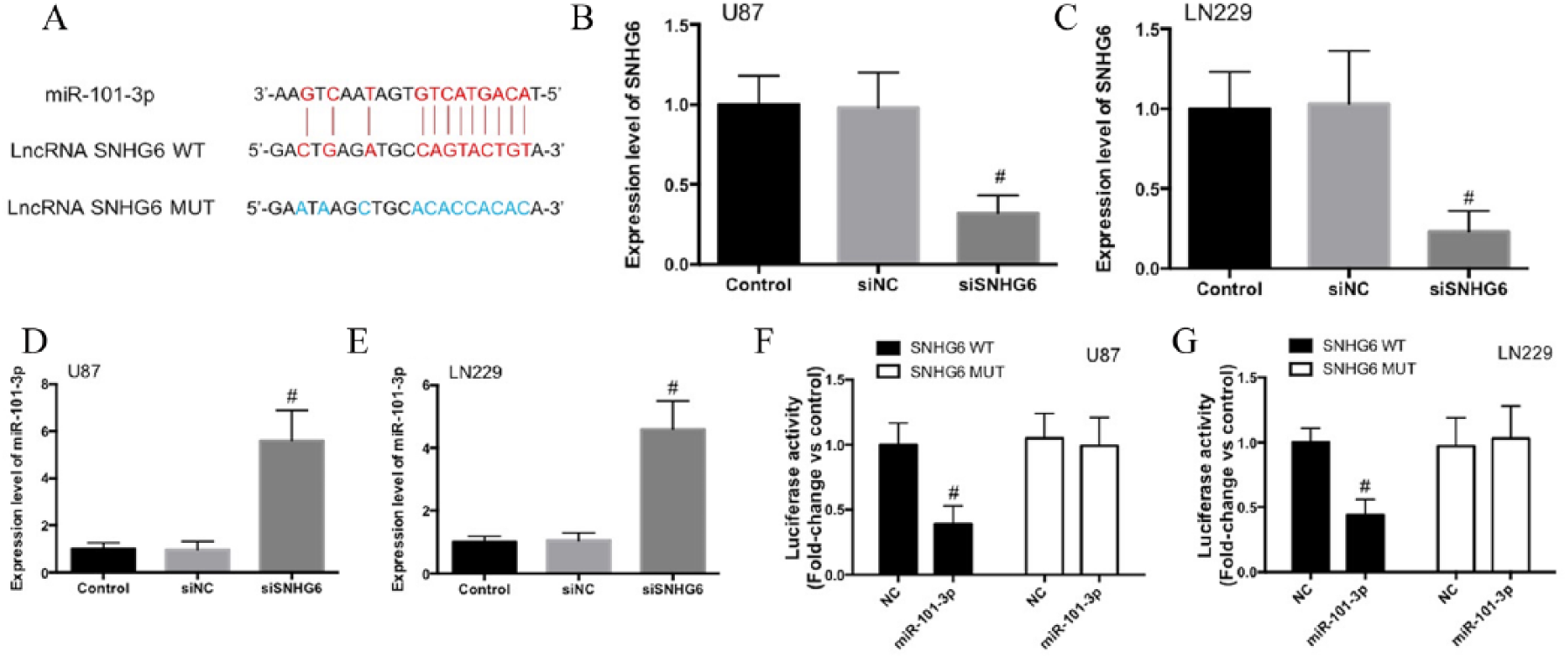
SNHG6 is a target of miR-101-3p. A) The predicted binding sites of miR-101-3p in SNHG6 (SNHG6-WT) and SNHG6 mutant (SNHG6-MUT) sequences. B) SNHG6 expression was analyzed by real-time PCR after knockdown of SNHG6 in U87 and LN229 cells. C) miR-101-3p expression was detected in U87 and LN229 cells after the transfection of siSNHG6. D) Luciferase activity was determined in HEK293T cells co-transfected with miR-101-3p mimic or NC-mimic and pGL3 luciferase reporters containing SNHG6-WT or SNHG6-MUT sequences.
SNHG6 promotes glioma malignancy through regulation of miR-101-3p
To examine the role of SNHG6 and miR-101-3p in the regulation of glioma malignancy, we transfected miR-101-3p mimic or anti-miR-101-3p to U87 and LN229 cells with stable knockdown of SNHG6, and the cell proliferation, migration, apoptosis, and expression of epithelial-mesenchymal transition (EMT) biomarkers were determined. Knockdown of SNHG6 remarkably decreased cell proliferation (Fig. 3A and B) and migration (Fig. 3C and D), enhanced E-cadherin expression, reduced vimentin expression (Fig. 3E and F), and increased cell apoptosis (Fig. 3G and H), indicating the reduction of glioma cell malignancy. The transfection of miR-101-3p mimic enhanced the decrease of cell proliferation (Fig. 3A and B) and migration (Fig. 3C and D), the increase of E-cadherin expression, the reduction of vimentin expression (Fig. 3E and F), and the increase of cell apoptosis (Fig. 3G and H) induced by knockdown of SNHG6. Anti-miR-101-3p reversed the the effects of si-SNHG6 on cell proliferation, migration, EMT, and apoptosis (Fig. 3). Taken together, these results indicated that SNHG6 regulates the tumorigenesis of glioma by modulating miR-101-3p expression.
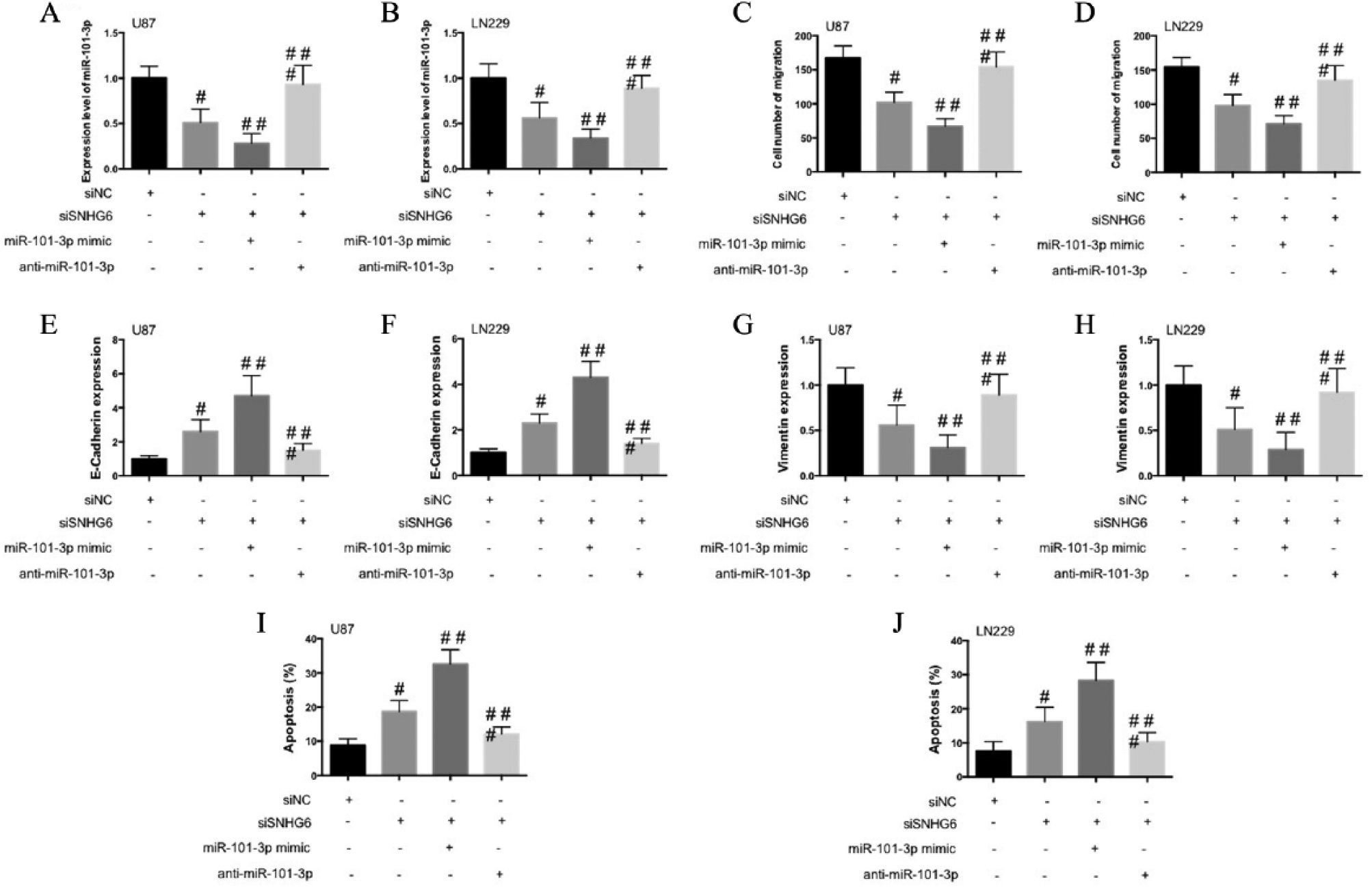
SNHG6 mediates glioma malignancy by negative regulation of miR-101-3p. A, B) U87 and LN229 cells were co-transfected with si-SNHG6 and miR-101-3p mimic or anti-miR-101-3p and cell viability was detected by CCK-8 analysis. C, D) The migration of transfected cells was determined by Transwell assay. E, F) mRNA expression of E-cadherin E, G) and vimentin F, H). I, J) Flow cytometry was used to explore the apoptosis of U87 and LN229 cells and percentage of apoptotic cells were shown.
SNHG6 promotes glioma tumor growth in xenograft mice
In the next step, we explored the role of SNHG6 in glioma in transplanted nude mice in vivo. Knockdown of SNHG6 remarkably reduced the increase of tumor volumes in xenograft mouse models (Fig. 4A). In tumor tissues, knockdown of SNHG6 increased the expression of miR-101-3p (Fig. 4B). Moreover, knockdown of SNHG6 increased the expression of E-Cadherin and reduced the expression of Vimentin in tumor tissues. These results indicated that knockdown of SNHG6 could suppress tumor growth in xenograft mouse models, which was related to the regulation of miR-101-3p and EMT.
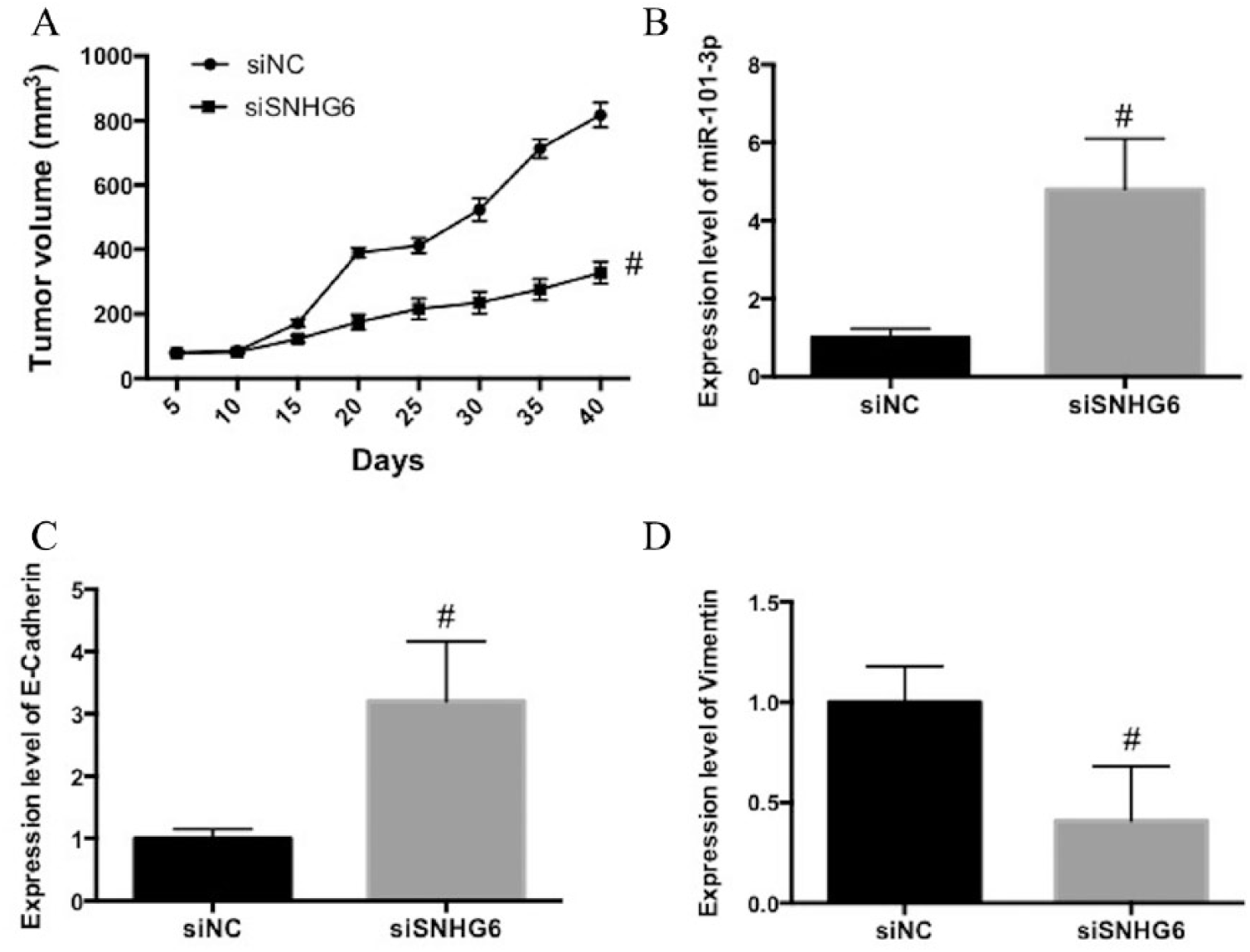
SNHG6 promotes glioma tumor growth in a xenograft mouse model. A) U87 cells stably transfected with si-SNHG6 and si-NC were subcutaneously injected into nude mice and tumor volumes were measured. B) mRNA expression of miR-101-3p. C) mRNA expression of E-cadherin. D) mRNA expression of Vimentin.
Discussion
snoRNAs have a battery of cellular functions, including chemical RNA modification, pre-RNA processing, and alternative splicing control. Increasing data have showed the function of snoRNAs in the regulation of cell proliferation and the development of tumorigenesis and metastasis (17). Among them, the role of lncRNA SNHG in modulating cancer cell proliferation and malignancy has received significant attention (11, 18-20). For example, lncRNA SNHG12 promotes cell growth and inhibits cell apoptosis in colorectal cancer cells (20). lncRNA SNHG16 has been found to contribute to breast cancer cell migration by competitively binding miR-98 with E2F5 (18). lncRNA SNHG20 predicts a poor prognosis for hepatocellular carcinoma (HCC) and promotes cell invasion by regulating EMT (19). In particular, lncRNA SNHG6 has been identified as being associated with the development of gastric cancer and HCC (14-16). In the present study, we explored the role of lncRNA SNHG6 in the regulation of glioma development and elucidated the possible mechanism.
We identified that lncRNA SNHG6 played an oncogenic role in the development of glioma. The expression level of lncRNA SNHG6 in glioma tissues or cells was higher than that in normal brain tissues or cells, and increased along with the malignant transition of glioma. Moreover, patients with higher lncRNA SNHG6 expression had shorter OS time than those with lower lncRNA SNHG6 expression. These results suggest that the upregulation of lncRNA SNHG6 is associated with the malignancy of glioma, and may serve as a prognostic marker for glioma patients.
The biological function of lncRNA is usually related to microRNA, and the interactions between lncRNA and microRNA plays important roles in various stages of tumor development (11, 19, 21). In this study, we identified that lncRNA SNHG6 had a direct binding site for miR-101-3p. Knockdown of SNHG6 increased miR-101-3p expression in glioma cells. The expression level of miR-101-3p in glioma tissues or cells was lower than that in normal brain tissues or cells, and decreased along with the malignant transition of glioma. Moreover, patients with lower miR-101-3p expression had shorter OS time than those with higher miR-101-3p expression. These results suggest that the downregulation of miR-101-3p is associated with the malignancy of glioma.
miR-101 has two precursor hairpin structures, which are transcribed from chromosome 1 and 9 separately in humans (22). Abnormal expression of miR-101 is associated with the progression of several cancers, including glioma (23), liver (24), prostate (25), and breast cancer (26). miR-101-3p is a dominant member of miR-101, and dysregulation of miR-101-3p is strongly related to the pathogenesis of some cancers. For instance, miR-101-3p may be a potentially reliable biomarker and could serve as a therapeutic target for esophageal squamous cell carcinoma caused by tobacco smoking (27). miR-101-3p is a key regulator of tumor metabolism in breast cancer via targeting adenosine 5′-monophosphate (AMP)-activated protein kinase (23). miR-101-3p suppresses cell proliferation and invasion, and enhances chemotherapeutic sensitivity in salivary gland adenoid cystic carcinoma by targeting proviral integrations of moloney virus 1 (28). miR-101-3p suppresses HOX transcript antisense RNA (HOTAIR)-induced proliferation and invasion through directly targeting the serum response factor in gastric carcinoma cells (29). Transfection of miR-101-3p mimic enhanced the decrease of cell proliferation and migration, increased the E-cadherin expression, reduced vimentin expression, and increased cell apoptosis induced by knockdown of SNHG6. Anti-miR-101-3p reversed the the effects of si-SNHG6 on cell proliferation, migration, EMT, and apoptosis. The results demonstrated that miR-101-3p is the target that is responsible for SNHG6-induced regulation of tumorigenesis of glioma. Moreover, miR-101 could inhibit proliferation, migration, and invasion of human glioblastoma by targeting sex-determining region Y (SRY)-box9 protein (30). Upregulated SNHG1 contributes to the progression of non-small cell lung cancer through the inhibition of miR-101-3p and the activation of Wnt/beta-catenin signaling pathway. These findings suggest that a more complex scenario, including other molecules, is involved in the development of glioma. Members of the SNHG family and downstream targets may function cooperatively, contributing to the progression of glioma. So-called targeted therapies have so far proved to be highly disappointing in glioblastoma (2017 phase III trial on rindopepimut), and therefore, multitargeted treatments—with these molecules as potential candidates—are probably the future in this highly elusive and variably mutated tumor.
In summary, SNHG6 plays an oncogenic role in glioma through targeting miR-101-3p. Based on our results and the findings of the role of SNHG6/miR-101-3p axis in hepatocellular carcinoma and gastric cancer, we suggest that the SNHG6-miR-101-3p pathway could be a common crucial regulator of tumor metabolism and progression. It has been demonstrated that SNHG6 could be a potential diagnostic and prognostic biomarker for glioma and other related tumors. In future studies, the level of SNHG6 in blood circulation should be examined to evaluate the diagnostic and prognostic value for tumor diagnosis and treatment. Overall, our study provides novel insights in the functions of lncRNA SNHG6 and miR-101-3p in the tumorigenesis of glioma.
Footnotes
Author contributions
Qiang Meng, Bao-Ying Yang, and Bei Liu are co-authors.
Disclosures
Financial support: No grants or funding have been received for this study.
Conflict of interest: None.
