Abstract
Objectives
Acute liver failure (ALF) is a rare life-threatening condition that leads to rapid deterioration of liver function. Although global awareness of ALF consequences is increasing, the precise molecular mechanisms associated with its rapid progression remain unclear. In the present study, we established a rat model of ALF using Lipopolysaccharide (LPS)/D-galactosamine (D-Gal) and explored the potential molecular mechanism of ALF.
Methods
Multiplexed isobaric tandem mass tag labelling combined with liquid chromatography-mass spectrometry was used to thoroughly screen for differentially expressed proteins in liver samples from LPS/D-Gal-induced ALF rat models.
Results
We identified 175 proteins, whose expression was altered by at least 1.5-fold, between the liver samples of ALF and control groups. Of these, 14 dysregulated proteins mainly participated in the regulation of neutrophil extracellular trap (NET) formation. Furthermore, rats with severe ALF showed elevated levels of cathelicidin antimicrobial peptide, myeloperoxidase, and fibrinogen gamma chain, consistent with NET formation. These findings suggest that the NET formation pathway may have contributed to the regulation of the clinical features and progression of liver injury in ALF rats.
Conclusion
To our knowledge, this study is the first to report a global differential protein expression profile of liver samples from rats with LPS/D-Gal-induced ALF. Our TMT-based quantitative proteomic analysis revealed molecular differences involved in NET formation between the ALF and control rat groups, potential therapeutic targets for ALF treatment as well as fundamental information for further detailed experimental research.
Keywords
Introduction
Acute liver failure (ALF), also known as fulminant hepatic failure, is a rare but severe condition with high morbidity and mortality risks, and is usually associated with rapid loss of hepatic function in patients without underlying chronic liver disease.1,2 Clinically, owing to its involvement in rapid and progressive deterioration of liver function, ALF affects the physiological status of almost every organ system by causing massive hepatocellular necrosis or infiltration, leading to encephalopathy and multi-organ failure.1–3 Although improvements in medical management have resulted in a substantially increased survival rate in patients with ALF, thus far, the mortality rate remains at approximately 40% without liver transplantation.4–6
Acute liver failure is attributed to various aetiologies and can occur because of hepatic injury due to drugs and poison, hepatitis virus, hepatic ischaemia, cancer, or other uncommon sources, with drug-induced liver injury being the leading cause of ALF worldwide. Accumulated drugs are metabolized into toxic agents in the body, leading to oxidative stress, inflammation, and apoptosis, which induce necrosis in hepatocytes and ultimately cause liver damage.2,5–7 Rat ALF models, induced by the combined application of lipopolysaccharide (LPS) and D-galactosamine (D-Gal) closely resemble clinical ALF, and therefore are one of the most commonly used experimental models to study ALF.5,8–10 Through drug-target interactions, LPS/D-Gal stimulation regulates liver macromolecular synthesis by modifying the liver transcription and translation machinery, and allows targets to meet the requirements of drug challenge.5,10 Therefore, liver tissue proteins involved in ALF evolution play essential roles in the progress of acute liver injury across a spectrum that includes apoptosis, autophagic cell death, and necrosis.11,12 The induction of LPS/D-Gal is associated with the Toll-like receptors, a family of pattern recognition receptors on Kupffer cells, which induce the transcription and expression of proinflammatory cytokines such as tumour necrosis factor-α (TNF-α), interleukin (IL)-1β, IL-6 and cyclooxygenase-2 (COX-2) that are associated with the mitogen-activated protein kinase (MAPK), nuclear factor-κB (NF-κB), and NOD-like receptor pyrin domain-containing protein 3 (NLRP3) signalling pathways.5,13,14 Previous research also indicates that the dysregulated proteins in the polyinosinic: polycytidylic acid [poly(I:C)]/D-Gal-induced ALF mouse model groups were primarily associated with oxidative phosphorylation, those in the LPS/D-Gal ALF mouse model groups were further enriched in metabolic pathways, and those in the CpG DNA/D-Gal ALF mouse model groups were involved in ribosomal activities. 15 These results indicate considerable differences in the molecular pathogenesis of ALF-related diseases between the rats treated with LPS/D-Gal and controls; however, the precise molecular mechanism remains elusive.
Comparative proteomic strategies involving tandem mass tags (TMTs) allow us to investigate the host factors involved in drug infection and can provide insight into alterations in signalling pathways, thus deepening our understanding of ALF pathogenesis.15–17 Therefore, in the present study, we established a rat model of ALF using LPS/D-Gal and examined the differential expression of proteins in pooled liver tissues of these rats using TMT-based liquid chromatography-tandem mass spectrometry (LC-MS/MS). Our results provide new insights into the potential molecular mechanisms underlying ALF and may provide fundamental information for future in-depth studies.
Materials and methods
Generation of the animal model and sample collection
Adult male Sprague –Dawley rats (6 weeks old, weighting 200 –220 g) were provided by the Laboratory Animal Center, Jiaxing University (Jiaxing, China). The rats were housed under normal laboratory conditions (23 ± 3°C, 12 h light/dark cycle) with free access to a standard pellet diet and water. Laboratory animal handling and experimental procedures were performed in accordance with the Animal Research: Reporting of In Vivo Experiments guidelines. The Institutional Animal Care and Use Committee of the Affiliated Hospital of Jiaxing University approved all experimental protocols, including procedures, sampling and animal care (LS2019-047). All efforts were made to minimise animal suffering.
The rat or mouse model of ALF induced within 24 h under these conditions, which, to some extent, resemble ALF in humans, and most symptoms of the tissue damage can be observed at least 6 h after LPS/D-Gal administration.5,7,8,18 After 1 week of acclimatisation, 72 rats were randomly divided into four groups representing LPS/D-Gal administration for different time periods: the control (n = 18), 6 h ALF (n = 18), 12 h ALF (n = 18) and 24 h ALF groups (n = 18), according to the International Guiding Principles for Biomedical Research Involving Animals. In each group, 18 rats were sub-grouped into three biological replicates to obtain repeatable accuracy. For the model groups, a combined intraperitoneal injection of 500 μL saline containing 8 μg LPS (Sigma-Aldrich, St. Louis, MO, USA) and 0.8 g/kg D-Gal (Sigma-Aldrich) as the optimal dose was used to induce ALF as described previously. 18 In the control group, rats were injected with the same volume of pyrogen-free 0.9% sodium chloride instead of LPS/D-Gal. Portal vein blood and liver tissue samples were collected for biochemical measurements and haematoxylin-eosin (HE) staining respectively. The overall technical route for this study is shown in Figure S1.
Sample preparation and protein digestion
Liver samples from the control and 12 h ALF group were divided into two groups: control (Group A, n = 18) and ALF rats (Group B, n = 18) according to standard operating procedures to minimise preanalytical variation. The total protein of liver tissues was extracted for each group for each group by mixing fresh tissues of equal weight from six individual samples of equal weights. To minimize individual differences in the rats, we used three repeated protein extracts per group.
Tissue preparation and protein digestion were performed as described previously.16,19,20 In brief, SDT buffer (4% SDS, 100 mM Tris-HCl, 1 mM DTT, pH7.6) was used for sample lysis and protein extraction. Protein digestion was performed using trypsin, according to filter-aided sample preparation procedure. For trypsin digestion, 200 μg of protein from each sample was sequentially digested with 4 μg of trypsin (Promega, Madison, WI, USA) in 40 μL of 25 mM NH4HCO3 buffer at 37°C overnight. Thereafter, the digested peptides were desalted in a C18 cartridge column (Sigma-Aldrich) and vacuum-dried for the remaining experiments.
TMT labelling and LC-MS/MS analysis
A 100 μg peptide mixture from each sample was labelled with the TMT 6-plexTM Label Reagent set per the manufacturer’s instructions for the 6-plex TMT Kit (Thermo Fisher Scientific, Waltham, MA, USA).16,19 TMTs 126, 127, 128, 129, 130 and 131 were used to label the control rat group samples (A) and the 12 h ALF rat group samples (B). Labelled peptides were fractionated using the High pH Reversed-Phase Peptide Fractionation Kit (Thermo Fisher Scientific). Thereafter, the collected fractions were desalted using C18 cartridges (Empore™ SPE Cartridges C18 (standard density), bed I.D. 7 mm, volume 3 mL, Sigma-Aldrich) and concentrated using vacuum centrifugation.
The obtained peptide fractions were subjected to nano-spray ionization (NSI) source followed by MS/MS on a Q Exactive mass spectrometer (Thermo Fisher Scientific) coupled with an Easy nLC 1200 chromatography system (Thermo Fisher Scientific). 16 The parameters used were as follows: survey scan range, 300–1800 m/z; automatic gain control, 3 × 106; maximum injection time, 10 ms; dynamic exclusion, 30 s; and normalised collision energy, 30 eV.
MS data and bioinformatic analysis
Following peptide separation, MS raw data were searched using the MASCOT engine (Matrix Science, London, UK, version 2.2) of the Proteome Discoverer 1.4 software against a human database provided by the Universal Protein Resource (http://www.uniprot.org/uniprot, released at 2021-04-27, with 32,230 entries) and a decoy database.16,19 For peptide identification, the following search parameters were used: max missed cleavages, 2; fixed modification, carbamidomethyl (C) and TMT 6 (N-term); variable modifications, oxidation (M) and TMT 6 (Y); peptide mass tolerance, ± 20 ppm; fragment mass tolerance, 0.1 Da. Additionally, all significant quantitative data for protein identification were reported with a 99% confidence level as determined by a false discovery rate threshold of 1.0% (Figure S2).
A protein was considered differentially expressed if the fold change was >1.5 or <0.67 with a p-value < .05 as determined using a paired t-test. Here, we used a volcano plot to display the differentially expressed proteins, where the x-axis represents log2 fold change and y-axis, the negative log10 of the p-value calculated from the two-tailed t-test. 16 Hierarchical clustering analysis of the differentially expressed proteins was done using the Java Tree View software (http://jtreeview.sourceforge.net).
The Gene Ontology (GO) annotation and pathway enrichment analysis of all the differentially expressed proteins were performed using the Blast 2 GO software (http://www.blast2go.com/b2ghome) based on functional annotations for biological processes, molecular functions, and cellular components. The protein pathways were retrieved using the Kyoto Encyclopedia of Genes and Genomes (KEGG) database (http://geneontology.org/) with the KEGG Automatic Annotation Server (KAAS, https://www.genome.jp/tools/kaas/).
Validation of the selected dysregulated proteins by western blotting (WB)
To verify the results of the proteomic study, proteins exhibiting substantially different levels in the two groups were subjected to WB as previously described.16,19 Briefly, 30 μg of protein from each sample was subjected to sodium dodecyl sulphate-polyacrylamide gel electrophoresis (SDS-PAGE) and subsequently transferred to nitrocellulose membranes (Millipore, Bradford, MA, USA). Subsequently, the membranes were blocked in phosphate-buffered saline with Tween 20 (PBST) buffer containing 5% bovine serum albumin (BSA), for 2 h at 20°C, and probed with the cathelicidin antimicrobial peptide (CAMP) (Affinity Biosciences), myeloperoxidase (MPO) (Affinity Biosciences), fibrinogen gamma chain (FGG) (Affinity Biosciences), citrullination of histone H3 (Cit-H3) (Abcam), and β-actin (Affinity Biosciences) antibodies overnight at 4°C. Following three washes with PBST, the membranes were incubated with appropriate horseradish peroxidase (HRP)-conjugated secondary antibodies (1:5000 dilution, TransGen Biotech, Beijing, China) at 37°C for 1 h. After washing in TBST buffer, the protein expression levels were determined by enhanced chemiluminescence and visualized using autoradiography.
Statistical analysis
Quantitative data were analysed using Student’s t-test between the two groups and expressed as mean ± standard deviation (SD). Results with p-values less than 0.05 were considered statistically significant. The analyses were performed using the Statistical Program for Social Sciences (SPSS) software (version 22.0; SPSS Inc., Chicago, IL, USA).
Results
Generation of the LPS/D-Gal-induced ALF rat model
The liver injury was determined by examining hepatic pathology and serum biochemical parameters in rats after the administration of a single dose of LPS/D-Gal (Figure 1 and Table S1). As shown in Figure 1(a), the LPS/D-Gal treatment led to obvious morphological changes in the liver, including hepatocyte necrosis, hepatic architecture destruction, inflammatory cell infiltration, and bleeding spot dissemination. The HE staining of the liver samples showed areas of widespread hepatocyte necrosis and extensive infiltration of inflammatory cells around the central veins in the 12 h ALF group compared to the control group. In contrast, the liver tissue sections from the 6 and 24 h ALF groups only exhibited some isolated areas of necrotic tissue, indicating that hepatocyte necrosis scores were significantly higher in the 12 h ALF group than in the 6 and 24 h ALF groups (Figure 1(b)). Additionally, some serum Indicators of the extent of liver damage showed marked differences among the four groups; alanine transaminase (ALT), aspartate transaminase (AST) and total bilirubin (Tbil) levels were significantly higher in the 12 h ALF group compared to the 6 h and 24 h ALF groups (Figure1(c) and Table S1). Therefore, the 12 h ALF rat model presents a reliable alternative to other available ALF animal models for studying the molecular mechanism of ALF and evaluating the potential of a novel therapy for ALF. Effect of lipopolysaccharide (LPS)/D-galactosamine (D-Gal) treatment on liver hepatic pathology. (a) Photograph of liver tissue, (b) Representative photomicrographs of haematoxylin-eosin (H&E) staining (100-fold), and (c) Serum levels of alanine transaminase (ALT), aspartate transaminase (AST), and total bilirubin (Tbil) in rats with LPS/D-Gal administration for different periods, respectively.
Differentially expressed proteins associated with ALF
Using the MASCOT engine, we identified 32,230 unique proteins across 36,959 peptides in the TMT6-plex labelling replicates. Among these, 4911 proteins were quantified (Table S2). To identify differentially expressed proteins, relative protein expression values were compared between the control and 12 h ALF rat groups. The distribution of molecular weight, peptide count, peptide length, and protein sequence coverage clearly showed that the overall proteome datasets of the control and 12 h ALF rat groups had no significant variability (Figure S3). The raw mass spectrometry proteomics data were deposited in the Integrated Proteome Resources (iProX) database and can be accessed at http://www.iprox.org/index with the dataset identifier PXD027868. Using hierarchical clustering analysis, 175 proteins with a mean expression fold change ≥ ± 1.5 (log2 = 0.58) were classified as differentially expressed in the liver of the 12 h ALF rat group compared to the liver of the control (Groups B vs. A) (Figure 2(a), Table S2). When the ratio of these 175 proteins was plotted on a heatmap, 142 and 33 proteins were found to be upregulated and downregulated, respectively, between the liver samples from the 12 h ALF group and the control, and these two sets of proteins were separated into distinct clusters (Figure 2(b), Figure S4). Bioinformatics analysis of differentially expressed proteins between the liver samples of the control and the 12 h acute liver failure (ALF) rat groups. (a) Volcano plot representing protein abundance changes. A total of 175 dysregulated proteins with fold change ≥±1.5 and p-values < .05 were identified. (b) Hierarchical clustering analysis of the 175 dysregulated proteins. (c) Subcelllar localization of 175 dysregulated proteins. (d) Gene Ontology (GO) analysis of 175 dysregulated proteins. The abscissa represents enriched GO function classifications, which are divided into three major categories: biological process (BP), molecular function (MF) and cellular component (CC).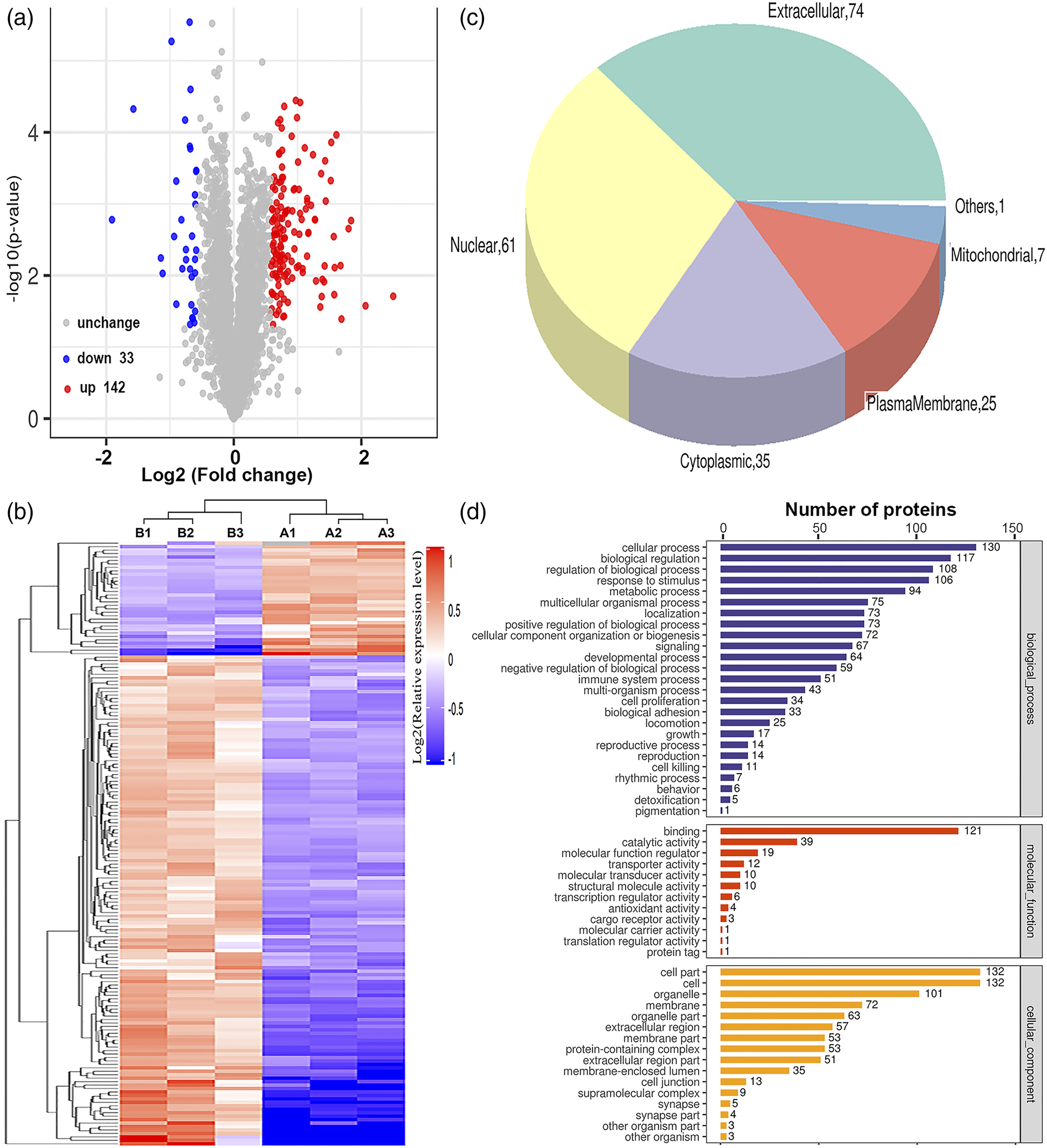
GO annotation was applied to classify the identified proteins according to their subcellular localisations with at least one term assigned to each protein. More than 42% of proteins were annotated as extracellular, with the other three main protein categories being nuclear (61), cytoplasmic (35) and plasma membrane (Figure 2(c)). Although extracellular, nuclear, cytoplasmic, and plasma membrane proteins were the most highly represented categories in the extracts, mitochondrial proteins were also readily identified, indicating that the protein extraction procedure was not strongly biased to a few cell compartments. As shown in Figure 2(d), the three most frequently occurring categories of biological process terms in this study were cellular process (n = 130), biological regulation (n = 117), and regulation of biological process (n = 108). We further investigated molecular functions involving these proteins using GO enrichment analysis and found that they were primarily associated with the GO terms binding (n = 121), catalytic activity (n = 39) and molecular function regulator (n = 19). Therefore, our findings suggest that many biological processes and molecular functions are extensively affected in rats with ALF, and they likely exhibit different molecular characteristics compared to controls.
KEGG pathway analysis of differentially expressed proteins
In order to further understand the cellular and biological processes affected, we turned to the protein pathways that involved the differentially expressed proteins. KEGG pathway-based enrichment analysis of the differentially expressed proteins was performed to further analyse the roles of protein expression alterations by the LPS/D-Gal treatment. The results showed that specific signalling pathways were involved in the molecular differences in host macromolecular synthesis between control and LPS/D-Gal-treated rats. As shown in Figure 3(a), dysregulated proteins mainly participate in the regulation of neutrophil extracellular trap (NET) formation (14 dysregulated proteins) (Figure 3(c) and Table 1), complement and coagulation cascades (13 dysregulated proteins), phagosomes (11 dysregulated proteins), NOD-like receptor signalling pathway (seven dysregulated proteins), focal adhesion (6 dysregulated proteins) and ECM-receptor interaction (five dysregulated proteins). Of these categories, the NET formation pathway was the top modulated signalling pathway for the enrichment of dysregulated proteins (Figure 3(b)). This finding suggests that the NET formation pathway may have contributed to the regulation of clinical features and progression of liver injury during the LPS/D-Gal treatment in ALF rats. Key signalling pathways involved in the lipopolysaccharide (LPS)/D-galactosamine (D-Gal)-induced acute liver failure (ALF) rat liver samples. (a) Kyoto Encyclopaedia of Genes and Genomes (KEGG) pathway-based enrichment analysis of 175 dysregulated proteins. (b) Neutrophil extracellular trap (NET) formation obtained from the KEGG pathway-based enrichment analysis. (c) Hierarchical clustering of 14 dysregulated proteins involved in NET formation. List of the typically differentially expressed proteins of liver samples involved in NET formation.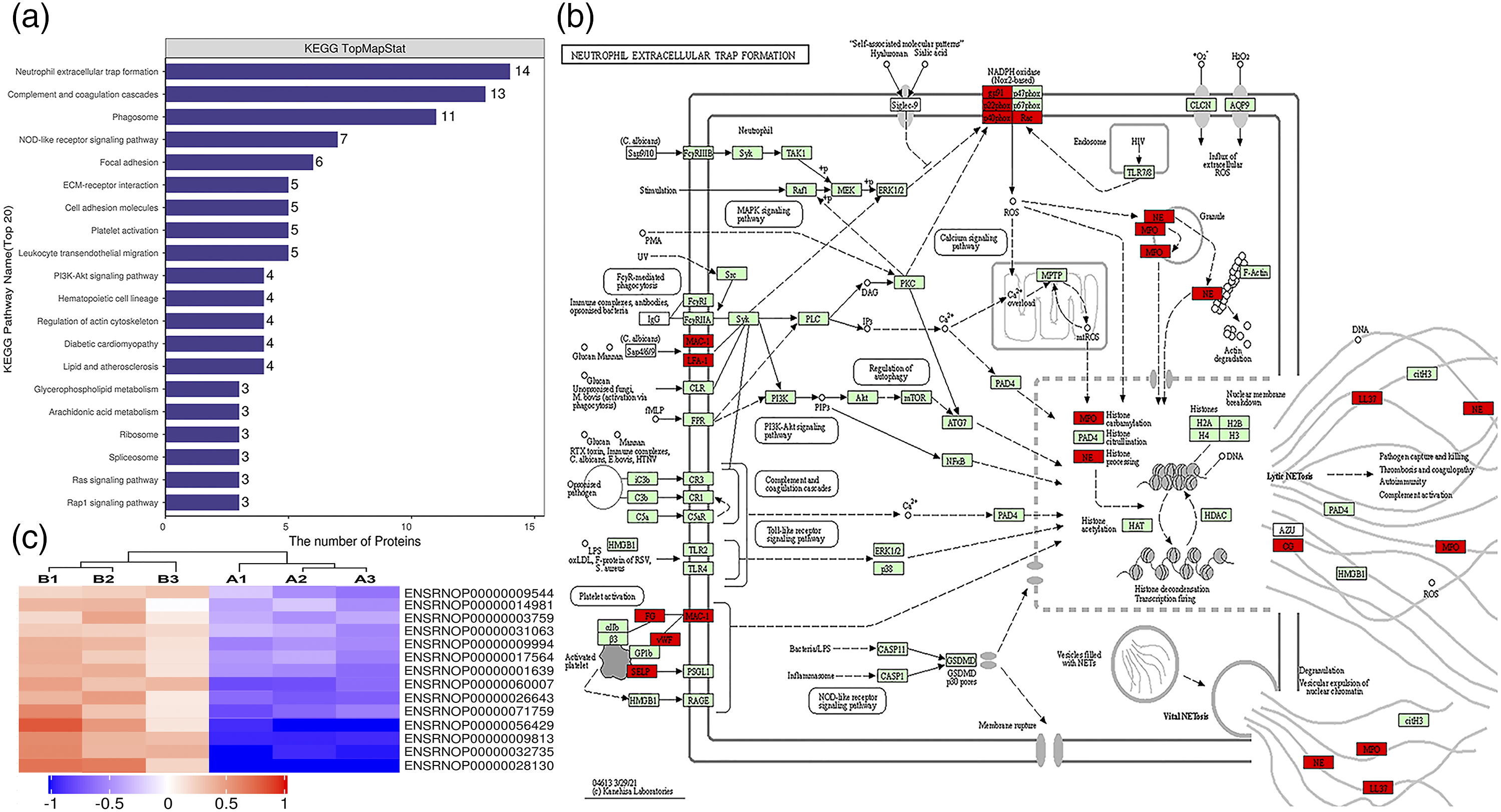

Verification of the selected differentially expressed proteins associated with NET formation
Based on the key modulated signalling pathways (Figure 3(b)) and hierarchical clustering analysis results (Figure 3(c)), CAMP (3.49-fold), MPO (2.02-fold) and FGG (2.70-fold) proteins were found to be differentially expressed between the 12 h ALF and control group rats. Expression changes in CAMP, MPO and FGG were further verified at the protein level by WB analysis after sample pooling. As shown in Figure 4(a), the protein levels of CAMP and MPO were significantly upregulated by 3.96-fold and 1.87-fold (n = 36 rats), respectively, while the protein expression of FGG was upregulated 2.94-fold (n = 36 rats) in 12 h ALF rat liver samples compared to the control samples. These results were consistent with the LC-MS/MS results. As the results of all analyses are comparable and consistent with each other, the three proteins, CAMP, MPO and FGG, might play important roles in inducing ALF in LPS/D-Gal-treated rats by regulating NET formation. Lipopolysaccharide (LPS)/D-galactosamine (D-Gal)-induced acute liver failure (ALF) in rats involved in neutrophil extracellular trap (NET) formation. (a) Validation of selected differentially expressed proteins of cathelicidin antimicrobial peptide (CAMP), myeloperoxidase (MPO), and fibrinogen gamma chain (FGG) in the liver samples ofrats after LPS/D-Gal administration for different time periods using western blotting analysis in the validation cohort. (b) Serum MPO-DNA levels, (c) citrullination of histone H3 (Cit-H3) levels, (d) tumour necrosis factor-α (TNF-α) levels, and (e) interleukin-6 (IL-6) levels were measured in rats with LPS/D-Gal administration for different time periods, respectively.
NET formation was significantly increased in LPS/D-Gal-induced ALF rats
To determine whether LPS/D-Gal contributes to NET formation, we measured the levels of serum MPO-DNA complexes, a specific marker of NET formation, in rats administered with LPS/D-Gal for different periods. Serum levels of circulating MPO-DNA complexes were significantly higher in 12 h ALF rats than in control rats (n = 36 rats, 5.98-fold, p < .01) (Figure 4(b)). Next, we performed WB to evaluate the expression of Cit-H3, a specific marker of NET formation. As shown in Figure 4(c), the liver tissue sample from the 12 h ALF rat group had significantly higher levels of Cit-H3 than the corresponding control samples, 6 h ALF and 24 h rat groups. Additionally, in this LPS/D-Gal-induced ALF rat model, we measured the serum levels of pro-inflammatory cytokines including TNF-α and IL-6, which play important roles in the pathogenesis of liver injury. As shown in Figures 4(d) and (e), the serum TNF-α and IL-6 levels in the 12 h ALF rats were higher than those in the control rats after the administration of LPS/D-Gal, suggesting that LPS/D-Gal can further drive NET formation and the concurrent tissue-toxic inflammatory response. These results are in good agreement with the KEGG pathway analysis results and the selected differentially expressed proteins CAMP, MPO and FGG.
Discussion
Although the overall incidence of ALF is gradually decreasing, it continues to be a major worldwide health problem because of its association with high morbidity and mortality risks.1–3 Patients with drug-induced ALF, account for a significant proportion of fatal hepatic injury cases, and previous studies confirm that ALF is associated with an increased risk of developing liver dysfunction and even hepatocellular carcinoma, compared to healthy individuals.11,15,21 While previous research focused on the disparities between ALF patients and healthy individuals, not much is known about the potential mechanisms and alterations in specific signalling pathways, in the context of drug-induced ALF. To our knowledge, the liver proteomic data presented in this study are the first to report molecular differences between rats with 12 h LPS/D-Gal-induced ALF and controls at the proteome level. These data provide not only confirmation of earlier studies, demonstrating that the ALF model induced by combined administration of LPS and D-Gal could be used to examine the mechanism of ALF, but also new information, which will facilitate further detailed investigations.
In the current study, we used TMT-labelling and LC-MS/MS to quantify the dynamic changes in the global liver proteome of pooled liver samples from two groups—control rat group and 12 h LPS/D-Gal-induced ALF rat group. In total, 36,959 peptides and 4950 proteins were identified, of which 4911 were quantified. Based on the identification criteria for dysregulated proteins (fold change ≥ ± 1.5, p < .05), 175 differentially expressed proteins were identified in the liver tissue samples of the 12 h ALF rat group compared to control rat group (Groups B vs. A). Many of them were associated with the regulation of NET formation, complement and coagulation cascades, and phagosome signalling pathways and mainly reflect the immune response against DNA, histones, and other intracellular proteins associated with liver injury.22–24
Interestingly, a significant outcome of this analysis was that many dysregulated proteins related to the NET formation signalling pathway may be used as novel candidate targets for ALF treatment. NETs, which are composed of extracellular strands of decondensed DNA together with various proteins, such as histones, neutrophil elastase, MPO, cathepsin G and perhaps as many as 30 other molecules, are associated with various human pathologies, in both infectious and non-infectious diseases, such as preeclampsia, rheumatoid arthritis, thrombosis, lung fibrosis, and cancer metastasis.25,26 The process of NET formation is called NETosis, and involves the extrusion of nuclear chromatin and cytoplasmic granule-derived proteins into the extracellular space followed by necrosis resulting in nuclear and plasma membrane rupture.27–29 NET formation is a regulated form of neutrophil cell death involved in several pathophysiological processes including host defence against pathogens, ischemia-reperfusion injury, autoimmunity, and tumour cell metastasis, which can be activated by multiple stimuli, including pathogens, cytokines and chemokines, and other physiological factors.25,28 For LPS/D-Gal treatment, activated NETosis is not merely a process reactive to inflammation and immunity but also critical determinant of the magnitude of liver injury and liver disease pathogenesis.26,27,30 Thus, our findings support the hypothesis that the degree of NETosis signalling pathway activation or inhibition determines the effects of NETosis on the hepatic manifestation of LPS/D-Gal-induced ALF in rats. Therefore, the NETosis signalling pathway is part of the immune defence mechanism and plays key role in liver injury associated with LPS/D-Gal-induced ALF.
Furthermore, our analysis revealed that the proteins CAMP, MPO and FGG were the most upregulated ones in the liver samples from the 12 h LPS/D-Gal-induced ALF rat group compared to the control. For example, CAMP (LL-37 in humans, CRAMP in mice/rats), a family of host-derived antibacterial polypeptides, also designated as host defence peptides, are natural broad-spectrum antibiotics that play crucial roles in regulating innate immune response, angiogenesis, and wound healing by exhibiting antimicrobial activities against both gram-negative and -positive bacteria, viruses, and fungi.31–34 Additionally, MPO, a heme peroxidase cyclooxygenase enzyme present in neutrophils and monocytes, plays a crucial role in host defence and inflammatory response, by catalysing the formation of reactive oxygen species involved in microbial killing.35–37 Consistent with the KEGG pathway analysis, which suggested the NETosis signalling pathway involved in inflammatory and immune disorders is to be one of the most important pathogenic factors of ALF, several proteins associated with those functions including CAMP, MPO and FGG were found dysregulated in our data. The NETosis signalling pathway plays key role in the development of LPS/D-Gal-induced ALF by regulating the expression of several proteins such as CAMP, MPO and FGG. In the current study, we observed that CAMP, MPO and FGG upregulation may increase the activation of the NETosis signalling pathway in the liver samples of LPS/D-Gal-induced ALF rats compared to those in control rats. However, the molecular mechanism underlying this remains unclear and requires further investigation.
Overall, we profiled liver proteins from the control and 12 h ALF rat groups and identified the NET formation signalling pathway for the prevention and treatment of ALF diseases by regulating potential protein targets such as CAMP, MPO and FGG. However, the main limitations of the present study were the small sample sizes and calculation and justification of the sample sizes selected in this study. Additionally, the TMT-based quantitative proteomic analysis was performed on relatively fewer liver samples and only differential protein expression was determined. It is necessary to continue investigations on the validity and elaboration of key dysregulated proteins through studies using biochemical analysis with larger sample sizes.
Conclusion
In this study, we used a TMT-based quantitative proteomic analysis approach and foundat least 175 proteins with altered expression. These altered proteins contribute to LPS/D-Gal-induced ALF, probably by regulating the activation of a variety of signalling pathways, including NET formation. In addition, we confirmed increased NET formation in rats with LPS/D-Gal-induced ALF, as indicated by increased specific marker levels and higher serum levels of pro-inflammatory cytokines including TNF-α and IL-6. Although quantitative proteomic analysis remains mostly descriptive, our findings provide mechanistic insights into LPS/D-Gal-induced ALF and suggest the potential application of dysregulated proteins associated with NET formation such as CAMP, MPO and FGG as candidate targets for ALF treatment.
Supplemental Material
Supplemental Material - Liver proteomic analysis reveals acute liver failure induced by lipopolysaccharide/D-galactosamine in rats involved in neutrophil extracellular trap formation
Supplemental Material for Liver proteomic analysis reveals acute liver failure induced by lipopolysaccharide/D-galactosamine in rats involved in neutrophil extracellular trap formation by Keyin Wang, Zhuolin Zou, Ting Zou, Dahai Wei and Min Deng in European Journal of Inflammation
Footnotes
Acknowledgements
Technical assistance for mass spectrometry analysis was supported by Applied Protein Technology (Shanghai) Co., Ltd.
Authors’ contributions
Dahai Wei and Min Deng made substantial contributions to conceive and design of the study. Keyin Wang performed most of the experiments. Zhuolin Zou and Ting Zou collected the samples and analyzed the data. Keyin Wang and Dahai Wei drafted the manuscripts. All authors read and approved the final manuscript.
Availability of data and materials
All data generated during this study are included in this published article. Total list of identified proteins has been uploaded as additional files. The mass spectrometry proteomics data have been deposited in the ProteomeXchange Consortium via the PRIDE partner repository with the dataset identifier PXD027868.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Scientific Foundation of Jiaxing City under Grant No. 2021AD30154, Zhejiang Provincial Natural Science Foundation of China under Grant No. LY20C010004, Zhejiang Provincial Medical Scientific Research Foundation of China under Grant No. 2021KY1108, 2019 Jiaxing Key Discipiline of Medcine-Lemology (Supporting Subject, Grant No. 2019-zc-02), and the Construction Project of Key Laboratory of Infectious Disease Involved in Viruses in Jiaxing City.
Ethics approval
Ethical approval for this study was obtained from * Affiliated Hospital of Jiaxing University OF ETHICS COMMITTEE OR INSTITUTIONAL REVIEW BOARD (LS2019-047)*.
Animal welfare
The present study followed international, national, and/or institutional guidelines for humane animal treatment and complied with relevant legislation.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
