Abstract
Erectile dysfunction (ED) is a multifactorial disorder that significantly impacts men's physical and mental health, as well as their interpersonal relationships, and traditional treatment options for this condition still face many challenges and limitations. This study aimed to identify key genetic factors associated with ED risk through Mendelian randomization analysis by integrating data from expression quantitative trait loci and protein quantitative trait loci across multiple cohorts. We also evaluated the roles of metabolic pathways using data from 1,400 plasma metabolites. Single-cell RNA sequencing (ScRNA-Seq) was used to analyze gene expression patterns of ED-related genes in various cell types, while molecular docking was employed to identify potential drug targets. Our findings indicate that DKK3 plays a protective role (OR = 0.8555, p = .0087), while SLAMF6 is associated with increased ED risk (OR = 1.2613, p = .0433). Metabolites such as piperine and choline phosphate mediate ED onset. ScRNA-Seq reveals reduced DKK3 expression in endothelial and smooth muscle cells and increased SLAMF6 expression in T cells, highlighting the roles of vascular homeostasis imbalance and immune dysregulation in ED pathogenesis. Molecular docking screens four small molecules, including icariin, luteolin, Danshenol A, and Danshenxinkun A as potential therapeutic agents. This study identified DKK3 and SLAMF6 as novel therapeutic targets for ED, provided a foundation for precision medicine based on vascular-immune regulation, and underscored the need for further mechanistic studies and clinical validation.
Introduction
Erectile dysfunction (ED) is a complex condition in men caused by a combination of psychological, neurological, endocrine, and vascular factors (Salonia et al., 2021; Xiong et al., 2024). The International Society for Sexual Medicine defines ED as the inability to achieve or maintain an adequate erection for satisfactory sexual intercourse No Authors (1993). The prevalence of ED has notably risen in conjunction with the aging global population and the increasing incidence of chronic diseases (Chen, 2025). Studies indicate that individuals with ED are at an elevated risk of experiencing depression and anxiety (Liu et al., 2018). Delays in seeking medical intervention, often due to social and psychological pressures, exacerbate both physical and mental health deterioration. Additionally, ED can contribute to strained interpersonal relationships and familial conflicts, thereby imposing a significant socioeconomic burden (Boddi et al., 2015; Elterman et al., 2021).
Treatments for ED primarily include phosphodiesterase type 5 inhibitors, vacuum erection devices, penile prosthesis surgery, hormone replacement therapy, and emerging therapies such as stem cell treatments and shockwave therapy (Hsu et al., 2025; Yafi et al., 2016). The most widely used method is oral phosphodiesterase type 5 inhibitors; however, this is a lifelong treatment, and its efficacy is limited by endothelial dysfunction, especially in patients with diabetes or severe atherosclerosis, where the response rate is less than 30% (Wang et al., 2021). New therapies such as low-intensity shockwave therapy, platelet-rich plasma injections, stem cell treatments, and botulinum toxin still lack high-quality trials and long-term data (Capogrosso et al., 2025). Additionally, vacuum devices and surgeries pose issues such as pain, infection risks, low compliance, and high health care costs (Bettocchi et al., 2021; Preto et al., 2024). More importantly, while current treatments alleviate symptoms, significant individual variability in efficacy exists, and they fail to address the underlying pathological mechanisms of ED, such as endothelial dysfunction, smooth muscle abnormalities, and immune microenvironment imbalance. It is crucial to uncover the molecular mechanisms of ED and develop novel targeted therapies for individualized clinical treatment.
Bioinformatics approaches such as machine learning, network pharmacology, and single-cell transcriptome integration have accelerated the discovery of therapeutic targets for diseases (Maciejewska-Turska et al., 2025; Saparov & Zech, 2025). Some algorithms overly rely on the statistical associations of differentially expressed genes without validating them through causal inference methods like Mendelian Randomization (MR; Davies et al., 2018). Isolated analyses of genomic or metabolomic data often overlook the three-dimensional regulatory networks among genes, metabolites, and proteins, as well as the effects of cellular heterogeneity and microenvironmental interactions. Key genes lack clear drug-binding domains or regulatory mechanisms, and there is a lack of druggability assessment frameworks (Sharma et al., 2024). Therefore, there is an urgent need to develop system biology approaches that integrate multi-omics data to comprehensively reveal the functional networks of ED treatment targets.
This study aimed to construct a gene–protein–metabolite regulatory network associated with ED by integrating druggable gene information, MR analysis, Summary data–based Mendelian Randomization (SMR) analysis, and multi-omics data, molecular docking techniques were employed to identify potential targeted drugs. These findings contribute to providing a theoretical foundation for the development of precision treatments for ED and offer new perspectives on shifting ED treatment from symptom management to mechanism-based intervention.
Materials and Methods
Study Design
The data used for this analysis were sourced from publicly available databases, so we do not need to conduct additional ethical reviews or obtain informed consent from participants. Figure 1 provides an overview of this study’s concept and process.

Detailed Design of This Research
Data Source
We obtained 4,479 druggable genes from prior research (Supplemental Material 1), which serve as a crucial foundation for drug development (Finan et al., 2017). We collected quantitative trait locus (QTLs) data related to these genes, the expression quantitative trait loci (eQTL) data were sourced from the eQTLGen Consortium (https://eqtlgen.org/). Protein quantitative trait loci (pQTL) data were obtained from three different research cohorts: the UK Biobank Pharma Proteomics Project (UKB-PPP; http://ukb-ppp.gwas.eu; Sun et al., 2023), the deCODE study (https://www.decode.com/summarydata; Ferkingstad et al., 2021), and the FinnGen project (https://www.finngen.fi/en/access_results; Pietzner et al., 2021). The GWAS data for ED were sourced from the IEU OpenGWAS under the specific ID ebi-a-GCST006956 (Bovijn et al., 2019). The data for single-cell RNA sequencing (ScRNA-Seq) were obtained from the publicly available database Gene Expression Omnibus ( http://www.ncbi.nlm.nih.gov/geo/), specifically using the GSE206528 dataset for ED (Zhao et al., 2022). Finally, researchers can access the GWAS summary statistics related to 1,400 metabolites through the GWAS catalog, with relevant IDs ranging from GCST90199621 to GCST90201020 (https://www.ebi.ac.uk/gwas/home; Chen et al., 2023).
IVs Selection
We adhered to the three core assumptions of MR analysis—relevance, independence, and exclusion restriction—guided by established practices in the field. A stratified selection strategy was employed to choose single nucleotide polymorphisms (SNPs) as instrumental variables (IVs), ensuring their validity: (a). Relevance Assumption: SNPs strongly associated with the exposure were selected, with a significance threshold of p < 5 × 10–8. This ensured that the IVs had robust predictive power for the exposure, mitigating weak instrument bias (F-statistic > 10). (b). Independence Assumption: Linkage disequilibrium clumping was applied (r2 < .001, clump kb > 10,000) to exclude correlated genetic variants, ensuring the IVs were independent of potential confounders. (c). Exclusion Restriction Assumption: We ensured that the IVs influenced the outcome solely through the exposure, excluding pleiotropic effects. This was verified through subsequent sensitivity analyses.
Given the limited number of IVs obtained under stringent thresholds in the GWAS data for 1,400 metabolites, and based on previous research experience, we relaxed the significance threshold to p < 1.0 × 10–5 (Sanna et al., 2019; Tan et al., 2025; Yang et al., 2025; Zheng et al., 2025). To further strengthen the results, sensitivity analyses and heterogeneity tests were incorporated to enhance interpretability. In conclusion, by integrating the three core assumptions of MR analysis with widely accepted methodological practices, we implemented a dual safeguard mechanism combining threshold-based screening and sensitivity verification. This approach ensures that our conclusions are robust, unaffected by genetic confounding, weak instrument bias, or horizontal pleiotropy, thereby providing a solid foundation for the reliability of our results.
MR Analysis
In this study, we primarily used the inverse variance weighting (IVW) method for MR analysis. If fewer than two IVs were available, we applied the Wald ratio method. We assessed the heterogeneity of the results using Cochran’s Q statistic. For sensitivity analysis, we employed several methods: MR pleiotropy residual and outlier detection (MR-PRESSO), MR Egger intercept test, and leave-one-out analysis. Specifically, the MR-PRESSO method carried out a global test to detect potential outliers that may cause pleiotropic bias. We reduced the impact of pleiotropy on the results by excluding SNPs that deviated from the normal distribution. A p-value < .05 indicated significance in the assessment of heterogeneity and pleiotropy. To ensure the accuracy and reliability of the analysis, we used the software packages such as “TwoSampleMR” and “MR-PRESSO.” These tools aided in validating the robustness and consistency of our two-sample MR analysis results.
SMR Analysis
To further validate the positive results in MR analysis, we employed the SMR and Heterogeneity In Dependent Instruments (HEIDI) methods (Zhu et al., 2016). In the SMR analysis, we utilized summary statistics of gene expression and genetic variation. Significant associations were considered to be those with p-SMR < .05, and p-HEIDI < .01 indicated that these associations are due to pleiotropic effects. By combining the SMR and HEIDI methods, we gain a deeper understanding and can validate the authenticity and robustness of the MR analysis results. This ensures that the identified causal relationships are not confounded by potential pleiotropic influences.
Mediation Analysis
In this analysis, druggable genes and metabolite data were treated as exposure factors and mediators, respectively, with ED serving as the outcome variable. Both the exposure factors and mediators must demonstrate significant causal relationships with the outcome. We also evaluated whether these druggable genes show significant associations with metabolites. If significant associations were found, we proceeded with a mediation MR analysis to examine the specific mediating effect of these metabolites.
In the first step, we calculated the total causal effect of druggable genes on ED (beta_all). Next, we assessed the causal effect of druggable genes on metabolites (beta1) and the causal effect of metabolites on ED (beta2). The mediating effect (beta_med) is calculated by multiplying the regression coefficients, and mediation proportion was calculated as follows: (beta1 * beta2) / beta_all. The total effect was divided into the mediating effect and the direct effect (beta_dir). The direct effect was calculated as follows: beta_all - beta_dir (Carter et al., 2021; Relton & Davey Smith, 2012).
Single-Cell Sequencing Analysis
We downloaded the ED scRNA-seq dataset from GSE206528, which contains three normal corpus cavernosum tissue samples and five patient samples, two of which have a history of diabetes. First, we used the Seurat package for quality control and normalization of the cells. Next, we performed dimensionality reduction with the RunPCA function and then clustered the cells using the FindNeighbors and FindClusters functions. For cell annotation, we referred to the method by Zhao et al. (2022) to annotate the cell populations. Following the annotation, we evaluated the expression of DKK3 and SLAMF6 in normal and ED tissues (including both diabetic and nondiabetic samples) and different cell subpopulations, and visualized the results with violin plots. We conducted pathway scoring using KEGG gene sets, and the results were displayed in a heatmap.
Protein–Protein Interaction and Protein–Chemical Interaction Network
The GeneMANIA database is a powerful bioinformatics tool. It constructs complex networks that generate significant hypotheses about gene functions and facilitate a thorough review of various gene lists. This process helps prioritize genes through detailed functional analysis. More information can be found on its official website (https://genemania.org/). We performed detailed meta-analyses of different protein expression factors using the advanced features of the NetworkAnalyst platform (http://www.networkanalyst.ca) to create a robust protein–chemical interaction network. This network was constructed using valuable resources from the Comparative Toxicogenomics Database (CTD; https://ctdbase.org/).
Molecular Docking
We downloaded the protein-coding information for DKK3 (UniProt: Q9UBP4) and SLAMF6 (UniProt: Q96DU3) from UniProt (https://www.uniprot.org/) and obtained their 3D structures from AlphaFold. AlphaFold, developed by DeepMind, is an artificial intelligence system that predicts protein 3D structures from amino acid sequences, with an accuracy that often rivals that of experimental results (Jumper et al., 2021). Information and molecular structures of small molecular compounds were sourced from the TCMSP (https://www.tcmsp-e.com/index.php) and PubChem (https://pubchem.ncbi.nlm.nih.gov/) databases. We used the PyMOL software to store protein and compound data in the PDB format, isolating the main ligands from the target protein and removing irrelevant water molecules, phosphates, and other inactive ligands (Warnecke et al., 2014). The format conversion of small molecular compounds was performed using the Open Babel software (O’Boyle et al., 2011). To perform molecular docking, we first used AutoDockTools 1.5.7 to convert the active ingredients and proteins from the PDB to PDBQT format and configured the parameters for the active site (Pinzi & Rastelli, 2019). We then employed AutoDock for docking, noting that the active ingredients show good target binding activity when the binding energy is −5.0 kJ/mol or lower. Finally, we validated the docking results using AutoDock Vina. This software generated binding results for the proteins and compounds, and we selected the result with the lowest free energy for visualization.
Statistical Analysis
We conducted data analysis using the R software (version 4.4.2; R Foundation for Statistical Computing, Vienna, Austria; https://www.R-project.org), and established that results would be deemed statistically significant only if the p-value < .05.
Result
MR Analysis Identified DKK3 and SLAMF6 as the Most Likely Target Genes for ED
During the QTL data screening, DKK3 and SLAMF6 were identified as key intersecting genes across four independent datasets (cis-eQTLs and cis-pQTLs) that have a significant causal relationship with ED. The IVW/Wald ratio–MR results indicated that both the eQTLs and pQTLs of DKK3 are causally linked to ED and significantly lower its risk (eQTLGen cis-eQTLs: OR = 0.9052, 95% CI [0.8213, 0.9976], p = .0445; deCODE cis-pQTLs: OR = 0.8555, [0.7615, 0.9613], p = .0087; UKB-PPP cis-pQTLs: OR = 0.8633, [0.7847, 0.9498], p = .0026; Fenland cis-pQTLs: OR = 0.8373, [0.7462, 0.9396], p = .0025). While DKK3 is associated with a reduced risk of ED, SLAMF6 has been found to significantly increase this risk (eQTLGen cis-eQTLs: OR = 1.2661, [1.0149, 1.5796], p = .0365; deCODE cis-pQTLs: OR = 1.2613, [1.0070, 1.5798], p = .0433; UKB-PPP cis-pQTLs: OR = 1.2525, [1.0031, 1.5638], p = .0468; Fenland cis-pQTLs: OR = 1.1448, [1.0019, 1.3081], p = .0468; Figure 2) (scatter plot and leave-one-out analysis are presented in Supplemental Material 2; details of sensitivity analysis are presented in Supplemental Material 1). All analytical processes followed the key assumptions of MR, and we performed thorough sensitivity analyses to confirm the reliability and robustness of the results. During this process, we did not observe any pleiotropic effects or variations, which further strengthens the credibility and scientific validity of our findings.

MR Analysis
SMR Analysis Between the Target Gene and ED
We conducted SMR analysis on the cis-QTLs data filtered through MR to further validate our conclusions. DKK3: (eQTLGen cis-eQTLs: p_SMR = .0371, p_HIEDI = .2404; deCODE cis-pQTLs: p_SMR = .0055, p_HIEDI = .0173; UKB-PPP cis-pQTLs: p_SMR = .0055, p_HIEDI = .0208; Fenland cis-pQTLs: p_SMR = .0058, p_HIEDI = .0151). SLAMF6: (eQTLGen cis-eQTLs: p_SMR = .0453, p_HIEDI = .3596; deCODE cis-pQTLs: p_SMR = .0444, p_HIEDI = .8594; UKB-PPP cis-pQTLs: p_SMR = .0477, p_HIEDI = .8029; Fenland cis-pQTLs: p_SMR = .0477, p_HIEDI = .7801) (Table 1). Together, the results from the SMR analysis and the preliminary analysis confirm a consistent direction and significance.
Summary of SMR Analysis for DKK3 and SLAMF6 Associated With ED Across Different QTL Datasets (eQTLGen, deCODE, UKB-PPP, and Fenland)

Mediation Analysis Based on 1,400 Metabolites
We aimed to explore whether metabolic factors influence the protective and promoting effects of DKK3 and SLAMF6 on ED. For this purpose, we conducted two-sample MR analyses of plasma protein levels and examined the relationships between 1,400 plasma metabolites and ED, aiming to reveal potential associations. Research findings suggest that certain plasma metabolites, such as piperine, methyl-4-hydroxybenzoate sulfate, and phosphocholine, along with the ratio of proline to trans-4-hydroxyproline, are linked to a higher risk of ED. Conversely, the plasma protein DKK3 decreases the levels of methyl-4-hydroxybenzoate sulfate and the proline to trans-4-hydroxyproline ratio, while SLAMF6 increases piperine and phosphocholine levels. We conducted a two-step MR mediation analysis to assess the mediating role of plasma metabolites in the causal relationship between DKK3 and SLAMF6 in ED. The results revealed that DKK3's overall effect in reducing ED is indirectly influenced by the levels of methyl-4-hydroxybenzoate sulfate and the ratio of proline to trans-4-hydroxyproline (mediation proportion: 8–11%, Table S6). In comparison, SLAMF6’s role in increasing ED is indirectly influenced by the levels of piperine and phosphocholine (mediation proportion: 7–14%, Table S6; Fig. 3; scatter plot and leave-one-out analysis are presented in Supplemental Material 2; details of the sensitivity analysis are presented in Supplemental Material 1).

Mediated Mendelian Randomization Analysis
Single-Cell Analysis
After thorough quality control of the ED and normal samples, followed by clustering analysis and annotation, we identified seven distinct cell populations. Among the identified populations, DKK3 was significantly expressed in endothelial and smooth muscle cells, while SLAMF6 was predominantly concentrated in T cells. Notably, DKK3 expression levels are significantly lower in all cell clusters from both diabetic and nondiabetic patients than those from the normal population (p < .001). In contrast, SLAMF6 expression levels are significantly higher (p < .05; Figure 4). This difference in expression levels is especially pronounced in endothelial cells and T-cell subpopulations, emphasizing the critical roles of these two cell types in ED. The KEGG analysis results between these two cell clusters are shown in Figure 5. In endothelial cells, the key pathways associated with high and low expression of DKK3 are as follows: WNT signaling pathway, VEGF signaling pathway, type II diabetes mellitus, asthma, and base excision repair. In T cells, the main pathways involved in high and low expression of SLAMF6 are as follows: antigen processing and presentation, chronic myeloid leukemia, natural killer cell-mediated cytotoxicity, NOTCH signaling pathway, and NEUROTROPHIN signaling pathway. The research results clearly indicate that DKK3 plays a key role in the development of ED by primarily targeting endothelial cells, while SLAMF6 is important in this pathological process by primarily targeting T cells.
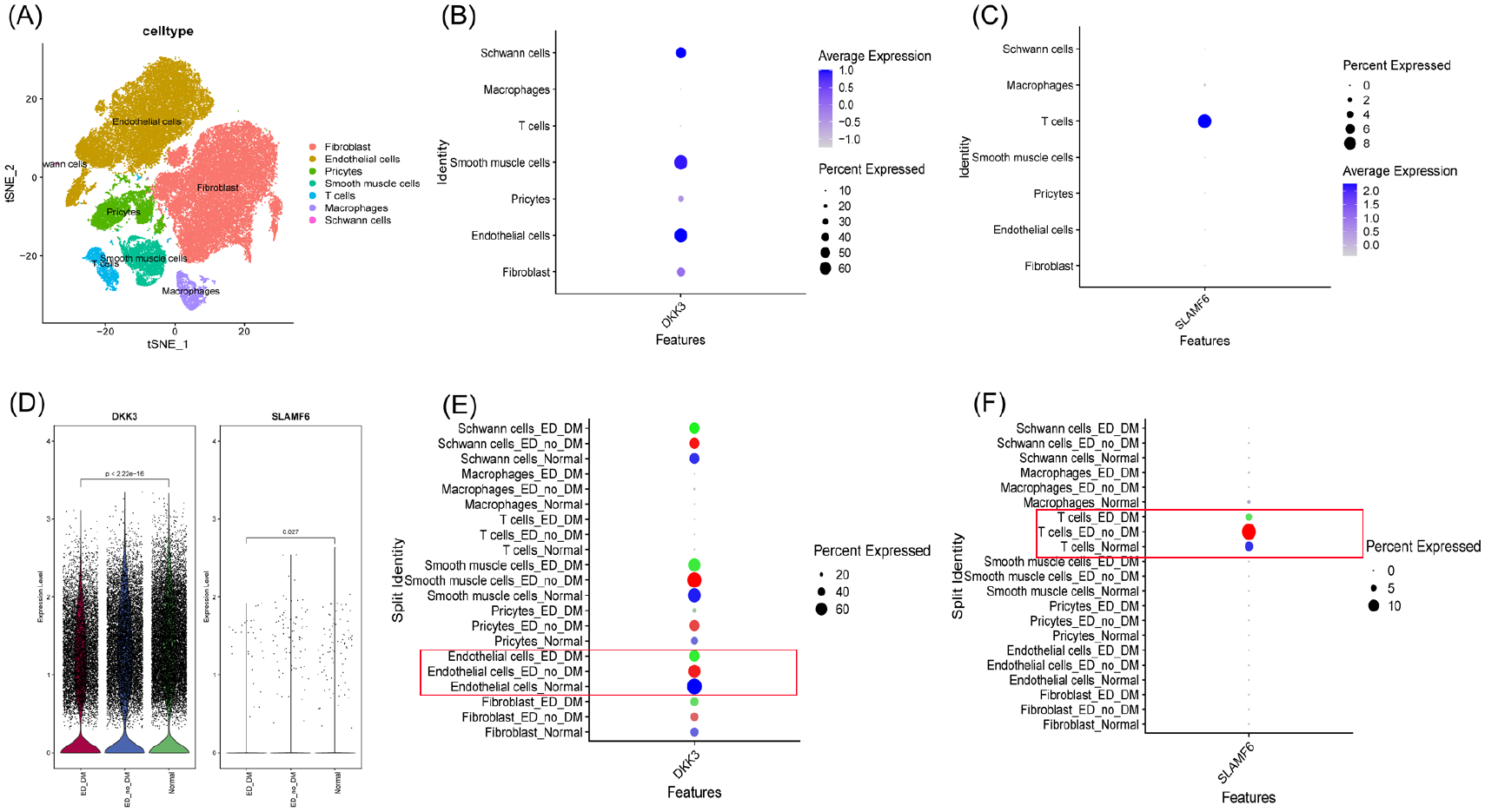
Single-Cell Analysis

Cell Subpopulation and KEGG Analysis
PPI and Protein–Chemical Component Interaction Network
First, we constructed a PPI network focused on DKK3 and SLAMF6 using the GeneMANIA database (Figure 6). Functional analysis through GeneMANIA revealed that these genes primarily participate in various key biological processes, including receptor inhibition activity, regulation of natural killer (NK) cell–mediated immunity, regulation of antigen receptor–mediated signaling pathways, negative regulation of the Wnt signaling pathway, positive regulation of lymphocyte-mediated immunity, and regulation of the canonical Wnt signaling pathway. We then investigated how DKK3 and SLAMF6 interact with various chemical components using the network analysis. Our findings from the CTD revealed that several chemicals, such as air pollutants, nickel, and copper, significantly interact with the target proteins. These interactions provide valuable insights into how environmental factors affect the biological functions of these genes.
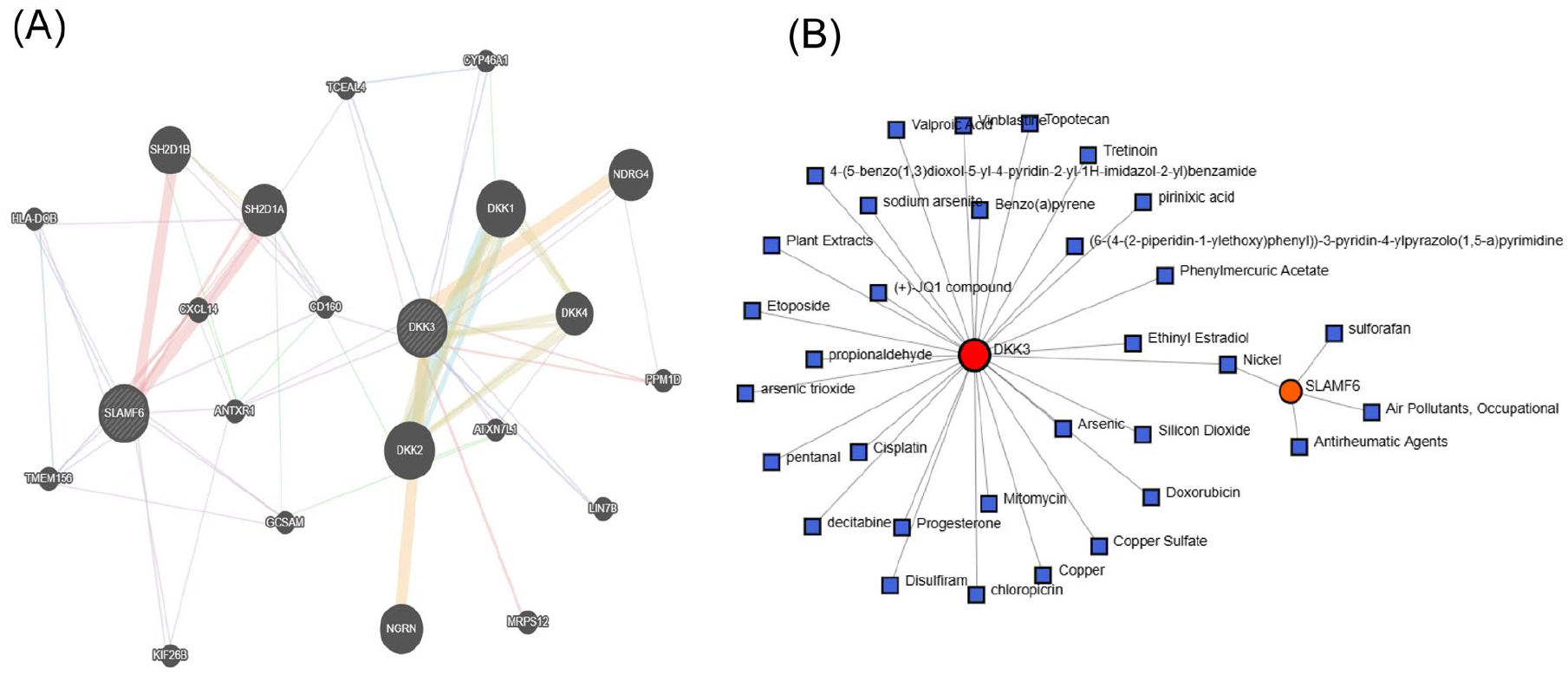
Networks for PPI and Protein–Chemical Interaction
Molecular Docking
In this study, we selected compound targets related to specific diseases from the TCMSP database, particularly focusing on four beneficial compounds from the traditional Chinese medicinal herbs: Epimedium and Radix Salviae. The selected compounds—Icariins, Luteolins, Danshenol A, and Danshenxinkun A—are key active ingredients for molecular docking with DKK3 and LSAMF6. We used a screening criterion of binding energy at or below −5.0 kcal/mol and confirmed the interactions between these key components and the hub proteins through molecular docking experiments. The results indicated that all selected compounds had stable binding capabilities with the target proteins, showing binding energies between −6.0 and −8.0 kcal/mol (Figure 7, Table 2). These results highlight a strong binding affinity between the key active ingredients of the anti-inflammatory and tonic compounds and the central targets. This further emphasizes their potential therapeutic applications in related diseases.

Molecular Docking. Binding Modes of Proteins and Ligands
Free Binding Energy of the Key Active Components of Tonifying Compounds and Key Target Protein
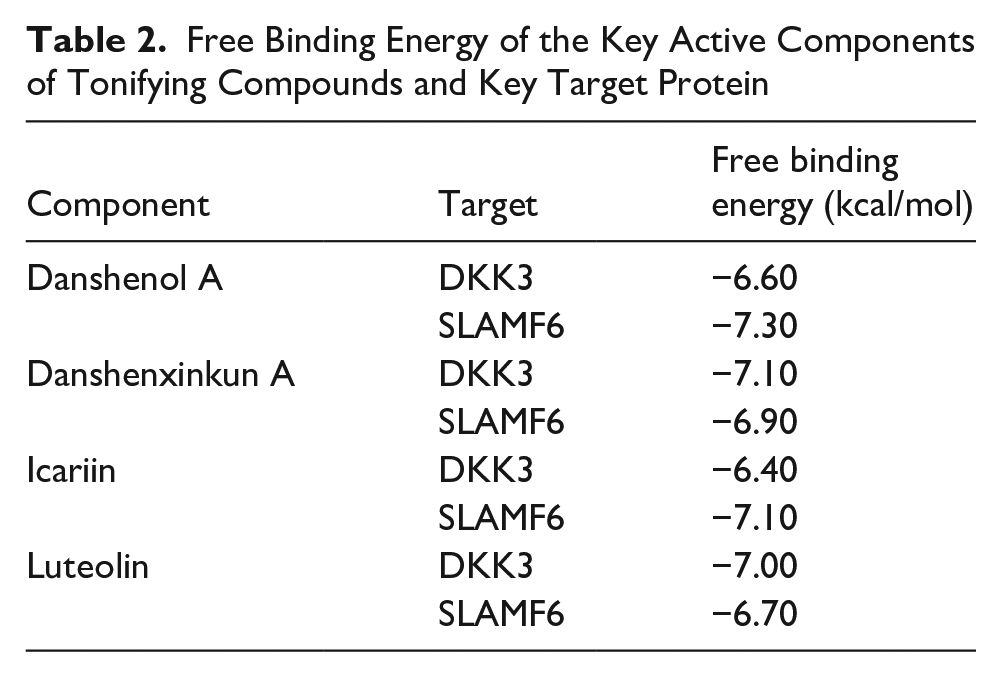
Discussion
In this study, based on 4,479 druggable genes, we used MR analysis to assess the causal relationship between these genes and ED by integrating eQTLs and pQTLs data from multiple cohorts. Finally, we identified DKK3 and SLAMF6 as potential therapeutic targets for ED. Through the specific expression patterns in ED single-cell data, the molecular mechanisms by which vascular homeostasis imbalance and immune dysregulation synergistically contribute to the pathogenesis of ED are elucidated. Additionally, molecular docking has identified the targeted potential of natural small molecules such as icariin, providing a theoretical basis for the development of precision therapies that combine vascular protection and immune regulation.
Through MR, SMR, and cross-validation across multiple pQTL cohorts, DKK3 was identified as a protective factor for ED, while SLAMF6 was determined to be a risk factor. Notably, DKK3 is an important member of the Dickkopf protein family and is classified as a secreted glycoprotein. The primary role of DKK3 is to engage in various physiological and pathological processes by regulating the Wnt/β-catenin signaling pathway. The structure of DKK3 allows it to bind to receptors in the Wnt signaling pathway. This binding inhibits their activity and affects various biological processes, including cell proliferation, differentiation, and apoptosis (Niehrs, 2006). Research shows that the expression of DKK3 is closely related to the activity of endothelial nitric oxide synthase (eNOS). Increased DKK3 levels promote eNOS expression and boost NO synthesis, which in turn influences endothelial function (Busceti et al., 2023; Kaltsas et al., 2024). The high expression levels of DKK3 in vascular endothelial cells are crucial for maintaining vascular homeostasis, consistent with our observations. Specifically, DKK3 contributes to both the formation and remodeling of blood vessels, as well as the regulation of vascular function (Baetta & Banfi, 2019; Yu et al., 2017). In our KEGG enrichment analysis of endothelial cells, we found that cells with low DKK3 expression showed activation of pathways such as Wnt and JAK-STAT, leading to endothelial damage, further supporting the conclusion that DKK3 is a protective factor for ED. Pathways such as asthma and type II diabetes mellitus were significantly enriched in endothelial cells with low DKK3 expression, indicating that DKK3 expression is closely associated with several pathological conditions, particularly diabetes-related ED, where its levels are significantly reduced (Song et al., 2020). This reduction in DKK3 levels suggests that it may play a role in the vascular complications associated with diabetes. Diabetic patients frequently suffer from endothelial dysfunction and vascular remodeling; the lack of DKK3 is thought to worsen these conditions. DKK3 may affect the blood flow in the penile corpus cavernosum by regulating how vascular smooth muscle cells change their characteristics (Karamariti et al., 2018). The phenotypic changes in smooth muscle cells are closely related to the contraction and relaxation capabilities of blood vessels. DKK3's regulatory role may influence blood supply to the penis, subsequently impacting erectile function. Thus, DKK3 is valuable for research in vascular pathology linked to ED. Future studies are expected to explore its potential applications in treatment.
SLAMF6 has been identified as a risk factor associated with increased ED susceptibility in our study. SLAMF6 is a key component of the Signaling Lymphocytic Activation Molecule Family and is a transmembrane glycoprotein primarily found in immune cells such as T cells, B cells, NK cells, and monocytes. This glycoprotein can modulate immune cell functions and their roles in immune responses by interacting with various cell surface receptors (Dragovich et al., 2019; Yigit et al., 2019). Researchers consider the abnormal expression and function of SLAMF6 to be closely associated with the occurrence and progression of various immune-related diseases. For example, excessive SLAMF6 activation may cause immune cells to attack the body’s own tissues, worsening the autoimmune diseases (Humbel et al., 2022). Chronic low-grade inflammation is recognized as a significant pathogenic mechanism of ED, with SLAMF6 potentially playing a crucial role by regulating T-cell activity and NK cell activity, which in turn influences the release of inflammatory factors and the progression of the inflammatory response. This effect may alter the hemodynamics and endothelial function of the penile corpus cavernosum, which can impact erectile function. Additionally, SLAMF6 might affect the fibrotic process in the penile corpus cavernosum by controlling macrophage polarization (Meng et al., 2022). Macrophages are essential in the inflammatory response. Their polarization state, either M1 or M2, directly influences tissue repair and fibrosis (Zhang et al., 2024). Thus, SLAMF6 may increase local inflammatory responses by encouraging macrophage polarization, which results in fibrosis and dysfunction of the penile corpus cavernosum. Similarly, in our study, we found that in the T-cell subpopulations of ED, cells with high SLAMF6 expression showed upregulation of immune-related pathways. This supports the role of SLAMF6 in regulating inflammatory responses, providing valuable insights into the immune mechanisms underlying ED. These findings also potentially reveal new targets for developing therapeutic strategies.
The high expression of SLAMF6 and the downregulation of DKK3 in ED emphasize the impact of dynamic gene interactions on the disease. DKK3 inhibits the Wnt signaling pathway and is linked to various diseases, including tumors and chronic inflammation. SLAMF6 is essential for immune regulation and intercellular interactions. The interaction between DKK3 and SLAMF6 through specific signaling pathways may further influence the development of ED. Our study also found that metallic nickel can simultaneously affect the expression of both DKK3 and SLAMF6, as demonstrated by the protein and chemical reaction data in the CTD database. Nickel can decrease DKK3 mRNA levels while increasing SLAMF6 mRNA levels (Correa da Rosa et al., 2015; Dhingra et al., 2014; Slotkin et al., 2010). Thus, as an environmental pollutant, nickel may influence the progression of ED by regulating DKK3 and SLAMF6 expression, which impacts associated signaling pathways. This further highlights the complexity and importance of gene interaction networks. Additionally, our findings indicate that various metabolites mediate the relationship between DKK3, SLAMF6, and ED, suggesting a common metabolic pathway among these molecules. Targeting both DKK3 and SLAMF6 in the treatment of ED could offer patients a more comprehensive approach. DKK3 enhances endothelial function and inhibits abnormal smooth muscle cell proliferation, thereby improving vascular function. SLAMF6 slows the pathological process of ED by inhibiting inflammation and enhancing immune cell activity. The combination of DKK3 and SLAMF6 may create a synergistic effect, enhancing vascular function and reducing inflammation, leading to more effective ED treatment. The effects and mechanisms of combination therapies still require further research to optimize treatment plans and determine the most effective strategies. Future studies should investigate how DKK3 and SLAMF6 interact and how they can be used together in ED treatment to offer patients more personalized and effective options.
In summary, this study offers valuable insights into the roles of DKK3 and SLAMF6 in ED, but some limitations must be acknowledged. First, using publicly available datasets may introduce inherent biases from the original studies, such as differences in sample populations and methodologies that could limit the generalizability of our findings. Additionally, although we used rigorous statistical methods to reduce potential confounding variables and variations, the possibility of confounding factors remains a concern, especially in mediation analyses involving metabolites. Finally, while the molecular docking results are promising, they are based on computational predictions and need experimental validation to confirm the biological significance of the observed interactions. These limitations highlight the urgent need for further research to validate our findings and investigate the underlying mechanisms in greater depth.
Conclusion
Our study suggests that DKK3 and SLAMF6 are promising molecular targets with significant potential in pioneering innovative clinical treatments. As research progresses, the complexity of their interactions will likely be further elucidated, which undoubtedly holds the potential to offer new hope and transformative possibilities for countless patients with ED.
Supplemental Material
sj-docx-2-jmh-10.1177_15579883251368307 – Supplemental material for Assessing Genetic Risk of DKK3 and SLAMF6 in Erectile Dysfunction: A Comprehensive Analysis Based on Mendelian Randomization
Supplemental material, sj-docx-2-jmh-10.1177_15579883251368307 for Assessing Genetic Risk of DKK3 and SLAMF6 in Erectile Dysfunction: A Comprehensive Analysis Based on Mendelian Randomization by Guangqiang Zhu, Chunlin Tan and Yugen Li in American Journal of Men's Health
Supplemental Material
sj-xlsx-1-jmh-10.1177_15579883251368307 – Supplemental material for Assessing Genetic Risk of DKK3 and SLAMF6 in Erectile Dysfunction: A Comprehensive Analysis Based on Mendelian Randomization
Supplemental material, sj-xlsx-1-jmh-10.1177_15579883251368307 for Assessing Genetic Risk of DKK3 and SLAMF6 in Erectile Dysfunction: A Comprehensive Analysis Based on Mendelian Randomization by Guangqiang Zhu, Chunlin Tan and Yugen Li in American Journal of Men's Health
Footnotes
Acknowledgements
The authors thank the participants of the Genome-Wide Association Studies and the Gene Expression Omnibus.
Author Contributions
Guangqiang Zhu: Writing—original draft, Software, Methodology, Data curation, Conceptualization, Visualization. Chunlin Tan: Writing—original draft, Methodology, Investigation, Formal analysis, Conceptualization. Yugen Li: Writing—review & editing, Funding acquisition, Validation, Conceptualization. All authors have reviewed and approved the final manuscript.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was supported by the City and College Technology Strategic Cooperation Project of Nanchong (22SXQT0215); the Project of North Sichuan Medical College (CBY21-QA38).
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
