Abstract
The treatment of erectile dysfunction (ED) remains a significant challenge. Mendelian randomization (MR) is being increasingly utilized to identify novel therapeutic targets. In this study, we carried out a genome-wide MR analysis on druggable targets with the aim of pinpointing latent therapeutic alternatives for ED. We collected data on the druggable genes and filtered out those associated with blood eQTLs, then performed two-sample MR and colocalization analyses using ED genome-wide association data to screen genes significantly linked to the condition. In addition, we carried out phenome-wide studies, enrichment analysis, protein network modeling, drug prediction, and molecular docking. We screened 3,953 druggable genes from the DGIdb and 4,463 from a review. Following data integration, 74 potential druggable genes were found to potentially regulate corpus cavernosum fibrosis. MR analysis of eQTL data uncovered five drug targets (TGFBR2, ABCC6, ABCB4, EGF, and SMAD3) significantly associated with ED risk. Colocalization analysis suggested a shared causal variant between ED susceptibility and TGFBR2, with a posterior probability (PPH4) exceeding 80%. Drug predictions utilizing DSigDB identified nolone phenylpropionate, sorafenib, and NVP-TAE684 as significantly associated with TGFBR2. Finally, molecular docking indicated strong binding affinities between these candidate drugs and the protein encoded by TGFBR2 (Vina score < −50). Through MR and colocalization analyses, the present study identified five potential drug targets for ED, with TGFBR2 showing remarkable relevance in blood. These findings offer valuable insights and potential leads for the development of more effective ED therapies, which may also contribute to cutting down the expenses involved in drug development.
Background
Erectile dysfunction (ED) is defined by insufficient penile erection and unsatisfactory intercourse (Najari & Kashanian, 2016; Woolf et al., 2023). According to the European Association of Urology Guidelines, the occurrence rate of ED climbs steadily with advancing age and shows significant variations across different ethnic populations. Specifically, among men aged 40–70 years, ED affects 52% of Americans, 30% of Europeans aged 40–79, and 63% of Asians aged 50–80 (Li et al., 2005; Salonia et al., 2021; Vollset et al., 2020). ED is a chronic disorder that brings about substantial physical, mental, and neurological ramifications. Projections indicate that by 2025, approximately 322 million men globally are likely to experience ED (Yafi et al., 2016). The onset of ED has been linked to multiple comorbid conditions and risk factors, including smoking, depression, penile trauma, low androgen levels, and prostate surgery (Bauer et al., 2020; Zeleke et al., 2021). ED management encompasses lifestyle modifications, optimizing treatment for comorbidities, and employing psychosexual or pharmacological therapies. The available treatment modalities include intracavernosal injections, oral phosphodiesterase 5 inhibitors (PDE5Is), hormone replacement therapy, penile prostheses, low-intensity extracorporeal shockwave therapy (Li-ESWT), stem cell injections, and others.
Mendelian randomization (MR) is a widely adopted approach for uncovering novel therapeutic targets. It achieves this by amalgamating summary data from expression quantitative trait loci (eQTL) and disease-related genome-wide association studies (GWAS) (Storm et al., 2021). By integrating GWAS data with molecular QTL (molQTL), such as eQTLs or protein QTLs (pQTLs), the identification of target genes linked to risk variants through causal inference can be achieved (Kreitmaier et al., 2023). By utilizing genetic data, MR replicates the process of a randomized trial, circumventing the requirement for direct drug testing (Davey Smith, 2007). In the present study, we conducted a genome-wide MR analysis focused on druggable genes to identify underlying treatment targets for ED. We first compiled data on the druggable genes, filtered those in blood eQTLs, and then conducted a two-sample MR analysis by integrating with ED GWAS data, with the intention of precisely identifying genes that have a strong connection to ED. Colocalization analysis was then performed to ensure result reliability. A phenome-wide analysis was executed on significant blood genes to investigate the associations between the treatment targets and a diverse range of traits. Moreover, enrichment analysis, protein network modeling, drug prediction, and molecular docking are carried out to offer directions for the advancement of more efficient ED therapies.
Method
Figure 1 offers a comprehensive overview of the present study.
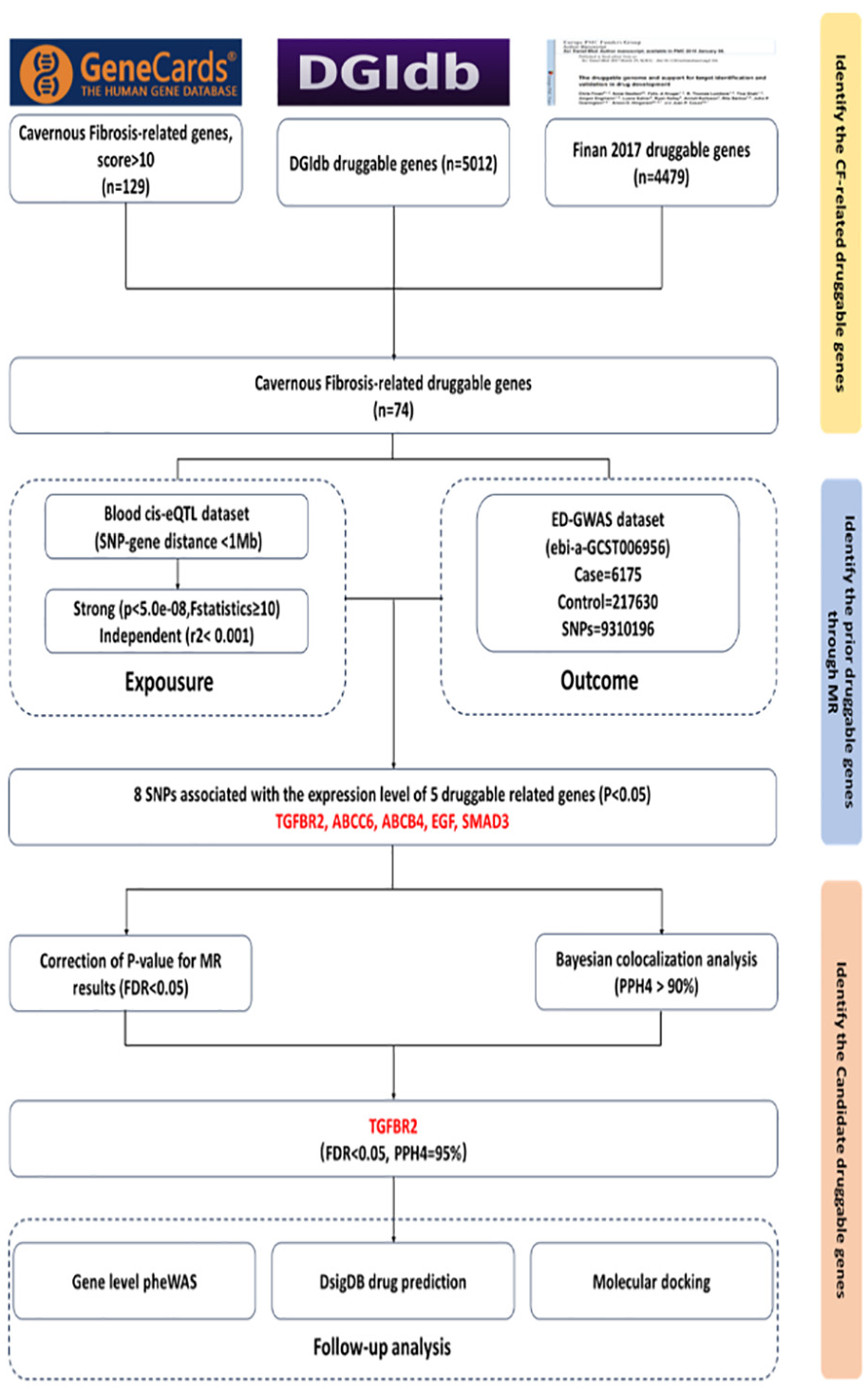
Offers an Overview of the Design of the Current Study
Druggable Genes
The data utilized in this study were obtained from the Drug-Gene Interactions Database (DGIdb, https://www.dgidb.org/) (Freshour et al., 2021) as well as a review article. DGIdb serves as a valuable resource, offering in-depth knowledge about drug-gene interactions and their possible therapeutic implications. We retrieved the “Interactions data” that had its most recent update in December 2023 from DGIdb, and incorporated the druggable gene list from Finan et al. (2017). In addition to this, genes related to cavernous fibrosis were obtained from the GeneCards (https://www.genecards.org/). To acquire these genes, we conducted a search for “cavernous fibrosis” on the GeneCards platform and selected those with a correlation score exceeding 10. Subsequently, we performed an intersection of the three gene sets to pinpoint the common genes, which were considered as potential druggable targets for cavernous fibrosis.
eQTL Data Sets
We searched the eQTLGen Consortium (https://eqtlgen.org/) (Võsa et al., 2021) and obtained cis-eQTL data in the blood for druggable genes. The alliance integrated 37 data sets, with a total of 31,684 individuals, the majority of whom were of European descent. With the eQTL data, one can identify genetic variants linked to gene expression levels. Situated within 1 Mb of the mid-point of each gene, these variants possess a minor allele frequency exceeding 0.01.
ED GWAS Data Set
We obtained the summary statistics regarding ED from a GWAS carried out by Bovijn et al. (2019). This particular study involved a cohort of 223,805 individuals of European descent, including 6,175 ED patients and 217,630 control subjects, and analyzed a total of 9,310,196 SNPs. To date, this data set represents the largest and most frequently utilized GWAS data on ED to date. For the subsequent analyses, we retrieved the Variant Call Format file of this data set via the IEU Open GWAS project (https://gwas.mrcieu.ac.uk/).
Mendelian Randomization Analysis
This “TwoSampleMR” package in R was employed for MR analysis (Hemani et al., 2018). The eQTL of druggable genes were selected as the exposure data. To construct the instrumental variable (IV), genetic variants strongly related to the expression of drug genes were extracted, specifically those located within a 1 Mb region flanking the coding sequence of the drug genes. To minimize the impact of pleiotropy and ensure compliance with the three MR assumptions, we implemented the genome-wide significance threshold (p < 5 × 10−8) and an F-statistic ≤ 10 to identify robustly associated IVs. Subsequently, to ensure the independence of each selected significant single nucleotide polymorphism (SNP) and to mitigate the effects of pleiotropy arising from linkage disequilibrium (LD), we established an LD coefficient threshold of r2 < 0.001, set the LD window width to 10 Mb, and employed the aggregation function of the “TwoSampleMR” package to derive IVs (Carter et al., 2019; VanderWeele, 2016). After identifying IVs, we extracted effect estimates of the same variant or its proxy from the GWAS data set for ED to facilitate data coordination. Within the framework of the random-effects mode, we utilized either the Wald ratio or the inverse variance weighting method to assess the association between exposure and outcomes. We used Cochran’s Q test to evaluate the heterogeneity of causal impacts across SNPs. Furthermore, the MR Egger intercept was employed for the assessment of SNP pleiotropy, with a p-value below .05 deemed statistically significant.
Colocalization Analysis
To minimize the confounding effects of SNPs originating from different genes on the disease, we employed the “colon” R software package (version 5.2.3) for colocalization analysis. This type of analysis is of great significance for validating potential co-causal genetic variants within the shared physical loci between eQTL and ED. For both the GWAS and blood eQTL data related to ED, SNPs positioned within a 1,000 kb region either upstream or downstream of the TSS of each potential druggable gene were omitted. P1 represents the probability that a SNP is linked to ED, P2 denote the likelihood that a SNP is a notable eQTL, and P12 signifies the joint probability of a SNP being both associated with ED and a significant eQTL. All probabilities were set to their default settings (P1 = 1×10−4, P2 = 1×10−4, and P12 = 1×10−5). The a posteriori probability (PP) is utilized to evaluate the degree of support for hypotheses, denoted as PPH0 to PPH4. PPH0 stands for the state of being independent of any trait. PPH1 implies an association with gene expression yet not with ED risk. PPH2 signifies an association with ED risk rather than gene expression. PPH3 demonstrates an association with both ED risk and gene expression, accompanied by distinct causal disparities. Finally, PPH4 represents the existence of shared causal variants between ED risk and gene expression. Given the limitations of colocalization analysis, subsequent analyses will concentrate on genes with a PPH4 value of 0.75 or higher.
Phenome-Wide Association Analysis
The Phenome-Wide Association Study (PheWAS) was carried out via the AstraZeneca PheWAS portal (https://azphewas.com/) to assess the pleiotropic effects and potential adverse reactions of prospective therapeutic targets. The initial study integrated data encompassing around 15,500 binary phenotypes and 1,500 continuous phenotypes (Dhindsa et al., 2023; Wang et al., 2021). This in-depth PheWAS analysis offers crucial perspectives for understanding intricate genetic traits and appraising the safety and effectiveness of drug targets.
Candidate Drug Prediction
Drug Signatures Database (DSigDB, http://dsigdb.tanlab.org/DSigDBv1.0/) (Yoo et al., 2015) is an extensive repository comprising 22,527 gene sets and 17,389 distinct compounds, and a total of 19,531 genes. In this study, we fed the previously identified important drug-capable genes into the DSigDB. This was done to assist in predicting potential drug candidates and evaluating the pharmacological activity of the target genes.
Molecular Docking
We utilized the cavity detection guide blind docking (CB-Dock2, https://cadd.labshare.cn/cb-dock2/), to assess drug candidates and their respective target binding energy and interaction model. CB-Dock2 is a server designed for the protein-ligand blind docking (Liu et al., 2022). The structural data of the candidate drugs were obtained from the PubChem compound database (https://pubchem.ncbi.nlm.nih.gov/) (Kim et al., 2022) and saved in the SDF format. The protein structural data were sourced from the Protein Data Bank (PDB, http://www.rcsb.org/) (Burley et al., 2023). Both drug structure files and protein structure files were uploaded to CB-Dock2 for molecular docking and subsequent visualization.
Results
Druggable Genome
A total of 3,953 druggable genes were retrieved from the DGIdb, and 4,463 were sourced from prior reviews. Moreover, 129 genes (with a score > 10) that were highly correlated with cavernous fibrosis were obtained from the GeneCards database. Upon data consolidation, 74 potential druggable genes, which might regulate corpus cavernosum fibrosis, were identified for further analysis (Figure 2).

Venn Diagram of Gene Sets for Druggable Genes and Cavernosum Fibrosis-Related Genes
Candidate Druggable Genes
After integrating the gene data from the abovementioned three parts, 74 druggable genes associated with cavernous fibrosis were acquired, and further blood eQTL data of these genes were obtained. We carried out an MR analysis and identified five crucial genes related to ED: TGFBR2, ABCC6, ABCB4, EGF, and SMAD3. Among them, TGFBR2 exhibited the most significant p-value, which remained less than 0.05 even after FDR correction. Among these five genes, TGFBR2 (OR = 1.22, 95% CI [1.09, 137]), EGF (OR = 1.20, 95% CI [1.03, 1.41]), and SMAD3 (OR = 1.17, 95% CI [1.01, 1.35]) might be pathogenic genes for ED. In contrast, ABCC6 (OR = 0.84, 95% CI [0.73, 0.96]) and ABCB4 (OR = 0.85, 95% CI [0.74, 0.97]) could potentially be protective genes against the disease (Figure 3).

A Forest Plot That Displays Five Genes, Which Are Significantly Linked to ED and Are Sourced From Blood
Colocalization Analysis
To evaluate the likelihood of shared causal variation between cis-eQTL and ED outcomes, we carried out colocalization analysis on the druggable genes that produced significant MR findings. The findings of the colocalization analysis indicate that there is a possibility that the susceptibility to ED and TGFBR2 share causal variation, with a posterior probability of PPH4 > 80%. Consequently, TGFBR2 has been recognized as an underlying target for drug development aimed at reducing the risk of ED (Table 1, Figure 4).
Colocalization Results of Five Significant Genes From Blood
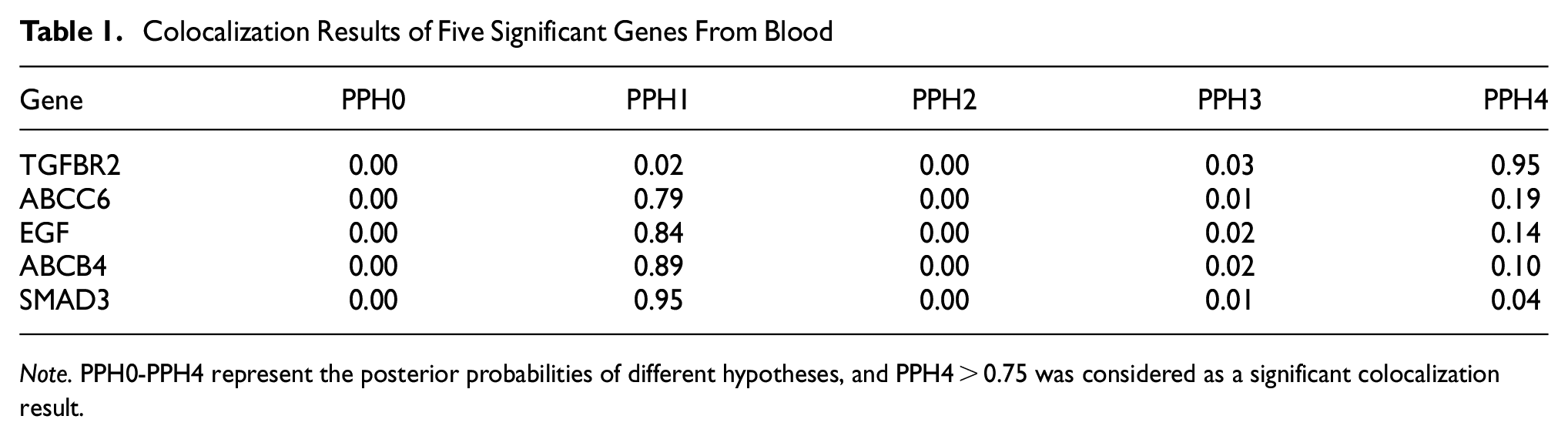
Note. PPH0-PPH4 represent the posterior probabilities of different hypotheses, and PPH4 > 0.75 was considered as a significant colocalization result.

Regional Plot of Colocalization Evidence of TGFBR2 and ED
Phenome-Wide Association Analysis
A comprehensive phenotype MR was conducted via the PheWAS portal and PheWeb database to unearth potential side effects related to targeting TGFBR2. It was shown by the results that there was no indication of a significant link between TGFBR2 and other phenotypes (p < 5e−10, Figure 5). These findings strengthen the validity of our research and suggest that using TGFBR2 as a therapeutic target may mitigate the likelihood of adverse drug reactions or unanticipated pleiotropic impacts.

Binary Traits PheWAS Association With TGFBR2
Candidate Drug Prediction
We employed the DSigDB to pinpoint potentially effective interventions. Subsequently, we curated a list of the top 10 interventions, ranked by adjusted p-values (Table 2). These findings revealed that Nandrolone phenylpropanoate, Sorafenib, and NVP-TAE684 were significantly correlated with TGFBR2 and EGF (Figure 6), indicating that these three drugs hold the potential to ameliorate ED via TGFBR2.
Candidate Drug Predicted by DSigDB

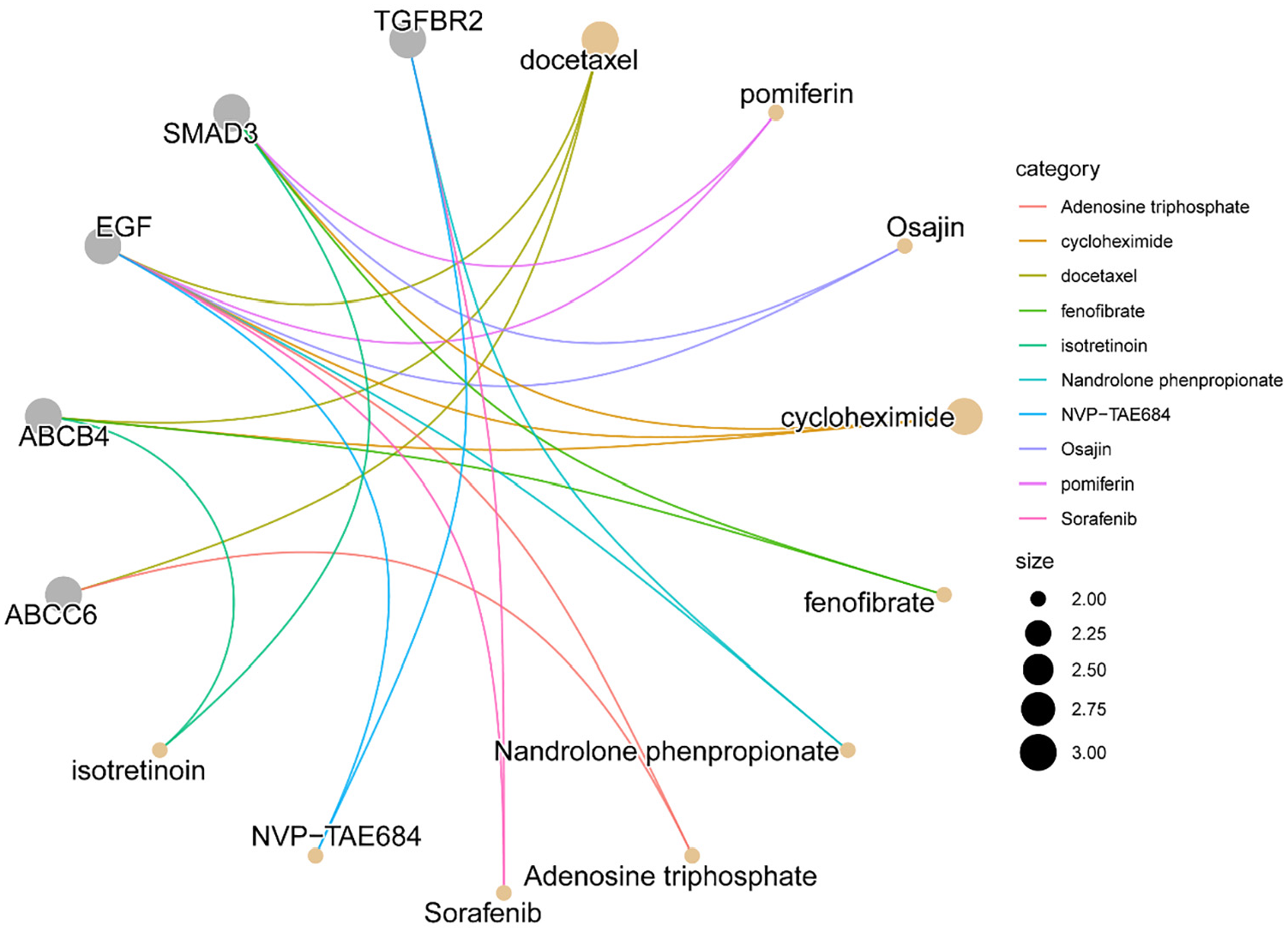
Cnetplot Displays Candidate Drugs Predicted by DSigDB
Molecular Docking
For the sequential docking of the three candidate drugs to the protein encoded by TGFBR2, we made use of the CB-Dock2 tool. Subsequently, these binding affinities of the molecules were assessed using the Vina score. The Vina score is based on the scoring function provided by AutoDock Vina software, which calculates the interaction energy and binding free energy between molecules to obtain a score. A lower score indicates a stronger binding ability. The results of molecular docking demonstrated that the three candidate drugs exhibited a strong binding ability to the protein encoded by TGFBR2 (Vina score < −50) (Table 3). The schematic diagram of molecular docking is presented in Figure 7.
Molecular Docking Results of TGFBR2 and Drugs


Molecular Docking Results of Available Proteins and Drugs. (A) TGFBR2 Docking Nandrolone Phenpropionate; (B) TGFBR2 Docking Sorafenib; and (C) TGFBR2 Docking NVP-TAE684
Discussion
In this research, we scoured the DGIdb and carried out an all-encompassing review of druggable genes, identifying 74 that were significantly associated with ED. By means of MR analysis, we pinpointed five pivotal genes related to ED: TGFBR2, ABCC6, ABCB4, EGF, and SMAD3, with TGFBR2 emerging as a possible drug target aimed at decreasing the risk of ED. To further validate the therapeutic potential of TGFBR2 as a targetable gene, drug prediction and molecular docking analyses were finally conducted.
Transforming growth factor-β (TGF-β) is crucial for regulating cell differentiation, proliferation, and apoptosis (Massagué, 2008). Two transmembrane serine/threonine kinase receptors, TGFBR1 and TGFBR2, are necessary for the TGF-β signaling pathway. Among them, TGFBR2 assumes a crucial function in the pathways associated with TGF- β and the process of signal transduction (Q. Zhang et al., 2023). Initially, the TGF-β ligand attaches to TGFBR2 on the plasma membrane, which sets off the generation of the TGFBR1-TGFBR2 complex. Following this, TGFBR2 causes TGFBR1 to be phosphorylated. Once phosphorylated, TGFBR1 goes on to activate Smad2 and Smad3 and add phosphate groups to them. Ultimately, the phosphorylated Smad2 and Smad3 associate with Smad4. The resulting complex then moves into the nucleus to regulate the expression of target genes (Ikushima & Miyazono, 2010; Schmierer & Hill, 2007). Research indicates that TGF-β1 impacts all stages of the fibrotic disease process, starting with the inflammatory phase, where infiltrating immune cells and macrophages prepare for the fibrotic phase, in which mononuclear cells and activated fibroblasts drive pathological matrix accumulation (W. Chen & Wahl, 1999).
Chronic and repeated injury gives rise to persistent tissue inflammation and the accumulation of extracellular matrix (ECM) within the internal structure of the cavernous body. This sequential process restricted relaxation of the cavernous body’s smooth muscle, results in insufficient arterial filling and venous occlusion dysfunction and ultimately culminates ED (Milenkovic et al., 2019). With the advancement of research, the fibrosis process of cavernous tissue has been increasingly elucidated. It has been noted that severe fibrosis of cavernous tissue is a critical factor in refractory ED (Fang et al., 2022). Fibrosis generally initiates with inflammation, which activates fibroblasts and other stromal cells. These activated cells secrete cytokines and prompt the synthesis and deposition of ECM. If the injury continues unabated, the inflammation escalates further. Consequently, the number of smooth muscle cells decreases, while the rate of ECM accumulation exceeds its degradation rate. This results in the generation of fibrotic foci, along with a decrease in tissue elasticity and compliance. Eventually, normal tissue cells are replaced, and their functions are impaired (Gonzalez-Cadavid, 2009; Milenkovic et al., 2019).
Moreover, vascular smooth muscle hyperplasia along with fibrosis occurs, which leads to the constriction of vascular channels and an elevation in resistance. Consequently, the blood sinuses experience inadequate blood filling. This insufficiency causes the pressure in the small veins beneath the tunica albuginea to be low, resulting in blood leakage from the veins. Ultimately, the penis fails to attain sufficient erectile hardness, giving rise to ED (Lopes et al., 2000; Ma et al., 2020). TGF-β is regarded as an essential molecule in the development of cavernous fibrosis and plays a major role in promoting the progress of fibrosis. When stimulated by factors such as inflammation, TGF-β in the ECM is activated and released in its active state. It binds to the cell-surface receptor, phosphorylates the receptor, and then combines with the activated Smad protein to form a complex. This complex traverses the nucleus pore and enters the nucleus to regulate the transcriptional activity of target genes and promotes the deposition of ECM (Castilla et al., 1991). Beyond the Smad pathway, TGF-β can also modulate cell behavior and matrix synthesis through non-Smad pathways (such as the MAPK pathway, RhoA-ROCK pathway, PI3K-Akt pathway, and Wnt pathway), thereby facilitating the development of fibrosis (S. Chen et al., 2019; X. Zhang et al., 2018). Therefore, TGF-β1 induces penile fibrosis by activating TGF-βRII.
Gene-based and targeted therapies have achieved remarkable progress in the treatment of cavernous fibrosis and are regarded as a promising future approach for the precise treatment of ED (Yu et al., 2018). H. B. Zhang et al. (2017) injected myocardium (Mycod) into the rat spongy tissues and subsequently conducted histological examinations of smooth muscle and phenotypic molecular tests. The findings revealed that Mycod injection maintained the contractile phenotype of smooth muscle cells in the spongy tissues, increased the ratio of smooth muscle cells to collagen, modulated fibrosis, and ultimately enhanced erectile function in rats with cavernosal nerve injury. Luan et al. (2020) established a diabetes model using homozygous transgenic rats carrying the hKLKL gene and control wild-type rats. These findings indicated that in the hKLK-carrying group, the level of apoptosis-promoting protein, as well as the expressions of TGF-β1 and collagen, were decreased, which verified the anti-fibrosis, anti-oxidative stress, and anti-apoptosis effects of hKLKL. Consequently, this treatment improved erectile function in diabetic ED rats. The abovementioned studies can effectively treat ED through their anti-fibrosis effects, which is consistent with our research findings.
Limitations
This research also has several remarkable limitations. First of all, while MR provides valuable perspectives on causal associations, it assumes a linear link between drug exposure and the relationship between exposure and outcome. However, this fails to comprehensively represent the real-world conditions of clinical trials. Second, the study’s generalizability is restricted because it mainly involves individuals of European descent. To improve its applicability, additional research and validation are essential to expand the findings to individuals with a wide range of genetic backgrounds. Third, while enrichment analysis is useful, it has its own drawbacks. Since it depends on a pre-defined set of genes or pathways, it might not incorporate all possible biological mechanisms or interactions. Fourth, the precision of molecular docking analysis is strongly reliant on the quality of both protein structures and ligands. Although this approach can pinpoint potential drug targets, it cannot ensure their clinical effectiveness. Therefore, additional experimental verification and clinical trials are indispensable for verifying the therapeutic potential of the targets that have been identified. Finally, since the absence of clinical data constitutes a shortcoming of this study, extra validation is required to ascertain the clinical importance of our findings.
Conclusions
Our study made use of MR and colocalization analysis to single out TGFBR2 as a potential drug target for ameliorating cavernous fibrosis in patients with ED. In addition, these results present promising outlooks for more efficacious ED treatment, and they may potentially cut down the costs related to drug development. The present research offers substantial contributions to the domain by highlighting the crucial role of TGFBR2 in treating ED and underscoring the need for future clinical trials of related drug targets.
Footnotes
Author Contributions
C.X. and Z.M. contributed to conception and design of the study. J.F., Y.L., and H.K. downloaded the database. Z.L. and Z.M. performed the statistical analysis. Z.L. wrote the first draft of the manuscript. Q.W. and L.Q. wrote sections of the manuscript. All authors contributed to manuscript revision and read and approved the submitted version.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was supported by the Shenzhen Science and Technology Program (grant no. JCYJ20220531092201002) and the Shenzhen Clinical Research Center for Traditional Chinese Medicine (grant no. SZCRC202508).
