Abstract
The National Animal Disease Center (NADC) conducts basic and applied research on endemic animal diseases of high priority that adversely affect US livestock production or trade. Experiments conducted at this center vary in range and scope with a subset involving synthetic or recombinant nucleic acids (DNA), microorganisms, and/or animals. Historically, the NADC used hard copy paper and filing systems to catalog and monitor these types of experiments, but to improve communication, tracking, searching, reporting, and documentation of Institutional Biosafety Committee (IBC) actions, this institution has transitioned to using a commercially available software tool to digitally manage protocols in our ongoing efforts to maintain excellence in regulatory compliance. In addition, similar to many other research institutions and universities, the scope of the IBC has expanded to include risk assessments on all work conducted at the center. This process has been streamlined using the Biosafety RAM open source software, developed by Sandia National Laboratories, and has stimulated productive discussions on best practices to safely conduct animal and microbiological experiments at the center. Although some initial challenges arose, successful implementation of these two software tools at the NADC has simplified the management of IBC compliance requirements and facilitated review processes at a high-containment government research facility.
Introduction
The National Animal Disease Center (NADC) is part of a multicenter United States Department of Agriculture (USDA) campus termed the
NADC is one of only a few similar research facilities in the world and is unique in its work with a range of large animal species, including wildlife such as bison, elk, and deer, in biosafety level-3 agriculture (BSL-3Ag) biocontainment. 1 The nature of diseases being studied requires that research activities be conducted under appropriate biocontainment and biosecurity conditions to prevent the spread of infectious diseases to nearby wildlife, livestock, poultry, and human populations. Containment facilities include laboratory and animal biosafety levels-2 and -3 (A/BSL-2, A/BSL-3) areas as well as a BSL-3Ag building 1 with flexible penning/gating emplacements that can be modified to accommodate different species and sizes of animals. The NADC is organized into four primary research units with an average of four research projects within each unit—viruses and prions, food safety and enteric pathogens, infectious bacterial diseases, and ruminant diseases and immunology. It is a world-class facility with a rich history and remarkable successes in animal disease research 2 and has been a leader in developing biosafety/biocontainment methods for livestock and poultry research.3 –6
We describe two software programs recently adopted to help streamline the workload for the NADC’s Institutional Biosafety Committee (IBC) and Institutional Animal Care and Use Committee (IACUC), although we focus on the former. We further discuss the benefits and pitfalls of implementation and use for each program. As biological safety, biosecurity, and the management and review processes that go along with a strong safety culture become subject to increasing external examination and perhaps regulation, the ability to quickly and conclusively demonstrate adequate (and even robust) review of research processes will be essential to the operational integrity of any large research organization of the future.
NADC’s Institutional Biosafety Committee
The NADC has an internal, standalone IBC to review experiments involving synthetic or recombinant nucleic acid molecules or organisms and conduct research project risk assessments per agency policies. The NADC’s IBC was formed approximately 10 years after the NIH guidelines were first published in 1976. 7 A total of five chairpersons have steered the course of 31 years of IBC operations, providing remarkable continuity. However, with the expansion of scope and complexity added to the IBC role, the chairperson duties now rotate on a more frequent basis than in the past. The committee consists of all personnel required under the current NIH Guidelines for Research Involving Recombinant or Synthetic Nucleic Acid Molecules (NIH Guidelines) to meet the needs of an animal research facility including a biosafety officer, veterinarians, researchers, and institutionally unaffiliated members from the local community. The vast majority of research conducted at the NADC, that is within the purview of the NIH Guidelines, falls under Section III-D, cloning into a Risk Group 2 agent. The IBC meets monthly and over the years has reviewed a diverse array proposed experiments. Hard-copy storage has created numerous inefficiencies when searching for a specific protocol, or when searching defined protocol topics, or even when compiling reports of currently active protocols in response to internal or external requests.
Additionally, hard copy management required all of the burdens of paper in the digital age—physical distribution of protocol packets, collection and documentation of handwritten comments, and the need to bear printing costs for documents that are only of transient use. Hard copies of protocols were not easily searchable by topic and storage became cumbersome. Ongoing scientific and technological developments (eg, gain-of-function studies, dual use research of concern, recombinant organisms with gene-drive characteristics, etc) will likely lead to a greater and greater workload for all IBCs, and an increased focus on management and review of such experiments will require sufficient documentation to easily and quickly respond to internal or external queries related to research. In addition, use of synthetic or recombinant nucleic acids or organisms in animal experiments requires review by both the IBC and IACUC and a mechanism for each committee to determine whether the requirements of the other have been met. These factors led the NADC to search for a digital solution.
IBC Protocol Management Software
In 2011, members of the NADC Senior Management Team began searching for an online management system to replace an older animal care and use protocol software program and desired a program that would also track research activities involving synthetic or recombinant nucleic acids or organisms. Efforts led by the IACUC and the IACUC compliance officer eventually identified a web-based commercial software application that would integrate our needs for an animal care and use protocol (ACUP) management software module and an IBC protocol module. After a long and arduous process to meet necessary USDA information technology (IT) security requirements, the fine-tuning of the web-based electronic protocol management system evolved over approximately a year and a half of weekly teleconferences with the vendor, the NCAH IT department, and end users to ultimately meet cybersecurity and technical institutional needs.
Specifically, the “out of the box” software had to be modified to conform to the cybersecurity and needs of the federal government relating to user access, levels of access, deployment, and data integrity/disaster recovery. Likewise, the needs of the end user or researcher included the ability to: import laboratory and barn locations where the work would be conducted, create and access institution-specific help instructions, add attachments to protocols, remove questions not applicable to this institution, develop an institution-specific protocol numbering system and animal ordering system, and other similar site-specific needs. Once all of the tailoring and modifications were completed, this version of software became known as the Research Animal Management System (RAMS).
Within RAMS, the researcher has access to the IBC or IACUC modules depending on whether the work proposed includes synthetic or recombinant nucleic acids or organisms and/or animals. The submitter is prompted to navigate through several screens, answering questions designed to capture personnel and protocol information as well as research locations. Within the IBC module, the project registration screen identifies if infectious agents, biological toxins, or synthetic or recombinant nucleic acids or organisms are to be used in the proposed experiments. Subsequent sections enable protocol details to be completed. Finally, questions relating to the NIH guidelines are asked, and then the investigator must determine the classification of the experiment according to those guidelines.
Once protocols are submitted, RAMS allows both the investigator and committee reviewer to interface during the review process. A general workflow for protocols submitted in RAMS is shown in Figure 1. Additionally, integration with IACUC is achieved through the inclusion of ACUP protocol numbers on IBC protocol submissions and vice versa. This provides an important assurance gate for reviewers to be sure that experiments conducted with animals that additionally include synthetic or recombinant nucleic acids or organisms have been considered by both the IBC and IACUC. This committee cross-checking feature was written into the specifications for the software requirements that were put out for bid by procurement staff. However, there were no vendors that offered tight integration of IACUC and IBC in a web-based format that worked on any computer platform.
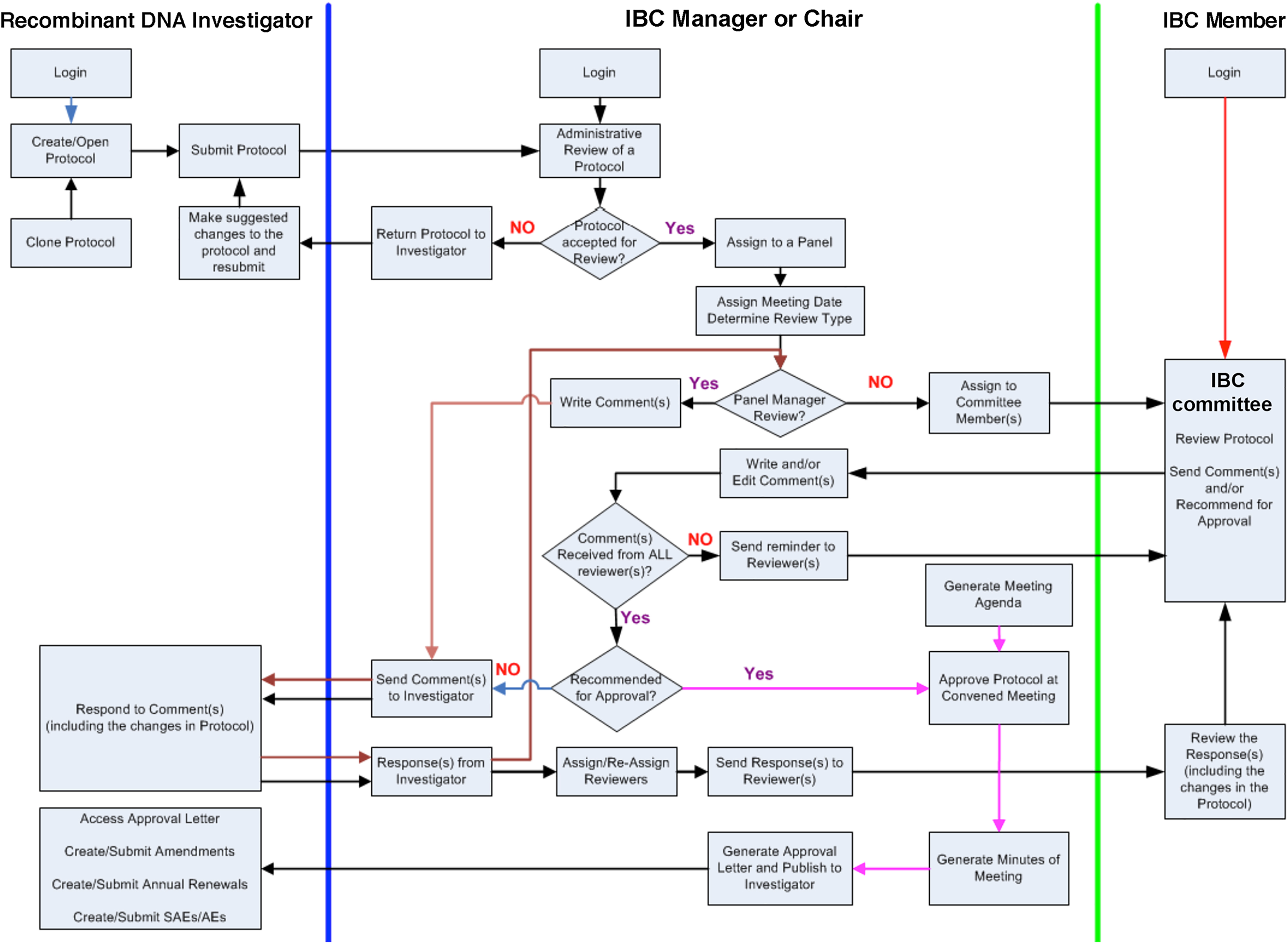
General workflow for recombinant or synthetic nucleic acid protocols submitted in Research Animal Management System (RAMS) showing the interactions of key personnel. The workflow is divided into the investigator, Institutional Biosafety Committee (IBC) manager or chairperson, and IBC member.
Collectively, this form of protocol management and review has greatly streamlined the IBC, IACUC, and investigator’s workload for IBC and IACUC submissions. On the investigator side, the ability to duplicate previously entered protocols, using a “clone” feature in RAMS, has greatly simplified the data entry process. With this clone feature, the researcher has the ability to quickly modify an existing protocol to submit as a new experiment for review by the IBC or IACUC. This feature is particularly helpful when personnel and protocol details in an existing, approved protocol are similar to those of a newly created protocol. Likewise, on the review side, each committee member can see when a protocol has been assigned to them, the deadline for completing a review, and the link to quickly access the protocol. Reviewer comments and investigator responses are permanently captured in the system and provide a rich source of documentation for how the review was conducted.
Moving to an electronic protocol management system has helped in the dissemination of protocols, documentation and timeliness of reviews, and ability to better archive and retrieve historical IBC and IACUC data in response to internal or external requests for information. This system also automatically tracks protocol renewal dates and notifies the investigator well in advance of expiration. A critical benefit is that the institutional official and other senior managers can be given direct electronic access to protocols, which was not previously possible. However, no system is perfect, and RAMS does have some issues. For example, it is not possible for the investigator to independently correct mistakes in a recently submitted protocol prior to its review by the IBC since protocols are locked once they have been routed to the committee. Instead, the IBC chairperson must be contacted for help in getting the protocol returned to the investigator for correction and resubmission. In our experience, this has happened frequently, and the process is cumbersome and frustrating to the investigator. However, despite this operational issue for the submitter, the protocol must be “frozen” after submission to preserve the integrity of the review process, so this feature must be maintained or addressed in some way. Nonetheless, it would be beneficial if the investigator retained rights to edit the protocol until the time a committee member accessed it for review (instead of upon submission and routing). Finally, there is a considerable upfront investment of time to program the software for generating automated renewal, expiration, and approval notices to investigators as well as automated committee and review assignment notices to IBC members.
Risk Assessment Software—Biosafety RAM
Performing risk assessments is a constant need in daily life activities and especially so in biological research. The NADC wanted to measure risk based on what is being done in an experiment and what potential hazards are present and catalog what methods and resources are in place to mitigate resulting risks. The goal was to perform risk assessments in a standardized and repeatable manner that could be compared across laboratories conducting research at the same institution. This process is not trivial, and a method to more easily, consistently, and comprehensively measure risk was sought.
The Biosafety RAM is an open source software package developed at Sandia National Laboratories (SNL) that enables a consistent platform for biological risk assessment regardless of the institutional environment.
8
The SNL collaborated with biosafety, infectious disease, and risk experts around the world to develop the software. The tool was designed to evaluate both the likelihood of accidental pathogen release or unintended infection as well as the consequences of such events after all institutional controls and policies have been considered. The tool only measures relative risk, not absolute risk. This software is particularly useful in several aspects of a formal risk assessment as defined by the The range and scope of the questions stimulate the respondent to fully assess all aspects of the risk in their work (agent and laboratory procedure hazards). In limiting the scope of each assessment to a research-focused area for the NADC, respondents consider who is doing the work and the specific facility features where the work is performed. By submission of the Biosafety RAM output to the IBC, review by an expert committee (inclusive of a biosafety professional) is assured. The Biosafety RAM graphical output is particularly useful for summary reviews or reviews with members of the public (should outreach initiatives be necessary).
Currently, version 2.5 of the Biosafety RAM program is used at the NADC. The assessment consists of a set of 105 questions with the investigator-provided responses collectively describing the locations, materials, procedures, and practices proposed for use. The program assigns a value to each answer provided, ranging from 0 to 4, which is used as the input for an assessment algorithm that outputs both a graphical display of the relative risks (with the likelihood of occurrence plotted against the consequence of occurrence) and a detailed, question-by-question, numerical report. All 105 responses are binned into a total of 19 categories of risk (Table 1), some pertinent to humans in the laboratory or in the surrounding community (Figure 2) and others pertinent to animals in the community. If both likelihood and consequence are scored at 1.25 or lower on the resulting scatter plot outputs (Figure 2), the biosafety risk is considered very low. Conversely, if both likelihood and consequence are plotted above 3.25, the biosafety risk is considered very high. In that case, the proposed locations, materials, practices, and procedures should be analyzed to determine which can be modified to reduce the degree of risk to an acceptable level. This analysis is facilitated by the numerical report, which provides a detailed, quantitative breakdown as to which questions feed into each risk category, the weight assigned to each question, and the resulting score based on the answer provided by the investigator.
Comparison of Analyzed Risk Categories with Graphically Displayed Risks.a
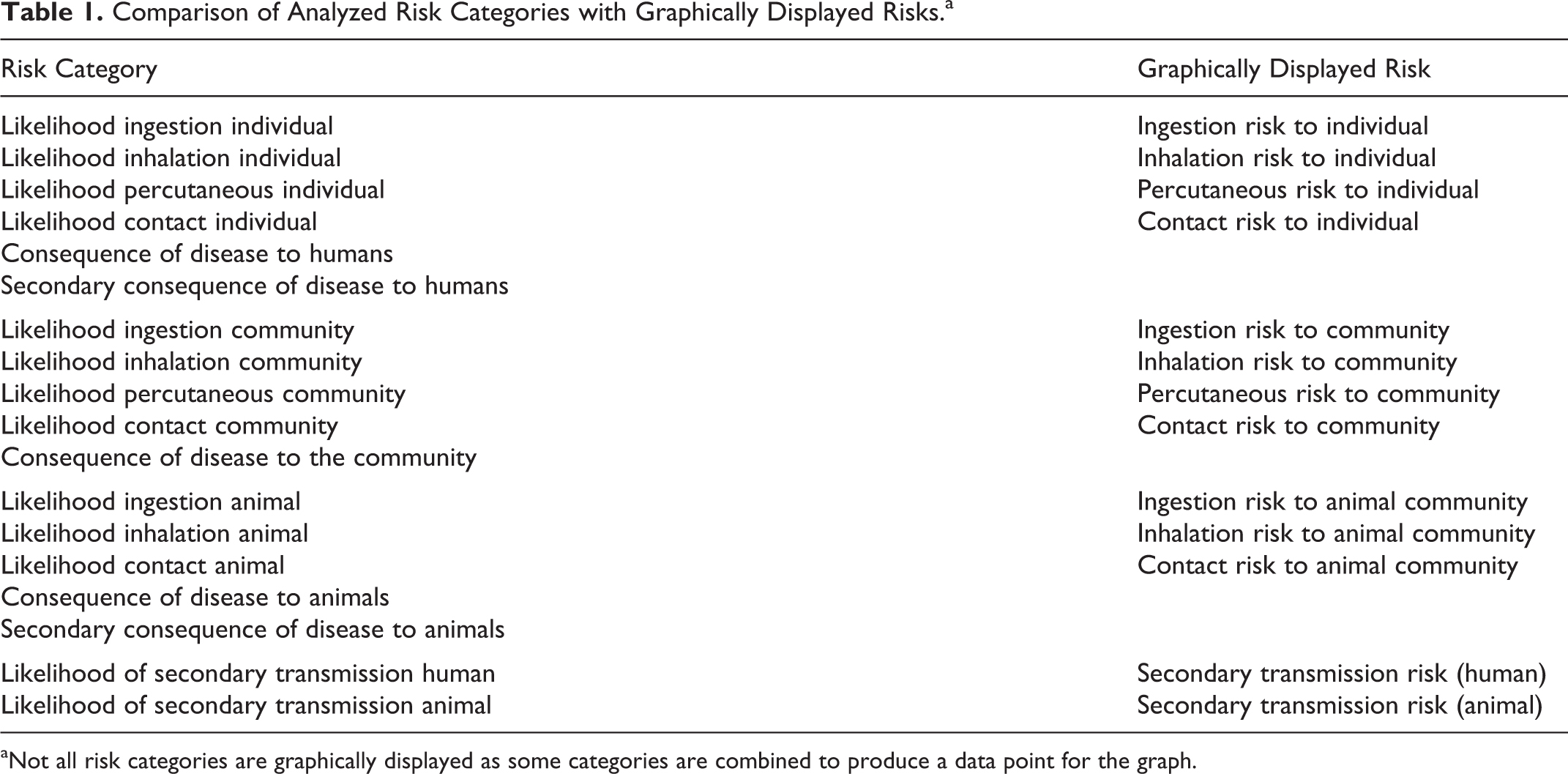
Not all risk categories are graphically displayed as some categories are combined to produce a data point for the graph.
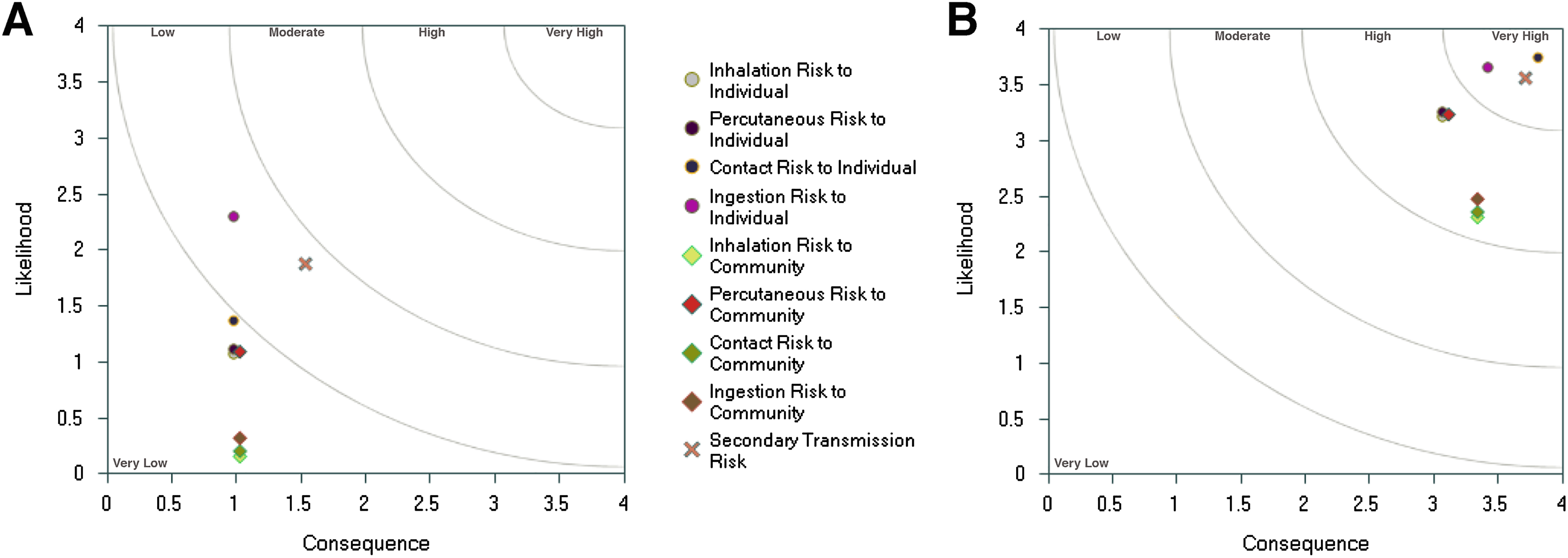
Graphical output from the Biosafety Research Animal Management System (RAMS) showing the degree of risk to humans in the laboratory and community for (A) a Risk Group 2
The Biosafety RAM tool attempts to create a framework for comprehensive risk assessment and facilitated review. The response to any question can be changed and the output reevaluated to determine if the alternative measures do indeed lower the risk of one or more categories. In this way, by breaking out the risk drivers in the algorithmic assessment and comparing different responses to related questions, the tool can be used to help researchers determine best methods to reduce biological threats in the laboratory. This allows respondents to examine what materials, practices, or facility features have the biggest influence on overall risk and target their safety and security efforts to the most efficient and effective mitigation strategies. It should be noted that the tool can also be used to determine if cumbersome control measures are excessive and unnecessary such that their elimination has little effect on risk. Caution should be used here since the determination of risk as acceptable, controllable, or unacceptable is dependent on the workers and institutional management.
The Biosafety RAM is a useful tool to greatly simplify risk assessment, but it is still a work in progress that would benefit from further refinement. For example, there appears to be a bug in the program that occasionally results in the appearance of “NaN” in one or more of the scored risk categories, which is a common abbreviation for “not a number.” This finding suggested that there was something wrong with the program’s handling of a combination of responses to questions that fed into a particular risk category. As a result, the risk could not be accurately measured for any category that showed NaN in the output. An analysis of this problem was conducted by compiling the scores for each of the 19 risk categories (Table 1) on past risk assessments submitted to the IBC. From this exercise, the problem categories that frequently gave a NaN and those categories that always provided a numerical risk score were determined. All of the responses to questions that fed into one of the selected problem categories (ingestion in animals) were then compiled. From this analysis, it appeared that certain combinations of responses led to the NaN designation for ingestion in animals category on the Biosafety RAM assessment. Although the appearance of NaN in the scoring of other categories was not comprehensively studied by us, this is a problem that will (hopefully) be addressed as newer versions are released.
Another limitation of the tool is that it does not consider the scope of the work to be performed or factor in an overall assessment of the training and competency of the personnel performing the work and/or the suitability of the facilities where the work is performed. Therefore, a full review and assessment of the Biosafety RAM output should be performed only by local officials with a high degree of detailed knowledge of the laboratory’s operations. The NADC IBC has circumvented one of these issues by requiring that investigators add a brief paragraph describing the scope of work to be performed on their risk assessment cover page. One of the value-additions of the time invested in completing such a risk assessment is to document and display a highly robust and defensible process to external reviewers; without such information, external reviewers will likely not be able to fully appreciate the safety and security provisions engaged to protect laboratory personnel and the surrounding community.
A beneficial feature of the Biosafety RAM is that the text of each question can be edited to add clarity for the respondent. For example, a question involving direct contact exposure was modified by adding the parameters of infection through intact skin or via the ocular or urogenital mucous membranes. However, this capability must be used with caution, ensuring that modifications do not affect the scoring algorithms or alter the intent of the questions. Should this occur, the assessment may be invalid. Therefore, modifications of the original text should be made only when there is a need to clarify some aspect of the question being asked. When a specific research situation does not fit neatly into the underlying assumptions of a question, the addition of a short, explanatory statement can also be helpful.
Conclusion
Inevitably, no matter how much work with the vendor is done prior to purchasing biosafety and compliance software, there will be a need to further modify either the forms or other aspects of any software package employed to manage protocol or risk assessment submission and review. Therefore, the ability to modify the software over time has been critical to the NADC’s implementation and use of these software tools.
A key benefit of the Biosafety RAM software as applied at the NADC is the inclusion of all scientists and project personnel in the risk assessment process. When each risk assessment is conducted, the engineering biosafety controls designed into a specific lab or animal biocontainment facility are considered and discussed, which serves to educate all parties involved about the overall facility features including air handling, waste stream handling controls, and personal protective equipment and other procedural methods in place to mitigate risks of the work down to an acceptable level.
Furthermore, implementation of these tools has enabled the NADC to better catalog, search, and create reports that detail risk assessments of research involving infectious agents, biological toxins, and synthetic or recombinant nucleic acids and organisms. This has enabled the center to more completely capture the scope of research actively being conducted at the NADC. The communication has also been enhanced between the IACUC and IBC committees with relevant protocol numbers from IACUC included on the IBC submission forms and vice versa. However, it is important to have a member on both committees to further facilitate communication. For NADC, this is accomplished by the biosafety manager.
Footnotes
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was funded by the USDA’s Agricultural Research Service.
