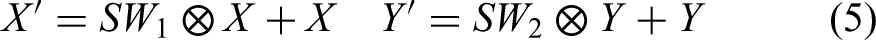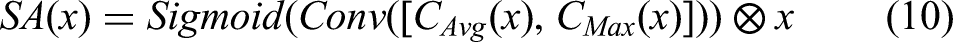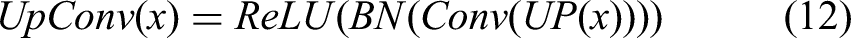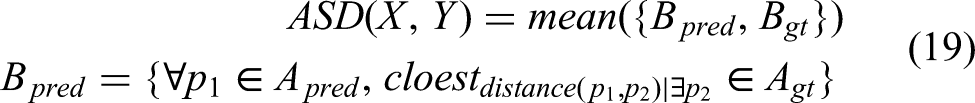Abstract
Introduction
Accurate delineation of the high-risk clinical target volume (HR-CTV) and organs at risk (OARs) is critical for cervical cancer brachytherapy. However, treatment planning is time-consuming, and prolonged waiting can lead to organ displacement and patient discomfort. Additionally, the steep dose gradients around HR-CTV amplify segmentation errors in HR-CTV and OARs. Therefore, achieving rapid and precise delineation of HR-CTV and OARs remains challenging. This study proposes a novel network model, MDA-TransUNet, for fast segmentation of HR-CTV and OARs in cervical cancer.
Methods
We applied MDA-TransUnet, a CNN-Transformer hybrid model, to segment the bladder, colon, rectum, small bowel, and HR-CTV on cervical cancer CT images. 122 cervical cancer brachytherapy patients’ CT images from three clinical centers were utilized for training and testing, with 80 cases allocated to training, 22 to testing, and 20 to external validation. Segmentation accuracy was quantified using the Dice Similarity Coefficient (DSC), Hausdorff Distance (HD95), and Average Surface Distance (ASD). Dosimetric differences were analyzed via paired t-tests.
Results
Compared to other methods, MDA-TransUnet achieved superior segmentation performance on the test dataset. The DSCs for the bladder, colon, rectum, small bowel, and HR-CTV were 94.54%, 79.27%, 79.27%, 88.90%, and 82.35%, respectively. Paired t-tests on five dosimetric metrics (D5cc, D2cc, D0.1cc, D90%, and Dmean) showed no significant differences. For OARs, the average difference in D2cc was less than 12%. For HR-CTV, the average difference in Dmean was less than 8%, and D90% was less than 11%.
Conclusion
This work demonstrates the superiority of MDA-TransUnet in segmenting OARs and HR-CTV for cervical cancer brachytherapy, with robust performance across multi-center datasets.
Introduction
Cervical cancer ranks as the fourth most common malignant tumor among women worldwide, with over 600,000 new cases annually, approximately 90% of which occur in developing countries with limited healthcare resources. 1 Its high incidence and mortality rates pose a significant threat to women's health. Precision radiotherapy, particularly combining external beam radiation therapy (EBRT) and brachytherapy (BT), is a cornerstone in treating locally advanced cervical cancer. 2 However, the unique anatomical location of cervical tumors—where high-risk clinical target volumes (HR-CTV) are closely adjacent to organs at risk (OARs) such as the bladder and rectum—necessitates a delicate balance between tumor eradication and normal tissue sparing during treatment planning.
Brachytherapy (BT), which involves placing radioactive sources directly into or near the tumor target, exploits the rapid dose fall-off with distance to deliver high local tumor doses while minimizing damage to OARs. 3 In cervical cancer treatment, BT has proven to significantly improve local control rates and patient survival. However, BT planning heavily relies on manual delineation of targets and OARs by physicians, leading to subjectivity, inefficiency, 4 and prolonged workflows. On average, radiation oncologists require 32 min to delineate HR-CTV and OARs for gynecological malignancies. 5 The demand for rapid yet precise planning creates high-pressure workflows prone to human error, while extended procedures exacerbate patient discomfort.
In recent years, with the growing adoption of deep learning, various neural network architectures—primarily convolutional neural networks (CNNs)—have been developed for segmentation in cervical cancer high-dose-rate brachytherapy (HDR-BT).6–9 The classic CNN-based segmentation model, U-Net, 10 and its variants employ symmetric encoder-decoder networks to automatically extract multi-level features through CNNs, significantly improving segmentation efficiency and accuracy.11–14 For example, Li et al 15 applied the adaptive deep CNN framework nnU-Net to segment the bladder, rectum, and HR-CTV on cervical cancer CT images. The nnU-Net method integrates three architectures—2D U-Net, 3D U-Net, and 3D cascade U-Net—to adaptively select the optimal architecture for each task. Zhang et al 16 developed a DSD U-Net model for automated delineation of the bladder, rectum, sigmoid colon, small bowel, and HR-CTV, achieving high accuracy as evaluated by the Dice similarity coefficient (DSC). Chang et al 17 proposed a hybrid network combining 3D U-Net and long short-term memory (LSTM) for HR-CTV and OAR segmentation, demonstrating superior performance over 2D U-Net.
Despite CNNs’ strengths in capturing local spatial features, they face limitations in modeling long-range dependencies between pixels. 18 Transformer architectures, enhanced by self-attention mechanisms, overcome this limitation by enabling global context modeling, showing promising potential in medical image segmentation tasks.19–23 To synergize the advantages of CNNs (local perception) and Transformers (global reasoning), hybrid CNN-Transformer architectures have emerged as a cutting-edge direction in medical image segmentation, exemplified by TransUNet. 24 Gu et al 25 pioneered the integration of Transformer's self-attention with CNN frameworks for segmenting the bladder, rectum, and colon, demonstrating significant effectiveness. However, their work focused solely on OARs segmentation in cervical cancer brachytherapy and did not extend the CNN-Transformer framework to HR-CTV segmentation.
Although deep learning-based automatic segmentation methods show promise in cervical cancer brachytherapy, existing studies still exhibit certain limitations. Firstly, most research relies on single-center datasets, struggling to adequately validate model generalizability and robustness in multi-center scenarios, and failing to effectively address challenges such as variations in imaging protocols and inconsistencies in contouring criteria across different centers. 26 Secondly, existing CNN-Transformer hybrid architectures primarily focus on the segmentation of Organs at Risk and have not yet been applied to the automatic segmentation of the High-Risk Clinical Target Volume. The HR-CTV is characterized by indistinct boundaries, variable morphology, and close proximity to OARs, making its accurate segmentation crucial for treatment planning. 27 Therefore, there is an urgent need to develop a novel segmentation method capable of simultaneously achieving precise segmentation of both HR-CTV and OARs while maintaining stable performance on heterogeneous multi-center data.
To address these challenges, this study proposes, for the first time, a CNN-Transformer hybrid network named MDA-TransUnet for segmenting both OARs and HR-CTV in cervical cancer brachytherapy. To validate the model's robustness across centers, this study integrated 122 cervical cancer brachytherapy patients’ CT images across three clinical centers for training and testing, providing robust support for evaluating the model's performance in heterogeneous scenarios. This design significantly differs from previous single-center studies and aligns more closely with clinical practical needs. Furthermore, we introduced a Multi-scale Adaptive Spatial Attention Gate (MASAG) and a Deformable Convolutional Attention Module (DCAM) into the CNN-Transformer framework to further enhance adaptability to organ deformation and multi-center variations. Compared to manual contouring, MDA-TransUnet achieved an average segmentation time of approximately 1.2 min per patient for both HR-CTV and OARs in cervical cancer CT images, representing a 26-fold speed increase. This holds promise for significantly reducing patient waiting time post-applicator insertion in clinical practice, alleviating patient discomfort, and mitigating the risk of organ displacement associated with prolonged waiting periods. MDA-TransUnet thus offers a novel technical solution for cervical cancer brachytherapy image segmentation.
Materials and Methods
Dataset
A total of 122 patients were enrolled in this Institutional Review Board (IRB)-approved retrospective study, including 52 patients from The Third Affiliated Hospital of Nanjing Medical University, 50 patients from The Affiliated Tumor Hospital of Nantong University, and 20 patients from The Affiliated Huaian NO.1 People's Hospital of Nanjing Medical University. Among these, 102 patients’ data (The Third Affiliated Hospital of Nanjing Medical University and The Affiliated Tumor Hospital of Nantong University) were utilized for training and testing, with 80 cases allocated for training and 22 cases for testing. 20 patients’ data from The Affiliated Huaian NO.1 People's Hospital of Nanjing Medical University served as an external validation set to further evaluate the model's generalization performance. This study was approved by the Medical Ethics Committee of The Third Affiliated Hospital of Nanjing Medical University (#2024KY213-01), the Medical Ethics Committee of The Affiliated Tumor Hospital of Nantong University (#2020-031), and the Medical Ethics Committee of The Affiliated Huaian NO.1 People's Hospital of Nanjing Medical University (#IIT2024101). Informed consent is waived for all participants with the approval of the Medical Ethics Committee. CT images were reconstructed using the Philips Brilliant Big Bore CT scanner (Philips Healthcare, Best, the Netherlands) with a matrix size of 512 × 512 and a slice thickness of 3 mm. All patients were treated using tandem and ovoid applicators (T + O). The HR-CTV, bladder, colon, rectum, and small bowel were manually contoured by experienced radiation oncologists (with over ten years of clinical expertise) using Monaco 5.40.01 (Elekta, Stockholm, Sweden). The radiation oncologists contoured the HR-CTV on CT images according to ICRU Report 89, with reference to MRI images obtained prior to the first brachytherapy session. 28
Data Preprocessing
The 3D-Slicer software and RT structure data were used to generate binary masks for HR-CTV and OARs for each patient. All binary masks were converted into one-hot encoded vectors with values ranging from 0 to 5. To mitigate overfitting, we implement strictly synchronized data augmentation: with 50% probability, applying random 90k° rotation (k∈{0,1,2,3}) followed by axial random flipping (horizontal/vertical); with 25% probability, performing random rotation between −20° to 20°; and with 25% probability, maintaining the identity transformation.
Network Architecture
In this section, we introduce our proposed MDA-TransUnet. Given the outstanding performance of TransUnet in medical image segmentation, we adopt TransUnet as the baseline model. For the encoder, we retain TransUnet's hybrid CNN-Transformer design: the CNN serves as a feature extractor to generate input feature maps, while the Transformer captures global and spatial relationships between features. The decoder consists of three key components: (1) UpConv blocks for feature upsampling, the Multi-Scale Adaptive Spatial Attention Gate (MASAG) to enhance feature representation, and the Deformable Convolutional Attention Module (DCAM) to robustly refine feature maps, as Figure 1.
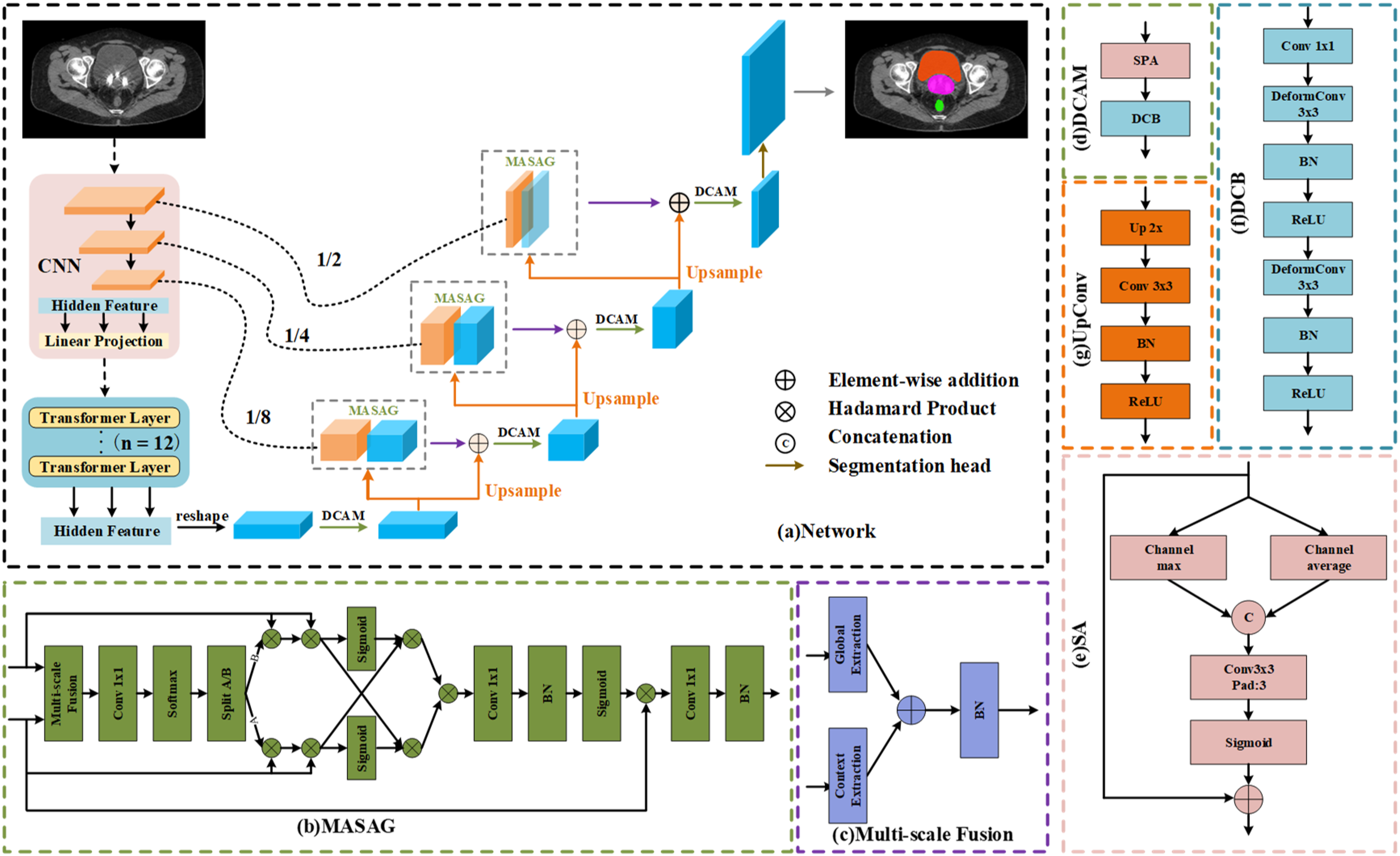
MDA-TransUnet Network Architecture Diagram. (a) Overall Framework of MDA-TransUnet, (b) Multi-Scale Adaptive Spatial Attention Gate (MASAG), (c) Multi-Scale Feature Fusion (MSF), (d) Deformable Convolutional Attention Module (DCAM), (e) Spatial Attention (SA), (f) Deformable Convolutional Block (DCB), (g) UpConv.
Multi-Scale Adaptive Spatial Attention Gate (MASAG)
We employ the Multi-Scale Adaptive Spatial Attention Gate (MASAG) module to enhance feature representation. MASAG aims to effectively integrate multi-scale information and guide the aggregation of spatial features, thereby improving overall segmentation performance. 29 Through a four-stage collaborative process—Multi-scale Fusion (MSF), Spatial Selection (SS), Spatial Interaction and Cross-Modulation (SICM), and Recalibration (RC)—MASAG progressively optimizes feature representations, addressing limitations of traditional methods in cross-scale information fusion and spatial weight allocation. The MASAG framework comprises the following stages:
Multi-Scale Feature Fusion (MSF)
The encoder feature maps (X) and decoder feature maps (Y) are fused through local and global context extraction branches:
Generates a two-channel weight map, as Equation (4):
Applies spatial weighting to encoder features (X) and decoder features (Y) separately, as Equation (5):
Spatial Interaction and Cross-Modulation (SICM)
This module facilitates feature interaction across spatial locations and enables cross-channel modulation of multi-scale information:
Encoder features are modulated by decoder global context, while decoder features are modulated by encoder local details, addressing the misalignment of low-level and high-level features in traditional U-Net skip connections, as Equation (6):
Final fused features are integrated via element-wise multiplication, as Equation (7):
The mutually modulated X'’ and Y'’ are fused via element-wise multiplication to generate the integrated feature U’. This multiplicative fusion more effectively highlights regions deemed significant by both feature maps.
Recalibration (RC)
Finally, the recalibration module adjusts feature map responses to emphasize meaningful information, as Equation (8):
This step refines the fused feature map through pointwise convolution, which is then activated by a sigmoid function to generate the attention map. Finally, the initial input X is recalibrated by performing element-wise multiplication with the attention map, followed by further processing through another pointwise convolution. The recalibrated feature map
Deformable Convolutional Attention Module (DCAM)
We employ the Deformable Convolutional Attention Module (DCAM) to enhance the network's adaptability to irregular shapes, size variations, and geometric deformations in medical imaging, challenges particularly prominent in cervical cancer brachytherapy segmentation tasks. By integrating the context-aware feature refinement of the Spatial Attention (SA) mechanism and the dynamic geometric adaptation of the Deformable Convolution Block (DCB), DCAM overcomes the limitations of conventional convolutional operations in modeling anatomical diversity. The DCAM module combines SA and DCB, as defined in Equation (9):
Spatial Attention (SA)
The Spatial Attention mechanism identifies and amplifies critical spatial regions in feature maps, emphasizing high-signal areas (eg, HR-CTV boundaries) while suppressing low-contrast or irrelevant regions and noise interference, as formulated in Equation (10):
Deformable Convolution Block (DCB)
We introduce deformable convolution to further refine features generated by SA and enhance the network's adaptability to geometric deformations—crucial for addressing anatomical heterogeneity in multi-center datasets. Unlike the fixed grid sampling in traditional convolution, deformable convolution dynamically adjusts kernel sampling positions by learning offset parameters, enabling the kernel to adaptively “warp” and align with irregular anatomical contours (eg, HR-CTV or colon boundaries), as detailed in Equation (11):
Upsampling (UpConv)
The UpConv layer progressively upsamples the features of the current layer to align the dimensions with the subsequent skip connection. Each UpConv layer consists of an UpSampling
Loss Function
Our method employs a multi-scale supervision strategy and a hybrid loss function to optimize model performance and mitigate class imbalance. The model generates predictions at four decoder levels (p1, p2, p3, p4), with each level contributing to the loss calculation. This approach reduces reliance on single-scale features by integrating multi-scale contextual information. The loss at each level is a weighted combination of Cross-Entropy Loss and Dice Loss, as defined in Equation (13):
Here,
The final loss is the weighted sum of losses from all four decoder levels, with equal weights assigned
Evaluation Metrics
Both geometric and dosimetric methods were employed for quantitative analysis. Geometric performance was assessed using the Dice similarity coefficient (DSC), defined as:
Here, smaller HD95 (95% Hausdorff Distance) and ASD (Average Surface Distance) indicate better shape agreement between segmentation results and ground truth contours, while a larger DSC (Dice Similarity Coefficient) reflects higher spatial overlap with the ground truth.
For dosimetric comparisons, dose-volume indices (DVIs) were utilized. For HR-CTV, we focused on Dmean (mean dose to the target) and D90% (minimum dose covering 90% of the target volume). For OARs, we evaluated D5cc, D2cc, and D0.1cc, where DXcc denotes the minimum dose to the hottest X cubic centimeters (cc) of the organ. Paired sample t-tests were performed to compare dosimetric differences, with p < 0.05p < 0.05 indicating statistical significance. All statistical analyses were conducted using Python 3.8.
Experiments and Results
Experimental Details
MDA-TransUnet was implemented in PyCharm on a computer equipped with an Intel® Core™ i7-10700 CPU and an NVIDIA GeForce RTX 3090 GPU. For fair comparison with other methods, all networks were trained under identical configurations. The model was optimized using the AdamW optimizer, which is better suited for complex multi-scale feature learning and improves convergence stability. The learning rate was adjusted via a cosine decay schedule, starting with an initial value of 0.0001 and a minimum of 1e-7. The batch size was set to 24, and training proceeded for 150 epochs.
Geometric Metric Analysis
As shown in Table 1, our proposed MDA-TransUnet achieves the highest Dice Similarity Coefficient (DSC) across all five target regions compared to other methods. The mean DSCs for the bladder, colon, rectum, small bowel, and HR-CTV are 94.54%, 79.27%, 79.27%, 88.90%, and 82.35%, respectively. Notably, for the small bowel (88.90%) and HR-CTV (82.35%), our method significantly outperforms suboptimal approaches (Trans-CASCADE: 87.55% for small bowel; EMCAD: 81.49% for HR-CTV). Regarding boundary precision, our method achieves the lowest average HD95 (95% Hausdorff Distance) among all OARs. For the bladder, small bowel, and HR-CTV, HD95 values are significantly reduced compared to suboptimal methods, demonstrating enhanced boundary control. In terms of Average Surface Distance (ASD), MDA-TransUnet delivers optimal ASD values for all OARs except the colon, with the bladder and HR-CTV showing an 8.5% reduction in ASD versus the second-best method. Compared to the classic TransUnet, our method achieves DSC improvements of 5.95% (bladder), 14.7% (colon), 9.05% (rectum), 11.67% (small bowel), and 4.93% (HR-CTV). These results validate that the Multi-Scale Adaptive Spatial Attention Gate (MASAG) and Deformable Convolutional Attention Module (DCAM) effectively mitigate limitations of skip connections while enhancing the network's ability to model local details and geometric deformations. Furthermore, Figures 2 and 3 visually compare segmentation results, confirming that contours generated by our method exhibit superior agreement with radiation physicist-delineated ground truth in shape, volume, and spatial localization.
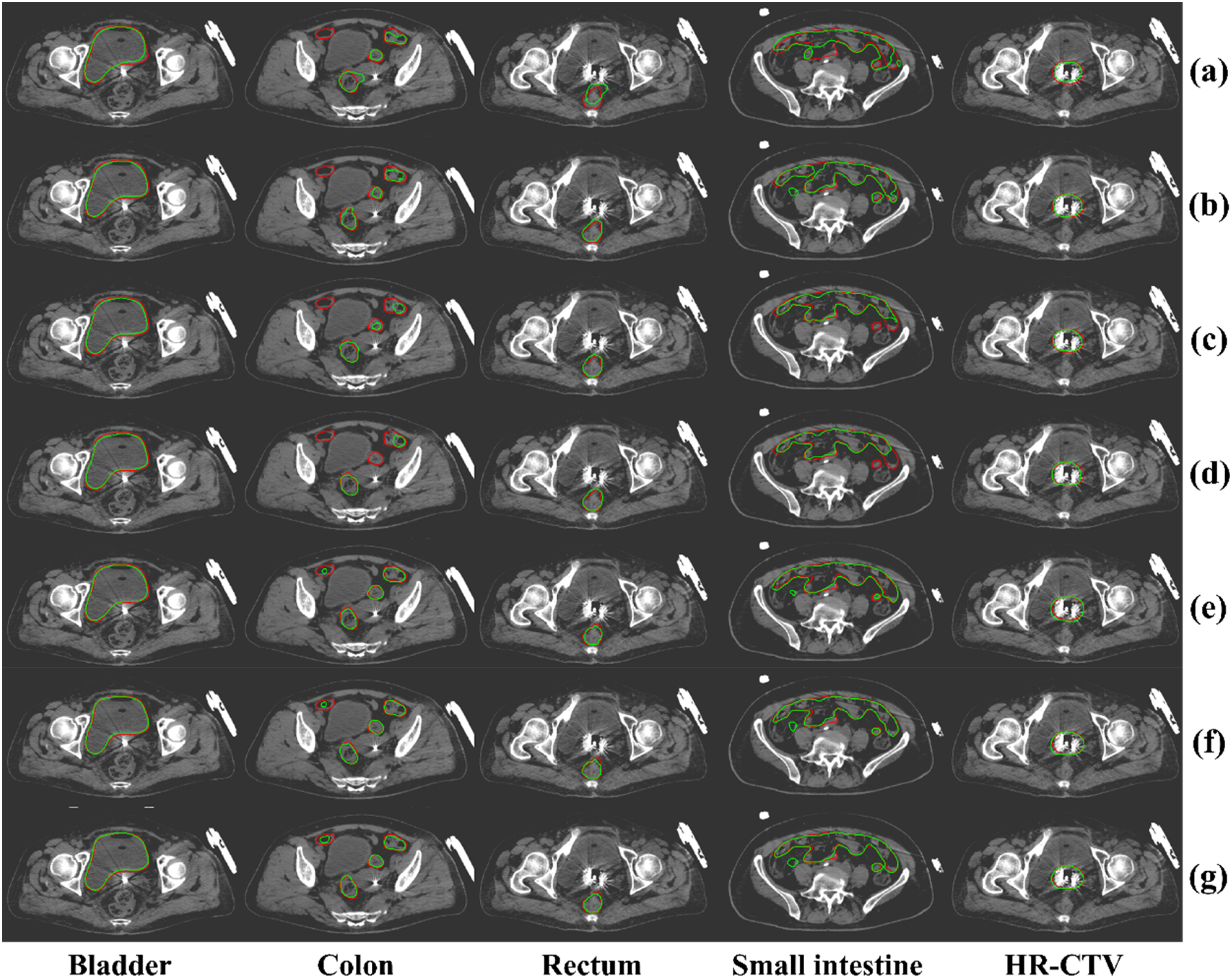
Visual Comparison of our Method with Other Methods, Where red Indicates the Ground Truth and Green Represents AI-Based Segmentation Results. Subfigures Correspond to: (a) TransUnet, (b) DLKA-net, (c) SelfRag-Unet, (d) Unet, (e) Trans-CASCADE, (f) EMCAD, and (g) our Method.
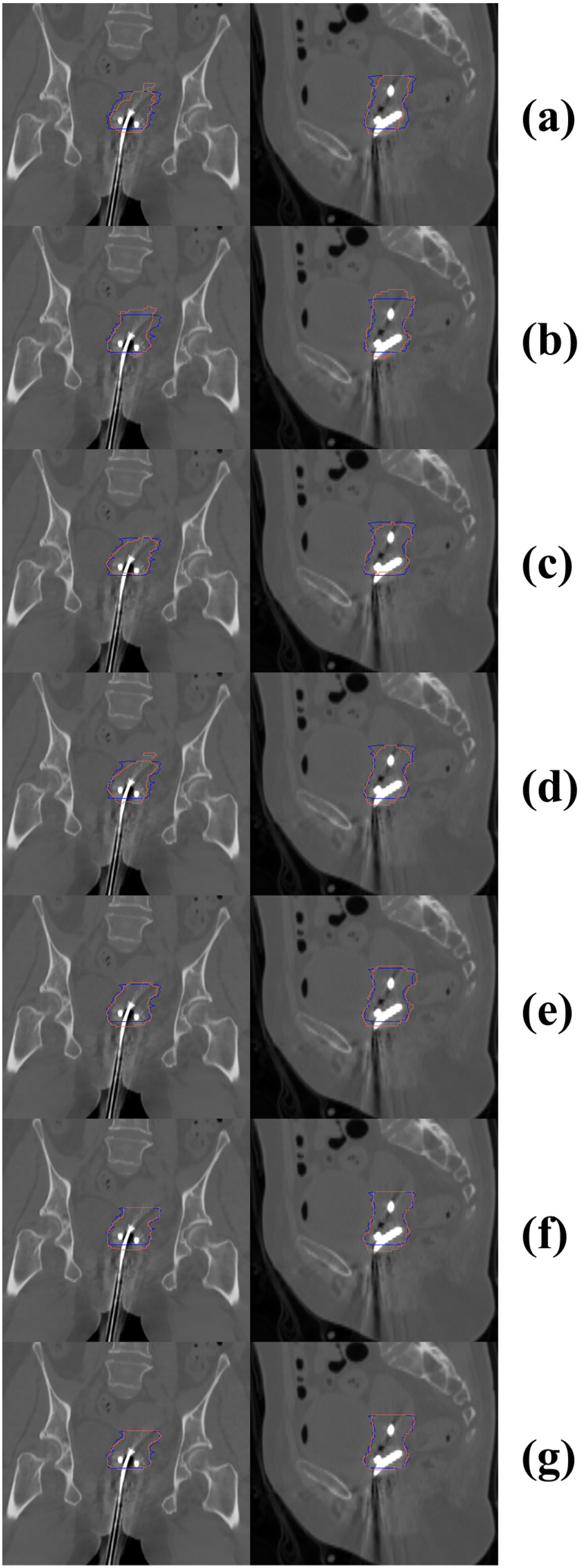
Visual Comparison of our Method Versus Other Approaches for HR-CTV Segmentation in Sagittal and Coronal Planes. Blue Contours Denote Ground Truth, While red Contours Represent AI-Generated Segmentations. Left Panels: Coronal Views; Right Panels: Sagittal Views. (a) TransUnet, (b) DLKA-Net, (c) SelfRag-UNet, (d) UNet, (e) Trans-CASCADE, (f) EMCAD, and (g) our Method.
Quantitative Comparison Between our Proposed Method and Other Methods, Where “our Study” Denotes our Approach.

Ablation Study
Ablation studies were conducted on the dataset to evaluate the effectiveness of different components in our proposed network. We incrementally removed key modules (MASAG and DCAM) and compared the results. As clearly demonstrated in Table 2, incorporating both MASAG and DCAM modules significantly enhances performance, particularly for HR-CTV segmentation, mitigating performance degradation caused by inter-center variations in contouring protocols and imaging parameters. The MASAG module improves feature representation quality and enhances cross-scale information fusion accuracy, thereby providing more reliable and focused feature inputs to DCAM. The deformable convolution operations in DCAM enable precise geometric adaptation at critical regions emphasized by MASAG (eg, organ boundaries). The resulting deformation-adapted features subsequently deliver more anatomically realistic information to MASAG during later decoding stages. This cascaded processing creates powerful synergies, proving particularly effective when segmenting challenging structures such as morphologically variable colons and boundary-ambiguous HR-CTV. The introduction of MASAG and DCAM collectively strengthens the model's capacity to capture anatomical details, accommodate multi-center heterogeneity, and handle geometric variations, achieving superior performance across all segmentation regions.
Ablation Study.

Dosimetric Analysis
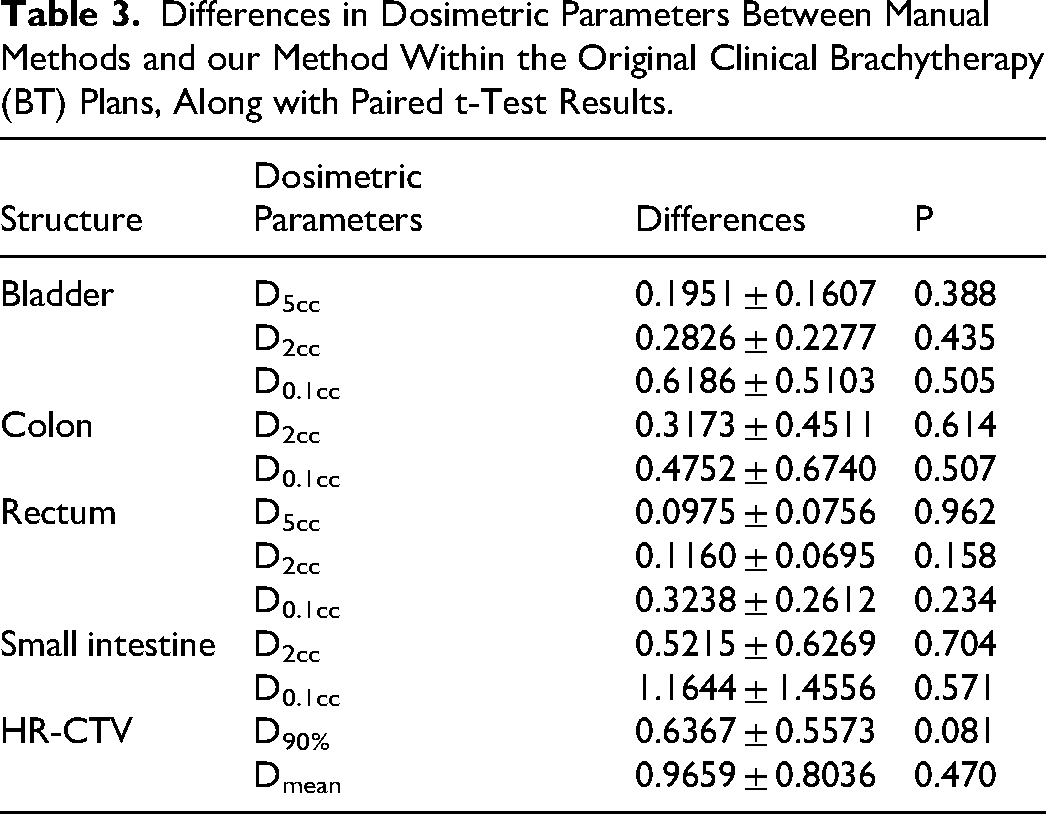
To evaluate dosimetric accuracy, we compared dose-volume metrics between manual and deep learning-based methods using paired sample t-tests, as summarized in Table 3. No statistically significant differences were observed across all dosimetric parameters. For OARs, the average differences in D2cc and D0.1cc were less than 12% and 15%, respectively. For HR-CTV, the average differences in Dmean and D90% were below 8% and 11%, respectively. Figure 4 illustrates dose distributions for a representative case, comparing our automated segmentation method with manual delineation.

Delineation of HR-CTV and OARs on CT Slices and Corresponding Dose Distribution Results. Blue Regions Represent the Standard Manual Contours; Green Lines Denote the Contours Segmented by our Method, and Colored Lines Indicate Dose Distributions.
Differences in Dosimetric Parameters Between Manual Methods and our Method Within the Original Clinical Brachytherapy (BT) Plans, Along with Paired t-Test Results.
External Validation
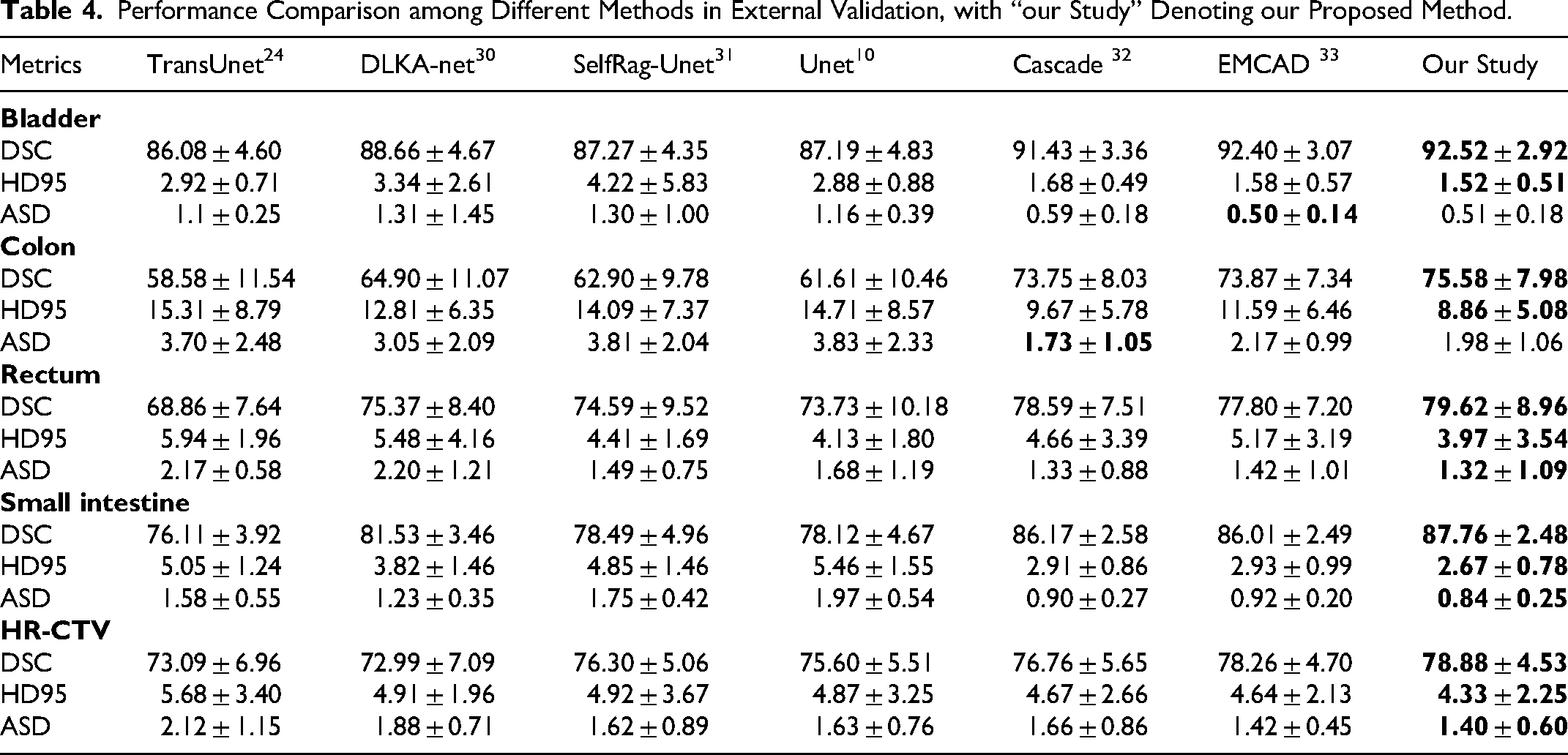
To evaluate the model's generalization capability on unseen data from medical institutions, this study additionally incorporated 20 cervical cancer brachytherapy patients’ CT images from The Affiliated Huaian NO.1 People's Hospital of Nanjing Medical University for independent testing (exclusively for external validation, not involved in training). As demonstrated in Table 4, MDA-TransUnet maintained optimal performance in the external validation set, achieving the highest DSC and HD95 values across all five target regions. Significant improvements were observed particularly for more challenging OARs (colon, rectum, and small bowel) compared to suboptimal methods. Although yielding second-best ASD values for the bladder and colon, it achieved the optimal average ASD across all target regions. These results demonstrate that the synergistic interaction between MASAG and DCAM effectively mitigates impacts caused by inter-center scanner variations and annotation discrepancies, confirming MDA-TransUnet's superior generalization capability.
Performance Comparison among Different Methods in External Validation, with “our Study” Denoting our Proposed Method.
Model Parameters
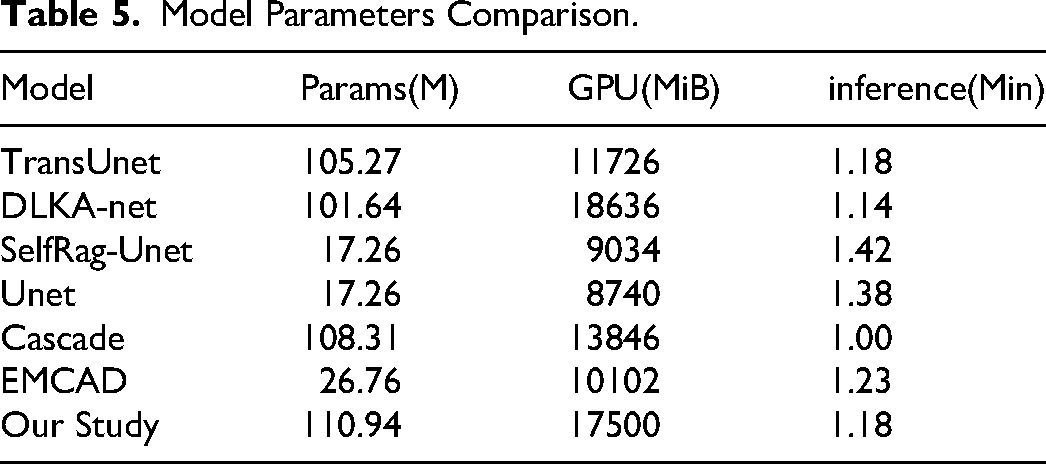
As presented in Table 5, MDA-TransUnet exhibits higher parameter counts and GPU memory consumption compared to other models, primarily attributed to increased computational complexity from the MASAG and DCAM modules, along with inherent requirements of the TransUnet encoder architecture. However, this computational overhead delivers substantial performance gains—particularly in handling multi-center data where it demonstrates superior generalization capability. While UNet and SelfRag-UNet achieve the lowest parameter/GPU footprints, their segmentation accuracy lags significantly behind MDA-TransUnet. Regarding inference speed, MDA-TransUnet matches similarly-sized models (TransUnet and DLKA-Net) while outperforming the smaller EMCAD architecture. Crucially, the 26-fold acceleration in segmentation reduces patient waiting time, mitigates organ displacement risks, and alleviates patient discomfort—demonstrating substantial clinical significance.
Model Parameters Comparison.
Discussion
The combination of external beam radiation therapy and high-dose-rate brachytherapy represents the standard of care for gynecologic cancer treatment, where HDR-BT has proven indispensable and strongly correlates with improved survival rates. Compared to conventional EBRT, brachytherapy's defining feature is its ability to deliver higher radiation doses to tumor regions near the radiation source while effectively sparing normal organs due to rapid dose fall-off. However, brachytherapy faces unique challenges. In HDR-BT, treatment planning must be completed rapidly after applicator insertion, often under time constraints that may introduce human errors. In recent years, deep learning-based methods have emerged as promising solutions to automate workflows, reduce patient wait times, and enhance comfort.
The purpose of this study is to investigate deep learning-based automatic segmentation methods for cervical cancer brachytherapy CT images. The proposed MDA-TransUnet achieves superior Dice Similarity Coefficient (DSC) across all target regions compared to other methods (Table 1). Our method performs best in bladder segmentation, largely attributed to its relatively regular anatomical structure and high contrast, facilitating clear boundary feature extraction. Compared to the bladder, the rectum, colon, and small bowel present greater segmentation challenges due to their anatomical and imaging characteristics. While the rectum maintains relatively stable positioning, its low contrast on CT scans makes boundary delineation difficult. The colon presents exceptional segmentation challenges due to its characteristically low contrast against surrounding adipose and soft tissues in CT images, combined with high deformability in both shape and position. The small bowel's position is substantially influenced by respiratory motion, peristalsis, and bladder/rectal filling status, while its appearance on CT images often blends with other soft tissues, complicating segmentation. Despite these challenges, MDA-TransUnet achieved the highest DSC scores for all three organs: 79.27% for colon, 79.27% for rectum, and 88.90% for small bowel. Gu et al 25 first combined CNN with Transformer in MFFUNet for segmenting brachytherapy OARs (bladder, colon, rectum), reporting DSCs of 92.65%, 61.86%, and 66.55% respectively. Comparatively, our MDA-TransUnet demonstrates significantly better performance on colon and rectum segmentation. In future work, we will explore more effective contrast enhancement techniques to further improve segmentation performance for both the colon and rectum. Additionally, the HR-CTV segmentation achieved 82.35% DSC, outperforming other methods despite variations in delineation protocols across centers.
Ablation studies (Table 2) demonstrate that incorporating MASAG and DCAM modules significantly enhances performance across all target regions. Particularly for structurally complex areas like colon, small bowel, and HR-CTV, DSC improved by an average of 2.48%. This validates the modules’ effectiveness in enhancing anatomical detail capture while mitigating cross-center variability, ensuring robust performance in multi-center scenarios.
Beyond geometric metrics, dosimetric evaluation remains crucial for automated segmentation. 34 Paired t-tests (Table 3) confirmed no statistically significant differences between our automated method and manual delineation across five dosimetric parameters: D5cc, D2cc, D0.1cc, Dmean, and D90%. Notably, despite poorer geometric accuracy in colon segmentation, dosimetric agreement with manual methods remained high, consistent with Wang et al's findings. 35 Dosimetric parameters like D2cc (representing the minimum dose received by the maximally irradiated 2cm3 of a volume) primarily depend on segmentation accuracy in high-dose regions (typically near applicators or targets). Sufficient overlap between automatic segmentation and ground truth in these critical areas can yield comparable dosimetric outcomes, even when contour discrepancies exist in low-dose regions. The Dice Similarity Coefficient (DSC) measures global volume overlap - shape variations in geometrically complex structures within low-dose, non-critical regions may reduce DSC without materially affecting dosimetry. Consequently, both geometric and dosimetric metrics are essential for comprehensive evaluation of automatic segmentation methods. Nevertheless, continuous optimization for improved geometric accuracy (particularly in high-dose regions) remains crucial, potentially further minimizing dosimetric differences while enhancing clinicians’ trust in automated results and overall model reliability.
To further assess generalization capability, 20 cervical cancer brachytherapy patients’CT images from The Affiliated Huaian NO.1 People's Hospital of Nanjing Medical University were independently tested (external validation). As shown in Table 4, MDA-TransUnet achieved optimal DSC and HD95 values across all five targets (bladder, colon, rectum, small bowel, HR-CTV) on this unseen dataset. For the challenging colon segmentation, it outperformed suboptimal methods by 1.74% in DSC and reduced HD95 by 0.81 mm. These results demonstrate robust adaptability to multi-center heterogeneity (eg, varying CT protocols and contouring practices). Though yielding suboptimal ASD for bladder and colon, the model achieved optimal average ASD across all targets, confirming boundary segmentation accuracy. Compared to other models, MDA-TransUnet exhibits superior performance stability and generalization capability on independent cross-center data. Our approach - incorporating data augmentation, multi-center training, and robustness-enhancing modules (MASAG, DCAM) - effectively mitigates inter-center annotation variations. Future efforts should establish refined contouring guidelines and cross-institutional consistency reviews to standardize ground truth.
This study has limitations. First, obtaining fully annotated multi-center cervical cancer brachytherapy CT datasets remains challenging, with inter-center variations in region-of-interest delineation. Second, the dataset remains relatively limited. Though multi-center data and independent external validation were employed, small sample sizes may introduce potential bias and constrain model learning capacity. Finally, while achieving excellent segmentation performance, MDA-TransUnet's high parameter count and computational complexity may hinder clinical deployment in resource-constrained settings. Future work will: (1) Expand datasets with multi-center cases, (2) Develop lightweight model compression strategies to enhance computational efficiency, and (3) Optimize clinical applicability.
Conclusions
This study proposes MDA-TransUnet, a deep learning method leveraging a CNN-Transformer hybrid architecture to automate the segmentation of HR-CTV and OARs in planning CT images for cervical cancer brachytherapy. Evaluation results demonstrate that our method outperforms other state-of-the-art (SOTA) approaches and exhibits no statistically significant differences from manual delineation across five dosimetric metrics. The proposed automated segmentation framework facilitates the automation of cervical cancer brachytherapy workflows, enhances treatment consistency, reduces post-applicator-insertion waiting times, and alleviates patient discomfort.
Footnotes
Abbreviations
Acknowledgements
Not applicable.
Ethics Approval and Consent to Participate
This study was approved by the Medical Ethics Committee of Changzhou No.2 People's Hospital Affiliated to Nanjing Medical University (Approval number: 2024KY213-01), the Medical Ethics Committee of Tumor Hospital Affiliated to Nantong University (Approval number: 2020-031) and the Third Affiliated Hospital of Nanjing Medical University (Approval number: ITT2024101). The requirement for informed consent was waived for all participants with the approval of the Medical Ethics Committees. The research was conducted in accordance with the principles embodied in the Declaration of Helsinki and local statutory requirements. All authors have reviewed research data.
Consent for Publication
Not applicable.
Author Contributions
CDZ conceived the study, designed the experiments, analyzed the data and wrote the manuscript. Data were collected by JJH, HJH, YB, STT, KY. NXY edited the manuscript. LC, QCJ, and XK supervised the data analysis. NXY reviewed literature, contributed to the manuscript and acquired the financial support for the project. ZH, SLT, HQJ, and CDZ performed the statistical analyses. All the authors accessed the study data and reviewed and approved the final manuscript.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work is supported by the National Natural Science Foundation of China (No.62371243), Jiangsu Provincial Medical Key Discipline Cultivation Unit of Oncology Therapeutics (Radiotherapy) (No. JSDW202237), Changzhou Social Development Program(Nos. CE20235063 and CJ20244020), and Postgraduate Research & Practice Innovation Program of Jiangsu Province (No. JX13614239).
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data Availability
The datasets generated and/or analyzed during the current study are not publicly available due to protection of individual patient privacy and the use of an in-house software but are available from the corresponding author on reasonable request.