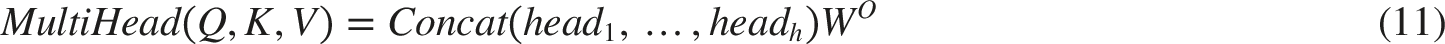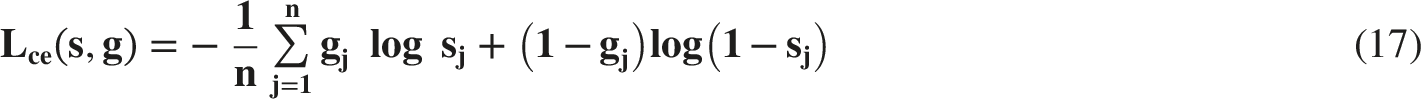Abstract
Purpose
A lightweight deep learning network SMA-Net was proposed to intelligently segment the skeletal muscle of the third lumbar (L3) level in patients with cervical cancer radiotherapy, and the segmentation performance of the network was evaluated.
Methods and Materials
A total of 160 eligible patients with cervical cancer admitted to the oncology department of our hospital from September 2021 to June 2024 were randomly divided into training set (N = 112), validation set (N = 16) and test set (N = 32) according to 7 : 1 : 2. The lightweight Mamba architecture is introduced into the UNet network, and the SAB and CAB attention mechanisms are introduced on the skip connection. The attention mechanism is used to suppress the irrelevant information in the image and highlight the important local features. The trained network is geometrically evaluated on the test set for segmentation performance, comparison of manual segmentation and predicted skeletal muscle area (SMA). Compare the parameters and computations of SMA-Net with existing networks.
Results
The dice similarity coefficient of SMA-Net network for skeletal muscle segmentation was 89.16%, the sensitivity SEN was 88.21%, the positive predictive value PPV was 90.13% and the 95% Hausdorff distance was 5.30mm. Manual segmentation is basically close to the predicted SMA. Our proposed network for cervical cancer patients predicted sarcopenia with 87.5% accuracy, 92.31% precision, 80% recall, 85.72% F1-Score, and 0.871 AUC. The calculation amount of SMA-UNet network is 1.50 GFLOPS, and the parameter amount is 1.24 M. The radiologist’s scores show that minor and no revision accounted for 93.75% on manual revision of skeletal muscle.
Conclusion
The lightweight SMA-Net proposed in this study can accurately segment L3 skeletal muscle and quickly calculate its area, which basically meets the clinical application and is convenient for clinical deployment. It is helpful for clinicians to quickly diagnose sarcopenia in patients with cervical cancer, save medical resources, reduce the workload of physicians, and improve diagnostic efficiency.
Introduction
Cervical cancer is a malignant tumor affecting the female genital system and ranks as fourth most common neoplasia in women worldwide. 1 Neoadjuvant chemotherapy and surgery or induction chemotherapy followed by chemoradiotherapy had been the standard of care in the management cervical cancer.2,3 Despite notable advancements in radiotherapeutic and chemotherapeutic approaches for cervical cancer, the tumor target and pelvic lymph regional leaking in the high radiation field also irradiate the bowel within the pelvic cavity and damage the bowel mucosa. Patients may compromise food intake, digestion, and nutrient absorption, hence aggravating the nutritional status of the patient. 4 These negative effects, such as weight loss, nutrient malabsorption, and even body composition changes including skeletal muscle5,6 ultimately lead to adverse clinical outcomes: treatment interruptions, infections, and longer hospital stays. 7 Sarcopenia is often defined as a reduction in skeletal muscle mass (SMM). Its occurrence and progression seriously affect the quality of life and mortality of cancer patients. 8 The skeletal muscle index (SMI) measured at the third lumbar vertebral (L3) level by computed tomography (CT) is a clinical accepted method for diagnosing sarcopenia. 6 SMI is the skeletal muscle area (SMA) normalized to the square of height and expressed in cm2/m2. SMI calculation requires the oncologists to manually segment the SMA. Accurate manual segmentation is not only time-consuming, labor-intensive but also prone to intra- and inter-observer variations among oncologists with different experience levels. In recent years, the advances in deep learning and computing resources provide novel opportunities to revisit these types of manual, time-consuming and routine tasks. In particular, deep learning has been shown to be particularly well suited to semantic segmentation tasks. 9 Thus, it might be both reasonable and necessary to develop a quick and effective deep learning-aided tool for automatically segmentation of skeletal muscle.
Although deep learning technology has made breakthroughs in the field of medical image segmentation (such as U-Net and its variants),10,11 It is still challenging to segment the skeletal muscle of cervical cancer automatically using DL because of the complexity of anatomic structure. Amarasinghe et al 12 constructed a 2.5 D UNet network to segment the L3 skeletal muscle of CT images for non-small cell lung cancer and acquired a mean Dice score of 0.92. Naser et al 13 used 2D and 3D ResUNet to intelligently segment the skeletal muscle at the third cervical vertebra level, and their Dice score reached 0.95 and 0.96, respectively. The automatic segmentation of skeletal muscle in radiotherapy CT for cervical cancer patients still faces unique challenges. The pelvic anatomical structure is affected by tumor invasion and radiotherapy edema, resulting in blurred muscle boundaries. In addition, fat infiltration caused by chemoradiotherapy may change the distribution of CT values of muscle tissue. These intelligently segmented convolutional neural networks (CNN) can only extract local features. The parameter of network calculation is too large and the calculation cost is high, which is not conducive to the clinical deployment and application of medical equipment. Recently, some researchers have introduced the state space model represented by Mamba into the classical UNet, which overcomes the limitation of the long-distance modeling ability of convolution, and achieves the dual advantages of high segmentation accuracy and low computational cost on the skin lesion datasets. 14 We thus proposed a lightweight network to segment skeletal muscle for cervical cancer radiotherapy using planning CT.
Materials and Methods
Study Design
Data of 160 cervical cancer radiotherapy patients from 2021-2024 at the our hospital were collected. The patients were manual annotated with skeletal muscle on L3 level. The planning CT images were obtained using a Brilliance CT Big Bore (Philips Healthcare, Best, Netherlands) according to a standard clinical acquisition protocol: 120 kVp, 120–300 mAs, 0.8×0.8 pixel spacing, 512×512 image size and 3 mm slice thickness. A series of curation and preprocessing procedures were performed to ensure that all data met the quality and standard requirement of the segmentation and predictive models.
Image Segmentation and Preprocessing
A total of 26246 CT scan slices were manually segmented for the training of the automated skeletal muscle segmentation network. Manual segmentation was finished by a junior radiologist in Eclipse 13.6 treatment planning system (Varian Medical System Inc, USA). The muscle and adipose tissue were separated by ‘Region-Growing’ tool through the Hounsfield unit (HU) thresholds −29 to +150 HU
15
for body composition. The finial manual segmentation results were reviewed and approved by a senior radiologist (above 20 years working experience). Skeletal muscle, including rectus abdominis, abdominal wall muscles, quadratus lumborum, psoas and erector spinae was segmented. Figure 1 shows the detail skeletal muscle segmentation in CT images. The augmentation techniques included random transformations, scale intensity range, random cropping (320×320), random flip (P=0.5), random rotation (±10°), and random intensity shift ([0,1]). Detailed segmentation diagram of the skeletal muscle
Development of Lightweight Segmentation Network
An illustration of our research workflow is shown in Figure 2. The main processes include six components: skeletal muscle manual segmentation in TPS, a preprocessing operations before models training, comparison of model prediction, and three-dimensional reconstruction. The evaluation includes quantitative geometric metrics and clinical assessment. The existing lightweight semantic segmentation networks usually utilize depth-wise, group-wise and factorized-wise to compress network size. Recently, the State Space Models (SSMs) represented by Mamba were introduced into the classical UNet convolutional neural network, which overcomes the limitations of the long-distance modeling ability of convolution, and achieves the dual advantages of high segmentation accuracy and lightweight network on the skin lesion dataset.
14
Ruan et al
16
firstly developed a medical image segmentation model based on Mamba architecture for the segmentation of skin diseases. Wu et al
14
analyzed the key factors in the influence of parameters in the Mamba architecture, and proposed a parallel visual Mamba layer called UltraLight VM-UNet to achieve the lowest computational load and excellent performance while keeping the total number of channels constant. The workflow of this work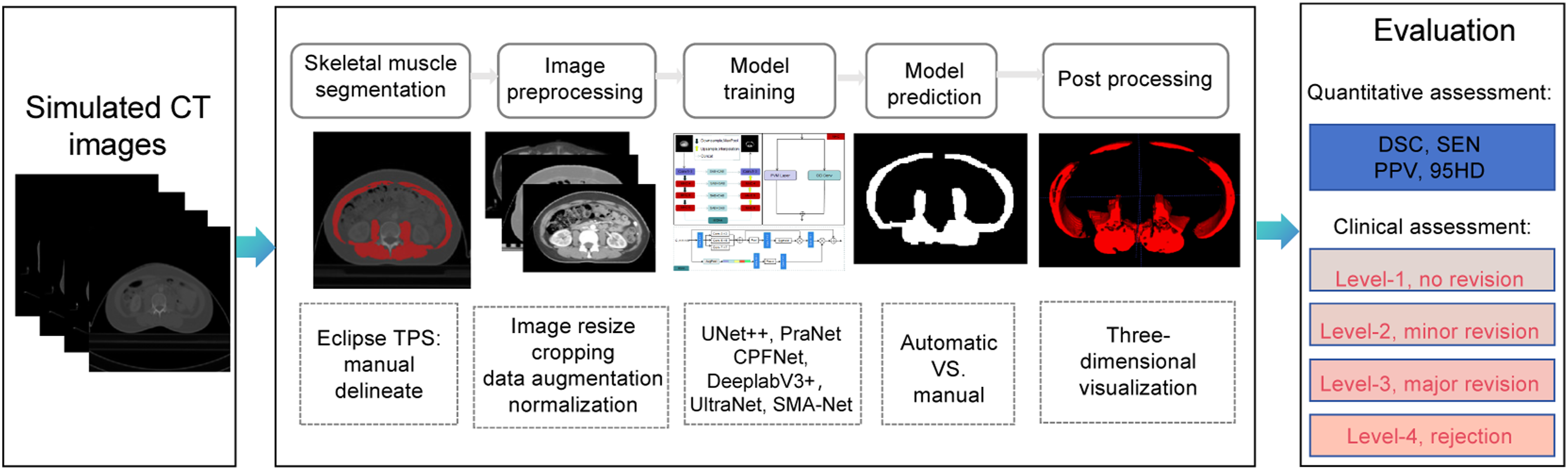
In our work, A new architecture called SMA-Net is proposed and presented in Figure 3, which is a typical U-Net variant. Based on UltraLight VM-UNet, we proposed Multi-Hybrid Convolutional (MHC) modules and introduced Multi-Scale Attention Aggregation (MSAA). MHC module is composed of a parallelized Parallel Vision Mamba (PVM) layer
14
and Omni-Dimensional Dynamic Convolution (ODConv).
17
The performances of U-Net-based segmentation models may be weakened if we concatenate these feature maps directly using skip connections. Self-Attention Block (SAB) and Cross-Attention Block (CAB) was used to eliminate semantic differences between different depth feature layers. Detailed segmentation network of the skeletal muscle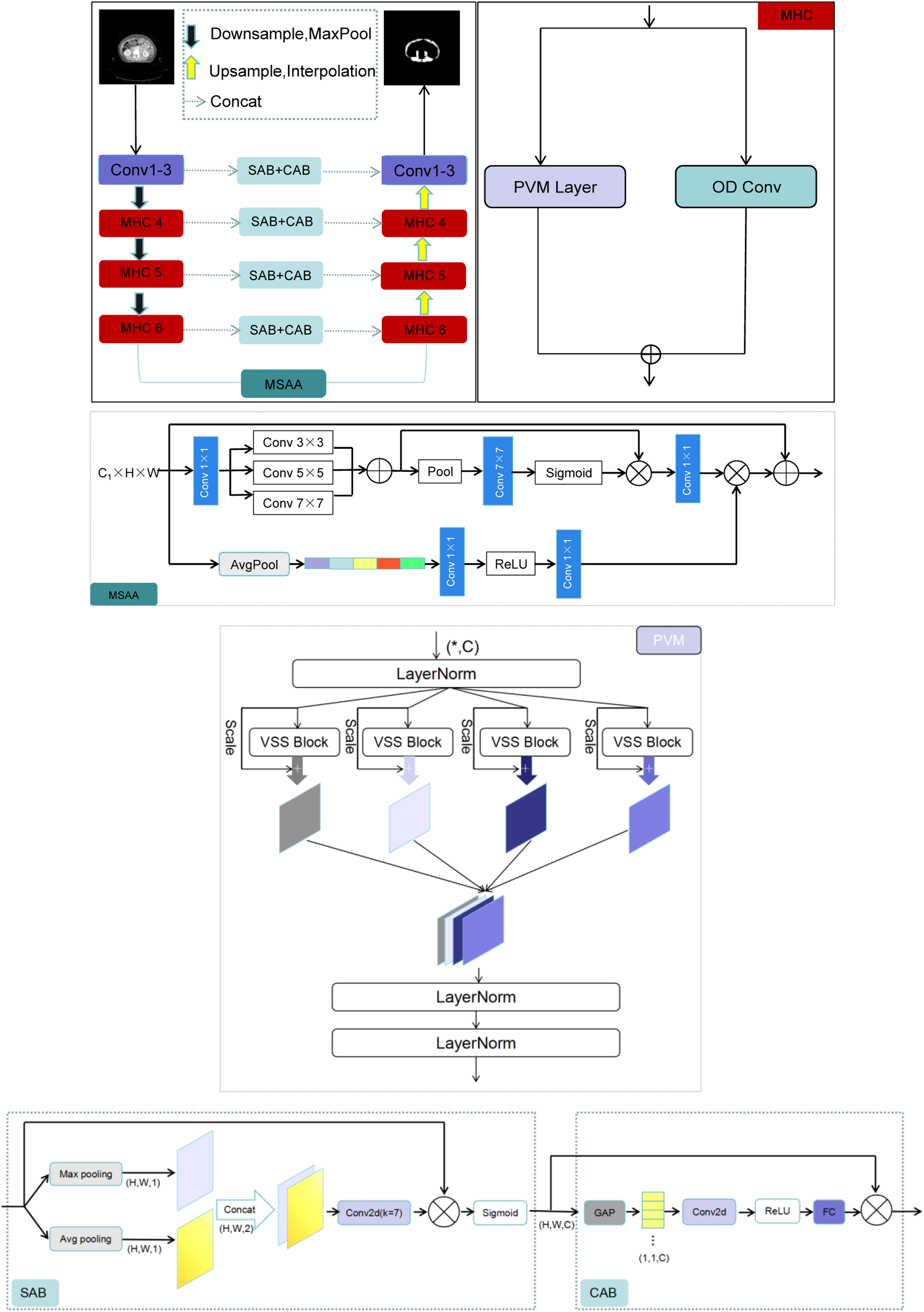
MHC Module
MHC module is composed of PVM and dynamic convolution (ODConv) in parallel. ODConv is an advanced dynamic convolution design. The aim is to improve the feature learning ability of convolutional neural networks by introducing a multi-dimensional attention mechanism. Unlike traditional methods that only focus on one dimension of the number of convolution kernels, ODConv learns complementary attention on four dimensions: space size, number of input channels, and number of output channels, thereby optimizing convolution operations. Both PVM and ODConv have lightweight features and both local&global feature extraction capabilities. A feature X with channel number C first passes through a LayeNorm layer and then is divided into
The core advantage of the MHC module is that it has the ability to capture long-range dependencies, strong local feature extraction capabilities, dynamic adaptability, and efficient computing characteristics. It breaks the limitations of traditional convolution and Mamba model, and achieves functional complementarity and performance leap.
MSAA Module
The deepest encoder and decoder are connected by MSAA module. In MSAA, spatial and channel dual paths are used for feature aggregation. The channel refinement starts with channel projection through 1×1 convolution, reducing the number of channels from C1 to C2. Multi-scale fusion includes convolution summation of different kernel sizes, such as 3×3, 5×5, 7×7. Subsequently, mean pooling and maximum pooling are used to aggregate spatial features, followed by 7x7 convolution and element-by-element multiplication with the sigmoid-activated feature map. The MSAA module aims to better capture multi-scale context information by aggregating features of different scales, which can enhance segmentation details, such as muscle edges, so as to improve the clarity of segmentation results.
SAB and CAB Module
The skip-connection path uses the SAB module and the CAB module.
14
The combined use of SAB and CAB allows for the fusion of multi-stage features of different scales of our network. An input sequence X∈Rn×d, where n is the length of the sequence and d is the characteristic dimension. Through three different linear transformations, Q, K, V three matrices are generated from the input X. Softmax is used to obtain the attention weight, and the attention weight matrix A is used to weight and sum the Value matrix V to obtain the output sequence Z. The specific operation can be expressed by the following equations. The detail introduction is from ref. 14.
The core contribution of PVM lies in its ingenious parallelization design, which effectively constrains the growth of Mamba model parameters, thereby achieving the ultimate lightweight of the model without significantly sacrificing performance. ODConv utilizes a novel multi-dimensional attention mechanism and parallel strategy to learn complementary attention of convolutional kernels in all four dimensions of the convolutional kernel space of any convolutional layer. Compared to traditional convolution, it can reduce redundant parameters and lower computational complexity. The MSAA module is designed to better capture multi-scale contextual information by aggregating features across different scales. The MSAA including Multi-Scale fusion module, spatial aggregation module and channel aggregation module. Multi-scale fusion involves summing convolutions with different kernel sizes, such as 3×3, 5×5, and 7×7. Subsequently, mean pooling and max pooling are used to aggregate spatial features, followed by a 7×7 convolution and element-wise multiplication with the feature map activated by sigmoid. Meanwhile, channel aggregation uses global average pooling to reduce the dimension to 1, and then generates a channel attention map through convolution and ReLU activation. This map is expanded to match the input dimension and combined with the spatial refinement map. Therefore, MSAA enhances spatial and channel features in subsequent network layers. By incorporating the MSAA module, the resulting feature map is enriched with refined spatial and channel information. The network was based on the open-source library Pytorch 1.13.0 framework, CUDA 117,Python 3.8.19 and the reference implementation of U-Net. The structure of SMA-Net consists of an encoder on the left side (used to extract image features) and a decoder on the right side (to reconstruct the segmentation map based on the extracted features). The size of the input CT image is 3 × 320 × 320. First, through a convolutional layer, the shallow features are extracted, and the image size becomes 32 × 320 × 320. Then, through a GroupNorm layer, which normalizes the input data so that the mean value of the data in each feature channel group is 0 and the variance is 1. This processing helps to reduce the scale difference between different feature channels, making it easier for the network to learn the correlation between features. Next is the down-sampling layer, we use MaxPool as the down-sampling method. The down-sampling does not change the number of channels of the feature image, but only changes the width and height of the image. At this time, the size of the image becomes 32 × 160 × 160. Then, an activation function GeLu was used to deal with the feature map. This smoothness of GeLu helps to reduce the gradient disappearance problem of the neural network during the training process, so that the network can better learn the characteristics of the input data, and can also accelerate the training convergence speed of the neural network. Repeat the above operation twice, the size of the image becomes 96 × 40 × 40.
The above output feature layer goes through three MHC. The Mamba architecture is known for its ability to process long sequences and global context information and its improved computational efficiency as a state space model. It can effectively model long-range dependencies in medical images, which is a key aspect for accurate segmentation. The visual state space (VSS) model represented by Mamba has become a promising method. The VSS structure is shown, in order to further improve the computational efficiency and eliminate the entire multiplication branch, because the effect of the gating mechanism is achieved by the selectivity of 2D selective scanning. Therefore, the VSS module is composed of a single network branch with two residual modules, which imitates the architecture of the ordinary Transformer module. The MHC architecture extracts deeper feature information, the number of channels is deepened, and the image size is 128 × 40 × 40. In this way, MHC is repeated twice, and the deepest feature layer is 256 × 10 × 10. In the skip connection path in the middle of the network, the network model uses the Self-Attention Block (SAB) and Cross-Attention Block (CAB) 14 to realize the fusion of multi-scale information. The decoding layer on the right side recovers the size of the network feature layer in turn. The feature of the network model is that the number of channels is set to 32, 64, 96, 128, 192, 256, respectively. This design helps to capture the details and context information of the image at different levels.
Model Training
The 160 annotated CT images were randomly divided into training set (N=116), validation set (N=16) and testing set (N=32) in the ratio of 7:1:2. The optimizer is Adam. The total loss function used in this study is a combination of the Dice loss function (Ldice) and Cross-Entropy loss function (Lce), which was shown below. The detail formula of Ldice and Lce is listed in ref. 18.
A cosine annealing scheduler were used (learning rate: 10−4, weight decay: 10−4). The training was terminated when the model loss did not decline in 100 epochs. The batch size was set as 16. Data augmentation techniques such as rotation, translation, flipping and cropping were applied. The pre-trained weights obtained from the large-scale trained on the ImageNet datasets. 19 The networks were run on a server with NVIDIA A100 graphics card and a 32 GB RAM.
Evaluation
The automatic segmentation performance was evaluated by the geometric metrics and clinical oncologist’s evaluation. The geometric metrics are dice similarity coefficient (DSC), sensitivity (SEN), positive predictive value (PPV), and 95% Hausdorff distance (HD95), which were seen below.
Where the TP, TN, FP and FN represents the number of true positive samples, true negative samples, false positive samples and false negative samples, respectively. h(C, P) = max{min||c-p||} (c∈C, p∈P) and h(P, C) = max{min||p-c||} (p∈P, c∈C). HD represents the distance between the surface point sets of the calculated true sample and the predicted sample. In order to eliminate the error, 95% HD (95HD) is used to eliminate the error effect caused by outliers. 95HD represents the largest surface-to-surface distance among the closest 95% surface voxels.
In addition, the automatic segmented results were examined and revised slice-by-slice by senior radiologists. The mean time of manual segmentation process for skeletal muscle takes 20 minutes to finish. Radiologists scored each automatic segmented result according to the following 4-level criteria 20 : Level-1, no revision (automatic segmentation results are acceptable). Level-2, minor revision (need for less than 10 minutes to revise it). Level-3, major revision (need for 10 to 20 minutes to revise it), and Level-4, rejection (need for more than 20 minutes to revise it).
Sarcopenia Diagnosis
The automatic segmentation results of SMA-Net from 32 testing patients’ mid-plane of SMA were used to predicted SMI. We developed an in-house software platform to calculate the SMI. In 2016, the CDC established an ICD-10 code for sarcopenia, making it a recognized medical condition. 21 SMI was used for sarcopenia diagnostic standard, according to the cut-off established for women (≤38.9cm2/m2). 22 The sarcopenia classification was evaluated by confusion matrix and receiver operating characteristic curve (ROC).
Results
Patient Characteristics
Baseline Characteristics of 160 Patients Datasets

Model Segmentation Performance
Comparison of the Automatic Delineation Performance in Testing Sets


Visual comparison maps of GT and different models. First column is different CT slices. Second to seventh column are the segmentation results of GT and different models
The best results are in bold. ↑ means higher value of this metric is better, and ↓ means lower value of this metric is better.
Ablation Experiments
Ablation Experimental Results of Our Network

Clinical Assessment
The automatic segmentation results by our proposed model were reviewed slice-by-slice by a senior radiologist. The radiologist manual revised the predicted results. Figure 5 shows the distribution of radiologist’s scores on the manual revision of skeletal muscle. In testing datasets, 3 patients (9.37%) were Level-1 (no revision). 27 patients (84.38%) were Level-2 (minor revision). 2 patients (6.25%) were Level-3 (major revision). Frequency counts and relative (%) distribution of each level for skeletal muscle testing in oncologist’s scores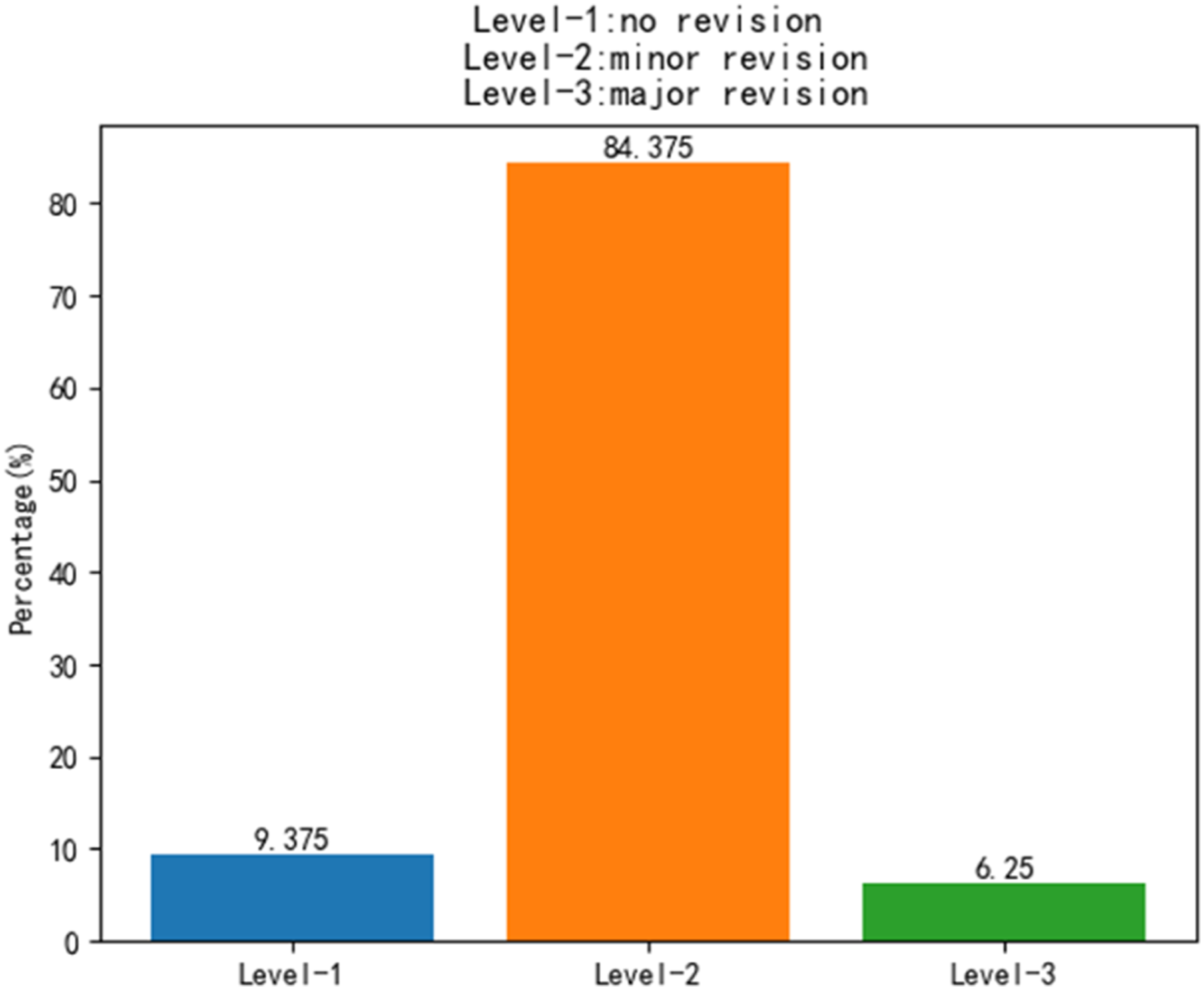
Sarcopenia Intelligent Diagnosis
The SMA derived from predicted segmentations showed near-perfect correlations with the true SMA in testing datasets, as shown in Figure 6.The sarcopenia diagnosis gold standard is L3-SMI ≤38.9 cm2/m2. The sarcopenia classification was evaluated by confusion matrix and receiver operating characteristic curve (ROC), as shown in Figure 7.The metrics of accuracy, precision, recall, F1-Score and area under curve (AUC) were calculated. Our proposed network for cervical cancer patients predicted sarcopenia with 87.5% accuracy, 92.31% precision, 80% recall, 85.72% F1-Score, and 0.871 AUC. The SMA of predicted results and true results The confusion matrix and receiver operating characteristic curve (ROC) in testing datasets

Model Complexity Comparison
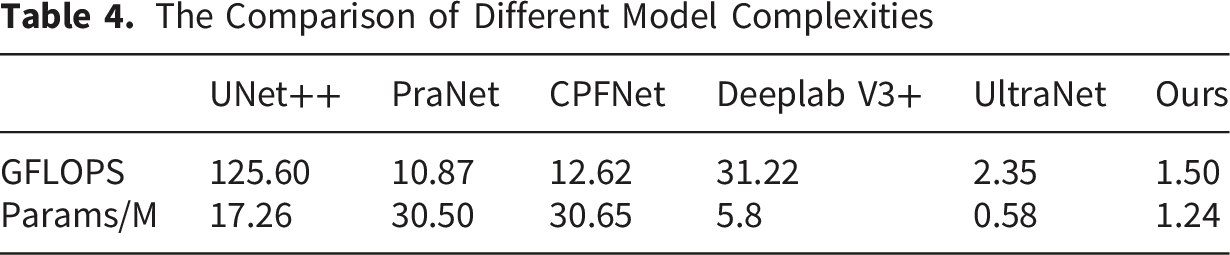
The Comparison of Different Model Complexities
Discussion
Skeletal muscle area (SMA) measurement can determine the nutritional risk of cancer patients and monitor the progression of malnutrition.23-25 At present, the planning CT is a necessary part of the radiotherapy process. In addition, studies have found26-28 that excessive loss of skeletal muscle may lead to poor survival outcomes in many types of cancer. SMI (SMA/square of height) is an important prognostic predictor. 29 Therefore, the measurement of SMA is of great significance in evaluating the nutritional status and prognosis of tumor patients.
Mamba’s state space model (SSM) achieves linear computational complexity in long sequence modeling through selective state mechanism, 30 which is more suitable for global context capture of high-resolution medical images. The advantage of the Mamba architecture is that it can efficiently process long sequences, which makes it five times faster than the traditional Transformer in reasoning speed. It uses state space to dynamically adjust the state of the model, allowing the model to selectively pass or forget information based on current data. It not only simplifies the commonly used state space modules, but also integrates linear attention-like blocks and multi-layer perceptions, so it can maintain high performance while reducing the complexity of the model. Ruan et al. 16 used the medical image segmentation model based on the pure Mamba architecture for the segmentation of skin diseases for the first time. Wu et al. 14 analyzed the key factors affecting the parameters in the Mamba architecture. Based on these findings, a parallel visual Mamba layer called PVM layer was proposed to achieve the lowest computational load and excellent performance while maintaining the total number of channels. In this work, we proposed a parallel structure including PVM layer and ODConv. The parameters were greatly reduced through two structures: dynamic sparsity, and parameter sharing, while using parallel complementarity to maintain model expression. Its combination of dynamic calculation and static efficient structure, which provides a new idea for lightweight network design. The SAB+CAB mechanism is used to suppress irrelevant information in the image and highlight important local features. The area of the segmentation result is calculated by the internal program, and the predicted area is basically consistent with the real area. The computational complexity of GFLOPS can evaluate the performance of computing devices, that is, the computational speed and efficiency of deep learning models when performing training or reasoning tasks. The number of parameters reflects the size of the occupied memory. Therefore, considering the lightweight and fast efficiency of clinical deployment, SMA-Net still has advantages over existing classical networks. About 93.75% automatic segmentation results need minor revision or no revision to meet clinical requirement, which shows that deep learning can assist doctors to greatly reduce the working time.
In recent years, deep learning methods have made important progress in intelligent segmentation of medical images. In the early stage, Burns et al. 31 divided 112 cases of abdominal CT into training set and test set, and used the deep learning algorithm UNet to intelligently segment the first to fifth lumbar (L5) skeletal muscles. The segmentation results on the test set were 0.879, 0.917, 0.930, 0.913 and 0.821, respectively. Naser et al. 13 used deep learning network to segment cervical skeletal muscle, and the DSC results reached more than 0.95. The DSC of SMA-Net network segmentation result in this study is 89.16 %, and the results of UNet++ 32 are also compared (DSC = 88.16 %), which mainly depends on the different data sets, resulting in poor generalization of the deep learning model. Furthermore, our ablation experiments showed that DSC could be improved from 87.56% to 89.16%. We comprehensively compare the proposed advanced deep learning segmentation framework with previous classical networks. It is found that the intelligent segmentation performance of the network is better than the classical network model published in the past, and the comprehensive best segmentation accuracy is obtained. The cross section of L3 skeletal muscle contains complex interfascicular fat infiltration texture, blood vessels and contrast changes between different muscle groups (such as rectus abdominis and external oblique). This requires that the model can not only capture long-distance context dependence to distinguish different muscle regions, but also adapt to local texture mutations. The MHC module uses visual Mamba and dynamic convolution in parallel. In the deepest part of the encoder, the feature map has low resolution but rich semantic information. However, there are differences in muscle size and proportion in images among different patients. A single-scale receptive field may not capture the global information of the entire muscle mass and its internal fine fat texture at the same time. We introduce MSAA, which extracts features under different receptive fields through parallel multi-scale convolution (or pooling), and automatically weighs the importance of each scale feature through the attention mechanism. For a large-size muscle, the model may give a higher weight to the large receptive field branch to integrate its overall shape; for the internal fine fat texture, the small receptive field branch will provide more important details. Compared with the radiologist’s manual segmentation, the working time is greatly saved. The doctor only needs very little time to modify on the basis of deep learning intelligent segmentation to achieve the purpose. In terms of clinical evaluation, the reason for the need for major revision is the misjudgment of the muscle group or the complete missed segmentation of the severe fat infiltration area. In some cases with low contrast, the model may incorrectly classify a part of the transverse abdominis as background or adjacent muscles. This error stems from an insufficient understanding of the context of the overall anatomical structure, resulting in the need for doctors to redraw almost the entire outline of the muscle. When the muscle is replaced by a large amount of adipose tissue, and its texture features are very similar to the background fat, the model may not be able to identify the muscle region at all, resulting in large missing segmentation. This challenges the robustness of the model to abnormal features under pathological conditions.
The results of this study show that the SMA-Net proposed in this study can accurately segment L3 skeletal muscle and quickly calculate its SMA, which basically meets the clinical application. It is helpful for clinicians to quickly diagnose sarcopenia in patients with cervical cancer. However, there are still some limitations in this study, such as limited sample size and lack of external validation. In the follow-up work, we can further combine multi-center to expand the sample size, and include more cervical cancer cases to construct a more comprehensive sarcopenia diagnosis network model.
Footnotes
Ethical Considerations
The study protocol was approved by the Ethics Committee of Huiyang Sanhe Hospital (No. Ethics [M] 2025-017). The requirement for patient consent was waived due to the retrospective nature of the study. In this project, the rights and interests of the subjects are fully protected and meet the requirements of the medical ethics committee. The research plan is approved. Approval Document No.: Ethical (M) 2025-017. The requirement for patient consent was waived.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Scientific and Technological Research Program of Chongqing Municipal Education Commission (KJQN202512819) and this project is financially supported by the Nuclear Energy Development Research Project under the State Administration of Science, Technology and Industry for National Defense (SASTIND), Grant No. HNKF202224(28).
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.