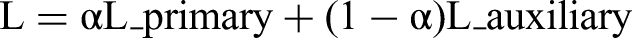Abstract
Introduction
This study evaluates the effectiveness of a lightweight vision transformer (EfficientFormerV2-S2) with a dual-output architecture for lung nodule classification, assessing its performance and generalizability across multiple datasets.
Methods
The study utilized datasets from three sources: Institution 1 (936 images), Institution 2 (280 images), and a public Zenodo dataset (308 images), comprising adenocarcinoma, squamous cell carcinoma, and benign lesions. Model evaluation included holdout validation, five-fold cross-validation, and benchmarking against the PneumoniaMedMNIST dataset. Comprehensive image preprocessing and augmentation techniques were implemented.
Results
The model demonstrated robust performance across all datasets, achieving test accuracies of 92.62 ± 1.65%, 97.14 ± 1.78%, and 95.74 ± 1.35% for Institutions 1, 2, and Zenodo respectively. Cross-validation results showed consistent performance with minimal variability (standard deviations <2%). On the PneumoniaMedMNIST benchmark, our optimized model achieved superior performance (accuracy: 0.936, AUC: 0.981) compared to ResNet18 and ResNet50 benchmarks.
Conclusion
The lightweight transformer-based model demonstrates excellent performance and generalizability across multiple institutional datasets, suggesting its potential for efficient clinical implementation in lung nodule classification tasks.
Keywords
Introduction
Lung cancer remains a leading cause of cancer-related mortality worldwide, with adenocarcinoma and squamous cell carcinoma (SCC) accounting for approximately 85% of cases.1,2 Accurate and early diagnosis is crucial for effective treatment planning, with medical imaging—particularly CT scans—serving as a primary diagnostic tool. Recent advances in artificial intelligence and deep learning have shown promising potential in enhancing diagnostic accuracy and efficiency, with transformer-based architectures emerging as particularly effective solutions for medical image analysis.
The interpretation of lung nodules on CT images presents significant challenges for radiologists, necessitating the differentiation between benign nodules and various cancer subtypes. Traditional computer-aided diagnosis approaches relied on handcrafted features or conventional deep learning architectures, which often struggled to capture the complex spatial relationships inherent in medical images. The emergence of transformer architectures has addressed many of these limitations by effectively modeling long-range dependencies and global context.
Transformer models have demonstrated remarkable performance across various domains, including medical image analysis. 3 Their self-attention mechanism enables effective modeling of relationships between distant image regions, which is particularly valuable for tasks requiring both global context understanding and fine-grained feature extraction. However, conventional transformers often have computational demands that limit their applicability in resource-constrained clinical environments.
Vision Transformers (ViTs) and Swin Transformers have revolutionized computer vision with their self-attention mechanisms and hierarchical architectures, respectively. Nevertheless, these models face notable limitations, particularly regarding their computational resource requirements. ViTs, for instance, divide images into tokens, leading to quadratic growth in token numbers with image size, which results in high memory consumption and computational overhead. This makes them less suitable for deployment in resource-constrained environments or real-time applications. Moreover, ViTs require extensive pre-training on large datasets, which can be challenging in domains with limited data availability, such as medical imaging. 4
Swin Transformers address some of these issues by introducing window-based attention mechanisms and hierarchical structures; however, their performance can degrade when transferring across different window resolutions. Additionally, training these models at scale demands significant GPU memory, further limiting their accessibility to researchers without high-end hardware.
Given these challenges, there is a pressing need to develop lightweight transformer architectures that balance efficiency and accuracy. Techniques such as adaptive tokenization and downsampling strategies have been proposed to reduce computational demands. For example, Swin Transformer V2 incorporates pooling layers to lower resource requirements while maintaining robust global dependency modeling. These advancements underscore the importance of exploring novel architectures that are computationally efficient yet capable of delivering state-of-the-art performance. Lightweight models are essential for expanding the applicability of transformers to edge devices, mobile platforms, and real-time systems, ultimately bridging the gap between cutting-edge performance and practical usability.5,6
EfficientFormer represents a significant advancement in transformer architecture optimization, offering improved computational efficiency and a reduced memory footprint compared to traditional transformer models.7,8 EfficientFormer, along with other lightweight transformers such as MobileViT, 9 addresses these limitations by incorporating innovative design principles that maintain performance while substantially reducing computational overhead.
Notably, the Swin Transformer10,11 shares similarities with EfficientFormer in its approach to efficient vision processing, employing a hierarchical structure and shifted windows for attention computation. However, EfficientFormer distinguishes itself through its dimension-dynamic mixing (DDM) mechanism, which adaptively adjusts the balance between spatial and channel dimensions, potentially offering greater flexibility in feature representation. 7 In the context of medical imaging tasks, particularly CT scan analysis, EfficientFormer's architecture may offer advantages in processing high-dimensional volumetric data more efficiently than its predecessors. Its ability to capture long-range dependencies while maintaining a compact model size could prove particularly beneficial for tasks requiring both global context understanding and fine-grained feature extraction, which are crucial in medical image interpretation. 12 However, comparative studies specifically evaluating EfficientFormer against other lightweight transformers in medical imaging contexts are limited, warranting further investigation to conclusively establish its efficacy in this domain.
Recent advancements in transformer-based approaches for lung nodule analysis have demonstrated promising results.13–19 For instance, Chen et al achieved high performance with a volumetric SWIN Transformer, obtaining 98.88% accuracy in distinguishing between benign solid lung nodules and different cancer types. 13 The emergence of lightweight architectures has been particularly noteworthy; for example, Cao et al's MSM-ViT demonstrated comparable performance to 3D models while using only a fraction of the computational resources, 14 and Maurya et al's C3-Transformer achieved impressive results with minimal parameters. 15
Additionally, Nabeel et al optimization strategies have shown significant impact, 20 with studies demonstrating improved lung cancer classification through systematic hyperparameter tuning and Yang et al hybrid optimization algorithms. 21 While efficient neural architectures have shown promise for multi-modal cancer classification [Uddin et al]. 22 These developments, combined with hybrid feature optimization techniques [Li et al],23,24 highlight the potential of efficient transformer architectures in medical imaging applications while emphasizing the need for further investigation into their clinical efficacy.
These developments highlight the potential of efficient transformer architectures in medical imaging applications while emphasizing the need for further investigation into their clinical efficacy.
In the current study, we present the application of EfficientFormerV2-S2 for lung nodule classification. Despite its potential advantages, applications of EfficientFormer in medical imaging have been limited. The proposed dual-output architecture combines transformer efficiency with auxiliary classification to enhance feature learning while maintaining clinical practicality.
Materials and Methods
Datasets
The study utilized a comprehensive dataset collection comprising data from three distinct sources. Each dataset was independently processed and divided into training, validation, and test sets to ensure robust evaluation of the model's performance across different data distributions.
Institution 1 (I1) contributed 274 cases (936 images), non-enhanced CT dataset, which contain 119 cases (377 images) of adenocarcinoma (ADC), 93 cases (357 images) of benign lesions, and 62 cases (200 images) of SCC.
Institution 2 (I2) provided 47 cases (280 images), with its non-enhanced CT dataset, consisting of 20 ADC cases (118 images), 18 benign cases (108 images), and 9 SCC cases (54 images).
Additionally, a public dataset from Zenodo (Jian et al 2024) 20 contributed 95 cases (308 images), comprising 172 ADC images, 103 benign images, and 33 SCC images.
To evaluate the model's performance and generalizability, it was also benchmarked using the pneumoniaMedMNIST dataset, which is part of the larger MedMNIST collection. 25 PneumoniaMedMNIST contains 5856 chest x-ray images with binary classes 0,1.
For each of the three primary datasets (I1, I2, and Zenodo), we employed a consistent data division strategy: 70% of the data was allocated to the training set, 15% to the validation set, and 15% to the test set. This division was performed at the patient level to ensure that images from the same patient appeared only in one of the three sets, thus preventing data leakage. Class distributions were maintained across all splits to ensure balanced representation of each nodule type.
Cross-Domain Validation Design
To evaluate model generalizability across different imaging protocols and patient populations, each dataset was acquired using distinct CT scanner configurations. Institution 1 utilized Siemens Definition AS + (128-slice and 64-slice) scanners, Institution 2 employed Canon Aquilion ONE Vision (640-slice) scanners, while the Jian dataset incorporated multiple manufacturers (GE, Siemens, UIH) with varying slice configurations and convolution kernels (B70f, B60f). Each dataset was trained and validated independently, with the others serving as implicit external validation sets, thereby providing robust assessment of cross-domain performance under different scanner vendors, protocols, and patient demographics.
Model Architecture
EfficientFormerV2-S2 is a hybrid CNN-Transformer architecture consisting of four stages: the first two stages employ MetaBlocks with depth-wise separable convolutions (32 and 48 channels respectively), while the latter two stages utilize DDM Transformer blocks (96 and 176 channels). The key innovation is the DDM mechanism, formulated as DDM(X) = γ × SpatialMix(X) + (1-γ) × ChannelMix(X), which adaptively balances spatial and channel mixing through a learnable parameter γ and employs pooling-based attention to achieve O(HW) complexity instead of the traditional O(H²W²) self-attention. The model processes 224 × 224 input images through 4 × 4 patch embedding, progressive downsampling across the four stages, and global average pooling before classification, achieving efficient deployment with 12.13 M parameters, 1.3G FLOPs, and 48.5 MB memory footprint while maintaining competitive performance through the combination of CNN inductive biases and Transformer global modeling capabilities.
We developed a dual-output transformer-based deep learning model for image classification, leveraging the EfficientFormerV2-S2 architecture as the backbone network. The model incorporates a primary classification pathway and an auxiliary classification branch to enhance feature learning and model regularization. The backbone network, pretrained on ImageNet, extracts hierarchical features from the input images. These features are then processed through two parallel classification heads: the primary classifier and the auxiliary classifier, each implemented as a fully connected layer. A dropout mechanism is strategically positioned between the backbone and classification heads to mitigate overfitting and improve generalization.
Network Components
- Backbone
EfficientFormerV2-S2 pretrained network
- Feature Dimension:
Adaptive feature space determined by the backbone architecture
- Classification heads:
Primary classifier: Linear layer mapping features to class probabilities
Auxiliary classifier: Parallel linear layer providing supplementary class predictions
- Regularization:
Dropout layer with configurable rate preceding both classification heads
As shown in Figure 1, the proposed model architecture incorporates a dual-output classification strategy built upon the EfficientFormerV2-S2 backbone.
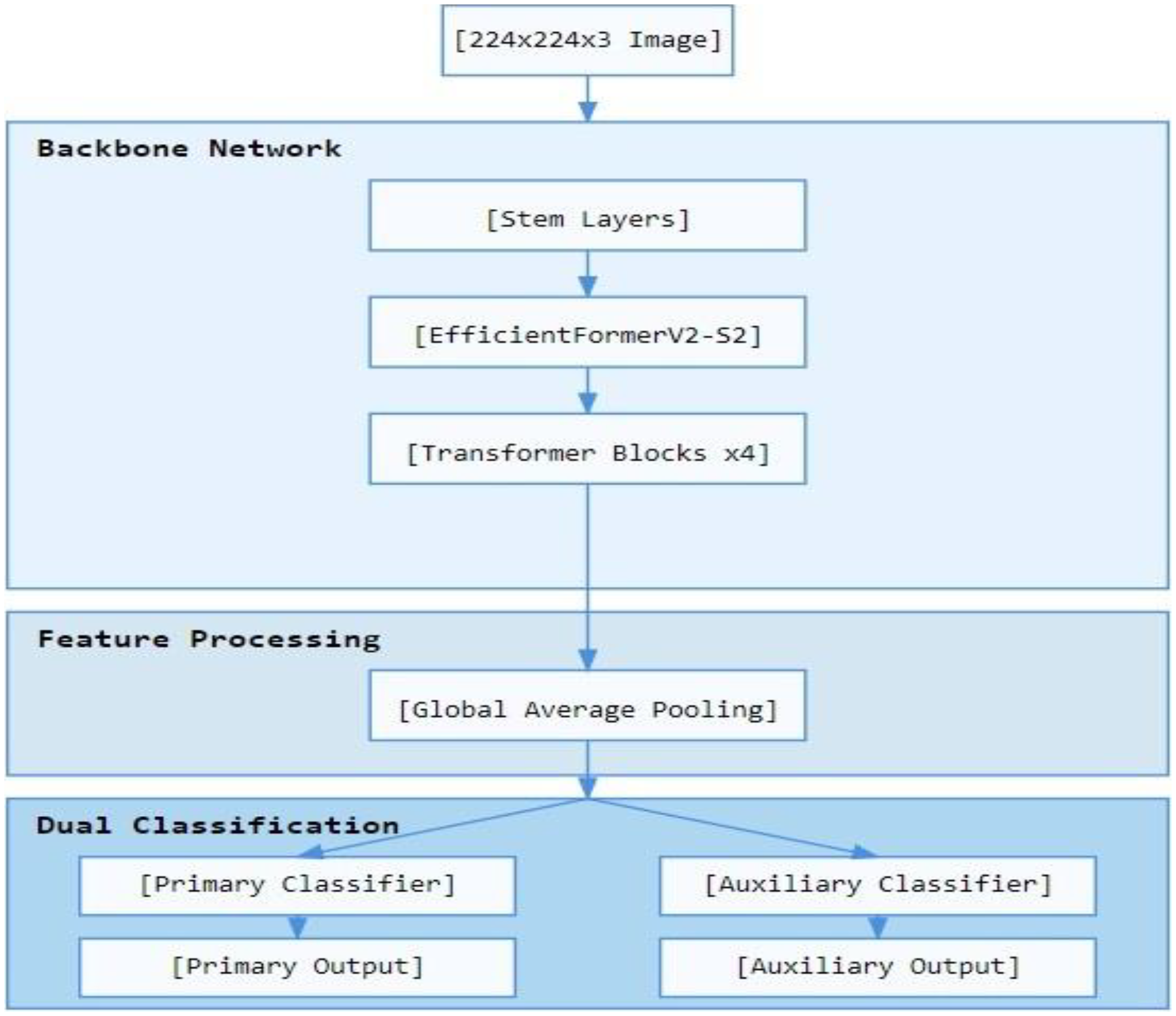
Schematic diagram of the proposed dual-output EfficientFormerV2-S2 model for lung nodule classification, showing the backbone network, feature processing, and dual classification pathways.
Image Preprocessing and Augmentation
Preprocessing DICOM images in 3D Slicer
26
by resampling the images to 1 mm thickness with 512 × 512 pixel resolution and adjusting window settings to lung-specific parameters (width −600 to 1500 HU), then generating 2D images representing different locations of each nodule saved in PNG format. To enhance model robustness and address potential data scarcity, we implemented a comprehensive image augmentation pipeline. Input images were standardized through resize operations and normalized using ImageNet statistics (mean = [0.485, 0.456, 0.406], standard deviation = [0.229, 0.224, 0.225]). The augmentation strategy comprised:
- Geometric Transformations: - Random horizontal flipping - Random rotation within ±15 degrees - Color Space Augmentations: - Brightness adjustment (±20%) - Contrast variation (±20%) - Saturation modification (±20%) - Hue shifts (±10%)
Ablation Study Design and Methodology
To evaluate our proposed EfficientFormerV2-S2 model, we conducted comparative analysis with state-of-the-art models on dataset I1, focusing on efficiency metrics including inference time, memory usage, and model parameters.
We tested different architectures, including ConvNeXtV2-Tiny, 27 ViT-Tiny-Patch16-224,28,29 EfficientNet-B3, 30 and SwinV2-CR-Tiny-NS-224, 11 as backbones to compare their performance and efficiency against our EfficientFormerV2-S2 model.
To evaluate the auxiliary classification head's contribution, we conducted ablation studies across all five backbone architectures. Models with(dual-output) and without (single head) auxiliary heads were trained under identical conditions: 224 × 224 input resolution, standard data augmentation (horizontal flip, ± 15° rotation, color jitter), AdamW optimizer (lr = 0.0001, weight decay = 0.01, batch size = 32), cross-entropy loss, and early stopping (patience = 10). Each configuration was trained five times with different random seeds (42, 1, 123, 456, 789) using a 70%-15%-15% stratified split, enabling statistical analysis with 95% confidence intervals.
Training Protocol
Configuration Optimization
We employed a hold-out validation strategy to identify optimal hyperparameter configurations. The dataset was partitioned into training (70%), validation (15%), and test (15%) sets, maintaining class distributions across splits.
Hyperparameter Optimization
We conducted a systematic evaluation of six distinct configurations, varying key parameters:
Mini-batch sizes: {16, 32, 64} Dropout rates: {0.3, 0.5} Weight decay coefficients: {0.01, 0.1}
Training Parameters
- Optimizer: AdamW with initial learning rate of 1e-4 - Loss Function: We implemented a custom dual-task loss function to optimize both the primary and auxiliary classification heads simultaneously. The loss function is mathematically formulated as: - L_primary is the cross-entropy loss for the main classification task - L_auxiliary is the cross-entropy loss for the auxiliary classification task - α is the weighting hyperparameter balancing the two objectives (set to 0.5)
where:
The equal weighting (α = 0.5) was chosen to ensure balanced learning between the two classification heads, encouraging the model to develop complementary feature representations. This approach helps prevent overfitting by requiring the network to optimize for the same classification task through two different pathways, effectively serving as a form of regularization. The auxiliary head provides gradient diversity during backpropagation, which helps the shared backbone learn more robust and generalizable features.
- Learning Rate Scheduler: ReduceLROnPlateau - Monitoring metric: Validation loss - Patience: 5 epochs - Reduction factor: 0.1 - Early Stopping: - Patience: 10 epochs - Minimum delta: 0.001 - Monitoring metric: Validation AUC - Mode: ‘max’ (monitoring for increases in validation AUC) - Implementation: Training was automatically terminated when the validation AUC failed to improve by at least 0.001 for 10 consecutive epochs
We specifically chose AUC as the monitoring metric due to its superior ability to differentiate between model performances and its enhanced interpretability in clinical contexts. Unlike accuracy, AUC provides a threshold-independent measure of model discrimination capability, making it particularly suitable for medical image classification tasks where class imbalance may be present.
The early stopping counter was reset whenever a new best validation AUC was achieved. Upon triggering early stopping, the model weights from the epoch with the highest validation AUC were restored for final evaluation. This approach ensured optimal model performance while preventing unnecessary computational expense and reducing the risk of overfitting to the training data.
Cross-Validation
The optimal configuration underwent 5-fold cross-validation to ensure robustness and generalizability. Each fold maintained consistent training protocols while providing independent validation of model performance.
Evaluation Metrics
Model performance was assessed using a comprehensive set of metrics:
- Area Under the Receiver Operating Characteristic Curve (AUC-ROC) - Classification Accuracy - Sensitivity (Recall) - Specificity - Loss Values (Primary and Auxiliary)
Reproducibility
- Random Seed: Fixed (42) for all random operations - Data Splitting: Deterministic partitioning using torch. Generator - Model Initialization: Consistent weight initialization across experiments
Model Visualization
For qualitative analysis of the model's attention patterns, we implemented Gradient-weighted Class Activation Mapping (Grad-CAM) 31 on randomly selected cases. This visualization technique was applied to the final transformer blocks of the EfficientFormerV2-S2 backbone, generating heatmaps that highlight regions contributing to the model's predictions. Grad-CAM was implemented post-training and used to provide examples of how the model classifies images.
Software Environment
The model was implemented using PyTorch 2.0 as the primary framework, with supporting deep learning libraries including torchvision and timm for the EfficientFormerV2-S2 implementation. Additional supporting libraries included NumPy for numerical computations, scikit-learn for metric calculations, Matplotlib and seaborn for visualization, and OpenCV (cv2) and PIL for image processing and Grad-CAM visualization. The entire implementation was conducted on Google Colab's cloud-based Python environment utilizing the standard GPU runtime, with Google Drive integration for data and model storage.
Results
The model performance was evaluated through different phases: comparative analysis with several state-of-the-art models, initial model optimization utilizing holdout validation, comprehensive cross-validation assessment, and benchmark comparison against established architectures using the PneumoniaMedMNIST dataset.
Ablation Analysis Results
The systematic ablation study revealed architecture-dependent effects of the auxiliary classification head across all evaluated models (Table 1). Under identical training conditions, the dual-output architecture showed varying improvements depending on the backbone. Visual Comparison of Efficiency Metrics Across Models are shown in Figure 2.

Visual Comparison of Efficiency Metrics Across Models.
Ablation Study and Efficiency Analysis of Neural Network Architectures with and without Auxiliary Classification Heads.

Performance evaluation encompasses predictive accuracy metrics (test accuracy, AUC, recall, specificity) and computational efficiency measures (parameter count, memory usage, inference time).
EfficientFormerV2-S2 demonstrated consistent improvements with the auxiliary head: test accuracy increased 0.28% (93.90% vs 93.62%), AUC improved 0.13% (98.03% vs 97.90%), recall enhanced 0.21% (94.82% vs 94.61%), and specificity increased 0.14% (96.64% vs 96.50%). These improvements came with negligible computational overhead (0.01 MB memory increase, unchanged inference time).
ConvNextV2-Tiny showed the most pronounced benefit, with notable accuracy improvements (93.19% ± 1.63% vs 91.77% ± 1.19%) and maintained high AUC performance (97.93% ± 0.70% vs 97.54% ± 0.55%). EfficientNet-B3 demonstrated minimal performance differences but improved training stability with reduced standard deviations. ViT-Tiny and SwinV2-CR-Tiny showed modest improvements with enhanced consistency across random seeds.
Computational overhead was minimal across all architectures: parameter increases ranged 579-4611 (< 0.02% increase), memory overhead was 2.3-9.2 KB, and inference time variations (±0.2-0.9 ms) were attributable to measurement variability rather than architectural differences.
Model Optimization with Holdout Method
Through the evaluation of multiple hyperparameter configurations, optimal settings were identified for each dataset. The optimization process involved testing various combinations of batch sizes: {16, 32, 64}, Dropout rates: {0.3, 0.5},Weight decay coefficients: {0.01, 0.1}, yielding dataset-specific configurations that maximized model performance.
For the I1 dataset, optimal performance was achieved with Configuration 3:
- Hyperparameters: batch size 32, AdamW optimizer, dropout rate 0.3, weight decay 0.01, learning rate 1e-4
The I2 dataset exhibited superior performance under Configuration 2:
- Hyperparameters: batch size 16, AdamW optimizer, dropout rate 0.5, weight decay 0.1, learning rate 1e-4
For the Jian et al(2024) dataset, Configuration 4 yielded optimal results:
- Hyperparameters: batch size 32, AdamW optimizer, dropout rate 0.5, weight decay 0.1, learning rate 1e-4
Five-Fold Cross-Validation Analysis
To establish the robustness and generalizability of our approach, we conducted comprehensive five-fold cross-validation using the optimal configurations. The analysis revealed consistent performance with minimal variability across all datasets:
Training Performance:
The model exhibited robust training characteristics across all datasets:
- I1 dataset: 97.34 ± 0.80% accuracy, 99.66 ± 0.17% AUC - I2 dataset: 97.35 ± 0.68% accuracy, 99.88 ± 0.05% AUC - Jian dataset: 98.98 ± 0.54% accuracy, 99.96 ± 0.04% AUC
Validation and Test Performance:
Cross-validation results demonstrated strong generalization capabilities:
- I2 dataset showed exceptional validation performance (99.52 ± 0.95% accuracy) - Test set performance remained consistently high: * I1: 92.62 ± 1.65% accuracy, 98.42 ± 0.60% AUC
* I2: 97.14 ± 1.78% accuracy, 99.87 ± 0.18% AUC
* Jian: 95.74 ± 1.35% accuracy, 98.59 ± 1.01% AUC
As shown in Figure 3, the five-fold cross-validation metrics (accuracy, and AUC) demonstrate consistent model performance across all three datasets. The model's classification performance across different datasets is illustrated through confusion matrices in Figure 4, demonstrating robust performance across all three institutions.

Five-fold cross-validation performance metrics across different datasets. Model performance metrics (accuracy, loss, and AUC) for (A) Institution 1, (B) Institution 2, and (C) Jian et al. (2024) dataset.
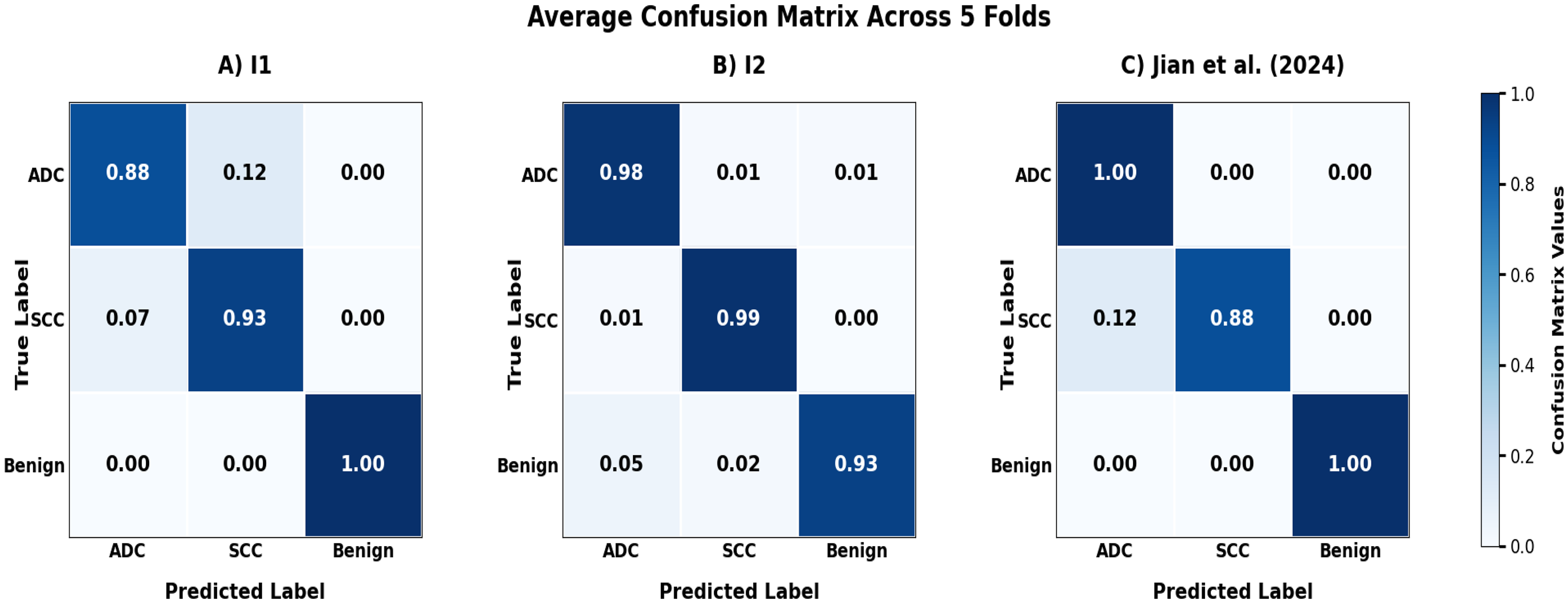
Average Confusion Matrices from 5-Fold Cross-validation Test Sets for (A) Institution 1 (I1), (B) Institution 2 (I2), and (C) Jian et al (2024) Dataset.
The consistently low standard deviations across all metrics (typically <2%) indicate remarkable stability in model performance across different data partitions. Table 2 presents the average over the five folds performance metrics obtained using the best configurations, with results averaged across all folds.
Five-Fold Cross-Validation Performance Metrics Using Best Configurations Averaged Across all Folds.

All reported metrics represent averaged performance across all five folds of cross-validation, with standard deviations indicating consistency across folds. The approach validates model robustness through comprehensive 5-fold cross-validation rather than reporting a single fold.
Confusion matrix and false positive false negtive cases analysis
Malignant Nodule Detection
Analysis of confusion matrices demonstrates clinically acceptable false negative rates for cancer detection across all institutions. Adenocarcinoma miss rates ranged from 1-9%, with Institution 2 achieving optimal performance (1%) and Institution 1 showing 9%. SCC detection rates were consistent, with false negative rates of 6-8% across datasets.
Benign Nodule Classification
False positive rates remained low across institutions, ranging from 4-9%. These rates represent manageable levels of unnecessary follow-up procedures while maintaining cancer detection sensitivity. Institution 2 demonstrated the lowest false positive rate (4%), followed by Jian dataset (7%) and Institution 1 (9%).
Inter-Cancer Misclassification
Misclassifications between adenocarcinoma and SCC occurred in 3-10% of cases. While these errors affect histologic-specific treatment selection, they represent lower clinical risk since both cancer types require immediate oncological management.
Clinical Significance
The model's performance profile supports clinical deployment as a diagnostic aid. False negative rates for malignant nodules (1-9%) fall within acceptable parameters for AI-assisted diagnosis, while controlled false positive rates (4-9%) minimize unnecessary interventions. The consistent performance across different institutional settings and scanner protocols demonstrates robust generalizability for clinical implementation.
Grad-CAM Visualization
To provide interpretability of the model's behavior, we applied Grad-CAM visualization to representative cases from the Institution 2 dataset. As shown in Figure 5, the visualization revealed varied attention patterns in correctly classified adenocarcinoma cases, with the model demonstrating both localized focus on specific lung regions and broader attention patterns, suggesting flexibility in feature detection strategies.

Grad-CAM Visualizations of Adenocarcinoma Cases from Institution 2, Showing Original CT Images with Radiologist Annotations (Red Circles), Attention Heatmaps, and Superimposed Views Demonstrating Varied Model Attention Patterns.
Benchmarking Using a Separate Dataset
We adopted the original hyperparameter configuration as described by Yang et al (2023). 25 The model architecture employs a pretrained EfficientFormerV2-S2 as the backbone in a multitask transformer configuration for binary classification. Training was conducted using the Adam optimizer with a learning rate of 0.001, batch size of 128, and dropout rate of 0.2. We implemented a custom MultitaskLoss function with α = 0.5 to balance the main and auxiliary tasks. The learning rate was managed using ReduceLROnPlateau scheduler (patience = 3, factor = 0.1) with early stopping (patience = 5) based on validation AUC. Input images were resized to 224 × 224 pixels, converted from grayscale to 3-channel format, and normalized using ImageNet statistics (mean = [0.485, 0.456, 0.406], std = [0.229, 0.224, 0.225]). Training was conducted for a maximum of 100 epochs with model checkpoints saved at peak validation performance.
Our model achieved enhanced performance with test accuracy of 0.889 and AUC of 0.969, surpassing the benchmark models ResNet18 and ResNet50, which achieved accuracies of 0.864 and 0.884, and AUC scores of 0.956 and 0.962, respectively as shown in Table 3. While the original MedMNIST paper implemented minimal configurations without image augmentation or regularization, we evaluated the PneumoniaMedMNIST dataset using the six hyperparameter configurations previously established in this work. The optimal configuration utilized [Adamw, batch size 32, dropout: 0.5 weight_decay: 0.1] and achieved a test accuracy of 0.936 and AUC of 0.981.
Comparative Analysis of EfficientFormerV2-S2 Against ResNet Benchmarks on PneumoniaMedMNIST Dataset.

Discusion
The field of lung nodule classification represents a critical area in medical imaging that has undergone rapid evolution, particularly in the transition from traditional radiomics to advanced deep learning approaches. This evolution is evident in the literature, from early approaches using CT-based radiomics signatures for nodule differentiation 32 to the emergence of deep learning solutions for binary classification challenges. 33 While these earlier approaches established fundamental methodologies, they highlighted a crucial limitation in the field - the predominance of binary classification models that may not fully address the complexity of clinical scenarios. The current study builds upon these foundations while addressing their limitations through a multi-class approach that better reflects clinical reality.
Recent used SWIN- transformer-based approaches in the field have shown varying levels of success, with some studies18,19,34,35 achieving promising results but requiring expensive voxel-level labels for relatively simple classification tasks. A recent advancement by Chen et al 13 demonstrated significant progress using a volumetric SWIN Transformer, achieving 98.88% accuracy in distinguishing between benign nodules, adenocarcinoma, and SCC, while eliminating the need for manual lesion outlining. However, this approach, though highly accurate, requires substantial computational resources due to its 3D volumetric processing nature. The current study explores an alternative direction by investigating the potential of a lightweight 2D transformer architecture, aiming to balance classification performance with computational efficiency for practical clinical implementation.
The current study's validation across two institutions and a public dataset revealed several key insights about the model's clinical applicability. First, the consistently high performance across these diverse datasets (two institutional and one public) suggests promising generalizability - a crucial factor for real-world implementation. The model's balanced performance profile, particularly between sensitivity and specificity metrics, addresses a common challenge in medical AI systems where high sensitivity often comes at the cost of reduced specificity. Notably, when compared to recent transformer-based approaches like Huang et al's TBFE model 36 and Cao et al's MSM-ViT, 14 the proposed model achieves comparable or superior performance while maintaining a more lightweight architecture. This balance between performance and efficiency is particularly relevant for clinical integration, where computational resources may be limited.
The comprehensive ablation analysis demonstrates that the auxiliary head provides consistent benefits across all architectures, with accuracy improvements ranging 0.14-3.40%. The auxiliary head functions primarily as a training stabilizer, reducing variance across random seeds rather than dramatically enhancing performance. Importantly, these gains come with negligible computational overhead: parameter increases remain below 0.02%, memory overhead under 0.02 MB, and inference time variations (±0.2-0.9 ms) within measurement uncertainty.
The auxiliary head enhancement maintains critical clinical requirements, with consistently low false negative rates for malignant nodules across both configurations. Balanced improvements across adenocarcinoma and SCC classifications indicate no subtype bias introduction.
However, overlapping confidence intervals between configurations for several models highlight the importance of statistical validation before claiming clinical superiority. While improvements are consistent, they often fall within seed-induced variability ranges identified in recent studies,37,38 emphasizing the need for rigorous statistical analysis in medical imaging applications where false conclusions carry high stakes.
The proposed model demonstrated robust performance across multiple metrics, with precision ranging from 93.80 ± 1.36% to 97.63 ± 0.70% and F1-scores from 93.75 ± 1.40% to 96.53 ± 1.13% across the datasets. These results demonstrate notable improvements over recent comparable studies in the field. Specifically, while Huang et al's TBFE model 36 achieved 97.06% overall accuracy, its lower recall (89.33%) and precision (90%) indicate potential performance imbalances. Cao et al's MSM-ViT 14 achieved 94.04% accuracy but utilized a more complex architecture. Additionally, while Mkindu et al's 3D-MSViT 39 and 3D-NodViT 40 achieved high sensitivities (97.81% and 98.39% respectively) on nodule detection, these approaches required more computational resources due to their 3D architecture. In contrast, the current study's lightweight 2D approach maintains competitive performance levels (sensitivity 93.79-96.34%) while offering practical advantages in terms of computational efficiency and clinical implementation potential.
The consistent high AUC scores (98.42 ± 0.60%, 99.87 ± 0.18%, and 98.59 ± 1.01% respectively) demonstrate robust discriminative ability across different institutional settings. This generalization capability is particularly noteworthy given the challenge of maintaining consistent performance across different medical imaging environments. The minimal variability in performance metrics (standard deviations <2%) further supports the model's reliability for clinical implementation.
The slightly higher AUC deviation in the Jian dataset (±1.01%) can be attributed to its non-homogeneous structure, incorporating multiple CT manufacturers (GE, Siemens, UIH) with varying slice configurations and convolution kernels (B70f, B60f), reflecting real-world clinical diversity.
Wang et al (2022) 17 demonstrated the successful application of a 2D transformer architecture for pulmonary nodule diagnosis, achieving an accuracy of 93.33% on the LIDC-IDRI dataset. Their work, along with our study, highlights the potential of 2D transformer-based approaches as an effective and computationally efficient alternative to more complex 3D models for lung nodule classification tasks.
The proposed model's effectiveness was further validated through benchmarking against established architectures using the PneumoniaMedMNIST dataset. In this comparison, the proposed EfficientFormerV2-S2 model achieved superior performance (accuracy: 0.936, AUC: 0.981) compared to both ResNet18 (accuracy: 0.864, AUC: 0.956) and ResNet50 (accuracy: 0.884, AUC: 0.962) benchmarks. 25 This performance improvement is particularly noteworthy as it demonstrates the model's capability togeneralize across different medical imaging tasks while maintaining its lightweight architectural advantages. The substantial improvement over these widely-used ResNet architectures further supports the viability of the proposed approach for practical clinical applications.
While the current study demonstrates promising results, several important limitations warrant discussion and suggest directions for future research. First, although the validation across two institutions and a public dataset provides evidence of generalizability, the model's performance across more diverse healthcare settings, including different CT scanner types and imaging protocols, remains to be established. The current datasets, while substantial, may not fully represent the complete spectrum of nodule presentations and variants seen in clinical practice. Additionally, the retrospective nature of the validation, while methodologically sound, does not fully address the challenges of real-time clinical implementation. Prospective clinical trials would be necessary to evaluate the model's performance in actual clinical workflows, including its impact on decision-making and patient outcomes. The current Grad-CAM analysis provides qualitative visualization only, without quantitative validation against radiologist-marked ROIs or statistical comparison of attention patterns.
Future studies could focus on extending the model to classify additional nodule types and pathological patterns, evaluating performance across different CT scanning protocols, and implementing the model in clinical workflows to provide valuable insights into its practical utility. Comprehensive interpretability assessment should include IoU analysis between model attention and radiologist annotations and statistical validation of attention patterns across correct versus incorrect classifications.
Conclusion
This study demonstrates the effectiveness of a lightweight dual-output transformer architecture for lung nodule classification. The model achieved consistent high performance across multiple institutions while maintaining computational efficiency. The balanced performance across accuracy, sensitivity, specificity, and AUC metrics, combined with minimal variability across datasets, suggests robust generalization capabilities. These results indicate that lightweight transformer architectures can effectively balance diagnostic accuracy with practical implementation requirements, potentially offering a more accessible path to clinical deployment. The success of this approach in maintaining performance while reducing computational overhead suggests promising directions for future development of efficient medical image analysis systems.
Footnotes
ABBREVIATION LIST
Ethical Considerations
This study was conducted as part of a national science project, with centralized ethics approval obtained from the Institutional Review Board of The Fifth Affiliated Hospital of Guangzhou Medical University [Institution 2 (I2)] (Ethics Committee approval number: KY01-2022-02-14).
Consent to Participate
The approval covers all participating institutions including The First Affiliated Hospital of Guangzhou Medical University [Institution 1 (I1)]. Written informed consent was obtained from all participants. The study was conducted in accordance with the Declaration of Helsinki and institutional guidelines for multi-site collaborative research. The public datasets from Zenodo and pneumoniaMedMNIST were already de-identified and publicly available, requiring no additional ethical approval. All institutional data was anonymized prior to analysis in accordance with relevant guidelines and regulations.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was supported by the National Natural Science Foundation of China (grant number 82272080).
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
AI Tools Disclosure
AI tools (Quillbot, Kimi AI, and Claude) were used for text enhancement, grammar checking, and rewriting portions of the manuscript to improve the clarity of the author-generated content. No AI tools were used for data collection, analysis, result generation, or scientific interpretation.